Imin midterm 2
1/41
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
42 Terms
Bacteria defenitions
- Human microbiome: all the microbes that live on & in us and keep us healthy
- Pathogen: microbes that cause diseae
- Infection: microbe that is potentiallu pathogenic replicating in or on host
- Disease: microbial infection that damages the host
- Pathogenicity: ability of an organism to cause disease
- Virulence: the extent to which a microbe can cause disease
how fast it replicated, how badly it damages the host
- Virulence factors: the adaptations & traits that promote virulence in a pathogen
- Opprtunistic pathogen: infections that result from oppurtunistic conditions
infect during compormised immune systems. Disruption in balance of normal microbes
- True pathogen; cause disease in healthy host
Obligate: a pathogen that must cause disease to be transmitted
Faculatative: can cause disease, but doesnt need a host to complete life cycles
Establishment of infection
- The presence & even growth of microorganisms on the host doesnt always lead to disease
- The interactions between host & microbe both contribute to the final outcome
- This relationship is influenced by virulence of microbe & defense mechanisms of the host
- There are many steps to establishment of infection that start with: colonization, persistance, & spread. Can be subdivided into
Exposure (by resperation, bodily fluids, food borne illness, sex)
Adherence
Invade host tissue
Evade immune system
Host damage (a hallmark of disease)
- Each step requires unique evasion strategies for the pathogen
- Louis Pasteur: developed ‘Germ theory of Infectious Disease’
Disproved the notion of spontaneous generation of microbes & miasma (cloud of disease) theory for spread of disease
Also explained why so many people died after surgeries where antiseptic technique was not practiced
- Robert Koch: studied the theory that microorganisms cause human disease. Showed cause & effect between pathogen and disease.
Developed Koch’s postulates. Pathogen:
Must be present in all cases of the disease
Must be grown in pure culture
Must cause disease from pure culture
Must reisolate from diseased host
Doesn’t work all the time
Staphylococcus epidermis
- Gram (+). Example of oppurtunistic pathogen.
- Normal component of the skin microbiome & important in protecting skin from unwanted pathogens
Encounter/exposure: ubiquitous component of skin microbiome
Host entry: most common pathogenic route of entry is nosocomial (hospital-borne)
It adheres to plastic catheters & implants
Adhesion & Invasion
tolerates changong osmolarity
adheres to plastics via teichoic acids
binds ECM
Adheres vis association w/ biofilms
Immune system evasion
biofilms provide resistance to AMPs & antibodies & mechanical removal
Extensive efflux pumps
Host damage: produces PSM (phenol-soluble modulins → highly hydrophobic)
Treponema pallidum
- believed to be the causative agent of STI syphillus
- Spirochete; has a endoflagellum where two or more flagella originate from opposite poles of the bacterium & are located in the periplasmic space
- We think an obligate human pathogen. Until recently, has been unculturable
- Endoflagella wrap around bacterial structure & prevent recognition by innate immune system
- Doesn’t have animal model. Antibiotics can treat it. The disease - 43% asymptomatic.
- Case study: clinical case of a women w/ a year long mouth ulcer
was about to go in for surgery to remove a presumptive tumour'
Did ELISA & pathological analysis - T.pallidum detected
Penicillin G managed the infection & resolves in 2 months
therfore alternative methods can be used to satisfy Koch’s postualtes
Basic bacterial structure
Peptidoglycan
- Repeating disaccharide of NAG & NAM
NAG (N-acetyl glucosamine): monosaccharide derivative of glucose
NAM (N-acetyl muramic acid): amide between glucosamine & acetic acid
- Structural componeny of Gram (-) cell wall
- NAG and NAM are cross-linked with oligopeptides at the lactic acid residue of NAM
- Gram (+) and (-) differ in their cross-linking peptide
- Dregree of cross-linking can reinforce cell strength
Gram (+): contain a pentaglycin crosslink along w/a tetrapeptide
Gram (-): contain directly linked tetrapeptide. D-Glutamate & D-Alanine are in 2nd & 4th position (D isomers of amino acids are used. VERY rare. used in bacteria to avoid protease digestion).
LPS (lipopolysaccharide)
- Major component of Gram (-) outer memb
- Highly antigenic → elicits a strong immune response
- Also inc the negative charge of the cell memb & helps stabilize the overall memb structure
requires association w/ divalent cations (Mg2+) to neutralize charge repulsion
- 3 molecular domains:
1) Lipid A: Inside outer memb. Phosphorylated glucosamine disaccharide w/ fatty acids (usually 6 acyl chains)
Amphipathic. Highly toxic and antigenic.
The hydrophobic fatty acid chains anchor LPS into the bacterial memb, the rest of the LPS projects from the cell memb.
lipid A domain is responsible for much of the toxicity of Gram (-) bacteria
Upon bacterial cell lysis by immune system, fragments containing lipid A are released into circulation causing fever, diarrhea, sometimes fatal endotoxic shock (Inc immune response ro release Lipid A → causes illness)
2) The core: attachs directly to Lipid A
Some diversity. Kdo (sugar) is most common.
Mostly sugar but can contain amino acids
3) O antigen: attached to core oligosaccharide
Highly diverse
Often determine strain type
switching/modifying O antigen is common in evasion strategies
- The LPS can come in Rough or Smooth or somewhere in between
degree of covalent additions is strain specific
O antigen complexes w/water to form a gel
Recall Griffith’s experiment (mice)
Smooth strains are generally associated w/ virulence
PRR (pattern recognition receptors)/TLRs/NODs
- Evolutionarily conserved germ-line encoded receptors expressed by cells of the innate immune system
- Structurally different – common function
- Detect PAMPs (pathogen associated molecular patterns)
include peptidoglycan (PG), Lipid A, teichoic acid, capsules (all unique to bacteria)
- Toll-like receptors: receptors of immune system that recognize PAMPs
expressed both on surface of innate immune cells & in endosomes
- When TLR is bound by ligand, it activates a signaling cascade that initiates the release of pro-inflammatory cytokines
therefore PAMPs recognized by PRRs (like TLRs) will elicit an immune response
- Different kinds of tLRs recognize different sets of microbial molecules (PAMPs)
TLR6: recognize PG on gram (+), Lipoteichoic acid on Gram (+), lipoproteins on Gram (-)
TLR4: recognizes LPS of Gram (-)
TLR3: found in endosomes (transitory compartment between golgi → memb → lysosome). Recognize phagocytozed pathogens
- NODs: (nucleotide-binding domain, Leucine-rich repeat - containing receptors)
- Sensors of intracellular PAMPS
- There is often overlap between TLR cascades and NOD cascades
- Different peptidoglycan components are recognized by different NODs
Helicobacter pylori
- O-antigen of H. pylori in some strains contain carbohydrates that resemble human Lewis antigens x and y
- Mimicrky protects from host immune response
- And – evasion of the NODs
- Invades stomach lining, but stomach has very good innate immunty from acid
- secretes a base to neutralize acid which liquifies mucous
- Motile helical bacteria that swims through liquified mucoud & invades stomach epithelium
- Modifies Lipid A to aid in evasion
TLR agonist → less virulent. Will activate TLR response
Dephosphorylation: reduces (-) charge to decrease sensitivity to AMPs
Remove (-) charge on phosphate by adding an amine
Removal of 2 acyl chains: hexacylated LPS (6 chains) is highly antigenic therfore removing 2 decreases degree of immune repsonse
TLR evasive → virulent. Evades immune system therfore more likley to cause disease
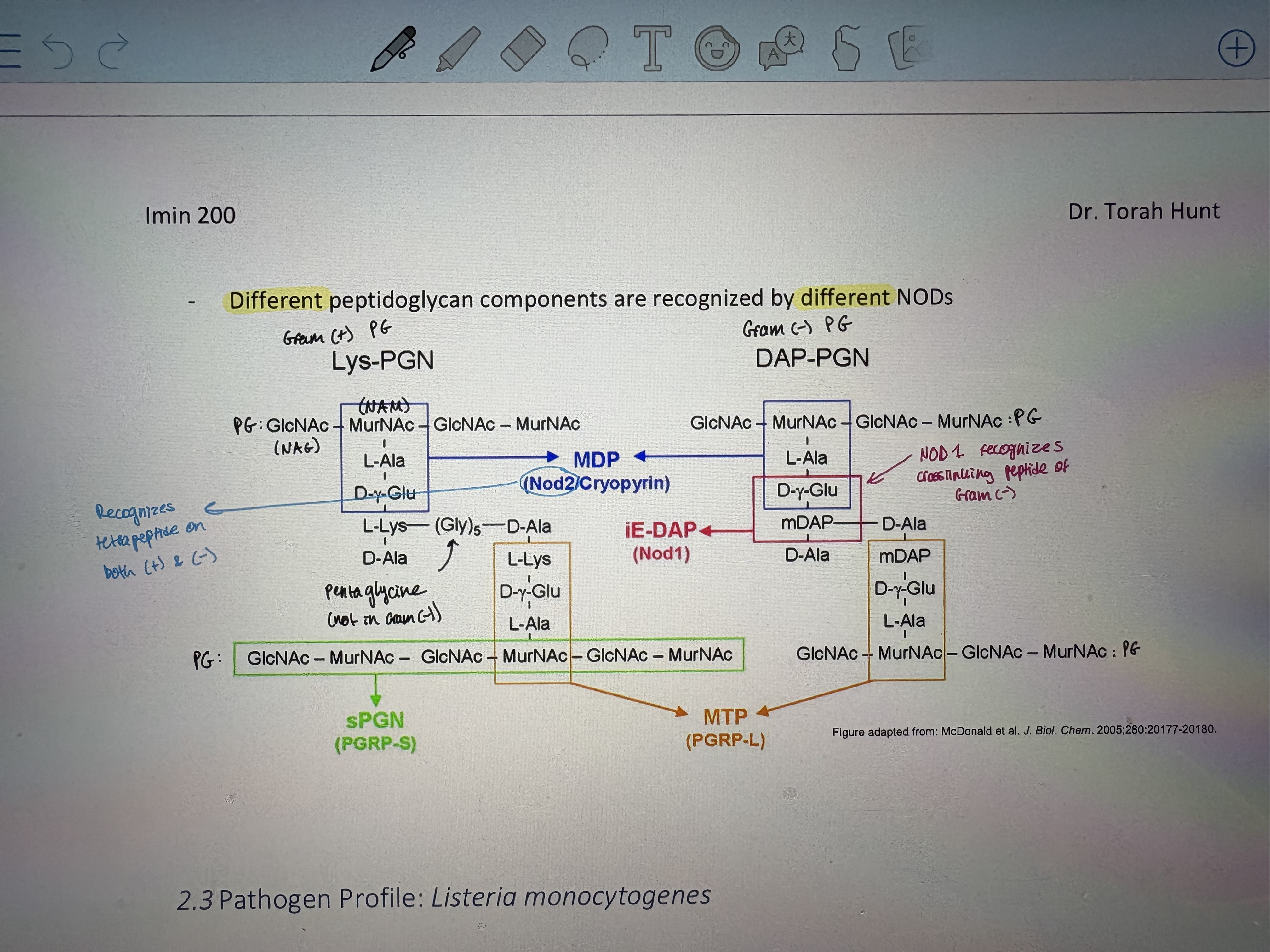
Listeria monocytogenes
- Gram (+). Causes listeriosis: a serious infection usually caused by eating contaminated food
- An estimated 1,600 people get listeriosis each year, and about 260 die. The infection is most likely to sicken pregnant women and their newborns, adults aged 65 or older, and people with weakened immune systems.
