L4: Mitosis and Cell division
1/80
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
81 Terms
Recap of 1aBoC: interphase
Cell growth
centrosome duplication
chromosome replication
establishmnet of sister chromatid cohesion
What happens when enter M phase
Radical remodelling of cytoskeleton
MT-based spindle drives chromosome segregation
contractile actomyosin ring powers cytokinesis
other cellular components segregated along the cytoplasm (e.g ribosomes)
Regulated vesiculation and fusion controls the segregation of endomembraneous structures
Phases of mitosis based on its cytological landmarks
Prophase
Pometaphase
Anaphase A and B
Telophase
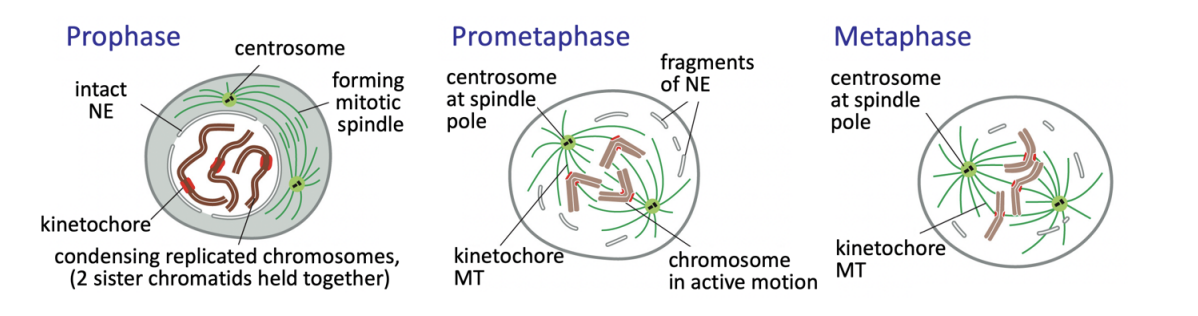
Prophase
chromatin condenses into well-defined chromosomes
two sister chromatids held together
Duplicated centrosomes (in most animal cells) split
migrate to set the spindle poles
As interphase MT network disassembles, centrosomes begin nucleating highly dynamic MTs
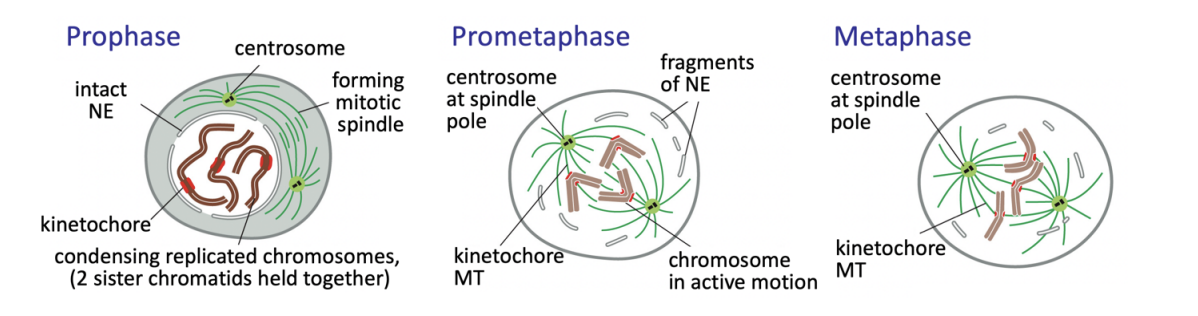
Prometaphase
Nuclear envelope (NE) breakdown complete
OPEN MITOSIS
MT capture at kinetochores assembled at the centromeric region of each chromosome begins
chromosomes then congress to cell equator→ metaphase plate
remains under tension
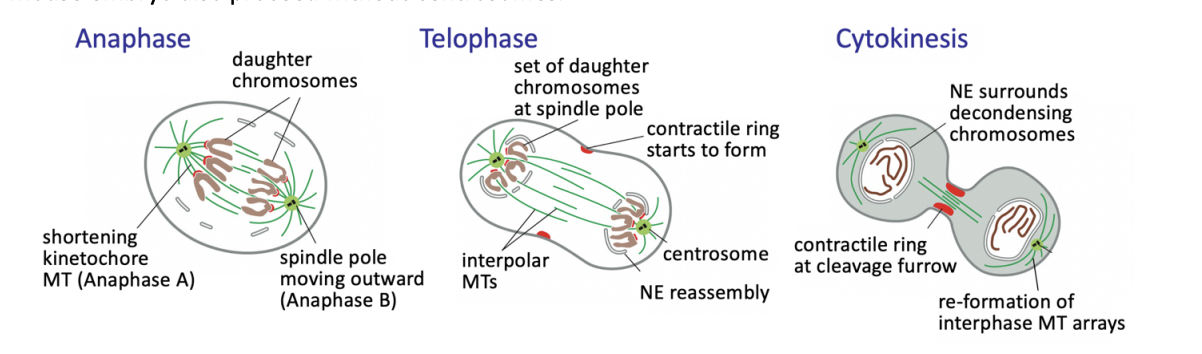
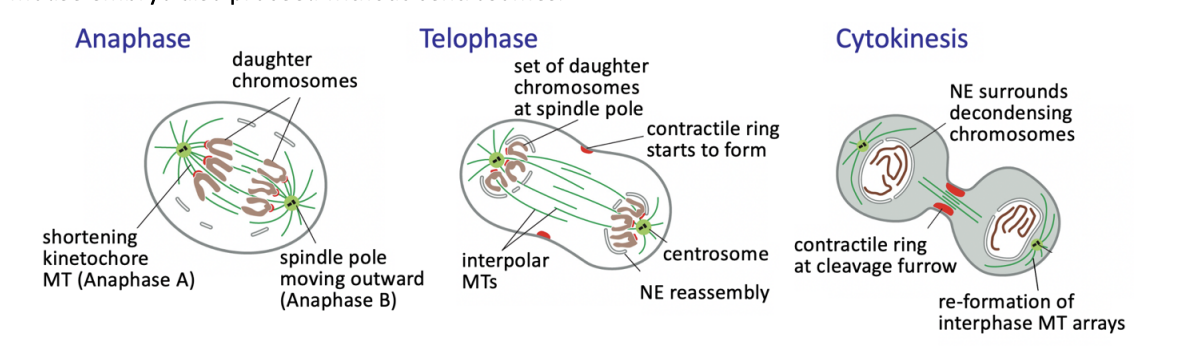
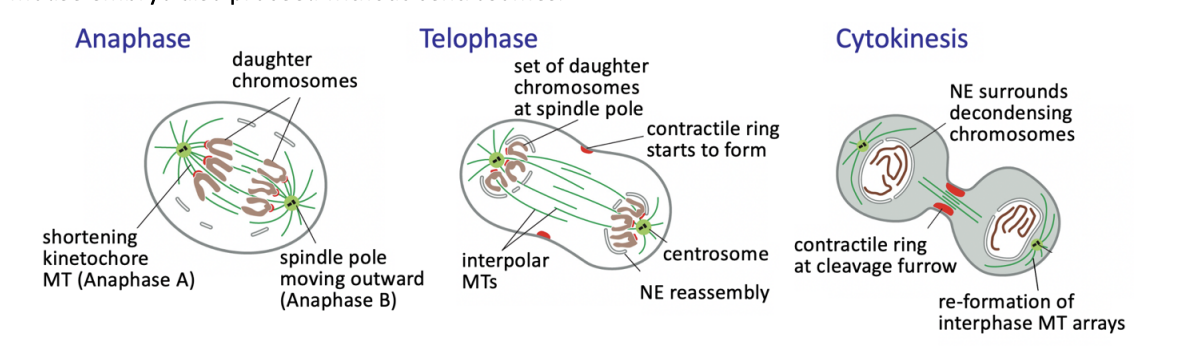
Anaphase
triggers loss of cohesion and separation of all sister chromatids
defines the birth of two new cells
from then on, each sister chromatid is referred to as a chromosome
Anaphase A
Kinetochore-MT shortening the chromosomes move poleward
Anaphase B
interpolar MTs grow and slide outwards
further separating the spindle poles
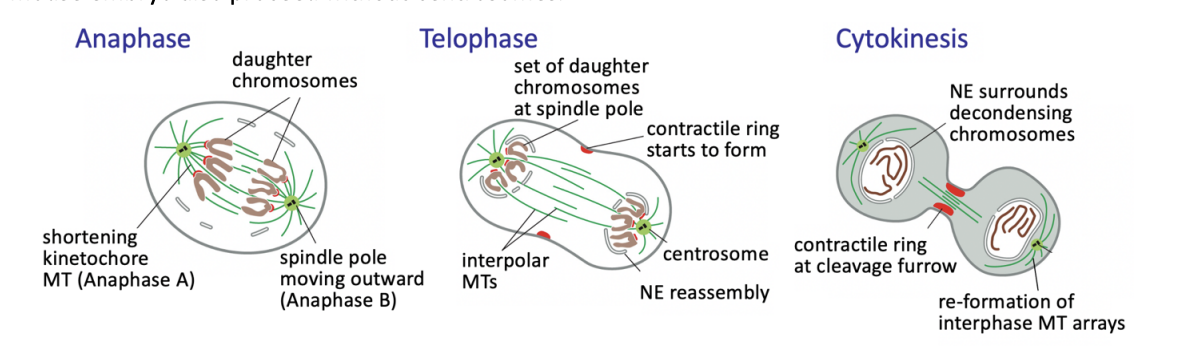
Telophase
chromosomes reach the poles
NEs refordm around them
chromatin recondenses
How is fungal mitosis different
Closed mitosis
mitotic spindle is formed within an intact nucleus
How is female meiosis in many animal species different
bipolar spindles are formed in the absence of centrosomes
What about the first mitotic divisions of the mous embryo?
also proceeed without centrosomes!
Cytokinesis: Animals
divide by constriction of an actomyosin contractile ring
Cytokinesis : Yeast
dividing by combining constriction of an actomyosin ring and the deposition of a septum
Cytokinesis: Plants
All of mitosis the same but just without centrioles
Cytokinesis
without contractile ringe
instead: deposition of specialised vesicle based structure
occurs at the site of division→ the cell plate
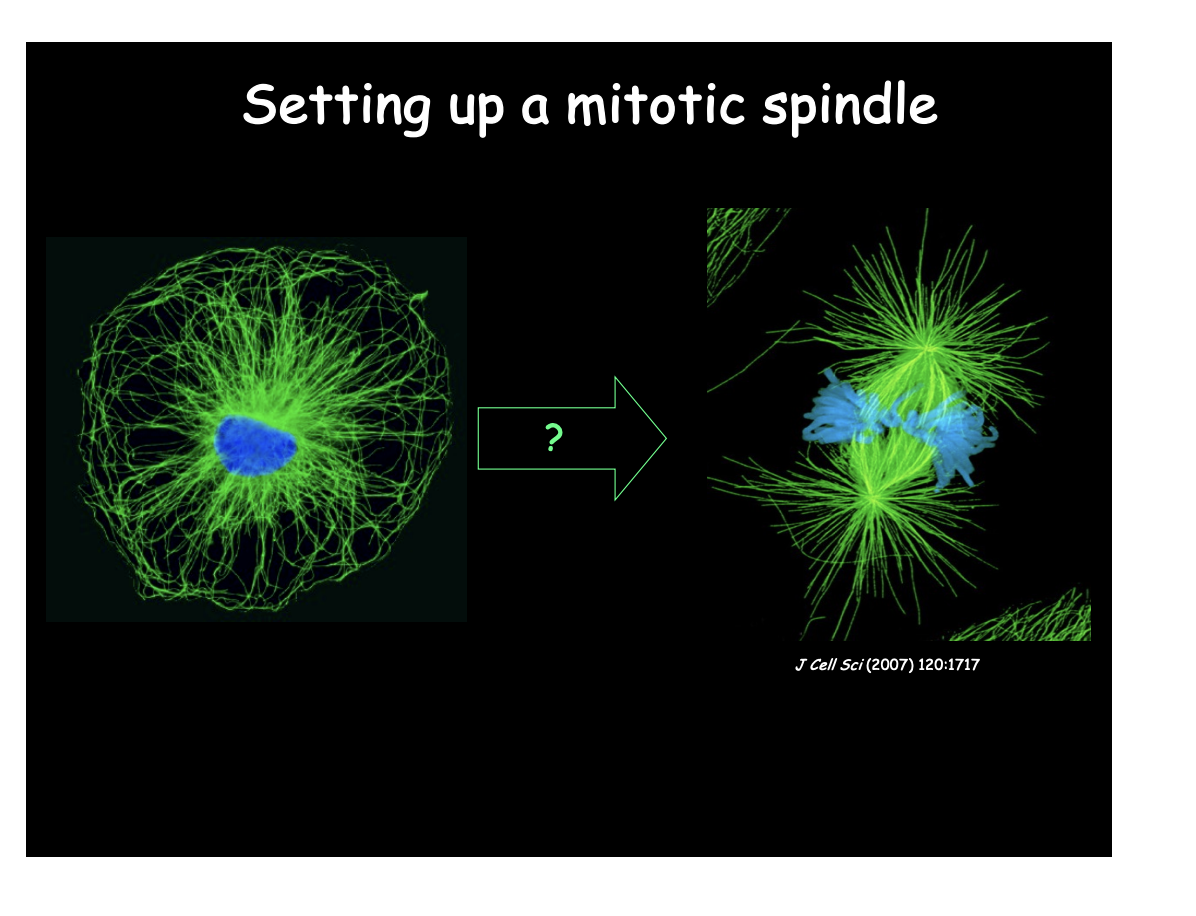
Focus of this lecture
How are these structures assembled?
