Topic 11- Translation
1/20
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
21 Terms
What is the Ribosome
what are they made of
where are they found?
Molecular machine made of 2/3 ribosomal RNA (rRNA) & 1/3 protein that builds proteins by translating genetic information from mRNA.
Large subunit
Small subunit
rNA- responsible for structure & function
proteins- help the rRNAs change shape as they catalyze chemical reactions
Ribosomes are found:
bound to ER (make proteins for secretion, membranes, or organelles)
floating in cytoplasm (make proteins used inside cell)
What are the 3 parts of the Ribosome?
what is Translocation
A site- where the new tRNA enters
P site- holds the growing protein chain
E site- where the empty tRNA leaves
mRNA shifts forward by a codon, empties first site and kicks the third out.
tRNA
function
composition
Brings specific amino acids to the ribosome during translation. (molecular bridge)
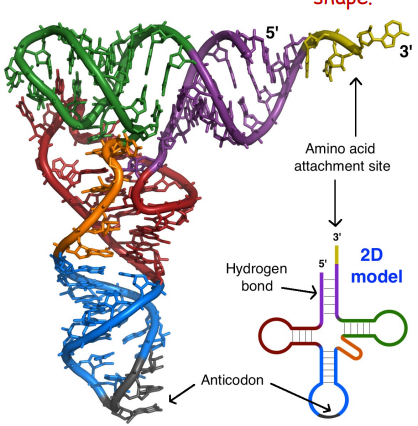
made of a single strand of RNA
Has Anticodon (complementary to mRNA codon)
complex 3D L , cloverleaf shape (due to base pairing)
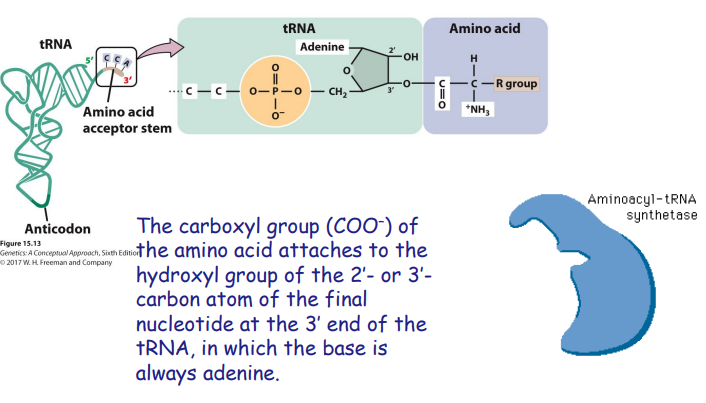
How does the right amino acid get linked to the right tRNA?
There's a different aminoacyl-tRNA synthetase for each of the 20 amino acids.
The tRNA and amino acid both attach to enzyme, enzyme links them together.
What is Wobble?
explain significance
Flexible base pairing between tRNA anticodon and mRNA codon that allows one tRNA to recognize multiple codons.
(flexibility at 3rd base of codon)
allows DEGENERACY of the genetic code
multiple codons → same aa → fewer tRNAs needed
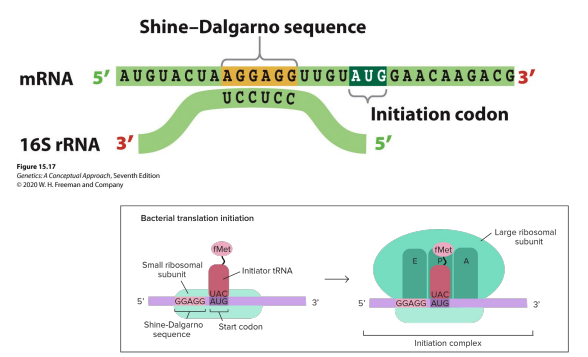
How does the ribosome know where to start in bacteria?(initiation)
what is the first amino acid used in bacteria?
The small ribosomal subunit of the ribosome base pairs with the Shine-Dalgarno Consensus Sequence (comes just before start codon) of the rRNA
Formylmethionine (fMet)
P site!
How does the Ribosome know where to start in Eukaryotes? (Initiation)
which site does Met bind to?
1) tRNA carrying methionine attaches to the small ribosomal subunit
2) with initiation factors, they bind to the 5’ end (5’ UTR) of mRNA (recognizing 5’ cap)
3) They “walk” along the mRNA → 3’ until they reach the first AUG in Kozak sequence
4) Large ribosomal subunit joins
P site!
What are Initiation Factors?
Specialized proteins that assist translation initiation
Help ribosome, mRNA, and tRNA find eachother
powered by GTP
What is circles-message amplification
If Poly-A binding protein coats the poly A tail, it interacts with cap binding proteins, circularizing mRNA
= translation of SAME mRNA
What are Stop codons
What are release factors
no tRNA
Recognized by release factor proteins that bind to A site and deconstructs subunit.
UAA
UAG
UGA
indicates termination
What direction are polypeptides formed?