- Is an intracellular pathogen.
can grow at 0-37 degrees (thermotolerant)
Present in soft cheeses and unpasteurized milk
Can survive w/in phagocytes
Attaches to host cell via D-galactose surface or cell surface
- The structure of Listeria monocytogenes (Lm) peptidoglycan (PG) has been recently characterized
- Has unusal immune evasion strategy against NLRs (NOD-like receptors)
50% of NAG is replaced w/ glucosamine = deacetylation
Allowed evasion of detection by NODs
previous to this → L.m. had a PG adaptation to allow evasion of NOD2, but this chnage made it sensitive to NOD1
secondary evasion of NOD1 involved deacytlation
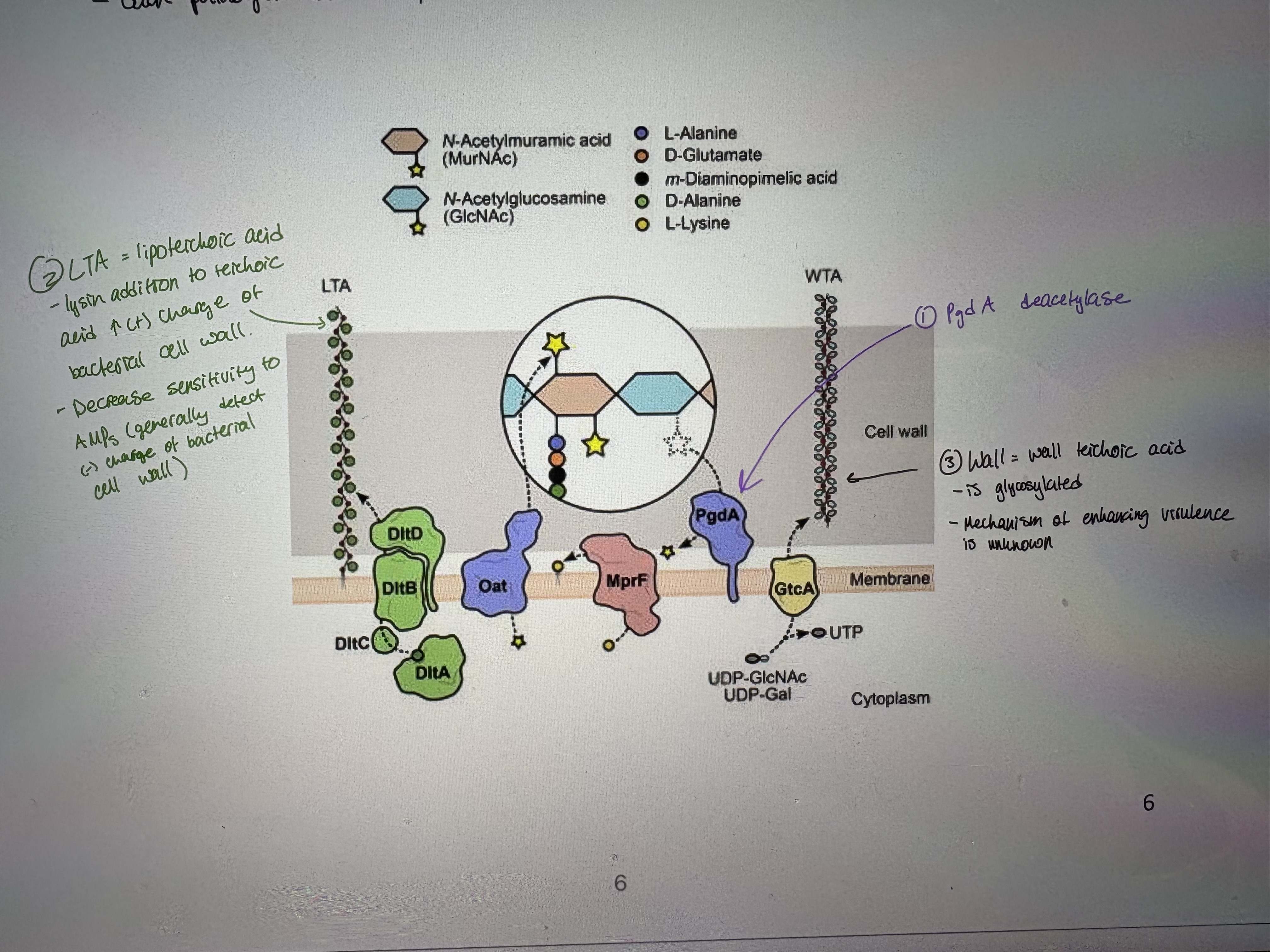
- To evade recognition and targeting by the host immune system, L.m. expresses surface proteins specialized in the introduction of fine modifications in cell envelope components (see pic)
Each pathogen has multiple evasion strategies
Lipid A modifications
- Some Gram-negative bacteria have evolved mechanisms to modify the structure of Kdo2-lipid A
most common sugar in core oligosaccharide
- Structure of Kdo2-lipid A from various types of bacteria
There are dozens of enzymes that aid in this modification
LPS = main outer memb component
E.coli K-12; non-pathogenic. Has 6 acyl chains
- Helicobacter pylori, Yersinia pestis and Vibrio cholerae have evolved different mechanisms of lipid A modification to aid colonization of their respective niches inside the host
Yersinia pestis
- Causative agent of the plauge. When residing in the flea vector, Y. pestis produces an endotoxic hexa-acylated lipid A that is a strong immunostimulant in humans.
- Discovered by Alexandre Yersin in 1894 (Was trained in the Pasteur school of thought)
- Has been used as a bioterror weapon
Unit 731 of the Japanese Imperial Army in WWII sprayed fleas infected with Y. pestis on the unsuspecting Chinese population. # of victims is unknown but presumably was high
- Y.pestis lives on a dlea resovoir and introduced to humans after flea bites
- Modifies Lipid A to aid in evasion
Temp is a virulence factor in Y.pestis (lives on fleas 21-27 degrees). Upon introduction to humans (21-37 degrees) this activates temp sensitive gene expression of virulence genes
Temp sensitive removal of 2 acyl chains. Less sensitive to TLR4 detection (less antigenic)
- Non-pili adhesion molecules:
1. Yad A: expressed in a temp-sensitive manner
outer memb protein that bind to collagen in host cell ECM
2. Invasin: binds to integrin proteins on host
Not found in Y.pestis (exists as a pseudogene)
Is present in Y.pseudotuberculosis & Y.entercolitica ; only expressed at 21-26. Facilitates phagocytosis in early stages of infection
Invasion binds beta1 integrin family members on mammalian cells
Vibrio cholera
- Water borne illness that exists in copepod reservoir (water borne invertabrate)
- Modifies Lipid A to aid in evasion
In non-virulent strain it has relatively normal LPS
In virulent strain → addition of glycine & a diglycine = decreases V.cholera sensitivity to AMPs & inc virulence
- Cholera toxin: 5B type toxin. 5xB subunits associate to form a barrel. Often heavily glycosylated (allows immune evasion, changes host range)
Both toxin genes are encoded in a prophage (intergrated viral genome). Only strains w/ prophae are pathogenic.
- Cholera toxin is secreted by the Type II secretion system
also secretes neuramidase enzyme. Removes salic acid off host cell receptor to facilitate toxin binding
- Toxin entry: CT binds GM1 (ganglioside receptor) on host intestinal epethelium. Toxin binding triggers RME (recptor-mediated endocytosis)
Bacterial toxins subvert transport pathways and ER functions for delivery into the host cytosol
For cholera toxin (CT), the KDEL (Lys-Asp-Glu-Leu) sequence (known to be an ER retrieval motif) is located at the carboxyl terminus of the CT A subunit and it binds to Erd2 in the Golgi
KDEL: 4 amino acids that are a signal that says “take me to the ER”
A subunit contains KDEL
At ER, A subunit escapes into cytoplasm → process requires host cell proteins. Erd2 in golgi reads KDEL, PDI unfolds A subunit to allow transport, Sec61 allows transport of A subunit into cytoplasm.
A subunit targets G proteins in cytoplasm
- Mode of action of toxin secretion: ADP-ribosylation of G protein, and constitutive activation of adenylate cyclase (AC).
Results in increased levels of cAMP within the host cell.
cAMP stimulates pkA (protein kinase A). Cause activation of massive chloride efflux.
Signal → GPCR → Gs (CT = active pathway in absence of a signal ie constitutive) → AC → ATP (ends) or cAMP → pKA → major chloride channel
Chloride efflux causes water to follow into lumen of intestine = diarrhea
AMPs (antimicrobial peptides)
- Potent members of the innate immune response
- Have been shown to kill Gram-negative and Gram-positive bacteria
- Divided into subgroups based on composition of secondary structures
- In general, recognize & associate w/ negative charges of bacterial cell walls
- Found in macrophages & lysosomes
- eg. Kalata B2: found in plants.
Shown to have; antimicrobial activity, insecticidal, anti-tumour activity
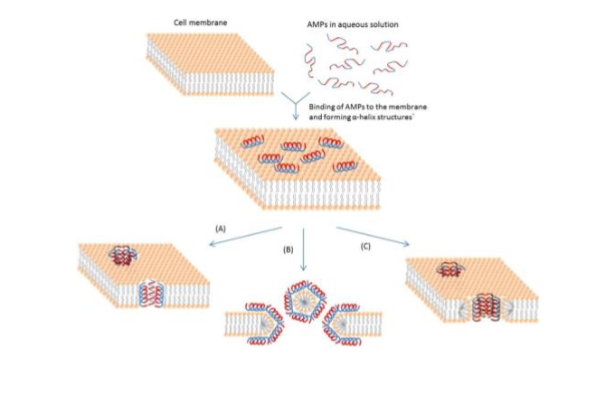
- Schematic representation of some action mechanisms of membrane-active AMPs.
