Neuro 3000 Unit 2 Flashcard Set
1/295
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced |
|---|
No study sessions yet.
296 Terms
Transgenic
any organism in which foreign DNA is introduced and stably integrated into the genome
What is the main goal of transgenesis
to express a foreign protein in specific parts of the brain
what is the main goal of gene knockout
gene knockout (see below) is to target and inactivate a specific gene, and determine what is wrong with the KO animal- IS A LOSS-OF-FUNCTION
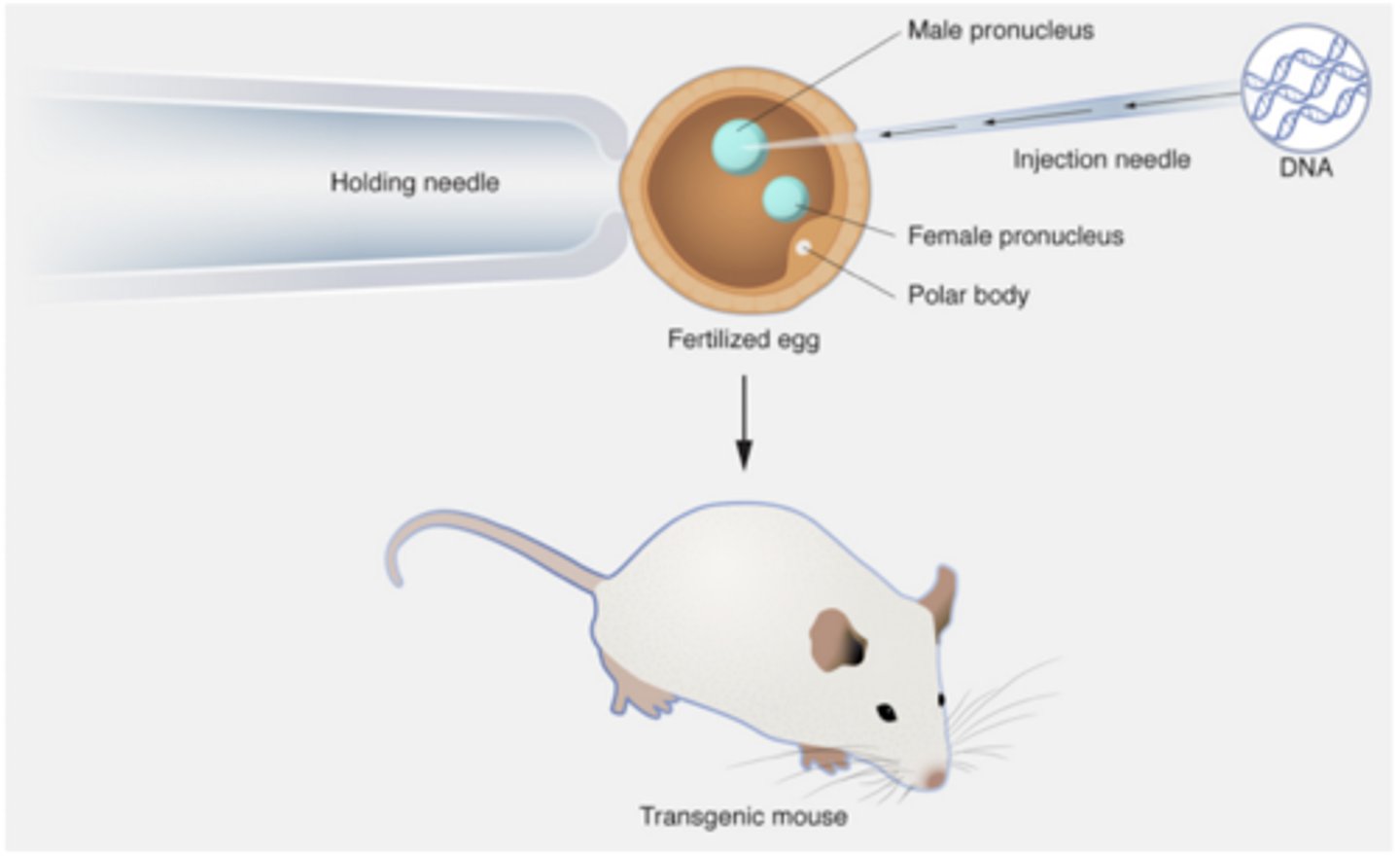
What does pronuclear injection do
DNA construct was injected directly into fertilized mouse eggs (zygotes)
Where does the pronuclear injected DNA integrate
Into a random location of the mouse's genome without a genetic locus being disrupted.
What is the more targeted approach to pronuclear injection
When injected into the ROSA26 gene locus, some of the problems associated with random integration are eliminated
What is the goal of pronuclear injection
to express some protein in a tissue/cell type of interest- CALLED GAIN-OF-FUNCTION as opposed TO gene knockout
A gene promoter
What does the promoter of the DNA construct do
It determines where the transgene will be expressed and a protein-coding sequence based on what traits the investigator wants to see expressed in the animal
The promoter of the natural gene can drive the knock-in, adding a loxP element into a gene (floxing it).