How do they work?
How is spatial and temporal coordination between the two achieved?
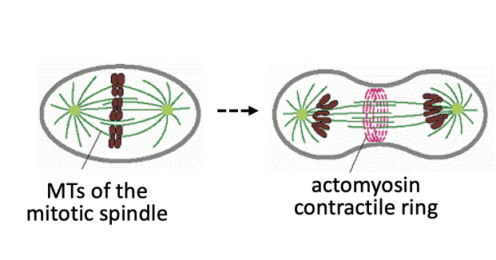
Overview: The spindle- an MT-based machine→ What is the spindle
Self organised bipolar array of microtubules
made up of three main classes of MTs
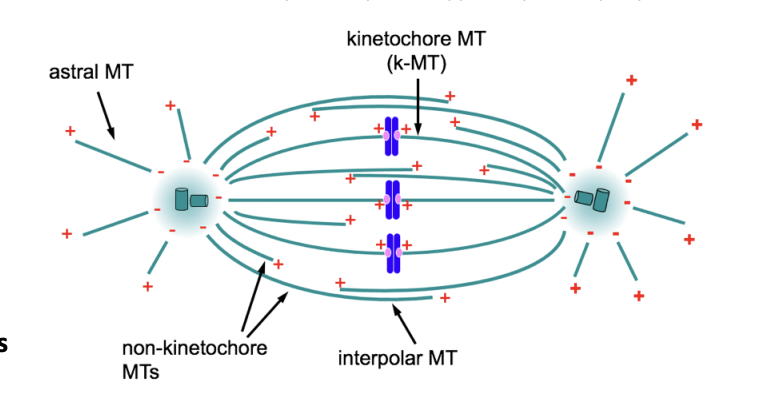
What three main classes of MTs
All
- ends focused at the spindle poles
typically set up by the centrosomes
BUT differ in regard to oreientation:
Kinetochore MTs (kMTs)
interact with kinetochores of mitotic chromosomes
Interpolar MTs
non-kinetochore MTs
interact at overlaps with non-kinetochore MTs emanating from the opposite pole
become cross-linked generating the spindle mid-zone
Astral MTs (aMTs)
extend toward the cell cortex
kinetochore MTs in yeast vs animals
Yeast
single kMTs attach each sister kinetochore
Animals
each kinetochore ends up attached to a bundle of 20 kMTs
→ Kinetochore fibre (k-fibre)
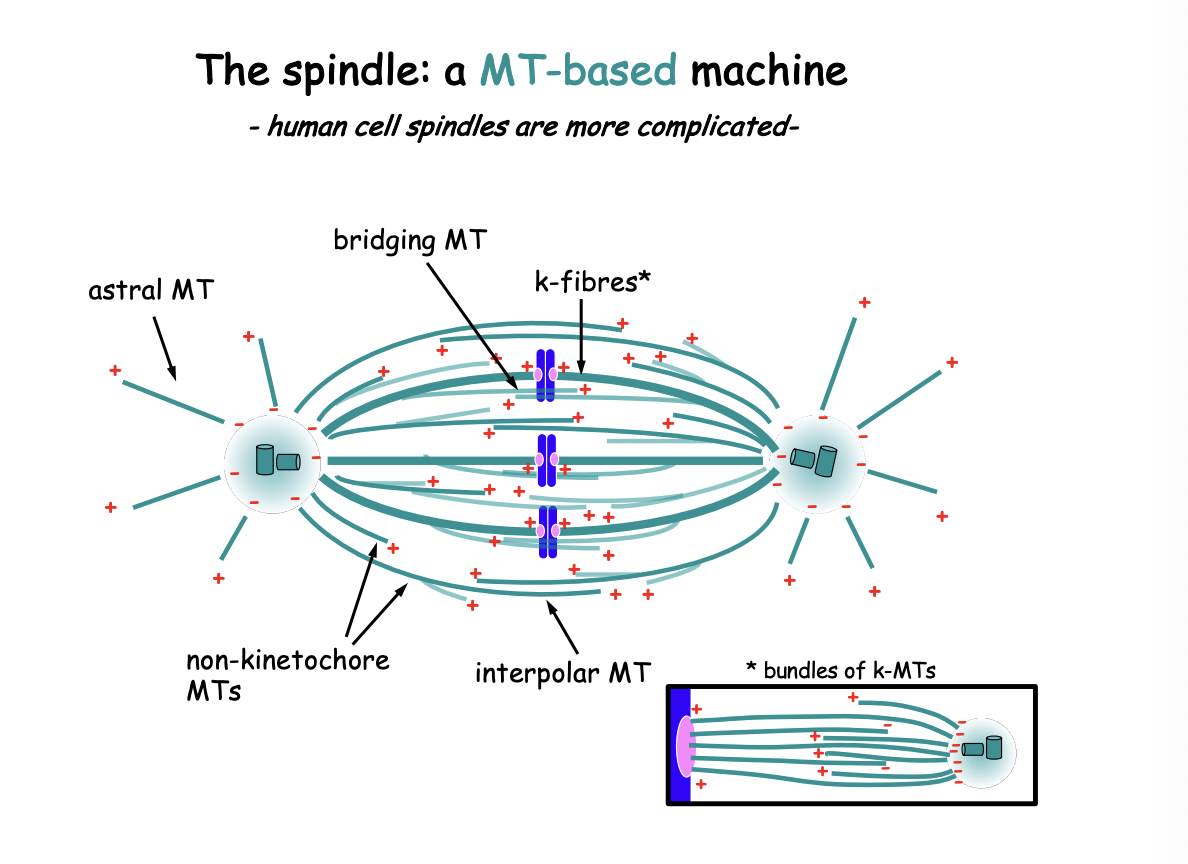
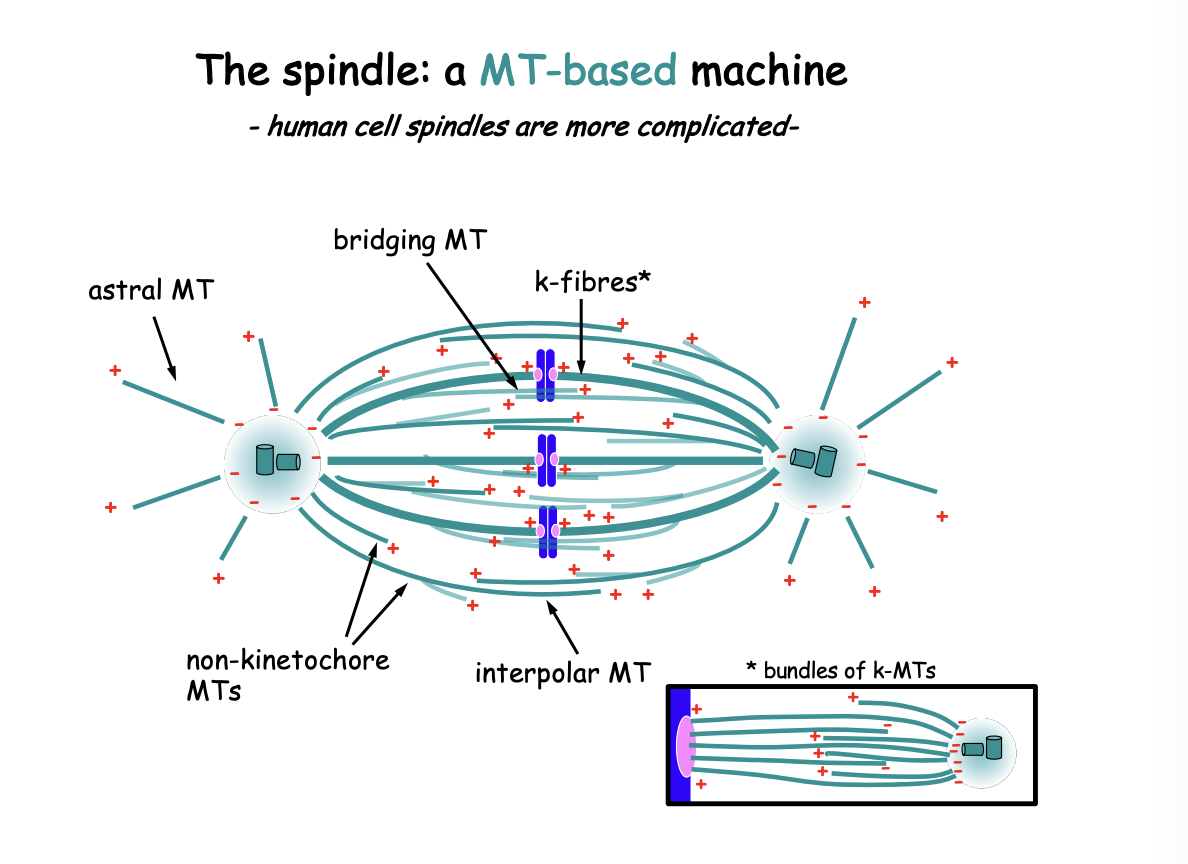
What has the 3MT population view been challenged by?
variety of advances in 3D microscopy analysis
E.g
3D reconstruction of entire spindles
What do these studies suggest?
mitotic spindle contains significantly more complex MT based modules:
may branch off from other MTs
not all (-) ends are buried near the centrosome
MTs may form antiparallel bridges between k-fibres
also contributing to spindle dynamics
Suggest human k-fibres contain BOTH MTs emanating from the centrosome AND MTs that do not reach the spindle pole
those may be crosslinked or interact through various MT-associated components
How and when do these MTs arise
When:
M phase
How:
from parallel pathways from MT nucleation:
centrosome-dependent pathway
centrosome-independent pathways
In both need:
Dynamic instability
large collection of MT interacting proteins
Search and capture of chromosomes
Once all chromosomes are bi-oriented→ they will essentially aligned at the spindle equator
Centrosome-dependent pathway: when does centrosome duplication happen
interphase
G1→ single centrosome is present with mother and daughter centrioles
S phase→ centrosome duplication begins
helped understand with super-resolution microscopy
semi-conservative
M phase→ each new centrosome in M phase carrying one of the original centrioles present in G1
Outline of the pathway (please check this slide!)