AMPs form pores in bacterial memb which disrupts p+ motive force (PMP)
3 models for AMP activity: A) Barrel-Stave model; AMPs polyerize & insert perpindicularly. B) Carpet model; small areas of memb get coated w/ AMPs. C) Torrodial pore; AMPs insert by first associate w/ phospholipid head
- Example of an AMP; Defensin → has high concentration in breast milk
Short peptides with a positive net charge and can undergo disulfide crosslinking to give peptides a circular shape
Agrees with hypothesis that defensins either depolarize the membrane or form pores in it
Still don’t know how defensins can cross outer membrane of Gram (-)
- Resistance to defensin/AMPs:
A) Modification to D-alanine in teichoic acid: dec negative charge of memb
B) Add acyl chains to LPS: harder for AMPs to penetrate
C) Add positively charged proteins
D) secrete negative charged proteins as a decoy
E) efflux pumps
F) destroy AMPs w/ proteases
Molecular mimcry
- Molecular mimicry is structural, functional or immunological similarities shared between macromolecules found on pathogens and in host tissues
plays an important role in immune responses to infection and in autoimmune diseases
- Infection may induce autoimmune responses which attack and destroy body tissues or organs
- Ex. Pathogen Profile: Campylobacter jejuni
Gram negative. Leading cause of bacterial food-borne diarrheal illness worldwide. No defined toxins that lead to disease
Evasion strategies include:
1. N-linked heptasaccharide glycosylation: makes cell surface proteins resistant to trypsin digestion
2. O-linked glycosylation: glycosylation of bacterial flagellum w/ 2 rare sugars (no longer “looks” like flagelin). Allows both evasion & adherence.
- Can cause Guillain-Barre syndrome; a potentially fatal paralytic autoimmune disorder
Modified LPS called LOS
LOS of C.jejuni mimics a ganglioside in human nervous system therefore immune syste accidentally attacks nervous system even after infection has cleared 9
Adherence
- Adherence to host cells surfaces is an essential first step in bacterial pathogenesis or infection
- For extracellular bacteria:
adherence allows bacteria to resist the mechanical clearing mechanisms of the host eg. urination or coughing
- For intracellular bacteria:
prerequisite for uptake (invasion)
- Adhesins: bacterial components that mediate interaction between the bacterium and the host cell surface
- Surface molecules on a pathogen, adhesins, bind specifically to complementary host cell surface receptors
- Bacteria can specifically adhere to essential cell components
can include interactions w/ cell surface proteins or sugars
Adhesins
- there are different types of pili (fragile hair-like adhesions. Frequently replaced therefore high potential for virulence)
- Pili attachment to surfaces, other cells, or other bacteria (are NOT flagella)
- Exist in both Gram (+) and (-) bacteria
1. P-pili and Type 1 pili
- Rigid, long and thin filaments that protrude off Gram negative bacteria
- Unique feature of Type I pili is they induce hemagglutination (RBC aggregates)
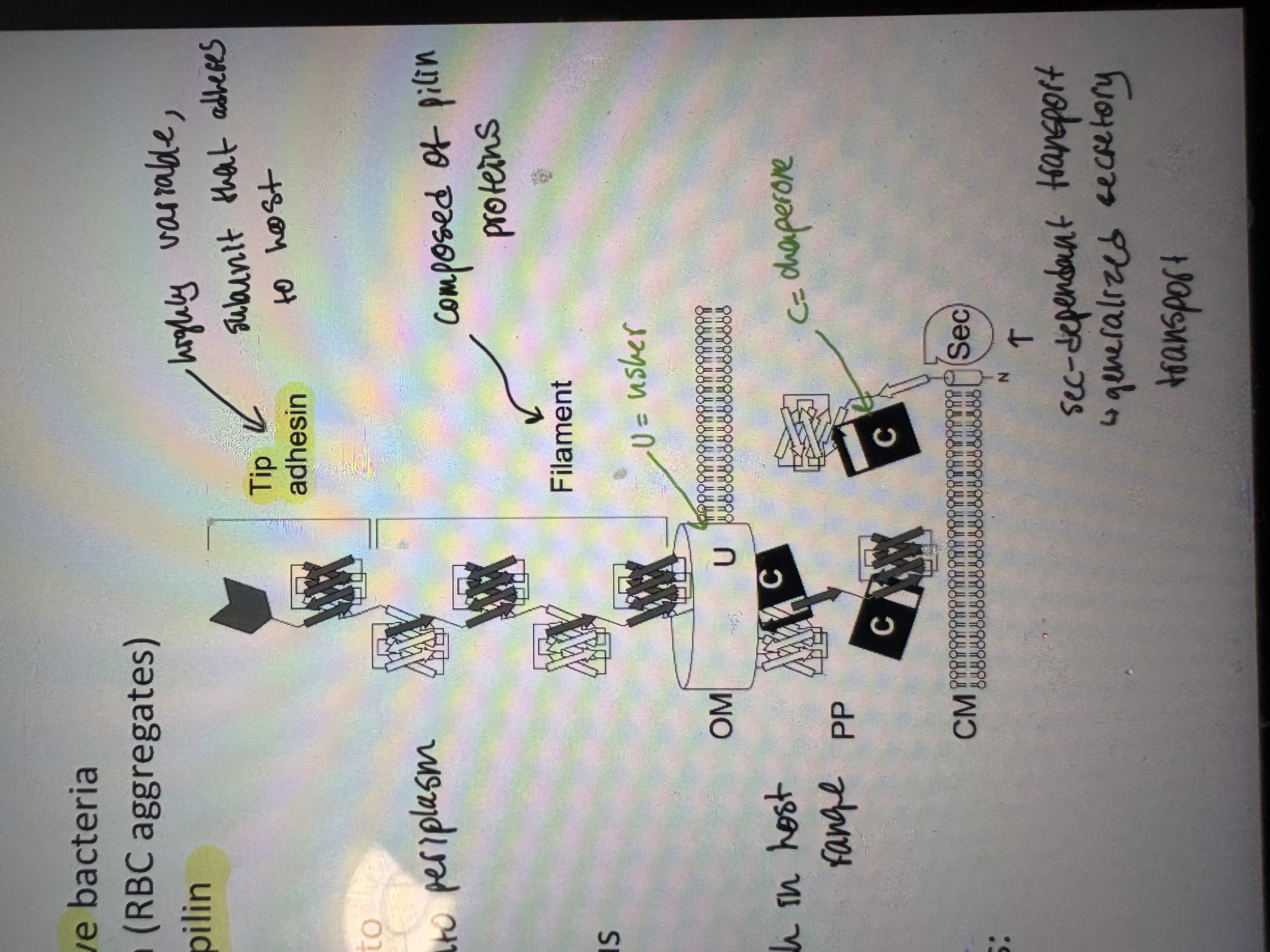
- Type I pili assembly – composed of polymerized subunits of pilin protein
- Tip adhesin attaches to host carbohydrates and contributes to tropism
pilin proteins are secreted by sec-dependant transport into periplasm
Assembled into a pilus via CU system as the transport across OM
- The chaperone/usher (CU) pathway assembles proteinaceous filaments on bacterial surfaces. eg Salmonella → 14 copies of operon therefore high variation which in host range
- These operons encode, at minimum, three different proteins: chaperon, pilin, usher
2. F1 pili
- Shorter, linear and flexible polymers that have a tendency to aggregate
- The F1 capsular antigen is expressed exclusively by Yersinia pestis
F1 pili in Yersinia is antiphagocytic (prevents phagocytes by macrophages)
- F1 capsule consists of a tangle of thin, linear Caf1 fibers
- Lots of variability in F1 pilus
allows immune evasion. Limited vaccine development.
- Pilus is under temp sensitive control
Not expressed at 20 (in flea), expressed at 37 (in humans)
3. Comparing P1, F1, Type 1
- Each pilus is built up with a different combination of subunits (Fim/Pap/Caf)
- Type I and P1 – assembled by the classic chaperone-usher pathway
Made up of flexible end tip and a rigid long rod
Share significant homolgy
- F1 – assembled by the Caf1 CU pathway
More flexible and numerous than type I and P1
Much longer. Caf1 is a unique protein subunit
Escherichia coli
- Gram negative common bacteria that can acquire pathogenicity due to acquisition of plasmids
- Uropathogenic E. coli is the most common cause of UTIs (urinary tract infections)
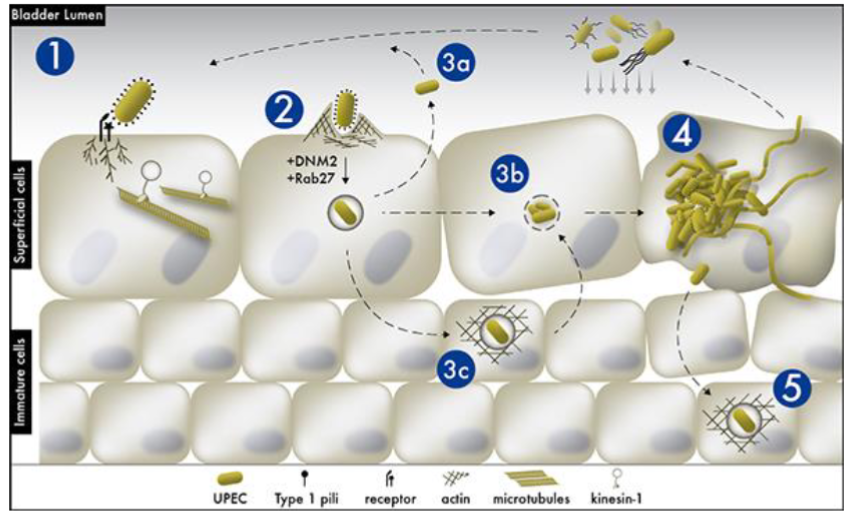
- Uropathogenic E. coli (UPEC) infection cycle:
Type 1 pili bind via tip adhesion to mannose on glycosylated urinary tract epithelium
E.coli adherence stimulates G-protein activation (Rab27) causing host cell actin reorganization
Three fates post-phagocytosis: 3a) replication w/in phagosome then exocytosis → slow, latent infection. 3b) destruction of phagosome allowing cytoplasmic replication → fast infection (secrete alpha hemolysin to lyse cells to liberate nutriants especially iron). 3c) phagosome gets encased in actin. Called a quiescent intracellular resevoir. Can remain in host cells for months. Flare up w/ stress (speeds up replication)
- Just eat cranberries? Chemical in cranberries that binds and blocks bacterial adhesions.
Works in lab, does not work in large scale studies
- Just eat Tums? Dec acidity in urine. Many dec infections.