What happens once the injected zygotes are implanted into living females
19 days later, transgenic mice are born- 5-10% of the injected zygotes will result in mice that have the new piece of DNA integrated stably into every cell of the animal
What are some uses of transgenic mice in pronuclear injection
Expression of neutral or Ca-sensitive fluorescent reporters to study cell physiology in living animals. Expression of a functional protein to disrupt development or function.
How is knockout mice different than transgenic mice
a specific gene in the genome is targeted for inactivation.
What does the DNA construct do in KO mice
the purpose is the target a specific gene by recombinational mechanism
How is DNA introduced into KO mice
Mouse Embryonic Stem (ES) cells are used to introduce the DNA construct and target the genes of interest.
What happens once the gene fo interest is targeted by the DNA construct
The modified ES cells can be implanted into an early stage mouse embryo where they give rise to all tissues, organs, and germ cells.
What was the first animal created using genetically modified ES cells?
a chimera: pluripotential cells mixed with a normal embryo- it has both copies of the gene inactivated.
How are ES cells created
by dispersing and culturing inner cell mass (ICM) cells from an E 4.5 day mouse embryo under special conditions
What can happen from ES cell developement
occasional cells will become immortalized- keep on growing- the pluripotential cells can generate all cell types and tissues (not extra-embryonic) when mixed with normal embryos
Global knockout
gene is knocked out in all cells of the animal
conditional knockout
gene is knocked out in just a small tissue or cell type of the experimenter's choice (spatial control)
inducible knockout
gene is knocked out in a tissue or cell type of the experimenter's choice (spatial control) and at the time of the experimenter's choosing (temporal control)
Conditional gene targeting
First: Express CRE recombinase (a protein from P1 phage, a virus that infects bacteria) under the
control of a cell type-specific promoter (transgenic Mouse 1)
Then: Make a second mouse using knock-out methodology to introduce loxP sites in a non-essential part of the gene (this is Mouse 2, actually a "knock-in")
Finally: Cross the two mouse lines to get a tissue-specific knock-out
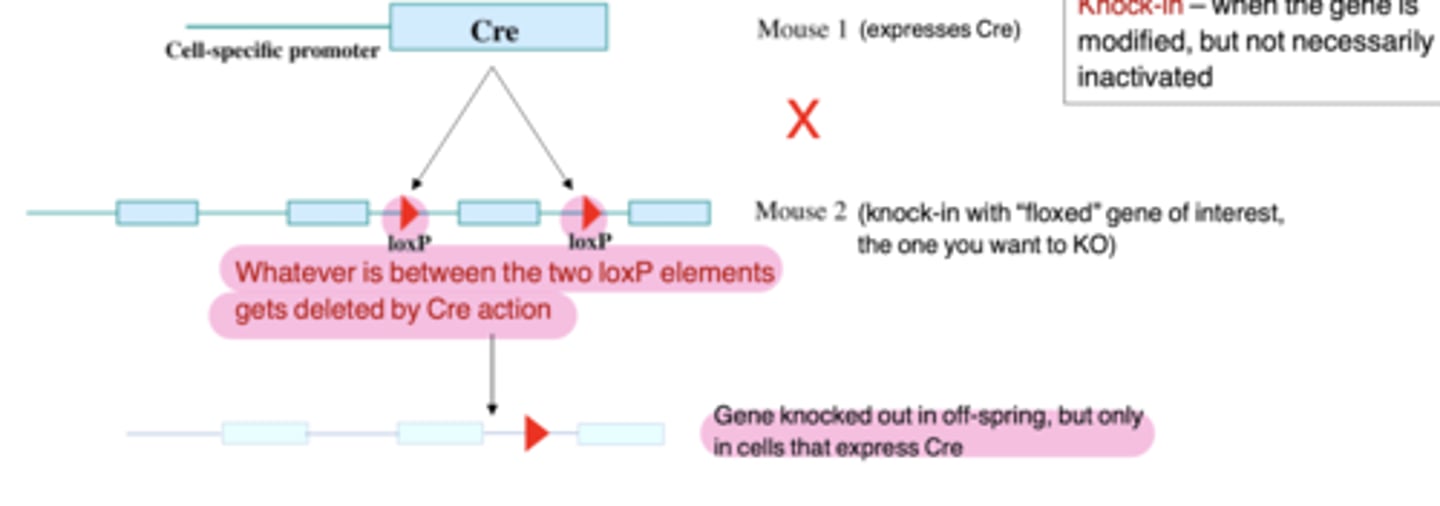
Knockout mice
when the gene is genetically inactivated
Knockin mice
when the gene is modified not necessarily inactivated
To change it in some way to express a modified version (amino acid to to study a ko=nown human mutation)
Where will the gene knockout occur?
In those cells in which the Cre recombinase is expressed. This is driven by the gene promoter that was chosen to link to Cre.