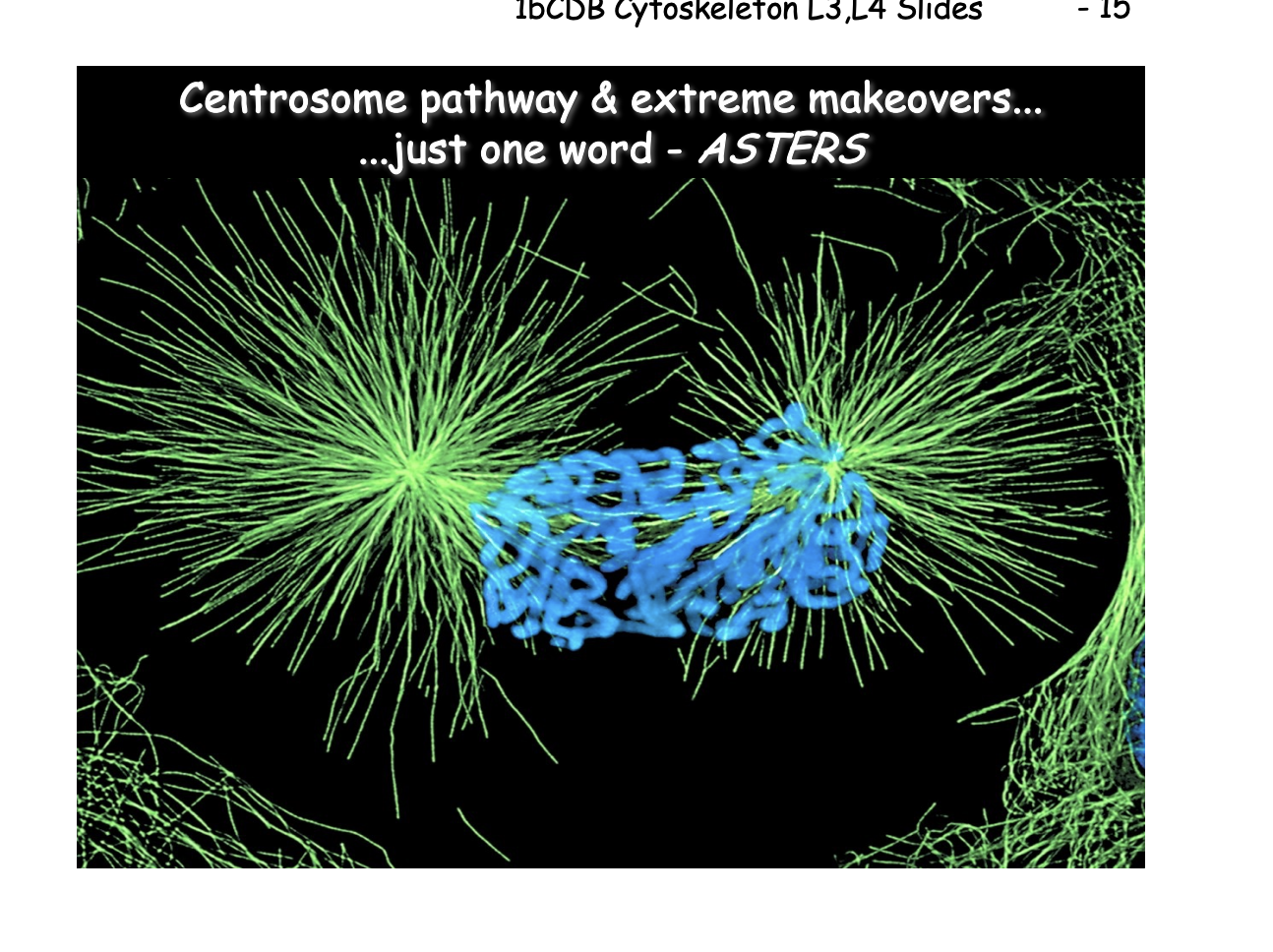
Increased MT dynamics, centrosome separation and aster formation
Antagonistic activities of MAPs and kinesin-13 control aster MT dynamics
Dynamic instability allows MTs to probe the 3D space of the cell
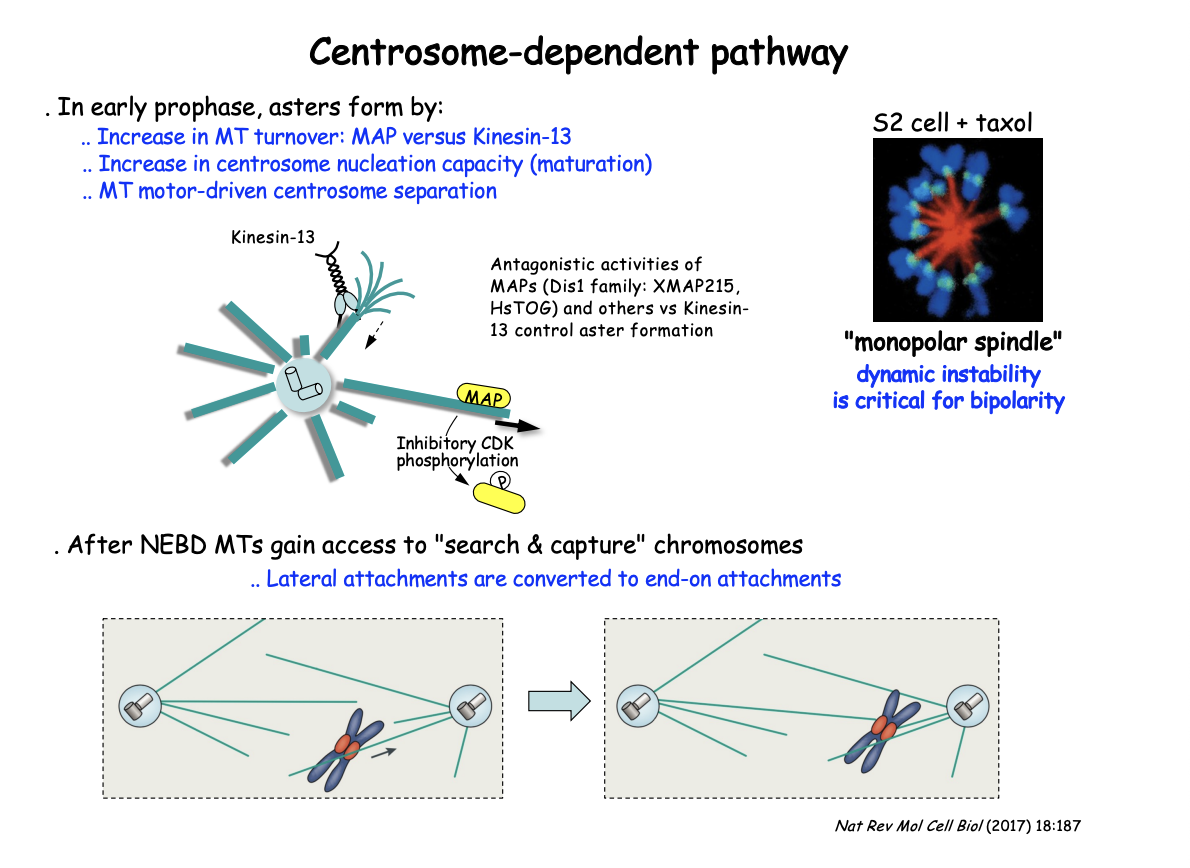
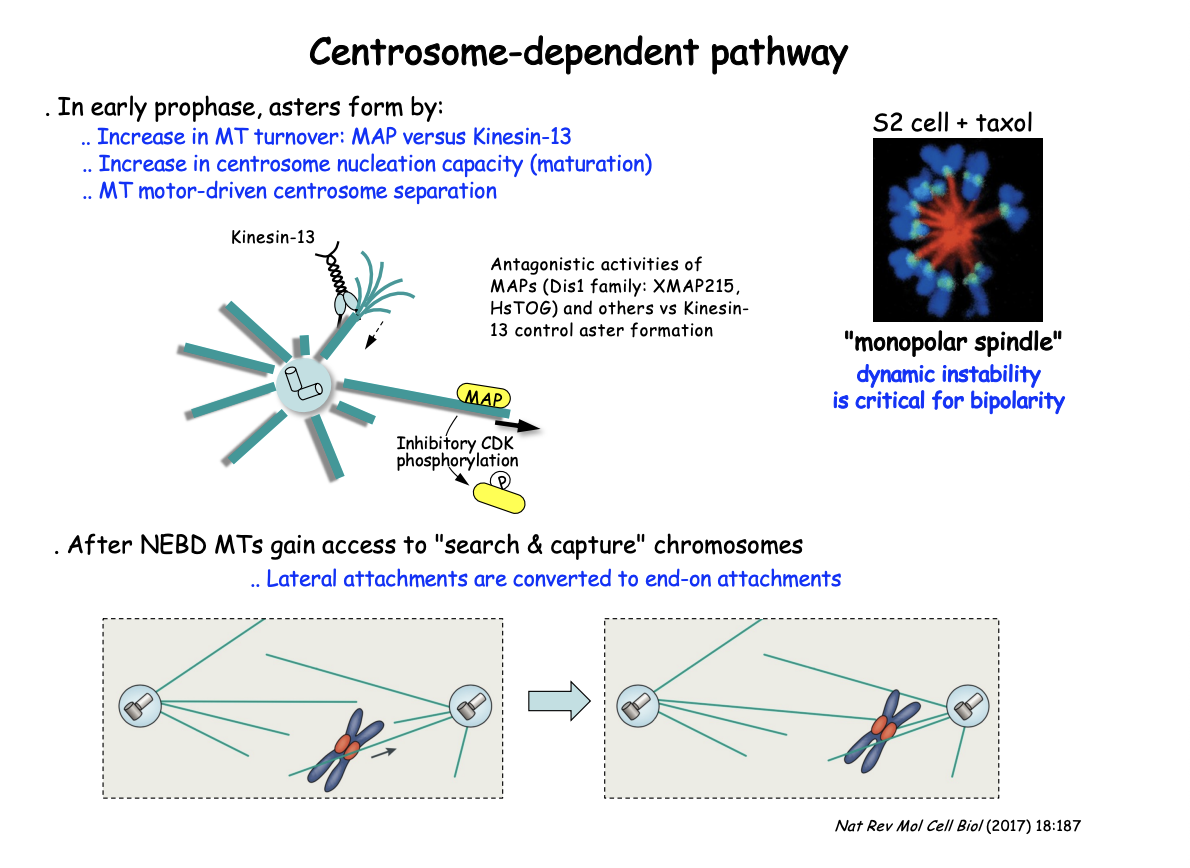
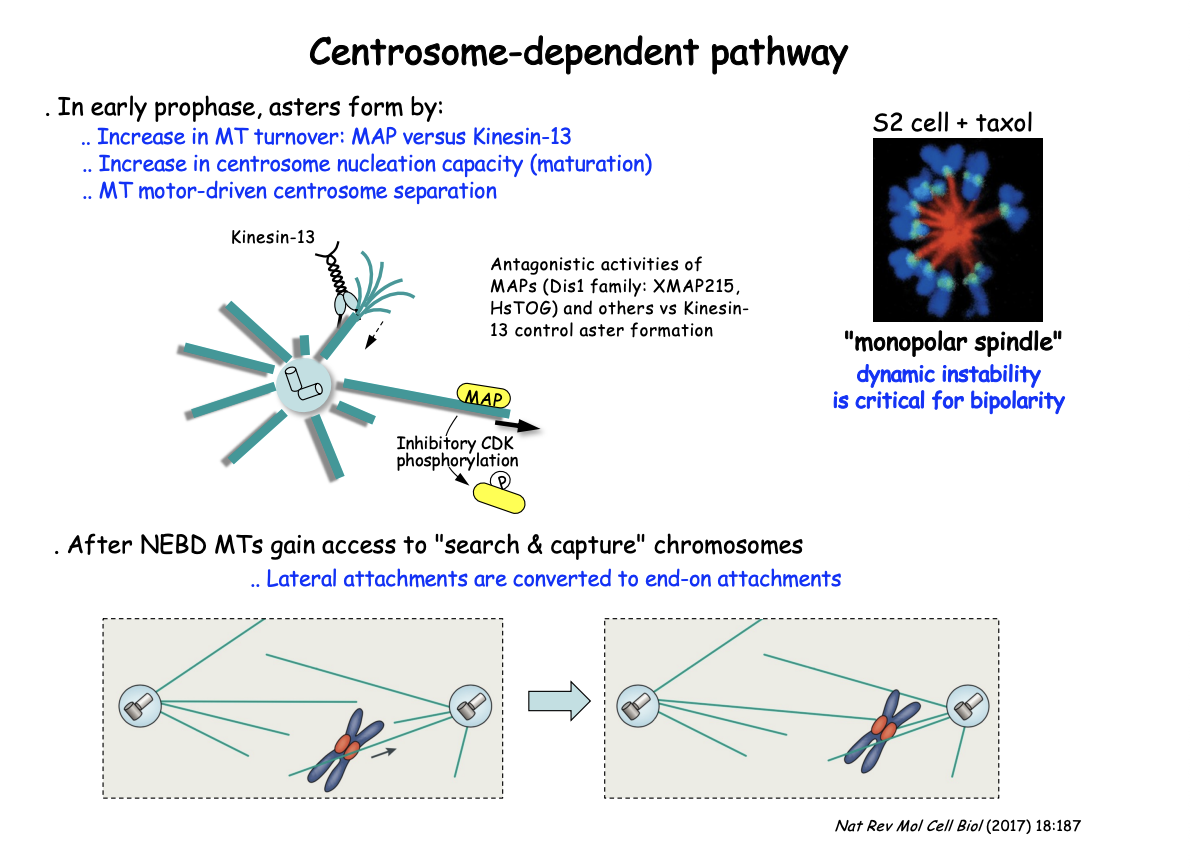
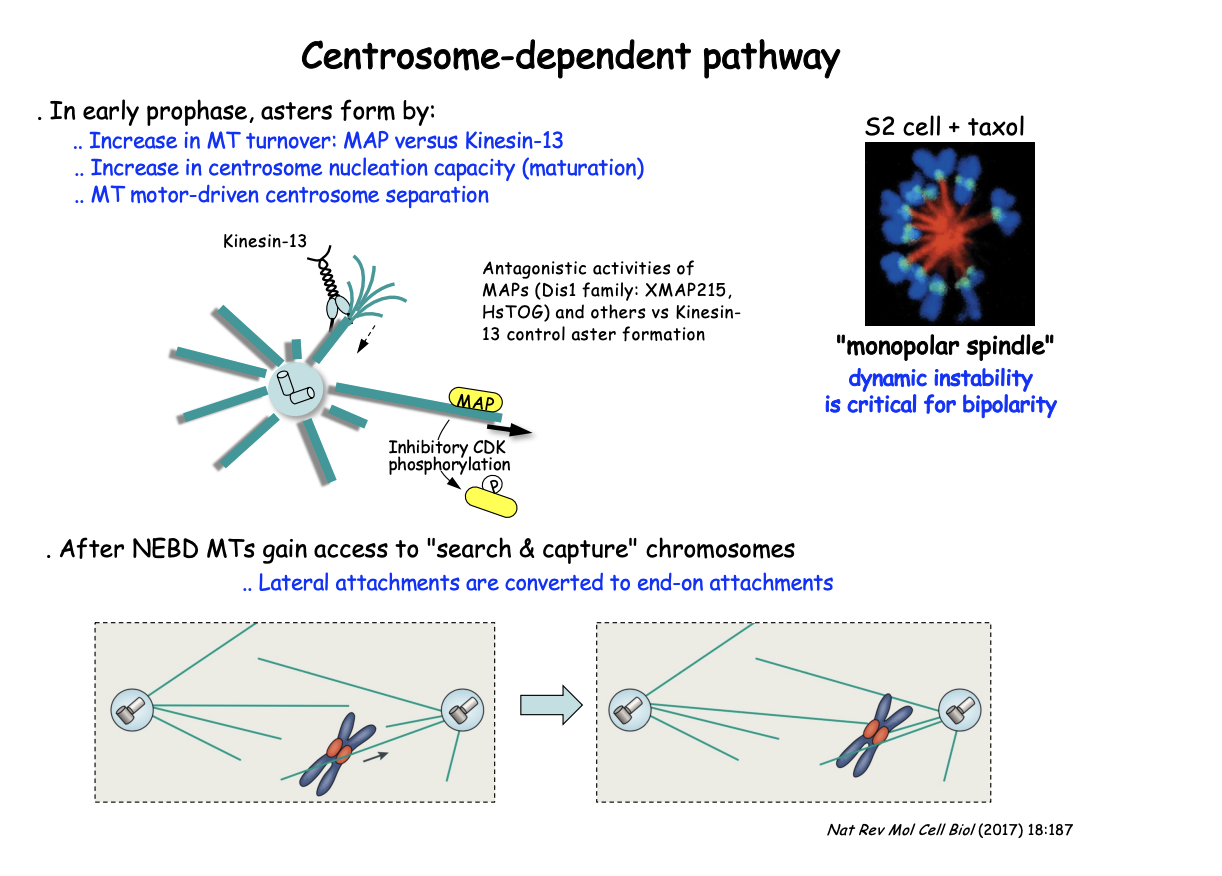
In early prophase, asters form by
increase in MT turnover: MAP vs Kinesin-13
Increase in centrosome nucleation capacity (maturation)
MT moto-driven centrosome separation
After NEBD MTs gain access to ‘search and capture’ chromosomes
lateral attachments are converted to end-on attachments
Increased MT dynamics, centrosome separation and aster formation
Dramatic increase in MT turnover during early prophase
centrosome separate (due to MT-based motors)
complete ‘maturation’→ progressively recruit more PCM increasing their nucleation capacity
Asters are formed
Astral MTs interact with the cell cortex to help aster separation
these interactions are powered by dynein/dynactin anchored at the cell cortex
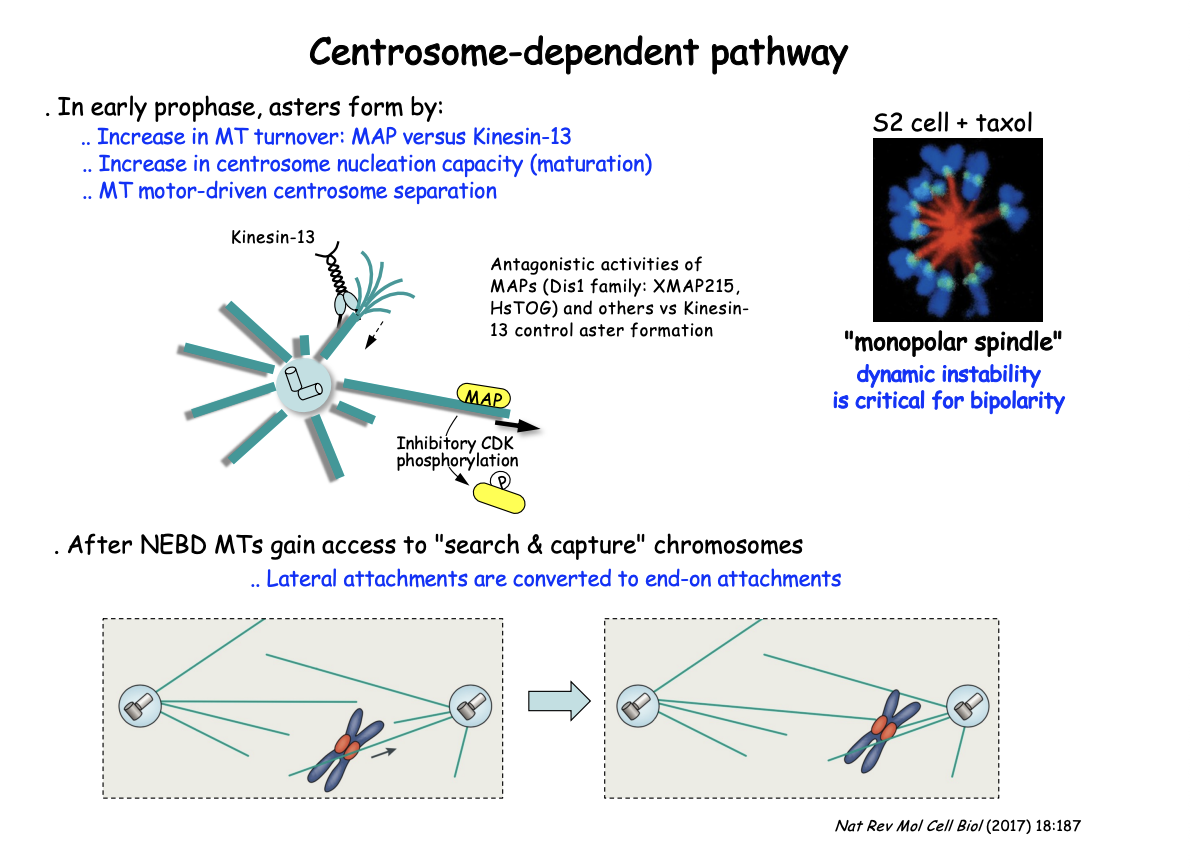
Antagonistic activities of MAPs and kinesin-13 control aster MT dynamics
Dis1 family MAPs (XMAP215, HsTOG) promote MT stability
BUT growth counteracted by the MT-depolymerising kinesin-13
This shifts in favour of MT depolymerisation by negative regulation of the MAP
through CDK-mediated phosphorylation as cells enter M-phase
This activity is demonstrated in vitro how
Exposing MTs to mitotic vs interphase cell extracts
to measure the ensuing MT dynamics
EXP: Add MT stabilising compound Taxol
RESULT: Prevents bipolarity