Type lV pili (aka fimbrae)
- Expressed by almost all Gram negative bacteria and some Gram positive
- Functions:
Twitching motility → Neisseria gonorrhoeae
DNA binding → Neisseria meningitidis
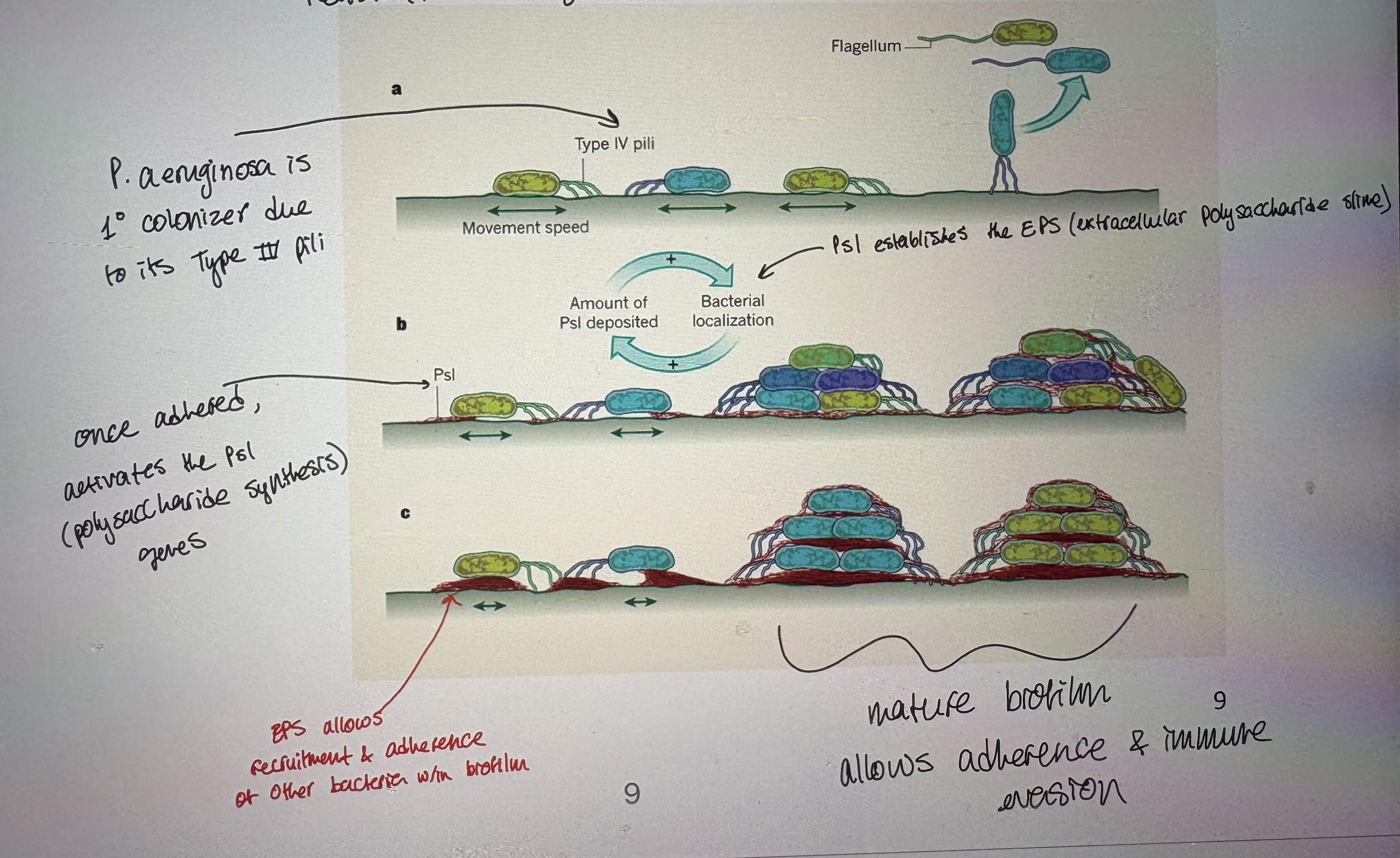
Biofilms:
Type IV pili are important adhesins, promoting initial attachment to a variety of chemically diverse surfaces
Biofilms are dense, multiorganismal layers of bacterial communities attached to surfaces
Attachment to surfaces is mediated by an EPS – extracellular polysaccharide slime
Primary colonizers of biofilms include Clostridia spp and Pseudomonas aeruginosa
EPEC (enteropathogenic E.coli)
- First pathotype of E. coli described
- Relies on Type IV pili to allow bacterial aggregation into microcolonies
- Allows LA (localized adherence)
- Then EPEC uses a Type III secretion system (more later) to induce expression of Tir in the host
Tir 1: translocated intimin receptor (host gene). Pathogen induces host gene expression.
- Once Tir is expressed, then the bacteria uses another adhesin; intimin
Adherence promotes A/E (attaching & effacing) pathology from bacteria
Upon adherence → bacteria cause host cell actin rearrangement; formtion of pedastal
Pathogencity is afforded by pathogencity (clusters of virulence factor genes) of pedastal
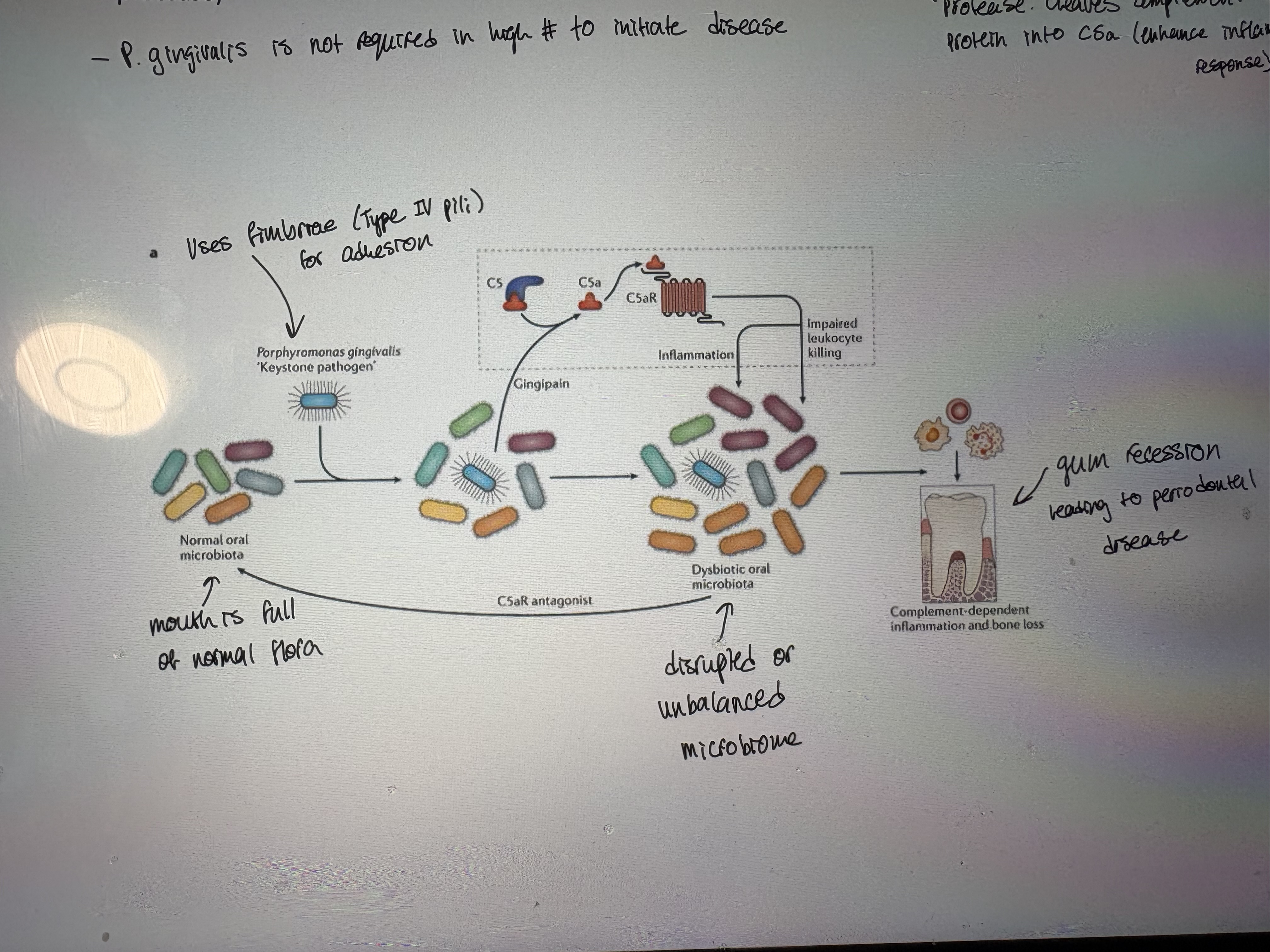
Porphyromonas gingivalis (Gram negative)
- Microbial dysbiosis on the teeth is facilitated by a shift in the microbiome towards Gram negative anaerobes
- Keystone pathogen causing periodontal disease – tissue destruction and bone loss in the teeth and gums
- P. gingivalis alone is insufficient to cause periodontal disease but instead facilitates an ecological change to a disease state
- Acts as a keystone pathogen by production of a virulence factor called gingipain (a cysteine protease → cleaves complement CS protein into CSa. Enhance inflammatory response)
- P.gingivalis is not required in high number to initiate disease
Neisseria gonorrhoeae
- Uses a Type IV pilus for motility along host epithileum
- Main causative agent of the STI gonorrhea
Estimated 86.9 million cases reported worldwide in 2017
Seeing an alarming rise in antibiotic resistance
- A Type IV pilus is absolutely required for colonization
Helps form microcolonies and biofilms and twitching motility
- Type IV Pili can retract while the pilus tip remains firmly adhered to the target surface leading to twitching motility
- Twitching motility is powered by extension of fimbriae, adherence, & retraction (depolarization)
Allows mvmt along urogential tract epithelium
- Motility may help engage Type ll secretion systems to facilitate toxin release
Neisseria meningitidis
- Gram (-) bacterium; aka meningococcus. An exclusively human pathogen
- Can cause meningitis and other forms of meningococcal disease
- One of few extracellular bacterial pathogens that can cross the blood brain barrier
- About 10% of adults are carriers of the bacteria in their nasopharynx
- It is the main cause of bacterial meningitis in children and young adults
- Causes developmental impairment and death in about 10% of cases
- Increased genetic variation and virulence in N. meningitides can be accomplished by natural competence and DNA uptake from Type IV pili
- Type lV pili: only fimbriae that can bind DNA
induces transformation (uptake of free DNA from environment)
binds DNA sequences containing DUS (DNA uptake sequences)
Inc variation has inc virulence
Pseudomonas aeruginosa
- Example of an opportunistic pathogen
- Primarily a nosocomial pathogen
- Rod shaped, Gram (-)ve, aerobic bacteria
- Cause: Respiratory infections in susceptible patients eg cystic fibrosis patients
- Is a successful oppurtunistic pathogen because:
Natural habitat; can grow in a wide temp range (4-37)
Metabolism; can grow on wide variety of substrates
Resistant to removal; associated w/ biofilm thus resistant to mechanical removal
Highly tolerant of chemical conditions. Resistant to many antibiotics.