Inducible Cre (CreER)
similar to Cre, but has been fused to the ligand binding domain of the estrogen receptor
What is different between CreER and Cre
CreER protein cannot move into the nucleus until the drug tamoxifen (an analog of estrogen) is provided, either injected into animals or in cell culture medium- this adds a temporal control for the experimenter- important and beneficial for neuroscience
Human gene therapy is based on what
MeCP2 Flox-STOP mice die at 15 weeks unless you turn the gene back on a few weeks earlier - this is called late-stage rescue- inspired viral gene therapy in humans
CRISPR is...
a system of clustered regularly interspaced palindromic sequences
what is Cas9
a nuclease enzyme that evolved in bacteria for the purpose of destroying invading viruses
What do CRISPR and Cas9 generate to target viral DNA?
Short stretches of RNA that exactly match the viral DNA.
What role does Cas9 play in the CRISPR process?
Cas9 allows RNA to hybridize with its complementary DNA sequence.
What effect do CRISPR and Cas9 have on the viral DNA sequence?
They generate short random deletions or insertions of several nucleotides.
What is the result of the modifications made by CRISPR and Cas9 on the viral protein?
They destroy the protein coding sequence of the virus.
uses of CRISPR/Cas9
Engineered for use in mammalian cells for the purpose of mutating genes of choosing
Non-homologous end-joining (NHEJ)
a way to generate short deletions or insertions
for targeted gene-inactivating small mutations (single nucleotide insertion/deletion)
gRNA is
20 nucleotide stretches that include a three nucleotide NGG sequence at its end, the protospacer adjacent motif, or PAM (see diagram)engineered into a full-length sequence
Homology directed repair (HDR)
Insert a much larger DNA sequence at the position of cleavage by Cas9 by injecting a donor DNA into the cells along with Cas9 and gRNA
For gene editing and repair or inserting large DNA fragments
CRISPR/Cas9 has the potential to
correct human diseases because CRISPR can be used to edit DNA in blood cells, which can easily be taken from a patient, treated with CRISPR while in culture, then put back into the patient
RNA interference (RNAi)
reduce the level of RNA by degrading it, which reduces the amount of protein that can be made
1. Introduction of RNAi Trigger:
2. Dicer Processing and RISC Loading:
3. mRNA Cleavage and Silencing:
Introduce small interfering RNA (siRNA) or short hairpin RNA (shRNA) into the target cells (through transfection, electroporation, or viral delivery methods). The RNAi trigger complements the target mRNA for specificity.
Once inside the cell, the introduced RNA is processed by the enzyme Dicer (cleaves long double-stranded RNA (dsRNA) into 21-23 nucleotide siRNA fragments). SiRNA molecules are then incorporated into the RNA-induced silencing complex (RISC), where they guide the complex to the complementary mRNA.
The RISC, guided by the siRNA, binds to the target mRNA and induces its cleavage or degradation (silences gene expression). If the mRNA is not immediately degraded, translational repression is involved.
Uses of RNAi
Improves motor and neuropathological abnormalities in Huntington's disease mouse model- have not shown any benefit in humans with Huntington's
Huntington Holiday Effect
Temporarily lowering the levels of the mutant Huntington protein (mHtt) in Huntington's disease can provide sustained benefits beyond the reduction period.
Gives the brain cells a "holiday" from the toxic protein, allowing them to repair and function better. Often achieved through the use of antisense oligonucleotide (ASO) therapies that target and degrade the mHtt mRNA.
Oligonucleotides are
Multiple nucleotides (subunits that make up DNA) are linked together in sequence. Chemically modified to be very stable when injected into animals
Antisense oligonucleotides (ASO's) are
short synthetic single-strand DNA molecules that the investigator designs, and whose sequence complements a short stretch of sequence on the mRNA (forms a double strand RNA-DNA hybrid)
Measuring changes in gene expression
Bulk measures (no cellular resolution):
Protein – Western blotting, ELISA
mRNA – qRT-PCR, RNA-Seq
Cellular level measures (with cellular resolution = you can measure changes in specific cell types):
Protein – Immunohistochemistry (IHC), Immuno EM
mRNA – In situ hybridization, single cell (sc) RNA-Seq
Western Blotting
bulk method to detect changes in the expression of a single protein- Starting with a bulk mixture of all proteins released after cell or tissue lysis and using an antibody, detect the specific protein of interest.
How to make an antibody
Inject a species (mice, rats, rabbits, goats, guinea pigs, pigs, chickens) with a foreign antigen (protein) from a different species then collect blood 1-2 months later and harvest the antibody
preparation of protein samples of comparison by western blot
Start with the same amount of brain tissue in each case
Once you extract and purify the proteins, you determine the concentration
Use the same amount of total protein from untreated and treated animals to run a western blot
Repeat with 3-5 treated animals and 3-5 untreated.
Run western blot - use antibody and chemiluminescent detection on x-ray film to quantify the relative intensity of your specific protein band of interest
Enzyme-Linked Immunosorbent Assay (ELISA)
Bulk measures of protein or other molecules
Indirect ELISA
- Antigen sample is added to plate
- blocking buffer is added to block remaining protein-binding sites
- A suitable primary antibody is added
- A suitable secondary antibody is added- an HRPO conjugate is then added which recognizes and binds to the primary antibody
- TMB substrate is added and converted by HRPO to a detectable form
Competitive ELISA
More sensitive and quantitative than western blotting
-Typically in a plate format with 96 wells-Each well is coated with antigen (same as the
molecule you need to quantify in your sample); the kit includes a primary antibody to the antigen.