centrosomes fail to separate leading instead to the cell assembling a ‘mono-polar’ spindle
SUGGESTS: shows the critical role of MT dynamics and turnover throughout this pathway
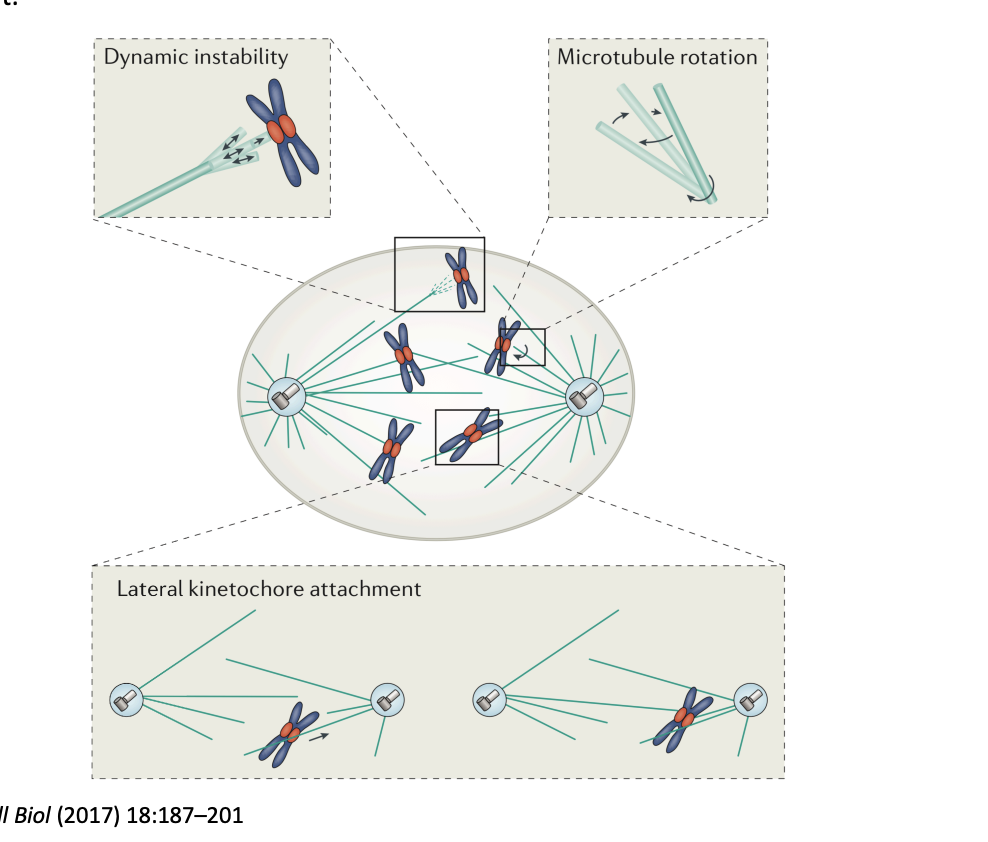
What does dynamic instability allow MTs to do
probe the 3D space of the cell
How is this done?
Search and capture
dynamic MT is nucleated by the centrosome
contacts kinetochore
captured
begins by lateral kinetochore- MT attachments
then are converted to end-on attachments
or reorient a kinetochore to favour an end-on attachment
dynamics suppressed
Why is it suggested that additional mechansims are at play?
Mathematical modelling
shows that unbiased ‘search and capture’ of kinetochores would significantly exceed the duration of spindle assembly observed in cells
THEREFORE: must be additional mechanism at play
What happens in particular?
density of MT ends generated by centrosomes (a critical determinant for efficient search and capture)
decreases with increasing distance from the poles
What do centrosome-independent pwathway do
ultimately promote MT end density near kinetochores
form sufficient MTs to permit the assembly of a functional bipolar spindles
even in the absence of centrosomes
both in vitro and in cells
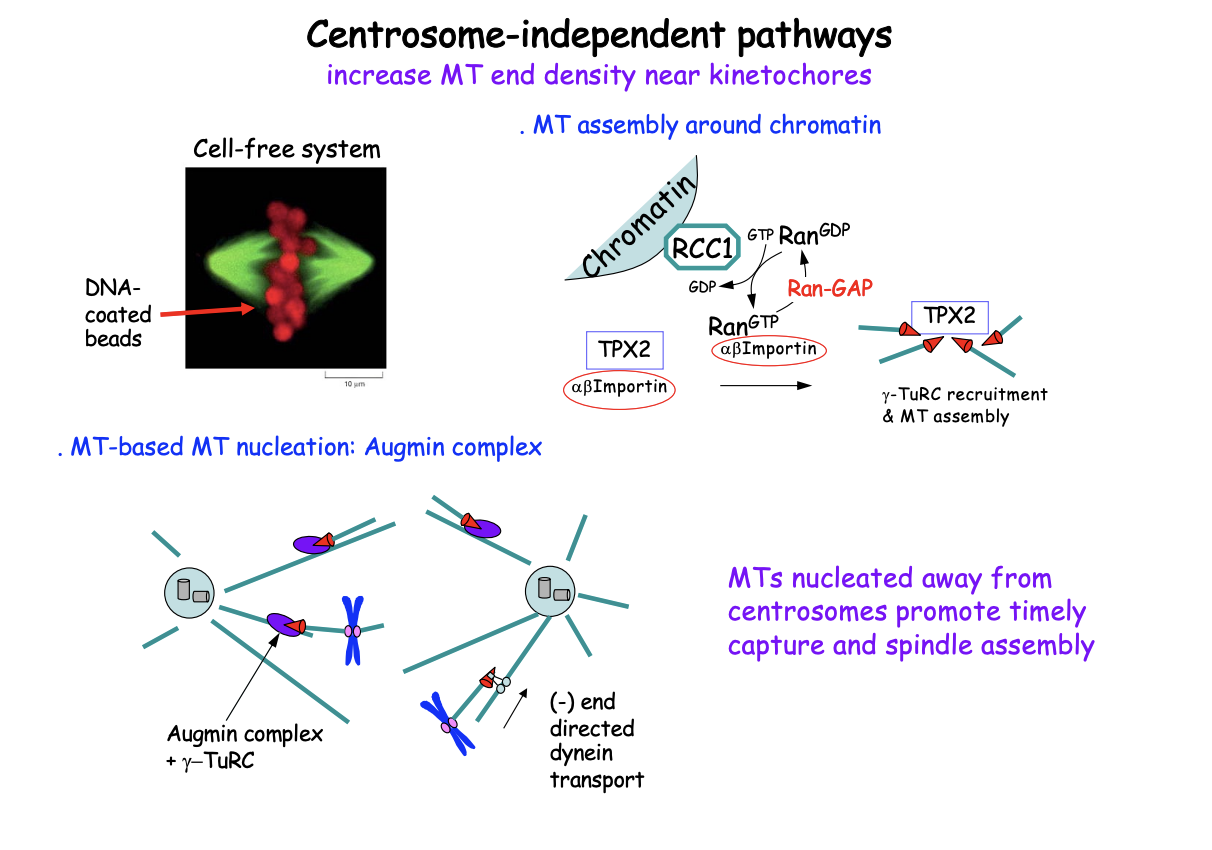
Centrosome-independent pathways: what demonstrated by
Cell-free systems
show how spindles assembly
involving chromatin and self-organisation drivenby motors
But can also be demonstrated in cell too
Bipolar spindle can assemble without centrosomes: in cell-free system
Bipolar spindle assembly is triggered by DNA-coated beads
convert into chromatin upon addition to xenopus egg extract in absence of centrosomes
What DNA can be used
even works with bacterial DNA!