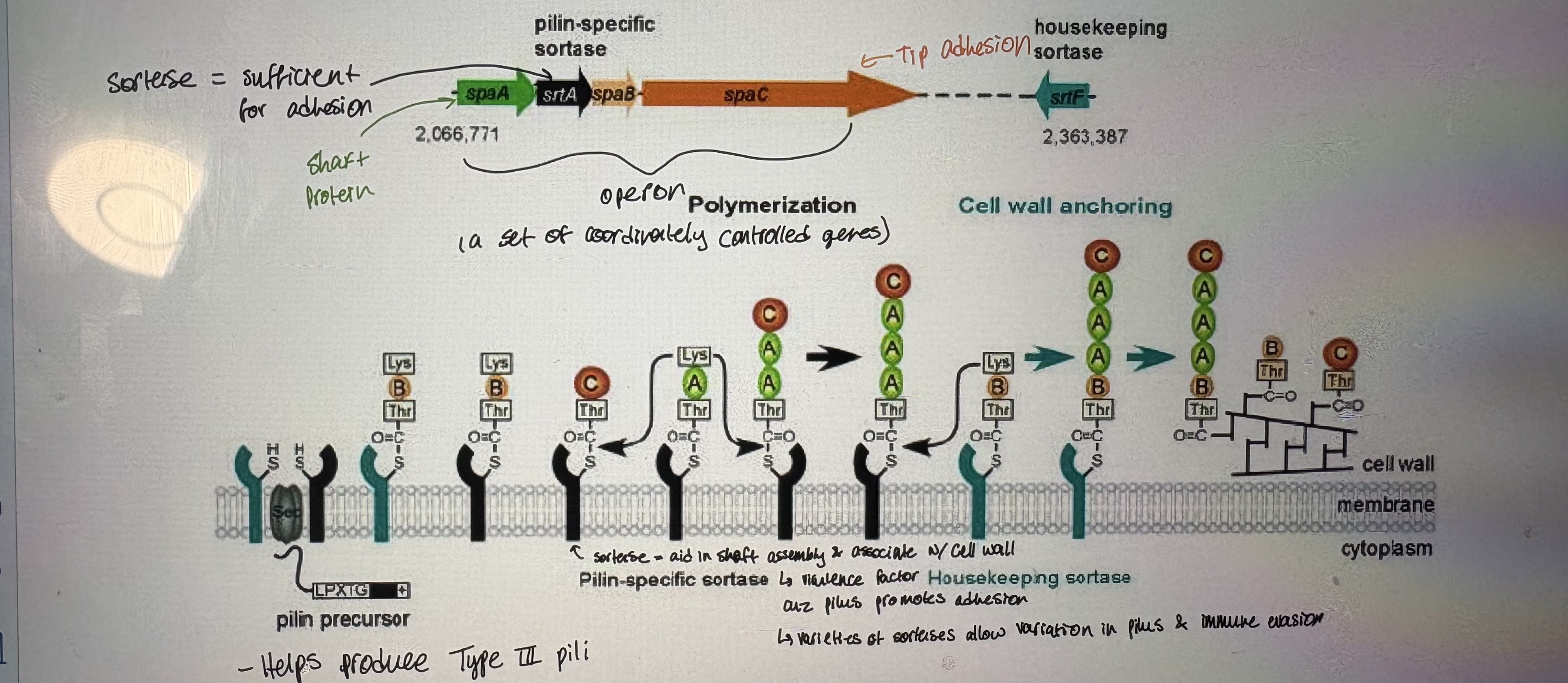
Sortase enzyme
- Virulence factor in Gram (+). Catalyzes transpeptidation step of pilin synthesis
- Important virulence factor of Gram positive pathogens
- Example: Streptococcus pneumoniae
- Unlike Gram negatives, Gram positive pili are formed by covalent attachment of pilin subunits to each other and to the peptidoglycan layer
- Sortase enzymes are a group of prokaryotic enzymes that promote the covalent anchoring of surface proteins to the cell wall envelope to enable each microbe to effectively interact with its environment
- Main functions:
Cleave surface proteins to allow rearrangements & modifications
Polymerize pilin synthesis
Sort proteins at cell surface and attach to cell wall
- Sortases found in Gram (+) bacteria.
Exceptions: a few Gram (-), a few archaea.
Corynebacterium diphtheriae
- Pathogen causes diptheria
- Symptoms: sore throat & fever, barking cough, grey patches on throat
- Genome: 6 Sortase genes (only one nessecary)
- Helps produce Type lll pili
- Pili & sortase genes are encoded in an operon
Non-pili adhesins (afimbrial)
- Many pathogenic bacteria rely on non-pili adhesins for host-cell entry
- Probably facilitate tighter binding of bacteria to host cells after initial binding by pili
- Common in foodborne Gram positive pathogens
- Some recognize cadherins and other components of the extracellular matrix of the host instead of carbohydrates
Listeria monocytogenes
- Once adhered, many pathogens use the cell’s own cytoskeleton to invade the target cell
- Listeria main adhesins are Internalin A (InlA) and Internalin B (InlB), and the receptors are E-cadherin and Met (a RTK growth factor receptor) (host targets), respectively
- Entry of L. monocytogenes into mammalian cells is a dynamic process
Both InlA & InlB are nessecary & sufficient for hos entry
Binding of pathogen triggers certain-mediated actin cytoskeleton rearrangment = facilitate pathogen entry
- InlA also induces cytoskeletal rearrangements in the host cell for uptake of L.monocytogenes by epithelial cells
- Triggers uptake of the bacterium by the host cells into vesicles that are not immediately targeted to the lysosome
Bacteria induces it’s own uptake and then quickly escapes the phagosome
Allows replication w/in cytoplasm. Manipulates host cytoskeleton (actin) to create polymerizing wave of actin to move bacterium across cell
Iron as a nutrient/in bacteria
- In the human host, iron is limiting because lactoferrin, transferrin, ferritin and hemoglobin bind most of the available iron
- Host cells sequester & limit iron availability because its an essential nutrient for pathogens eg. lactoferrin; chelates iron at mucosal surfaces, ferritin; stores iron w/in cell.
- Consequently, the cellular concentration of the ferric ion is too low for microorganisms to survive by solely using free iron for survival
- Provides nutritional immunity: if host “hides” nutrients → becomes one of our first lines of defense against infection
- The sequestration of nutrient iron is the most significant form of nutritional immunity
- Microorganisms can overcome iron nutritional limitation in the host by procuring iron:
Extracellularly; get iron from our extracellular stores eg. transferrin | lactoferrin
Intracellularly: from host hemoglobin. Many pathogens produce hemolysins to lyse RBCs
- Nutritional immunity can be overcome in the host via two general mechanisms (bacterial molecules used to access iron):
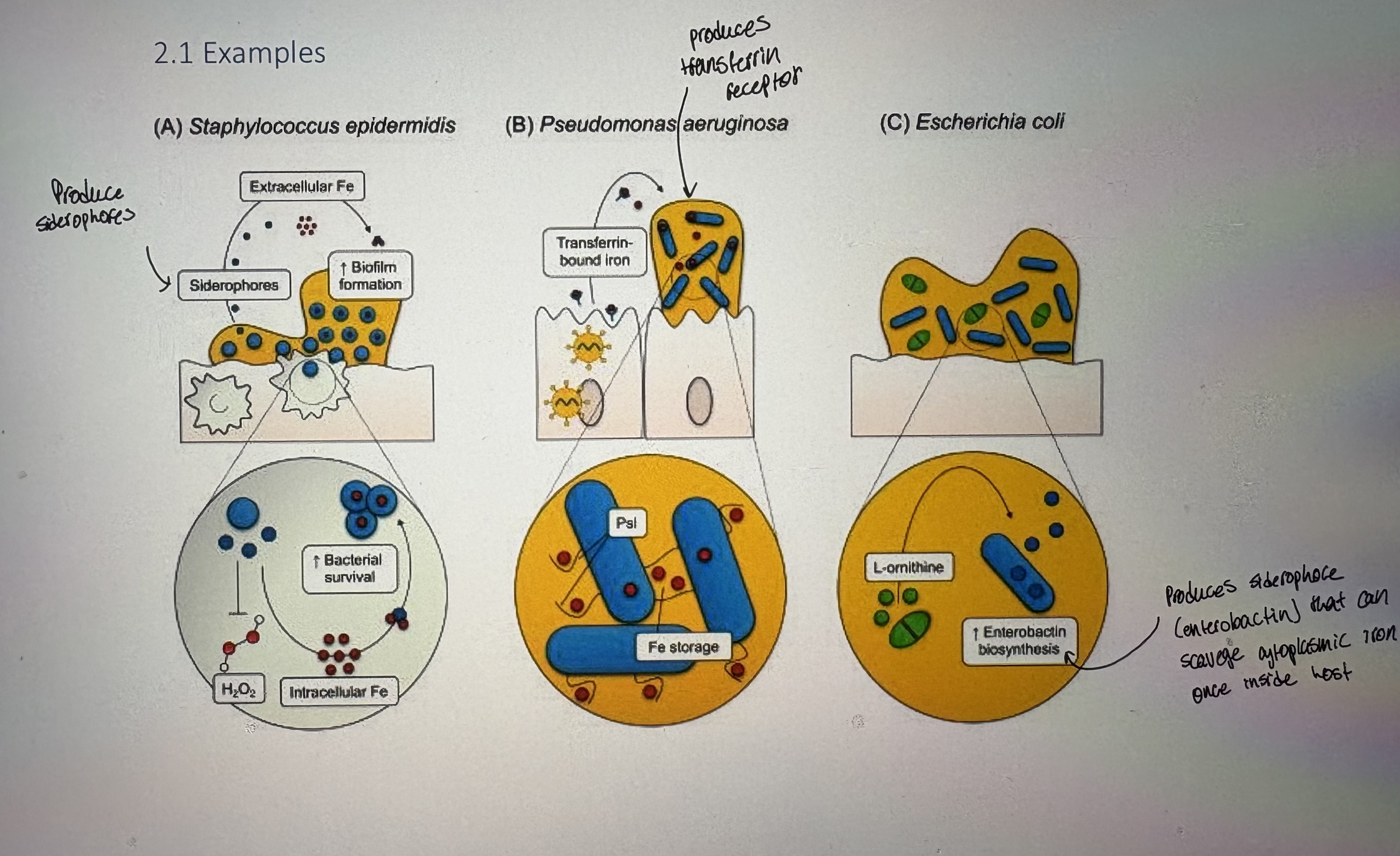
Siderophores
Bacterially produced molecules that chelate iron w/ high affinity
Usually bind free iron (low availability)
Because siderophores are secreted by bacteria, the microbe must also have ways to take up iron-bound siderophore
Receptor-mediated iron acquistion from host proteins
bacteria can take up heme, lactoferrin, etc. BUT host proteins need to be stripped of iron.