- The kit comes with the primary and secondary antibodies and the detection reagents
-The antigen in your samples ‘ competes’ for binding to the antigen that is coated on the well and decreases binding of the antibody (so less color reaction, means more antigen in your sample)
Quantitative (q)RT-PCR (qRT-PCR)
About 10,000 genes are expressed at some level in a given brain region of interest; RNA extracted from that tissue will be a complex mixture of all the RNA transcribed from all of the expressed genes- the level of expression is determined by the transcriptional rate of that gene. Highly expressed genes will make up a higher percentage and signal of the total RNA than the lower expressed gene.
What is PCR
A method to amplify the signal of the RNA of interst
Cycle of PCR is
Denaturation (heating), primer annealing (cooling), and primer extension (DNA synthesis)
Primers in PCR are used to
amplify a stretch of the sequence of interest
How many cycles are there in the typical process of specificity via PCR
20-30 cycles- 1-million fold (exponential amplification
Reverse transcriptase
RNA back to DNA because DNA is much more stable than RNA
qPCR
Powerful quantitative PCR method- low throughput (can only quantify one RNA at a time).
How long does it take to detect baseline signal in qPCR before exponential growth occurs
around 20 cycles
Preparation of RNA samples for comparison by qPCR
Start with the same amount of brain tissue in each case
Once you extract and purify the RNA, you determine the concentration
Use the same amount of total RNA to run 2 separate qPCR reactions- one for uncreated and one for treated
Repeat with 3-5 treated animals vs untreated animals
Use comparative CT method to quantify expresso=ion differences across conditions
What does a leftward shift in real-time qPCR mean
means that treatment increases expression of that gene
A rightward shift in real-time qPCR means
treatments decreases expression of that gene
1- cycle difference/2 cycle difference is how much fold difference in fluorescence
2-fold/4-fold (exponential)
RNA - Seq ) bulk measure of RNA
High throughput measures of changes in many RNAs at the same time
Process of RNA -Seq
1. Isolate total RNA from cells or tissue (controls vs experimental conditions); no cellular resolution
2. Use an equal amount of RNA from each tissue to generate short cDNA fragments (50-100 bp), which are then sequenced ("reads")
3. Reads are aligned to the genome by a computer program, which identifies the gene that corresponds to each read across samples 4. Data can be analyzed as raw read counts or normalized to total reads obtained in each condition; the number of reads directly correlates with gene expression level
Immunohistochemistry (IHC) method of...
to analyze protein expression at the cellular level- can only analyze 1 protein at a time
IHC method
-Fix the brain with formaldehyde
-Cut tissue slices and incubate with antibody in solution
-Antibody attaches to antigen wherever it is present
-Visualize where the antibody is using a tagged secondary antibody
-Technique is quantifiable if you use fluorescence microscopy
In Situ Hybridization
method to analyze RNA expression at cellular level- can only study 1 RNA at a time
ISH method
- for mRNA
- Use an RNA or DNA probe that is complementary to the RNA you want to detect in the tissue
- The probe is synthesized using chemically modified nucleotides that can be detected (fluorescence or colorimetric)
- Apply it to the tissue and wash away excess
- Visualize under a fluorescence microscope or
visible-light microscope
Single cell (SC) RNAseq
new method that allows high- throughput analysis of changes in gene expression in individual cell types
benefits of SC RNA seq
BULK AND HIGH THROUGHPUT
Can identify changes in many mRNAs at the same time
Allows you to measure changes in gene expression between two different conditions, such as controls vs drug-treated animals. This can be done for each different cell type.