What is required for spindle self-assembly
Motor proteins
How was the role of motor proteins probed
Cell-free systems
by determining the impact of specific inhibitors or specific antibody-based depletion
Findings:
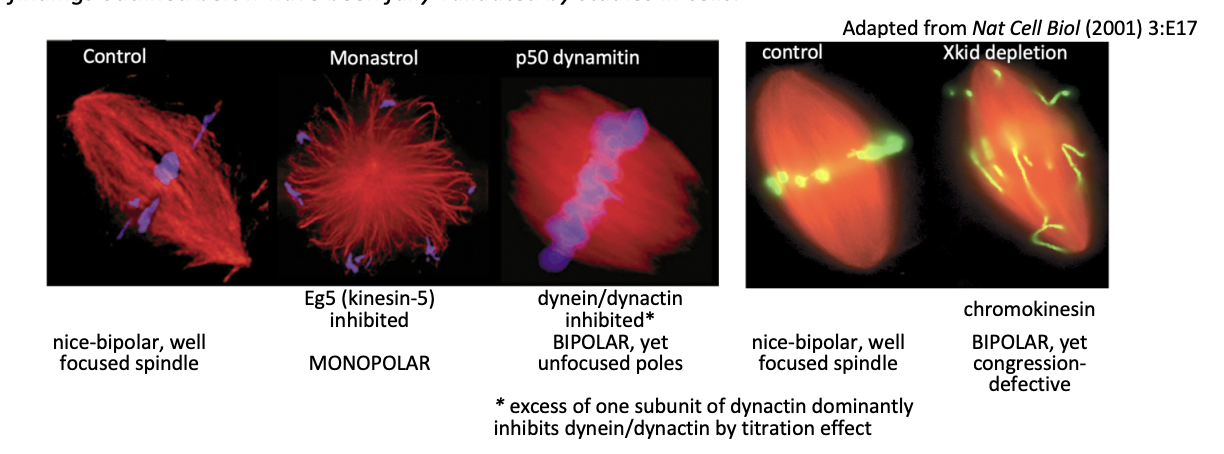
What Motors are used and what are they used for to help self organise MTs into bipolar arrays
Chromokinesins (Kinesin-4 and 10) for MT nucleated near chromatin
Bind chromatin as cargo
+ end directed
push parallel MT (-) ends away on each side on the chromatin mass
resolved the initial array around chromatin into 2 half spindles
Kinesin 5
bipolar, tetrameric + end-directed
crosslinks and slides antiparallel MTs past each other
organising the two halves into bipolar spindle
supports the MT overlap at the mid-zone
Dynein
Multimeric(-) end directed
moves along parallel MTs
focuses the poles by bringing (-) ends close together
note: picture→ black arrows indicate the direction of movement of an MT
other arrows mark the direction of motor movement
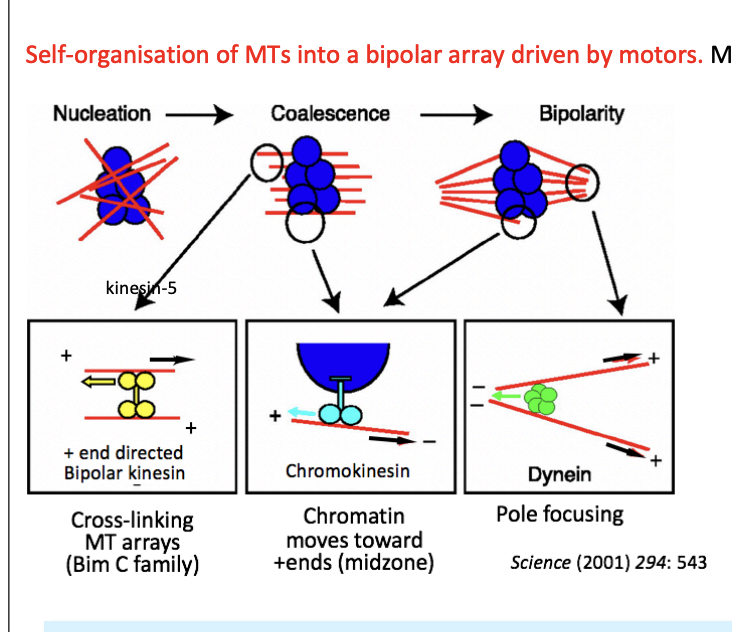
Centrosome-independent spindle assembly relies on what
alternative pathways for MT nucleation
the key molecular players and sites have been identified
They also require gamma-TURCs for their function
Molecular basis for centrosome-independent pathways: three pathways
Chromatin-dependent nucleation of MTs
MT-dependent nucleation
MT nucleation near kinetochores
Chromatin-dependent nucleation of MTs
RanGEF RCC1 binds to chromatin while the Ran GAP is in the cytoplasm
Creates a gradient of Ran GTP near chromatin
Ran GTP causes the release of TPX2 from importins (see michaelmas stuff)
Free TPX2 promotes MT nucleation (via- gamm-TuRCs) and stabilty
Favours polymerisation around chromatin