- Both Gram-negative and Gram-positive bacteria can sequester iron from transferrin, lactoferrin, hemoglobin, and siderophores
Gram (+):
Heme can be directly brought into the cell
Iron-siderophore complexes (brought in as a complex)
ABC transporter family can bring in iron-containing molecules (ATP mediated). Often, iron is stripped during transport liberating iron for a nutrient for bacteria
Gram (-): Iron-containing proteins need OM-transport into periplasm then periplasmic proteins strip iron then CM transporters internalize iron.
Heme receptors on the OM
Secrete scavenger binding proteins
Bring in iron-siderophore complexes
- Bacteria have OM receptors that are specific to host iron-containing proteins ie lactoferrin receptor specifically for lactoferrin.
- Exs in pic
- Mammals respond:
Siderocalin is a mammalian lipocalin-type protein → binds siderophores = should prevent pathogen from obtaining iron
Siderophores → Fe scavengers from bacteria. Siderocalin → siderophore binding molceules from host
Some pathogens have evolved mechanisms to evade siderocalins
Borrelia burgdorferi
- Bacterium that causes lyme disease
- B. burgdorferi resembles other spirochetes - has an OM & IM with a thin layer of peptidoglycan in-between
- Can live without iron. How?
eliminates Fe-requiring proteins
Substitute FE w/ Mn (manganese → chemically equivalent to Fe)
Bacterial toxins
- A poisonous substance produced by bacteria.
Promotes infection by causing host cell damage = disease
- A toxin is generally specific to a particular bacterial species
Ex. Only Clostridium tetani produces tetanus toxin. Only Corynebacterium diphtheriae produces diphtheria toxin
- Endotoxins: LPS.
Gram (-) Lipid A is an endotoxin (component of bacterial structure that is toxic)
Even small amounts can activate immune system → response: fever, inflammation, & can induce shock
- Exotoxins: secerted toxins with high potency. Mostly in Gram (+) bacteria
specific to a paticular cell structure or function in the host
Producted in virulent stains
3 main types: 1) AB toxins 2) Pore forming toxins (PFTs) 3) Superantigens (SAgs)
AB toxins
- The AB toxins are two-component protein complexes secreted by a number of pathogenic bacteria
- They interfere with internal cell function
- Many protein toxins that act intracellularly (within the host), consist of two components:
One having enzymatic activity (called “A”, for activity)
One binding at the cell surface, which leads to internalization (called “B” for binding)
- Mechanism of action:
ADP-Ribosylation. A subunit has enzymatic activity.
ADP-ribosylation is the addition of one or more ADP-ribose moieties to a protein.
ADP-ribosylates various target proteins
Diff toxins have diff targets of ribosylation
Ribosylation = covalent, post-transcriptional modification (is bulky; gets in way of function)
Enzymes are active in host cytoplasm inactive host proteins
- Two major classes:
Single peptide chain AB toxins eg Clostridiodes botulinum
Multisubunit AB-type toxins eg Vibrio cholera (in some species A and B are encoded by diff genes)
A-B: components are synthesized seperately but associate during secretion
A+B: components are synthesized seperately but associate after secretion
5B (or AB5): components are synthesized seperatley & the complex 5 x B subunit eg CT = cholera toxin
Clostridiodes botulinum
- Produces single peptide chain AB toxins
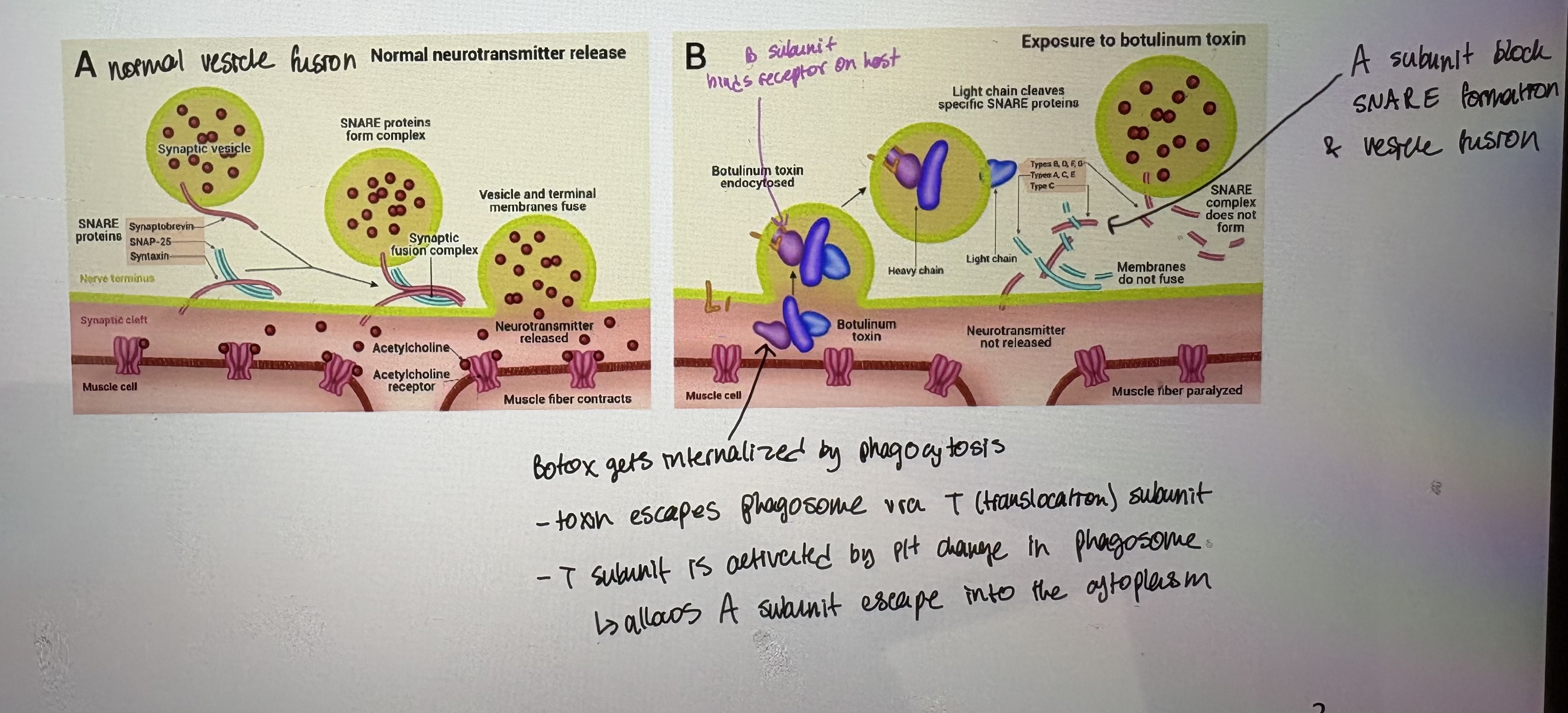
- BoNT (Botox) attacks neurons in humans and animals and is responsible for paralytic symptoms
- Toxin genes likley spread by horizontal gene transfer in pathogenicity islands
- Toxin binds to neurons & prevents neurotransmitter vesicle fusion & release of neurotransmitter
- AB toxin is internalized by the B-subunit binding to the cell
Pore-forming toxins (PFTs)
- Protein exotoxins
- Frequently cytotoxic; they create unregulated pores in the membrane of targeted cells
- Two major types:
alpha PFTs: pore forming toxins that use alpha helical domains
beta PFTs: rich in beta sheets. Require olgimerization.
- PFTs insert a transmembranous pore into a host cell memb
Facilitated by the acidic pH near the surface of the host membrane
All cells have a slightly acidic pH near surface
- Ex. LLO- Listeriolysin O from Listeria monocytpgenes (beta PFT → allows host cell specificity)
Binds to membranes with high cholesterol content
Forms the largest known membrane pores
Beta-hemolytic
Superantigens (SAgs)
- Are a class of antigens. Causes massive immune repsonses & nonspecific targeting
Cause non-specific activation of T-cells resulting in polyclonal T cell activation and massive cytokine release
SAgs are produced by some pathogenic viruses and bacteria as a defense mechanism against the immune system
- Bacterial SAgs are secreted pyrogenic (cause fever) exotoxins
- Exs.