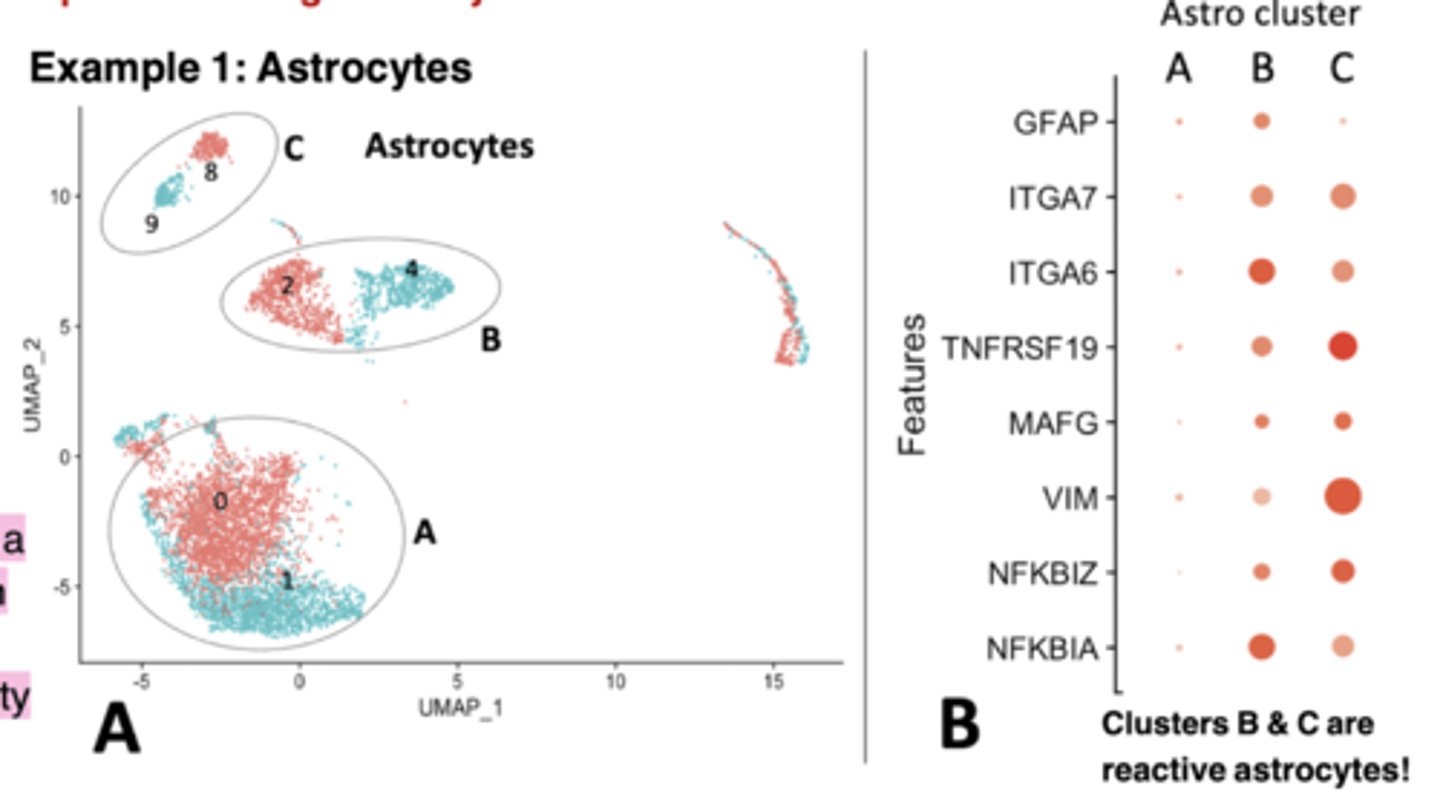
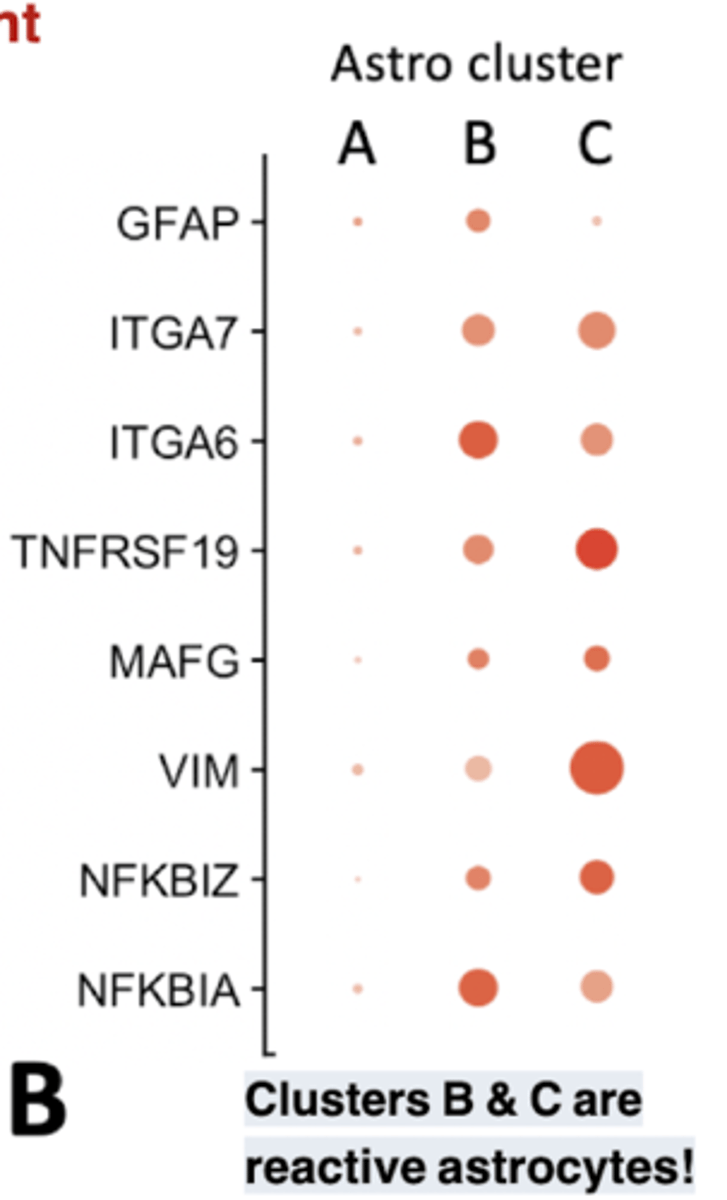
Astro cluster plot
each dot represents a single cell, and cells are clustered based on similarity of gene expression (principle component analysis (or PCA) dimensionality Clusters B & C are reduction)
reactive astrocytes! Bigger dots means increased expression.