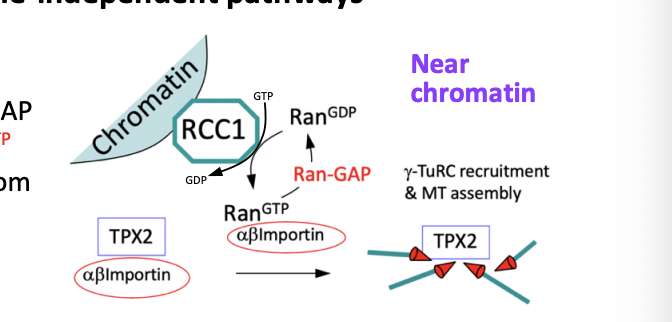
MT-dependent nucleation
Octameric Augmin Complex also recruits gamma-TuRCs to the side of a pre-existing MT
nucleating a new MT (MT branching)
This contributes to MT amplification
new MTs are transported along pre-existing MTs to join the spindle
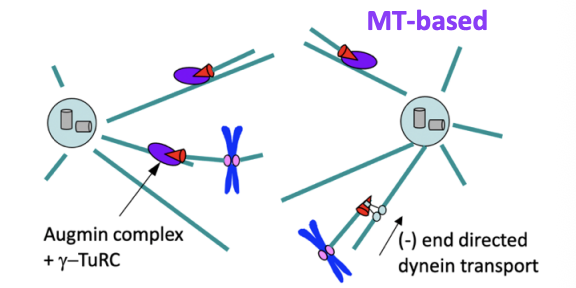
MT nucleation near kinetochores
may be another mechanism for initiating kinetochore capture
through non-centrosomal MTs that join the spindle ensemble via dynein-mediated transport or interactions with other cross linkers
Bipolarity at metaphase: overall what sets the length of the metaphase spindle (pole-pole distance)
balance of pulling and pushing forces
leads to dynamic alignment of chromosomes held under tension at the metaphase plate
Balance of pushing and pulling: kinesins and dyneins
Overlapping ‘interpolar’ MTs are cross linked by
+ end directed kinesin 5
- end directed kinesin-14
antagonistic forces determine distance between the poles
dynein/dynactin +NuMA complexes
crosslink MTs and focus the poles by (-) end directed movement
Forces from cell cortex to pull the poles apart via astral MTs
Dynein/dynactin
- end directed
generates forces from cell
Draw chromosomes away from the spindle poles
Chromosome associated + end directed kinesin-10
interactions between chromosome arms and spindle
This force disappears in Anaphase A
Due to:
proteolysis
relocation of the chromokinesin
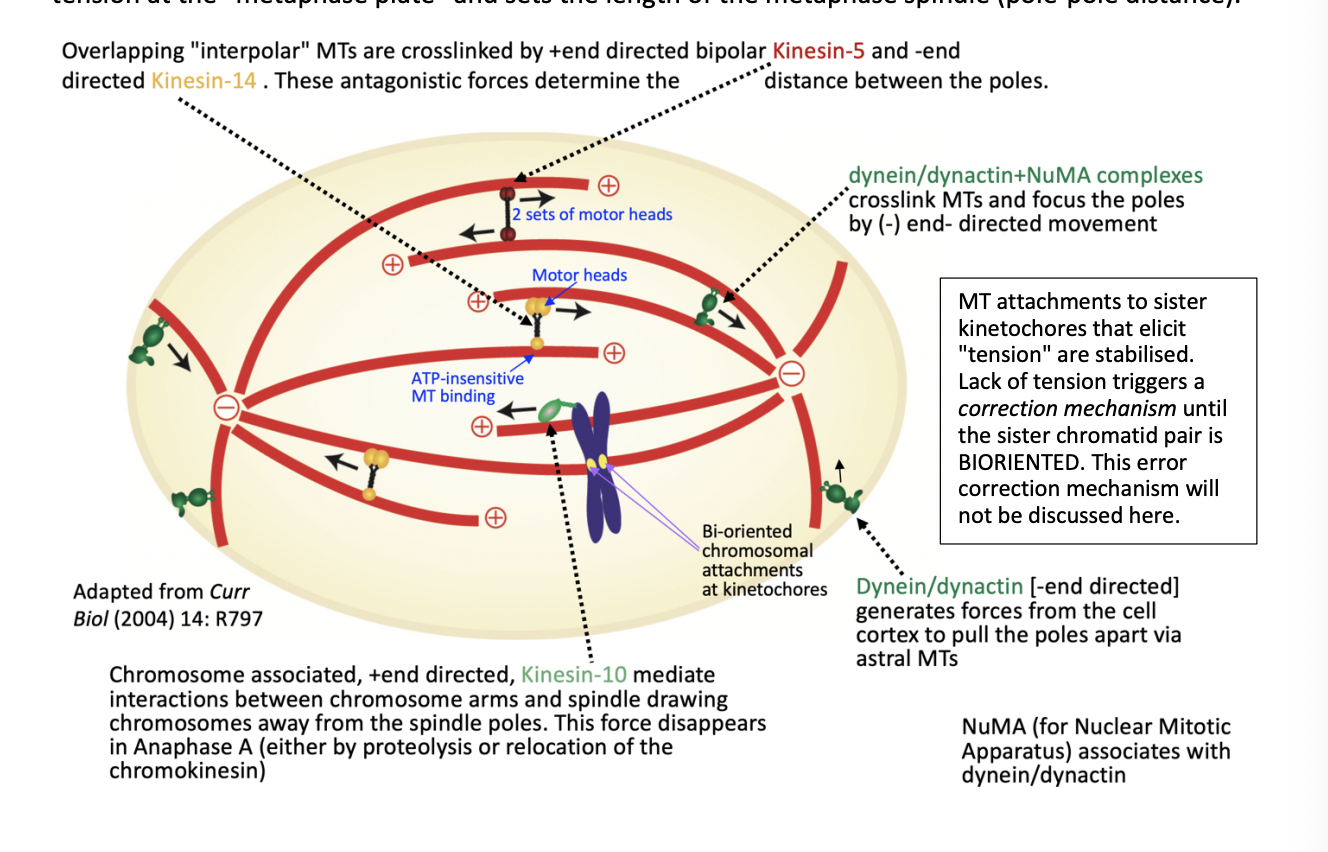
What are NuMA
Nuclear mitotic Apparatus
associates with dynein/dynactin
Error correction mechansism
MT attachments to sister kinetochores that elicit tension are stabilised
lack of tension triggers a correction mechanism until the sister chromatid pair is BIORIENTED