Staphylococcal enterotoxins serotypes A-E, G, & H
Group A Streptococcal pyrogenic exotoxins A-C
Staphylococcal exfoliatin toxin
Staphylococcal TSST-1
- 1970s – young woman died of TSS – toxic shock syndrome due to super-absorbant tampon use
S. aureus colonized the vagina and was virulent due to SAg production
- SAgs bind MHCII of macrophages and receptors on T-cells (TCRs) without entering the cell
- SAgs stimulate pairing of macrophages & Tcells
Causes massive production of inflammatory cytokines = leads to shock
SAgs can activate ~20% of Tcell population
Secretion systems
- zthere are different types of secretion systems used by bacteria
Named T1SS through T6SS
- Function:
Protection
Transport of extracellular components
Communication
Release of virulence factors; most pathogens have evolved specialized secretion systems for virulence factors
- Mechanisms differ between Gram-negative & Gram-positive bacteria
- In Gram-negative bacteria, secretion can occur in:
1-step process: across both CM & OM eg. T1SS, T3SS, T4SS, T6SS
2-step process: export into the periplasmic space via the universal Sec or Tat pathways 1) across CM via Sce or Tat 2) across OM via T2SS or T5SS
- In Gram-positive bacteria:
Secreted proteins are commonly translocated across the CM by the Sec pathway or Tat pathway
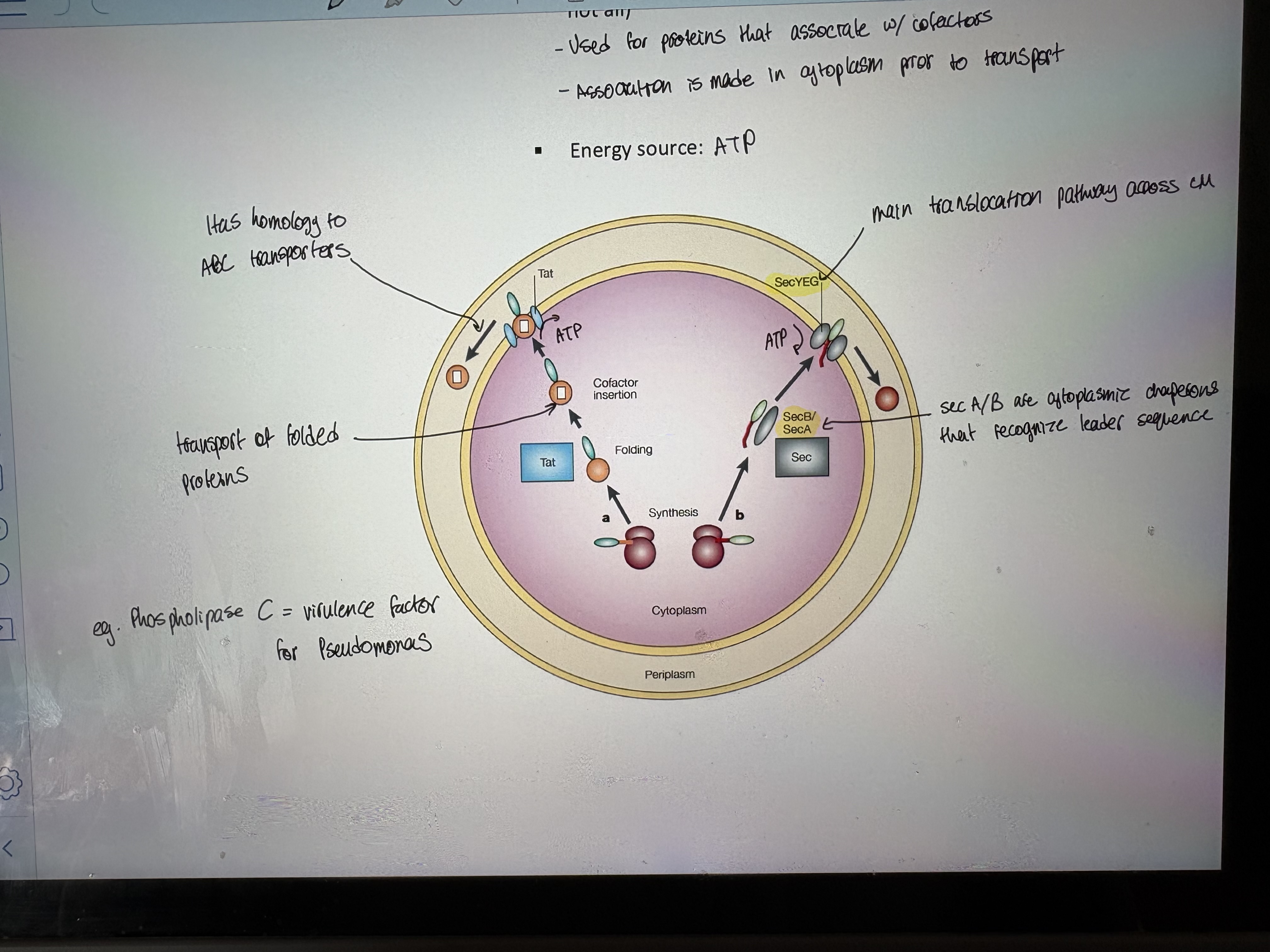
Sec system translocates unfolded proteins
o Common to all Gram (+) and (-). Translocated proteins have a hydrophobic N-terminal leader sequence (signal for secretion). Requires ATP
Tat systems transport folded proteins (see other pic)
Found in both Gram positive and negative bacteria (but not all). Used for proteins that associate w/ cofactors. Association is made in cytoplasm prior to transport
Energy source: ATP
- Overview in Gram (-): many secretion systes are encoded by operons. Allows easy sharing via horizontal gene transfer
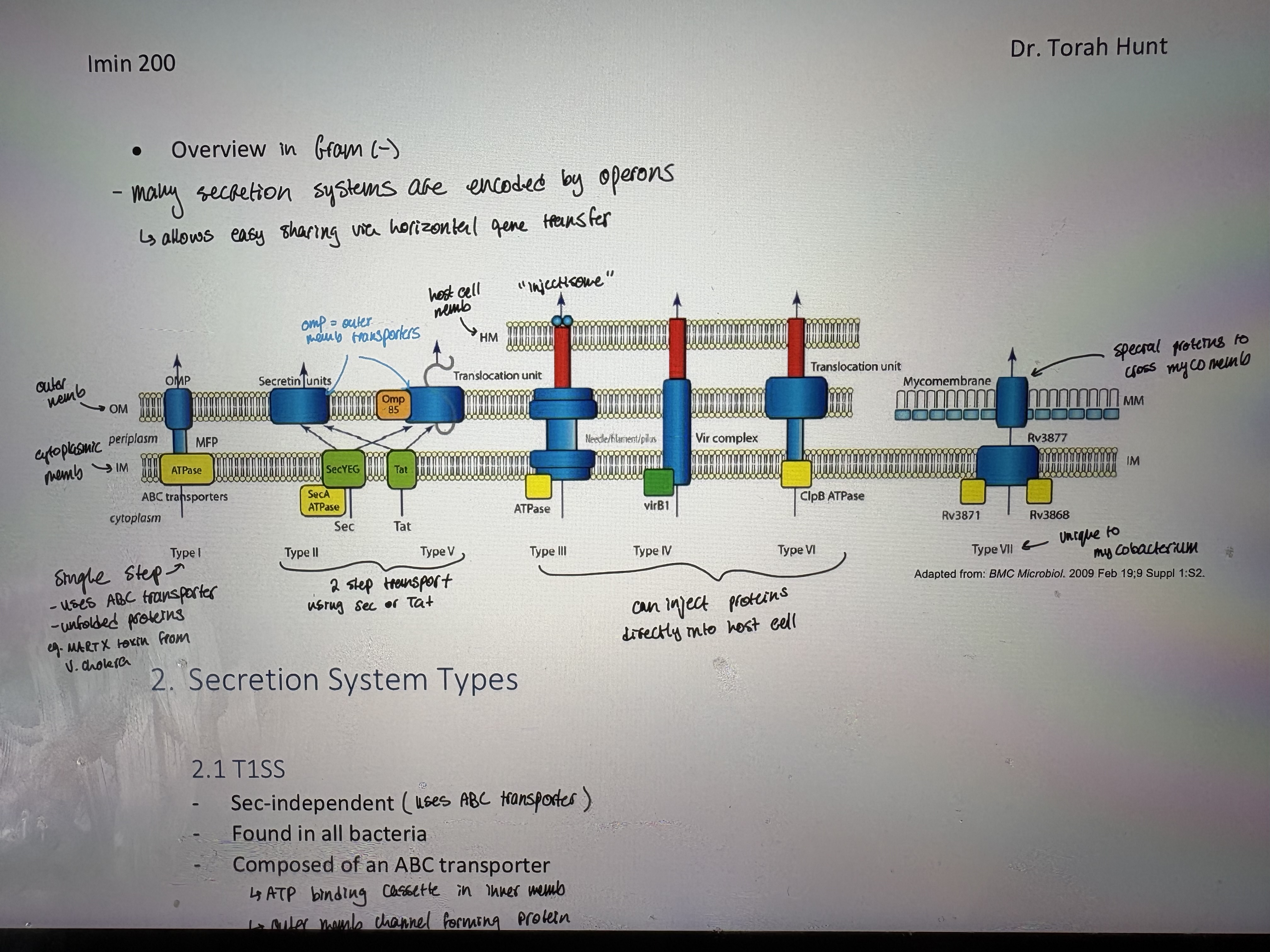
T1SS/T2SS
- Sec-independent (uses ABC transporter)
- Found in all bacteria
- Composed of an ABC transporter
ATP binding cassette in inner memb
Outer memb channel forming protein
Periplasmic fusion protein
- Unfolded proteins are transported directly from cytoplasm to extracellular environment = one step secretion
- Example virulence factor:
α-hemolysin (a PFT) from E. coli
MARTX toxin of V. cholera (“cluster bomb”. Anti inflammatory prevents immune response) → a very large protein. Cleaved after transport by autoprocessing cysteine protease domain. All cleavage products are toxins.
see pic
- The type ll secretion system (T2SS) is also known as the Sec-dependent system
- 2 step secretion:
Many proteins that pass through the T2SS must first reach periplasm via the Sec pathway
- The T2SS is evolutionary related to the pilus biogenesis machinery
- Virulence determinants secreted via the T2SS include:
ADP ribosylating toxin of enterotoxigenic E.coli
Cholera toxin (CT)
Exotoxin A of Pseudomonas
T3SS
- The type III secretion system (T3SS):
Found in many Gram-negative bacteria – pathogenic & non-pathogenic
- The machinery of the T3SS is termed the injectisome
The injectisome delivers unfolded effector proteins across bacterial and host membranes into the cytosol of host cells
- Once inside, they may modulate a large variety of host cell functions, including immune and defense responses
T3SS is a large multiprotein complex that can span 3 membranes
Inject proteins directly into host cytoplasm
- The structure and function of the injectisome have been well studied in the animal pathogens Salmonella typhimurium & Yersinia pestis where it was first described
- Genes for T3SS are often located in gene clusters on large pathogenicity islands suggesting these systems may have moved horizontally between species. See pic.
- T3S effectors are euk-like bacterial proteins
Interfere with a wide range of cellular processes
Display sequence, structural or functional resemblance to proteins found in higher organisms
- Example toxin that uses T3SS; YopE from Yersinia
Acts later in infection cycle. Yersinia switches from phagocytic in early infection to antiphagocyte → temp sensitive gene control
Injectosome introduces YopE to host cell cytoplasm. Causes actin cytoskeleton arrangements to prevent passage of endosome to lysosome.