Optogenetics: Use of light-gated channels from algae and other marine species
can be done in cell culture, or using transgenic or knock-in approaches in whole animals
a technique that controls neural firing via light- light-activated ion channels that are inserted into the cell membrane if expressed in neurons of interest
Advantages of optogenetics via Use of light-gated channels from algae and other marine species
Selectivity: ionic and light wavelength
Temporal Precision- Msec Resolution
Abilities of optogenetics via Use of light-gated channels from algae and other marine species
Excite (channelrhodopsins) or inhibit (halorhodopsins) specific neurons in living animals
ChR2 comes from algae; NpHR comes from salt-loving halobacteria
Different rhodopsins used for different times/patterns of stimulation
Can use methods to express rhodopsins in specific cells at specific times (cell-specific promoters in transgenic mice, virus injection, etc)
Use of optogenetics to show that the locus coeruleus (main source of norepi) controls the sleep-wake cycle (arousal)
Wait till animals are in NREM sleep, then hit them with blue light, and the animal wakes up, falling out of REM sleep.
The same light stimulation has no effect in animals without ChR2 expression
Viral vectors used in neuroscience research
Tool to express foreign proteins in the brain without using transgenic mice
where a protein of interest is expressed by injecting the virus directly into the brain
Advantages of Viral vectors used in neuroscience research
the gene is only in the cells near the injection site.- investigator control. Can be used for gene therapy (ex. intrathecal injection)
Once inside, the virus-driven expression of a protein is stable because it is engineered to be.
Abilities of Viral vectors used in neuroscience research
can incorporate conditional or inducible
gene expression/inactivation in the virus (expressing Cre or CreER from the virus, and injecting the virus into a brain region in a “floxed” or "edited" mouse with loxP elements inserted in a gene of interest – would target
the KO just to the injected brain region)
What are electrical synapses also known as?
Gap junctions
What proteins form gap junctions in electrical synapses?
Connexins
what directional are gap junctions and what does this mean
They are bidirectional
This makes cells electronically coupled
This allows for fast communication
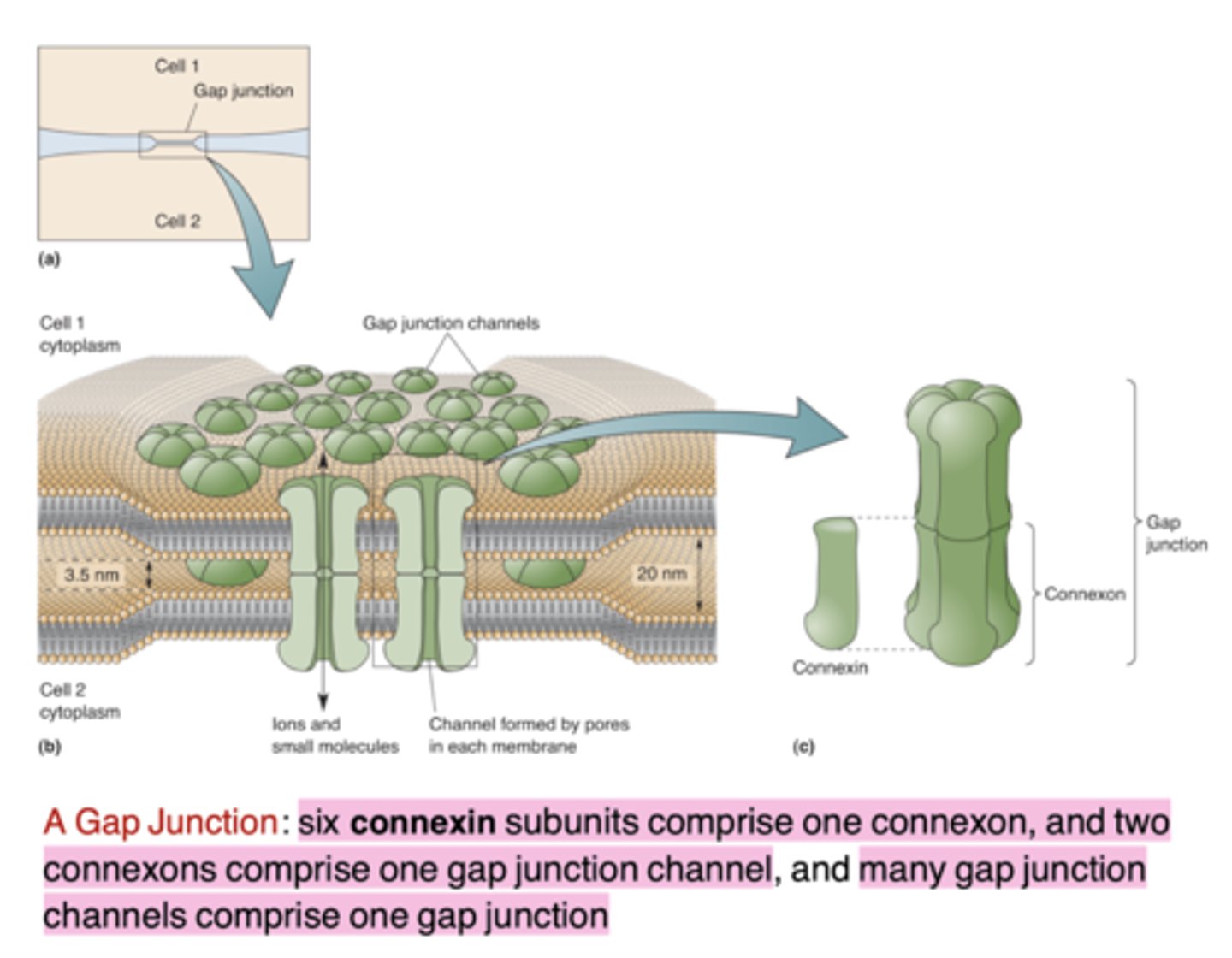
Gap junctions are usually...