N → C
NH3+ → O=C-O-
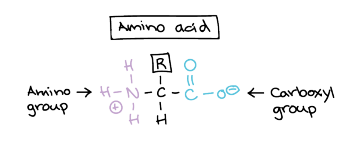
Amino Acid Structure
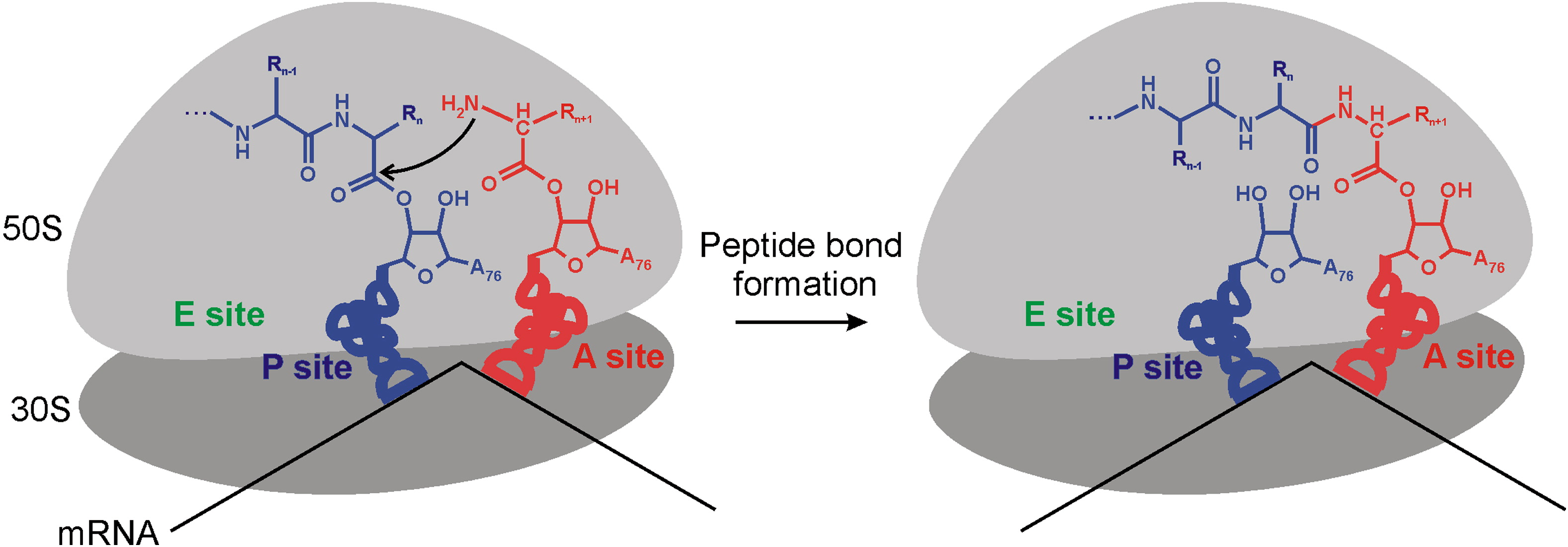
In elongation, the creation of peptide bonds between amino acids is catalyzed by…
rRNA in the Large subunit
peptide bonds = covalent bonds between amino acids
Constitutive expression
Genes continuously expressed under normal cellular conditions.
Not regulated, expressed continually
What is an operon?
Cluster of genes in prokaryotes controlled by a single promoter and transcribed into one mRNA
Components:
Promoter (RNA polymerase binding site)
Operator (regulatory switch)
Genes (encode proteins)
Function: Allows genes to be turned ON/OFF together
Allosteric Protein
give example!
Changes shape in response to environment
eg. Repressor protein
Negative Inducible Operon
(give example)
Catabolic pathways (breaks down molecule)
Repressor binds to DNA
Presence of molecule inactivates repressor → comes off DNA
eg Lac Operon
Negative Repressible Operon
(example?)
Anabolic Pathways (makes molecules)
Repressor does NOT bind to DNA
Presence of molecule activates repressor → binds to DNA
eg. tryptophan in e-coli
Define Catabolism and Catabolite Repression
Catabolism: breaking down lactose → glucose
Catabolite Repression: favoring the use of glucose over metabolims of other energy sources
(represses metabolism of other sources in presence of glucose)
cAMP
“Hunger Signal”
Turned on when low Glucose → turns on ALL metabolism
Global regulator that coordinates 100s of genes (pos control mechanism)
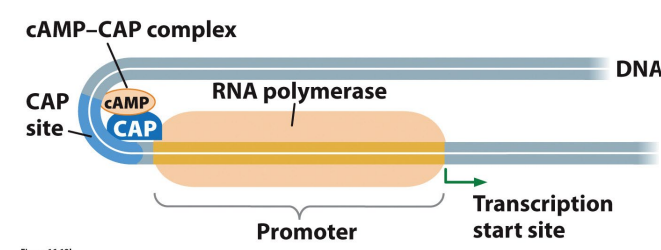
What is CAP
The positive effect of cAMP is activated by Catabolite Activator Protein (CAP).
cAMP binds to CAP'
together CAP–cAMP complex binds to a site slightly upstream from the lac gene promoter.
improves binding of polymerase and increases transcription rate 20-fold.
The binding of the cAMP–CAP complex to DNA produces a sharp bend in DNA that activates transcription.