Dynamic MTs at the metaphase spindle: even though MTs are stabilised in a spindle, the MTs are actaully…
Under continuous treadmilling
POLEWARD FLUX:
addition at the (+) ends (near spindle equator
is balanced by
Tubulin loss at (-) ends near the centrosomes
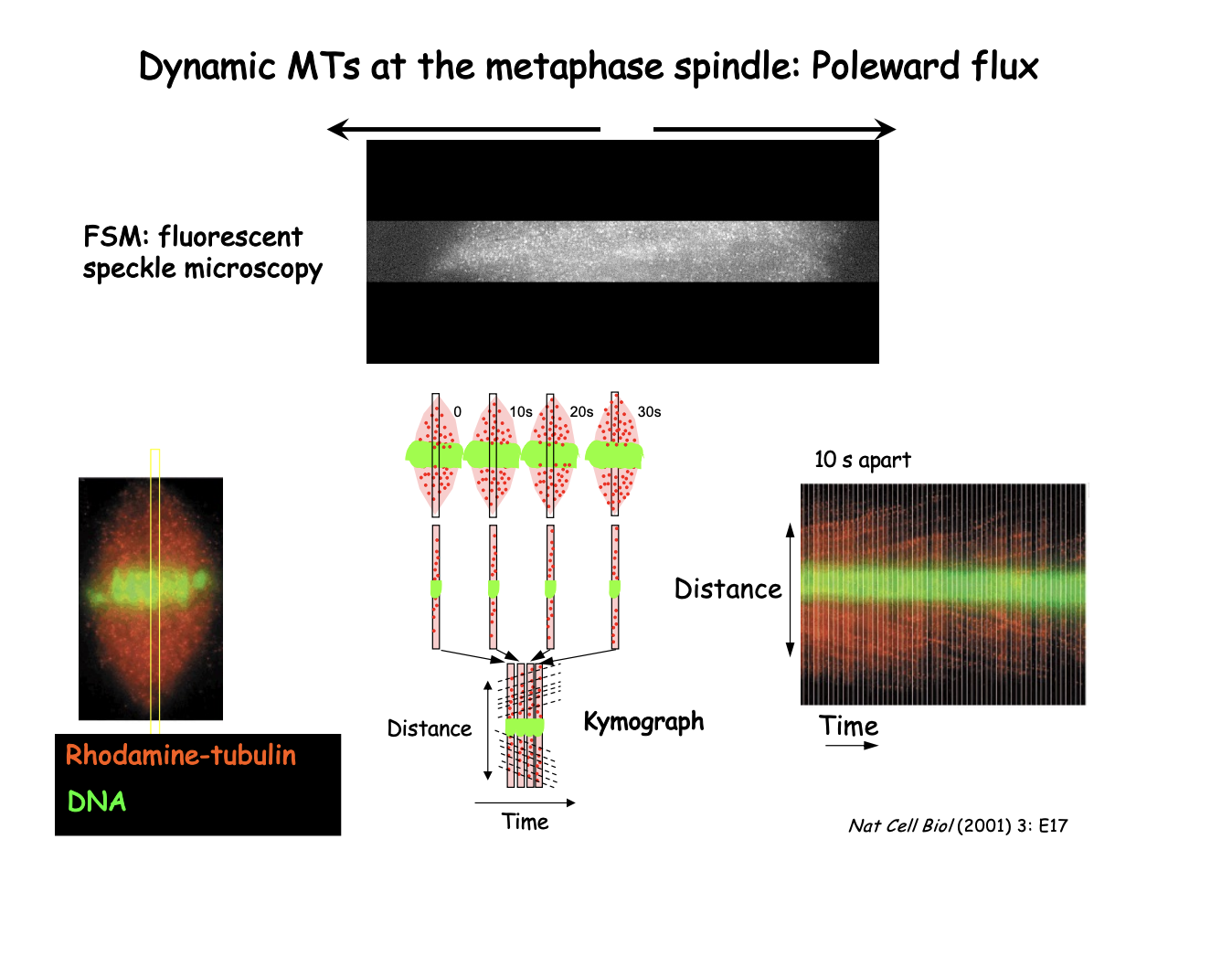
Evidence for poleward flux using what
Flouresencet Speckles Microscopy
FSM
How FSM shows Poleward flux: Xenopus extract spindle
Low density of Rhodamine-labelled tubulin
generates speckles that can be followed as fiduciary marks
First (left)
Whole spindle pictures
Kymograph (Right)
series of strips from the centre of spindle taken at 10s intervals
displayed side by side
diagonal streaks represent individual speckles moving towards the pole as a result of flux
Slope of the line→ reflects flux veolcity
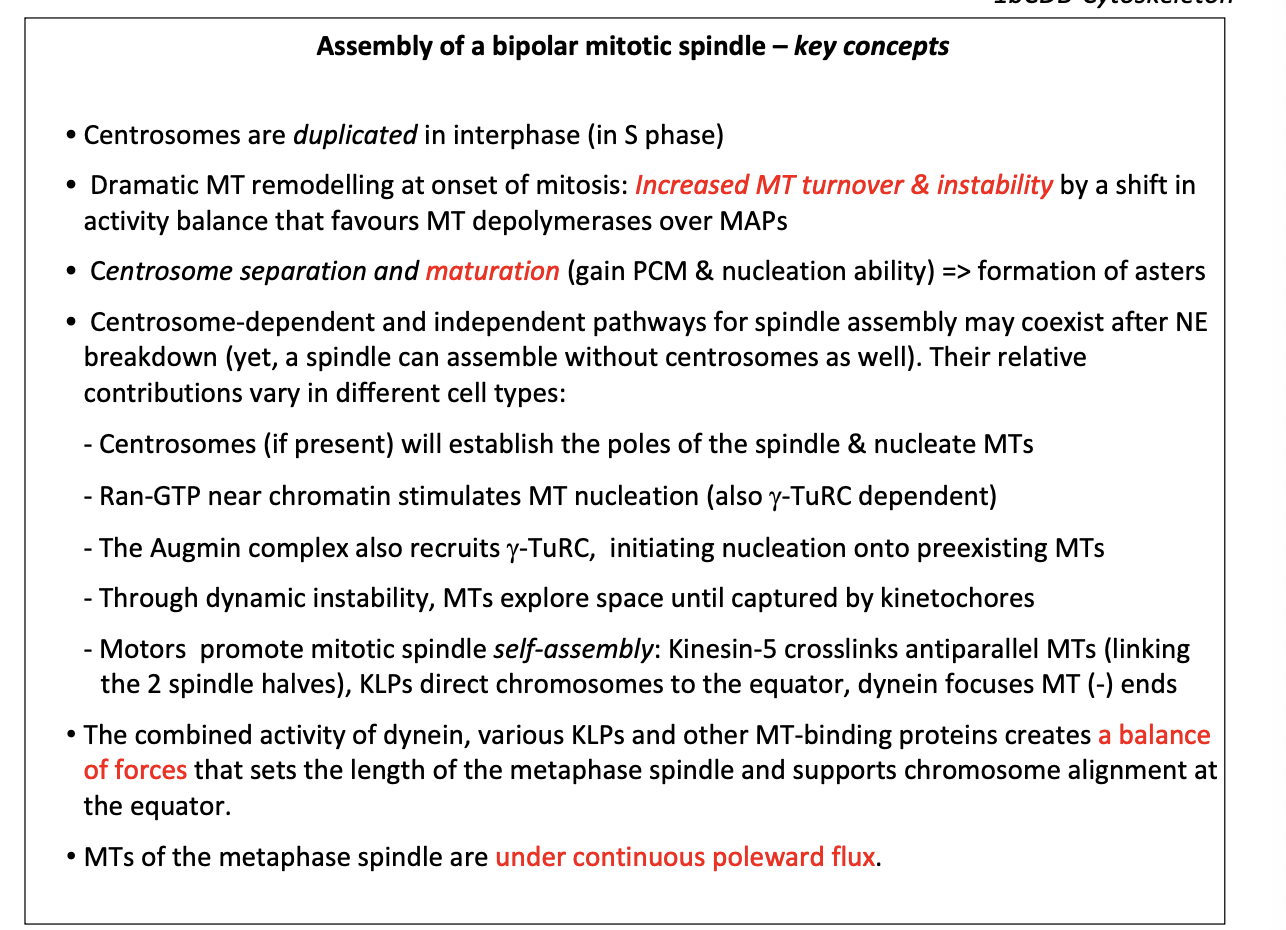
Assembly of bipolar mitotic spindle key concepts

Anaphase onset is under surveillance by
a checkpoint→ Spindle Assembly Checkpoint
see 1a BoC notes
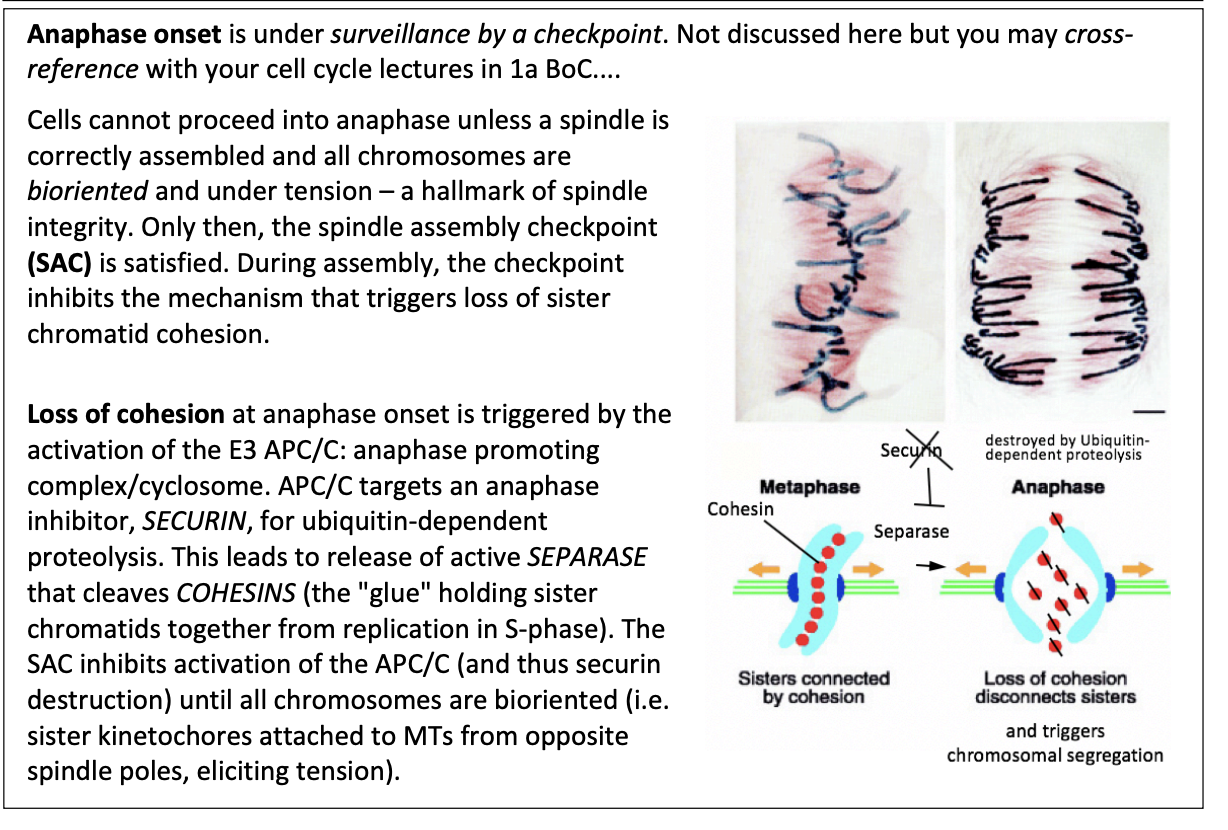
Cell cannot proceed into anaphase unless…
spindle is correctly assembled
all chromosomes are bioriented under tension
hallmark of spindle integrity
If this is done then…
spindle assembly checkpoint (SAC) is satisfied
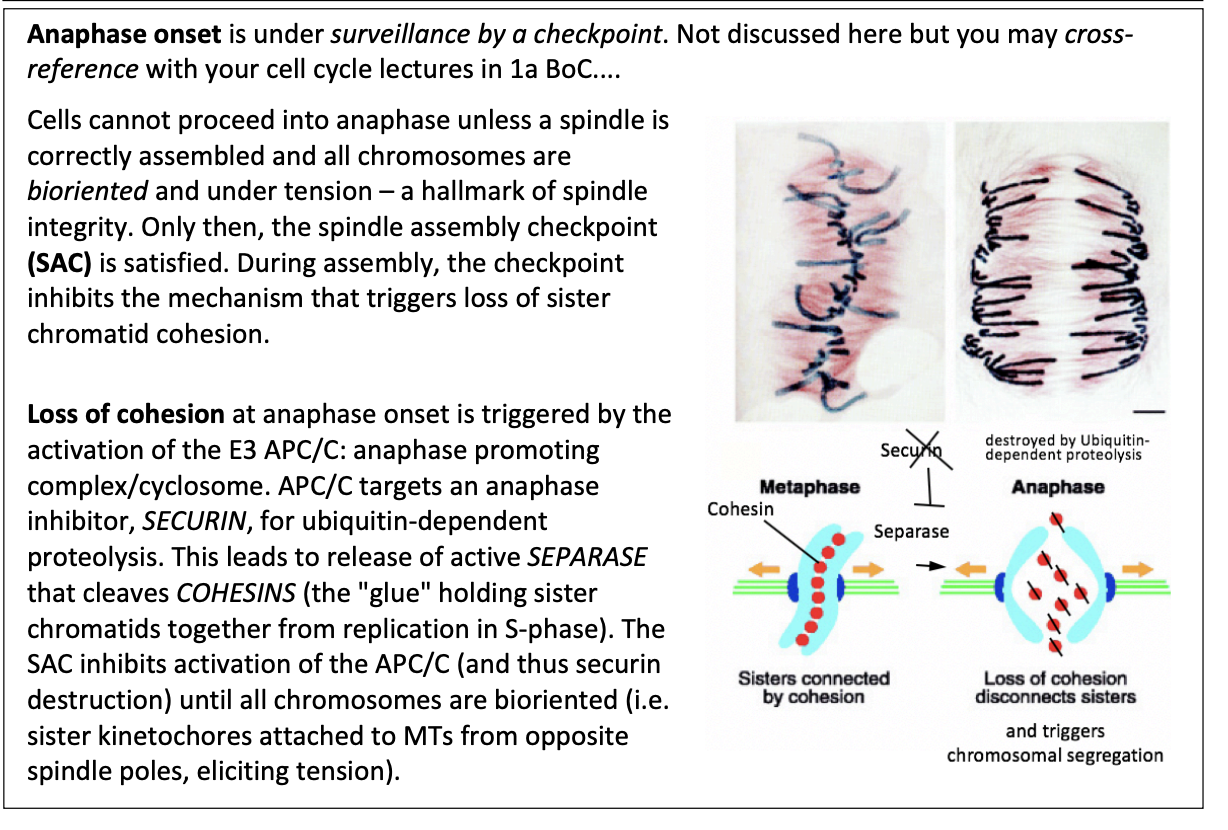
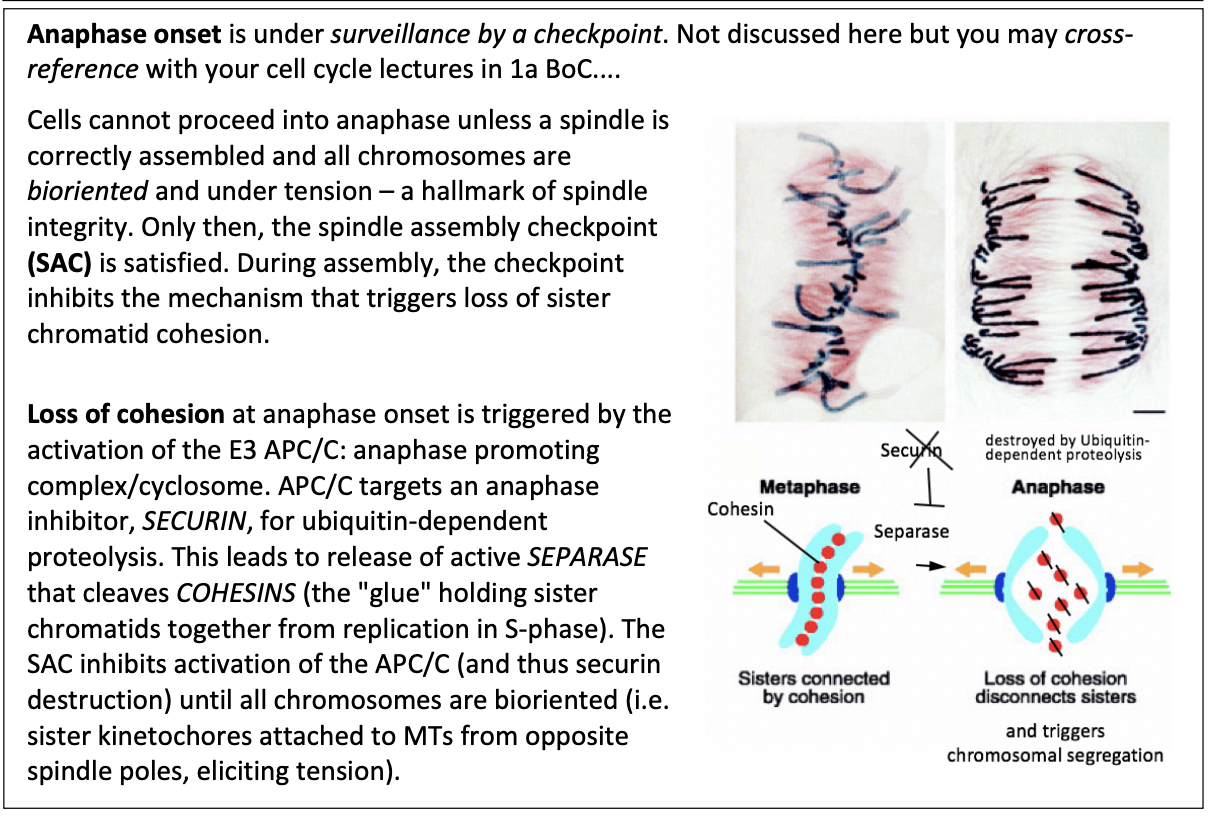
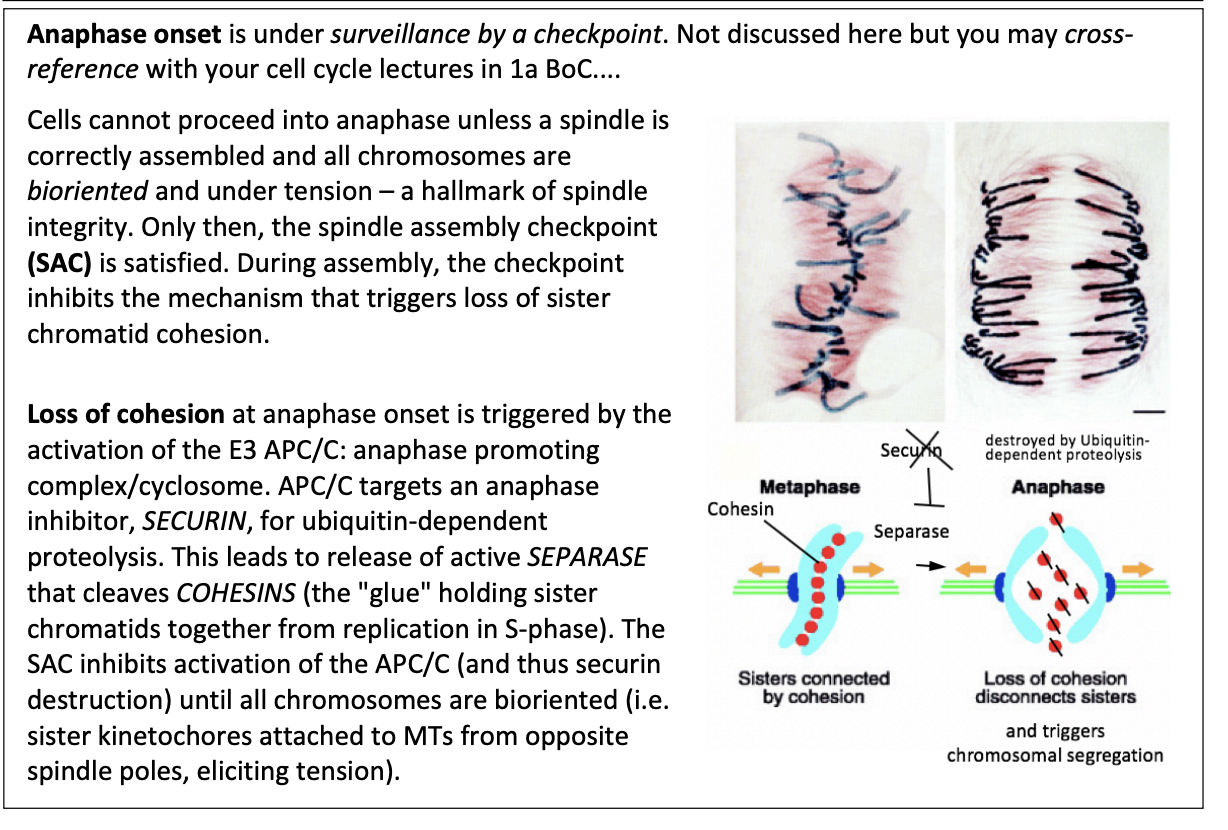
During assembly, the checkpoint inhibbits…
the mechanism that triggers loss of sister chromatid cohesion
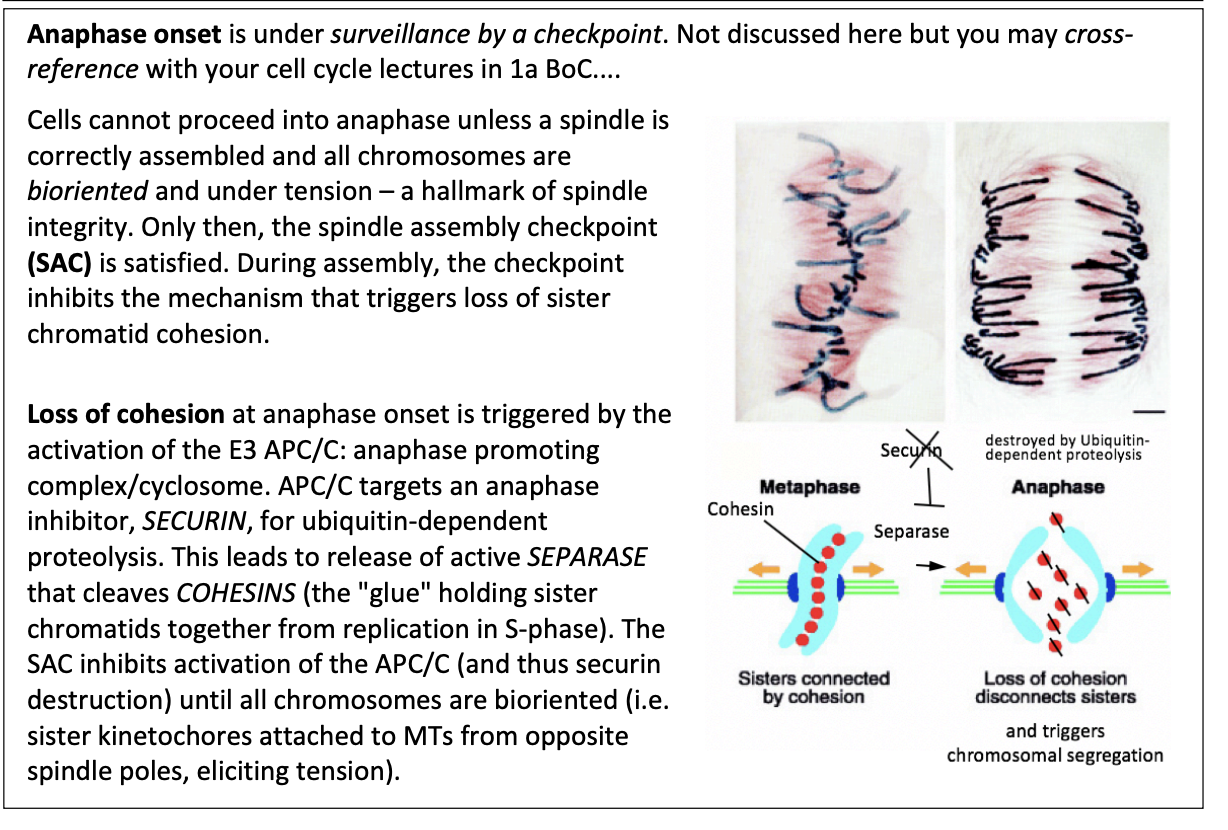
This loss of cohesion is triggered by what
Activation of E3 APC/C
anaphase promoting complex/cyclosome
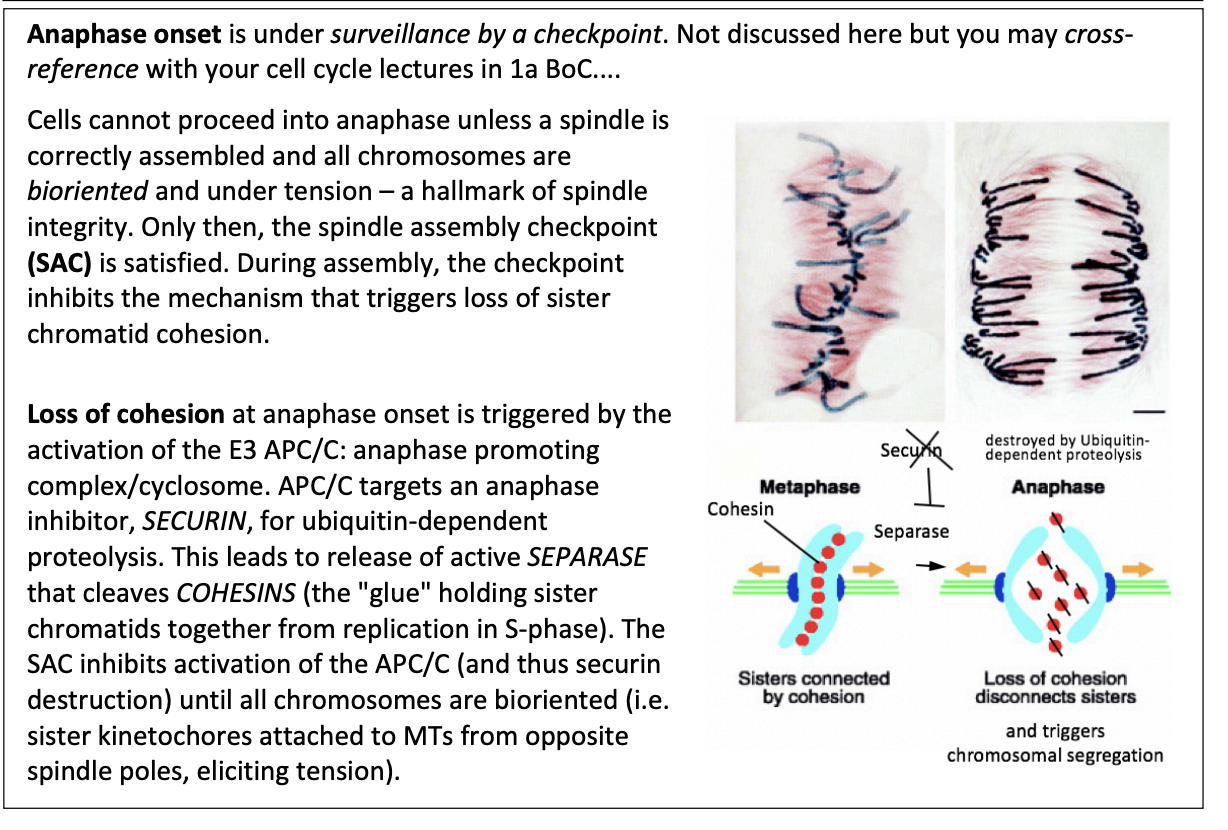
How does this work?