What is the primary function of electrical synapses?
Electrical synchronization of cells
Why are electrical synapses important in the brain?
They are important for the function of selected brain circuits.
Inferior olivary neurons use electrical synapses in what way
they utilize gap junctions and have synchronized Vm oscillations and action potentials- without the junctions the synchrony goes
what did Otto Loewi
experiment on a frog and discovered that ACh slows down the heart
won a Nobel prize with Dale.
What type of synapse are most synapses
chemical
What are presynaptic axons
They can contact different parts of postsynaptic cells, such as axosomatic, axospinous, etc.
Astrocytes play what role in CNS chemical synapses
they are often present and participate in regulation of signaling
What are active zones
The presynaptic site has them
Where synaptic vesicles filled with neurotransmitters (NTs) are found.
How many active zones can be in a synapse
one or many at a single synapse
where are vesicles
they are "docked" at active zones
What are large secretory granules (aka dense-core vesicles)
they are some distance away from active zones and carry peptide neurotransmitters
what are peptide neurotransmitters
neuromodulators that are trafficked long distances from the soma into the axon
postsynaptic specializations =
postsynaptic density (PSD)
where the neurotransmitter (NT) receptors are located
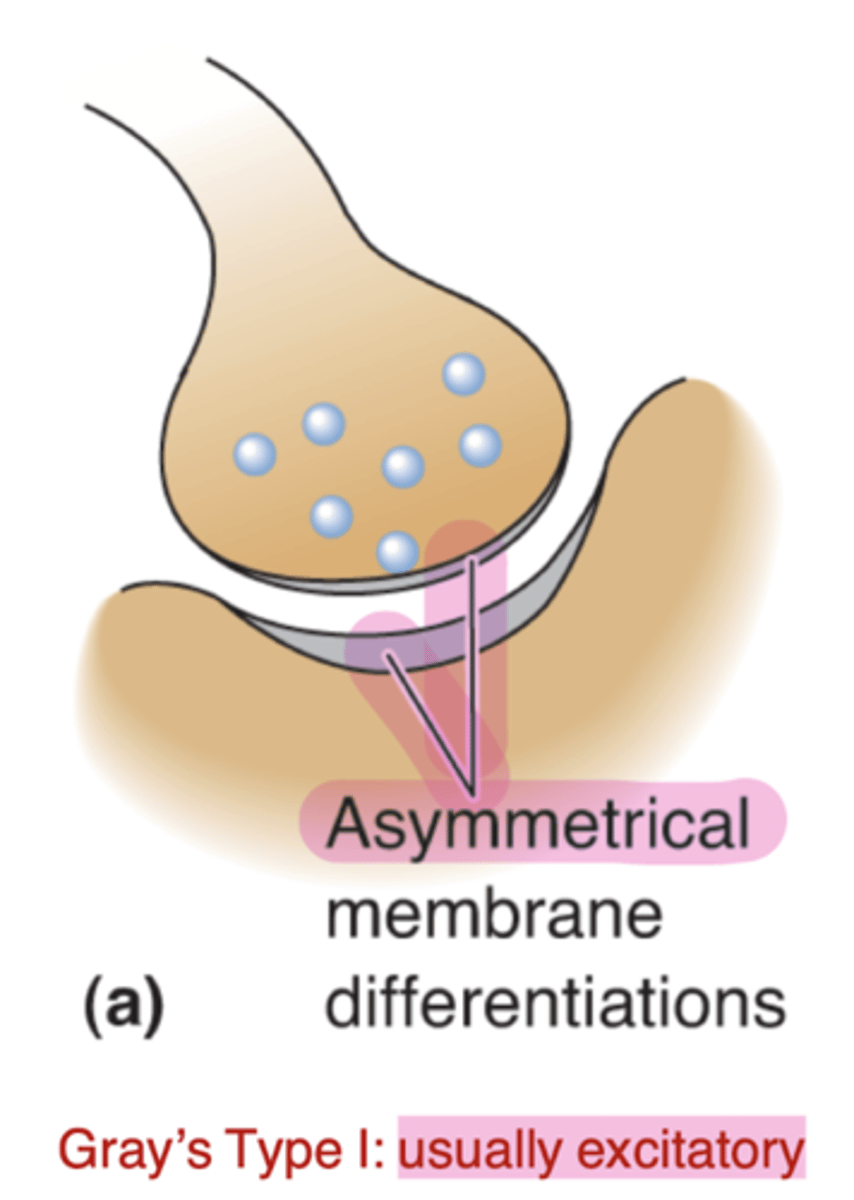
Gray's type I
Asymmetrical membrane thickness at synapse, usually excitatory
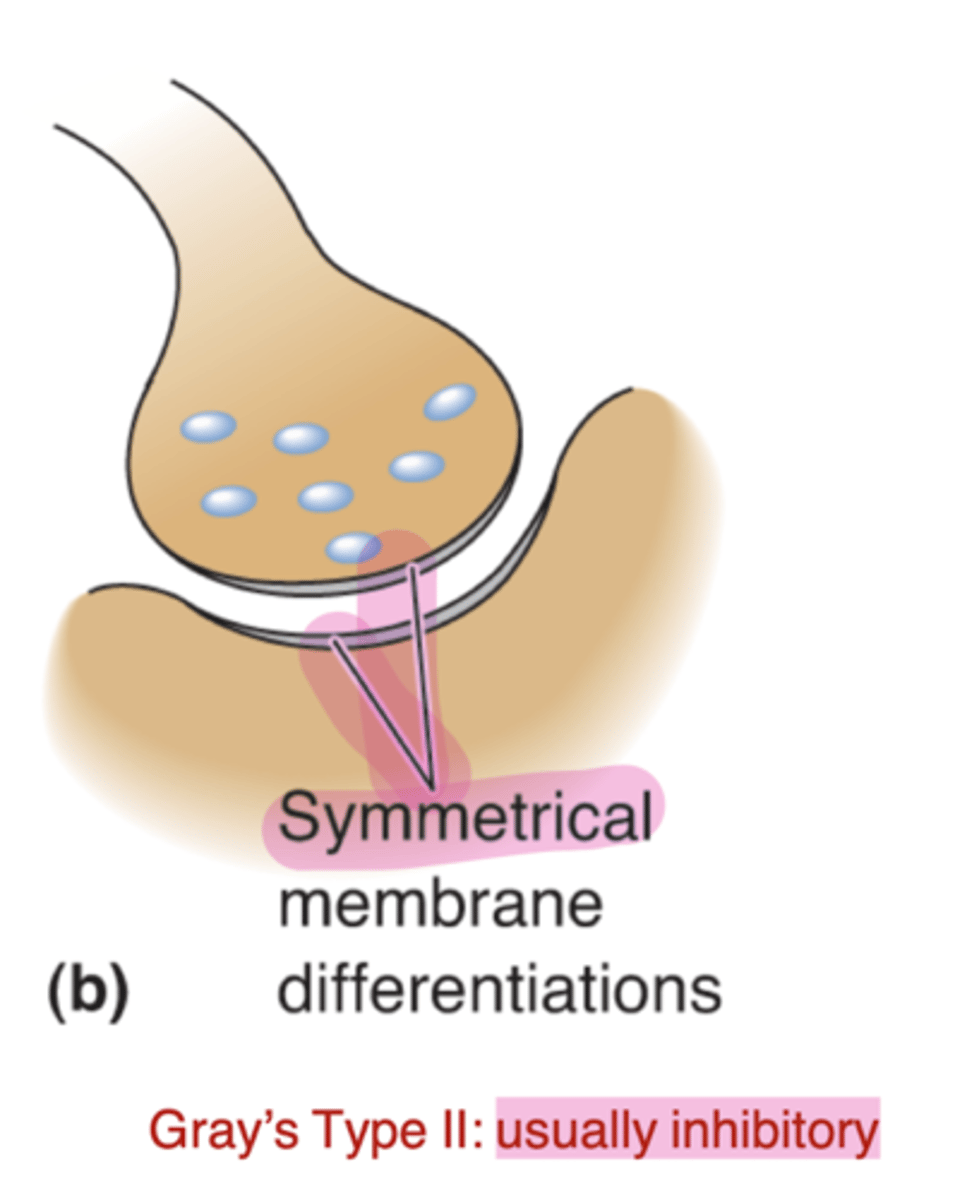
Gray's type II
Symmetrical membrane thickness- usually inhibitory
where are Neuro Muscular Junctions (NMJ)
between motor neurons and muscle
Why are NMJs easier to study than CNS synapses
they are fast, large, and reliable due to the size of the motor endplate and they have a large number of active zones