Once inside endososmes = Yersinia switches gene expression to antiphagocytic
Adhesions = phagocyte
T5SS/T7SS
- The T5SS is the most widespread secretion pathway for the transport of molecules across the outer membrane of Gram-negative bacteria
- Called an autotransporter
A very large number of proteins are secreted via the T5SS including adhesins, toxins and proteases
Example virulence factor: IgA proteases of Neisseria gonorrheae and Neisseria meningitides. These proteases cleave the hinge region of immunoglobulin A
- Notes: Type V is often involved in housekeeping secretion therefore hard to use this as a therapeutic target w/out inhibiting good bacteria (also Type l)
- IgA1: allows immune system evasion inactivating immunoglobulin A
- Translocater domain: secreted by Type V
- The type VII secretion system (T7SS) is a specialized secretion apparatus that is required for the virulence of mycobacteria (i.e.: Mycobacterium tuberculosis)
In 2003, the first T7SS was identified (it was named ESX-1)
- T7SS gene clusters have since been identified in several Gram-positive bacteria:
Staphylococcus aureus, Listeria monocytogenes, Bacillus subtilis
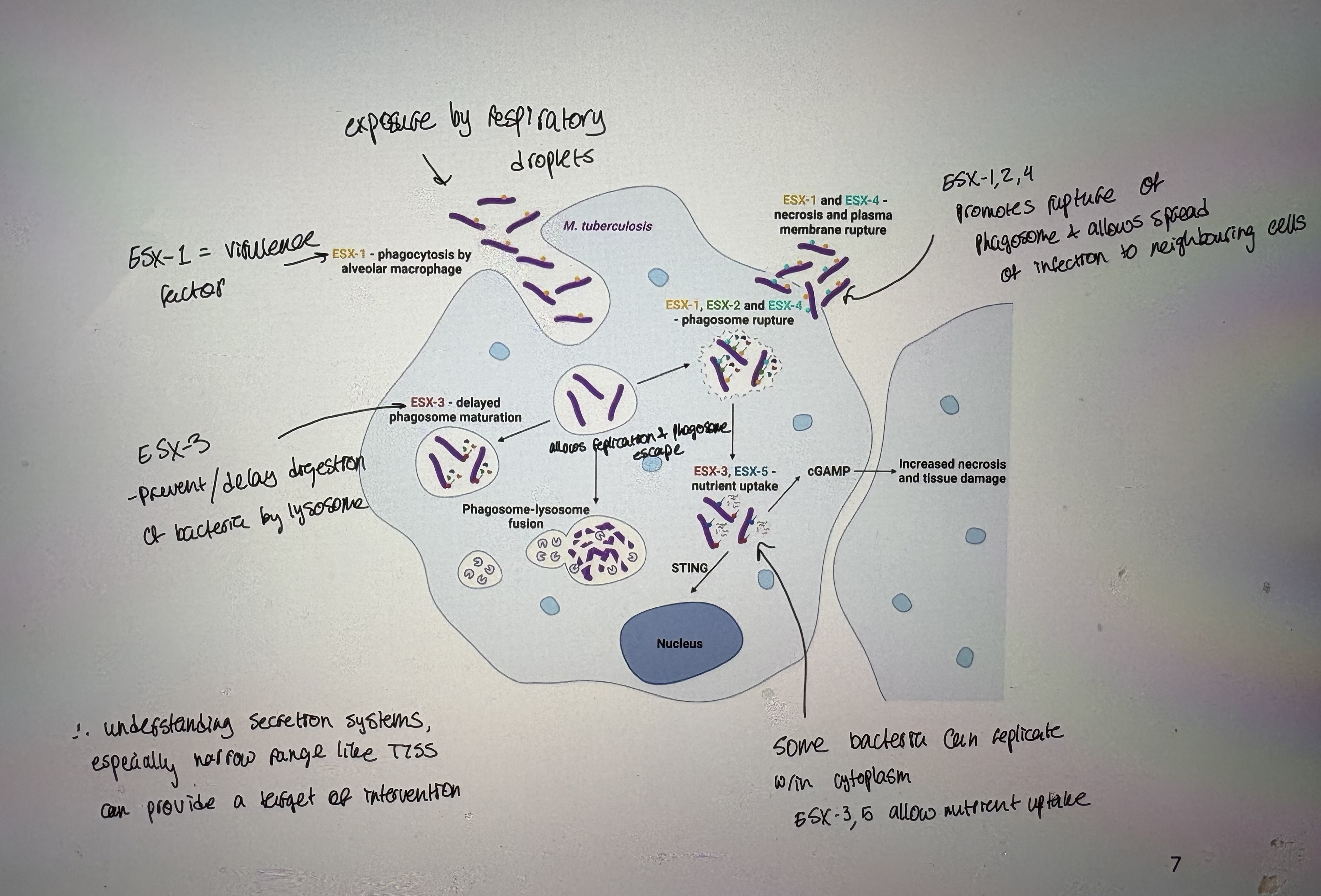
Mycobacterium tuberculosis
- Causatative agent of tuberculosis
- The cell envelope of mycobacteria consists of:
cytoplasmic memb
periplasmic space → peptidoglycan & arabinoglycan
mycomembrane → waxy mycolic acid outer layer
Composed of genes ESX1- 7 = most are virulence factors contributing to tuberculosis
- The T7SS allows the bacterium to escape destruction in phagocytotizing cells
secretes various virulence factors
- ESX-3 gene: prevents/delays digestion of bacteria by lysosome and nutrient uptake
Antibiotics; mechanism of action
- General principle behind antibiotics is that they must target only bacterial cells
- There are broad spectrum (target many species) vs narrow spectrum (target fewer species) antibiotics
- Advantages of narrow spectrum antibiotics
The narrow-spectrum antibiotic will not kill as many of the normal microorganisms in the body as the broad spectrum antibiotics
Problem w/ narrow spectrum → you have to wait for a positive culture & snesitivity test
need to target bacterial specific → bacterial ribosome, peptidoglycan
Classic antibiotics taregt peptidoglycan eg penicillin
Many treat unique prokaryotic ribosomes
- However, resistance has developed for all known antibiotics eventually
- Why does resistance develop?
Resistance has always been a part of the microbial arms race. It is NOT unique to pathogens.
Other cells produce antibodies to kill off competition therefore antibiotic resistant mechanisms are prevalent in nature
Overuse in prescriptions & agriculture → has given low dose exposure in environment. Have given weak selection for strong resistance (low risk oppertunities evolve)
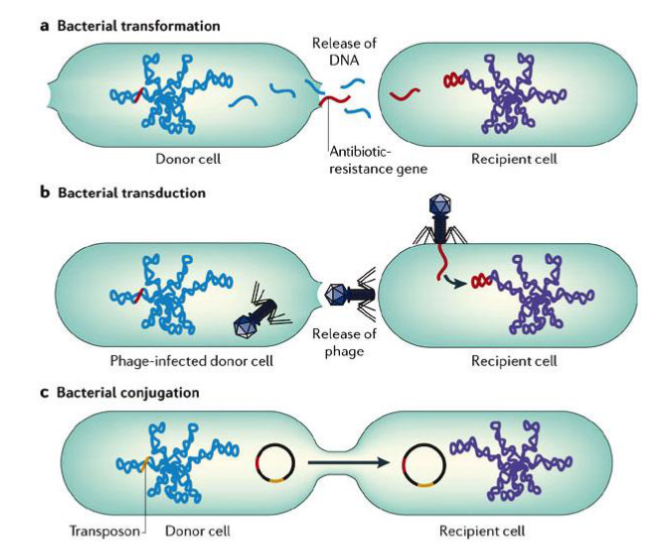
Antibiotics; sharing resistance
- Once resistance evolves by natural selection & random mutation then mutations can spread by horizontal gene transfer
- Transformstion = uptake of free DNA from environment
- Transduction = viral-mediated exchange of bacteria DNA (very random)
*requires sequence homolgy (similarity) to integrate into genome
- Conjugation = direct transfer of DNA between related species using a conjugated pilus
plasmid transferred may contain drug-resistance gene eg. R plasmid → resistance plasmid
- The population of bacteria, before exposure to an antibiotic, is composed of genetically variable bacteria.
- Upon repeated short exposure to antibiotic, natural selection takes place with survival of resistant bacteria
- Within a mixed population of bacteria there is existing variation
therefore exposure to an antibiotic if high dose → most bacteria will die
If there is low or incomplete dosages → creates a weak negative selective pressure → provides oppertunity for inc resistance
- What can we do to fight antibiotic resistance?
Decrease unnecessary exposure to antibiotics
Combination drug therapy:
Treating a bacterial infection with multiple antibiotics
Attacking multiple components of the bacterial cell simultaneously may weaken it enough to clear the infection
Identify new drug targets:
Essential and unique components in the bacterial cell
Develop new antimicrobials or chemically alter old drugs to make them more effective against MDR pathogens
Development of vaccines against multi-drug resistant pathogens
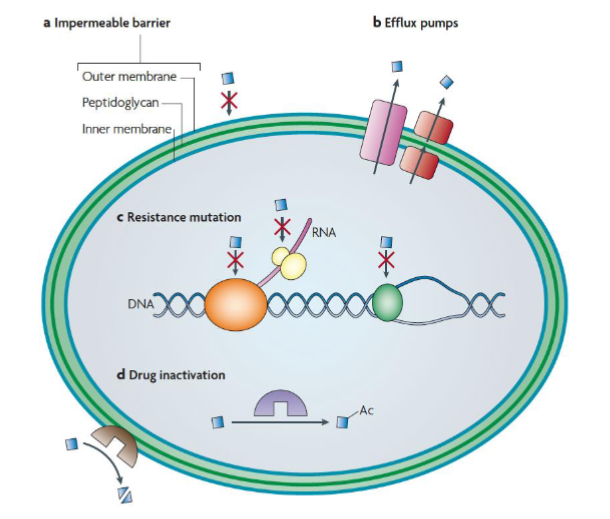
Antibiotics; mechanism of resistance
1. Impermeable barrier:
- Some bacteria are intrinsically resistant eg. mycobacterium, bacteria in a biofilm
2. Multidrug resistance efflux pumps:
- These pumps secrete antibiotics from the cell. Some transporters can pump antibiotics directly outside the cell, whereas others (red) secrete them into the periplasm (Gram-negative bacteria).
- There are five families of multidrug-resistance efflux pumps:
ABC superfamily: general efflux pump eg. E.coli resistance to chloramphenicol
MFS: major facilitator family eg. tetracycline resistance
MATE: multidrug and toxic compound extrusion family
SMR: small multidrug resistance eg. Ecoli
RND: resistant module superfamily. Only in Gram (-) eg P aeruginosa
3. Resistance mutations:
- These mutations modify the target protein, for example by disabling the antibiotic-binding site but leaving the cellular functionality of the protein intact.
- Specific examples include mutations in:
gyrase: mutations in gyrase have allowed resistance of fluoroguinolones
small ribosomal subunit: allowed resistance streptomycin
4. Inactivation of the antibiotic:
- Inactivation can occur by covalent modification of the antibiotic:
- such as that catalysed by acetyltransferases (purple) acting on aminoglycoside antibiotics
- or by degradation of the antibiotic, such as that catalysed by β-lactamases (brown) acting on β-lactam antibiotics
Ac = acetyl group (-CH2-CH3). Covalent addition inhibits antibiotic function.
ESKAPE pathogens
- A group of highly antibiotic resistant bacterial pathogens which are responsible for a large portion of hospital acquired infections worldwide:
1. Enterococcus faecium (VRE)
- VRE = vancomycin-resistant Enterococci. Causes a range of illnesses eg blood infections
- Vancomycin binds to the D-ala-D-ala peptidoglycan peptide preventing transglycosylation and transpeptidation
- Vancomycin cleaves crosslink in peptidoglycan → destabilizes cell wall making bacteria sensitive to osmotic lysis → resistance required via mutation in PG synthesis = change to D-alanine in linker
2. Staphylococcus aureus (MRSA)
- MRSA = methicilin-resistant Staphylococcus aureus
3. Klebsiella pneumoniae
4. Acinetobacter baumannii
5. Pseudomonas aeruginosa
- Causes lung infections. Used to be sensitive to pencillin.
- Pencillins contain β-lactam ring. Resistance mutation = β-lactamases cleaves & inactivates antibiotic.
- β-lactamases are enzymes produced by bacteria & provide resistance to β-lactam antibiotics such as penicillins.
6. Enterobacter spp