APC/C targets an anaphase inhibitor SECURIN for ubiquitin-dependent proteolysis
leads to release of active SEPARASE
cleaves COHESINS (the glue holding sister chromatids together from repication in S phase)
What does SAC do then?
Inhibits activation of APC/C
thus securin destruction inhibited
until all chromosomes are bioriented
sister kinetochores attached to MTs from oppsoite spindle poles
eliciting tension
Anaphase: What is anaphase A
Initial period of anaphase in which chromosomes move toward the respective poles along a spindle of constant length:
kMTs shorten
kinetochore acting like a ‘pacman’ at the + end
while MT poleward flux continues
result: chromosomes move toward the poles
while pole-to-pole distance remains constant
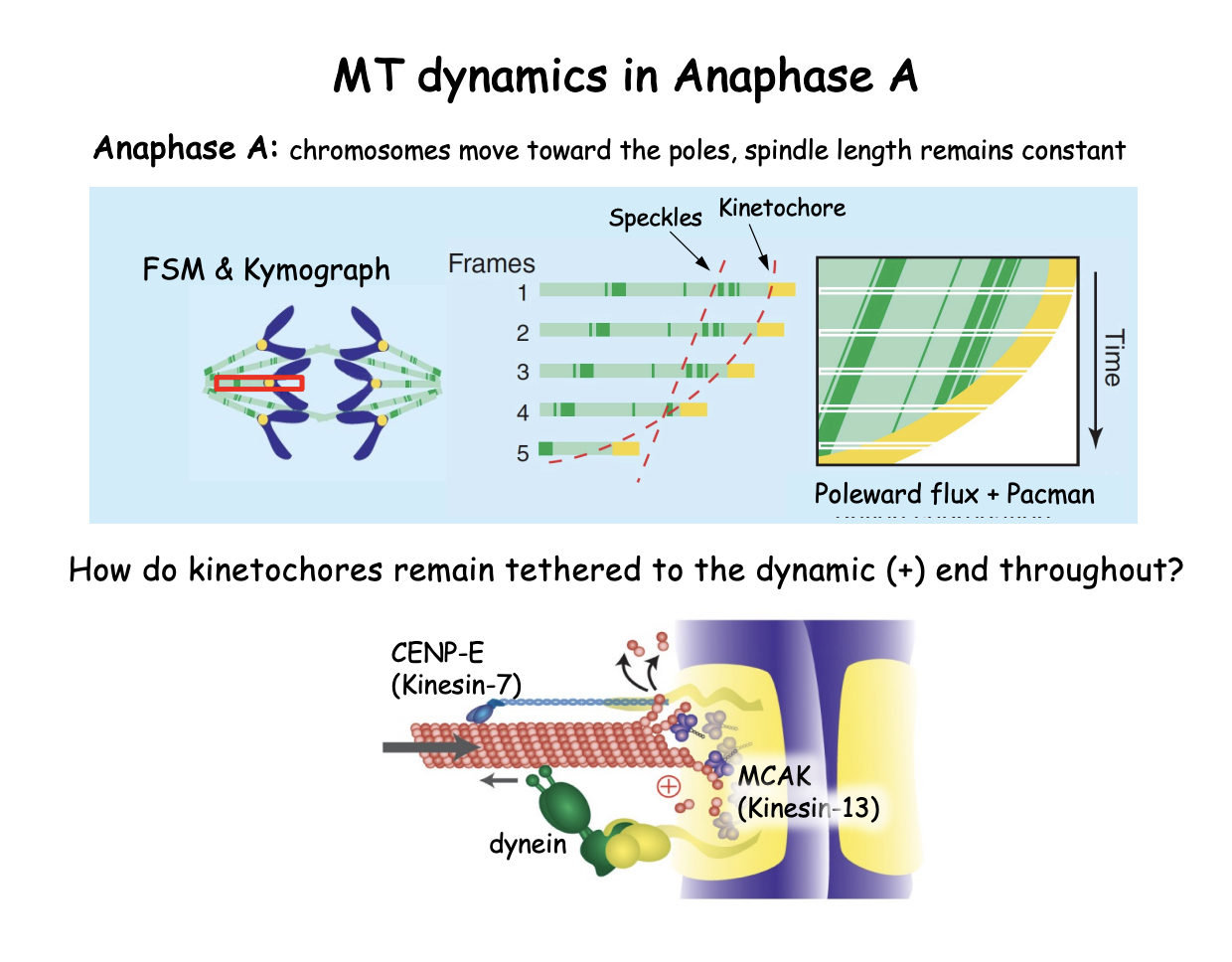
FSM followingg 2 fluorescnet items reveals the underlying MT dynamics of this:
Lines of constant slope→ reflecting the flux of MT subunits
kMTs also shorten over time accelerating the kinetochore mark that overtakes MT speckles
SHOWS: + end ‘pacman’ style depolaymerisation of kMTs overlapping with poleward flux
What has also been proposed about how kMTs move towards poles
chromosomes attachmed to short k-MTs may also move toward the poles by synein-drievn transport
short kMT as cargo
Evidence of this: Laser ablation
Exp: Laswer ablation to cut spindle MTs
RESULT: severed MTs linnked to chromosomes are delivered to the pole by dynein walking along an intact MT
Remaining questions to answer
How does MT depolymerization at the kinetochore end (+end) drive chromosome movement?
How are kinetochores retained despite MT shrinnkage?
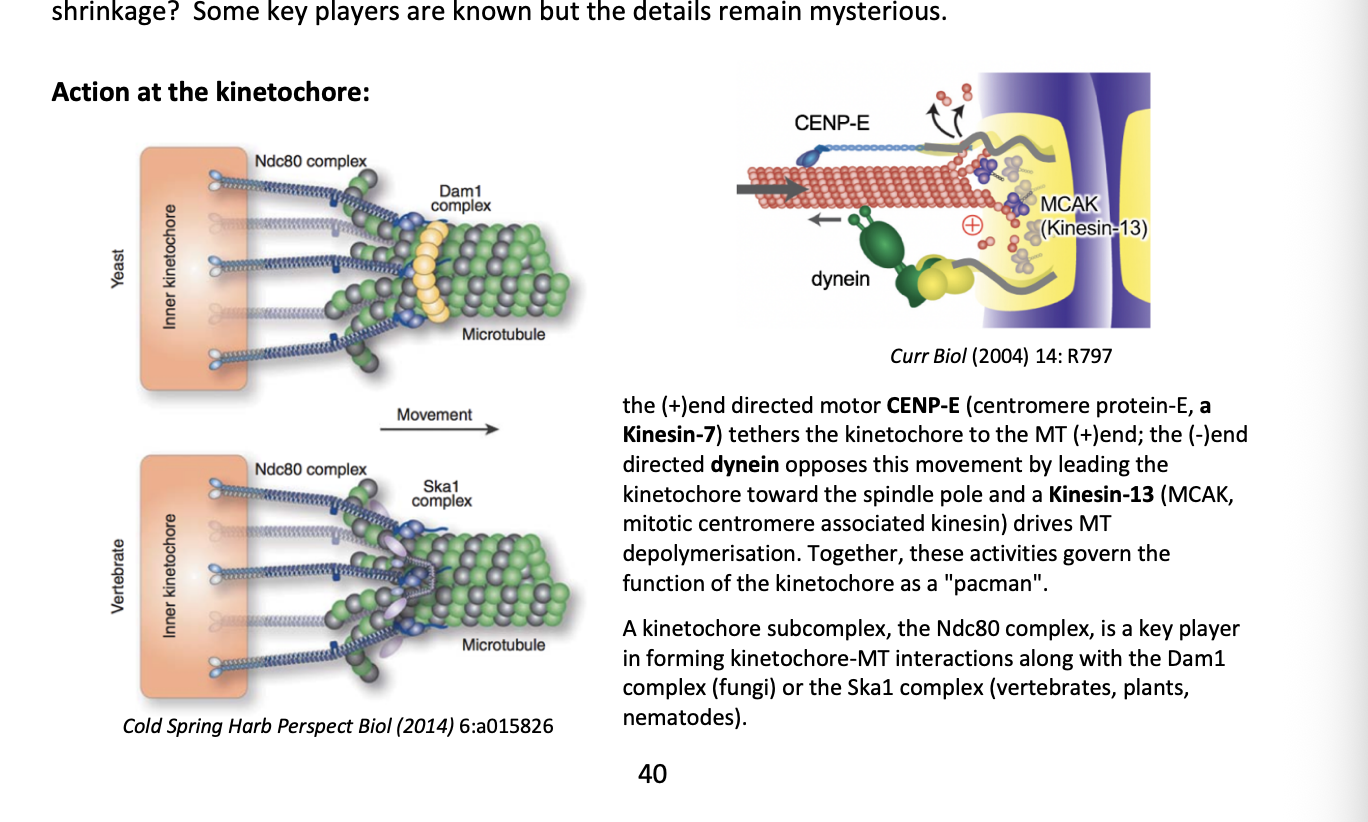
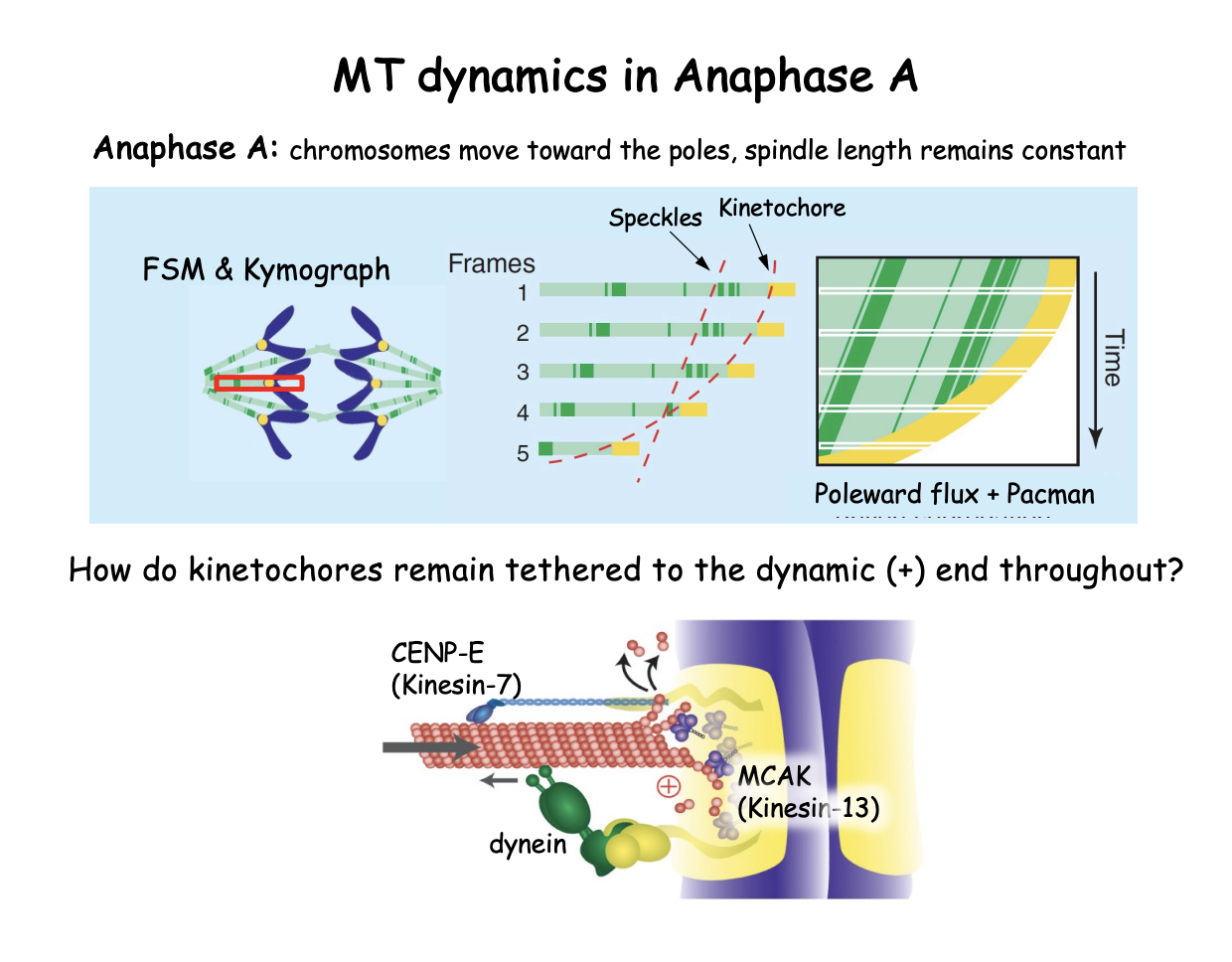
What govens the function of the kinetochores as a ‘pacman’
+ end directed motor CENP-E
(centromere protein E, a Kinesin-7)
tethers the kinetochore to the MT (+) end
- end directed dynein
opposes this movement by leading the kinetochore toward the spindle pole
Kinesin-13 (MCAK→ mitotic centromere associated kinesin)
drives MT depolymersation
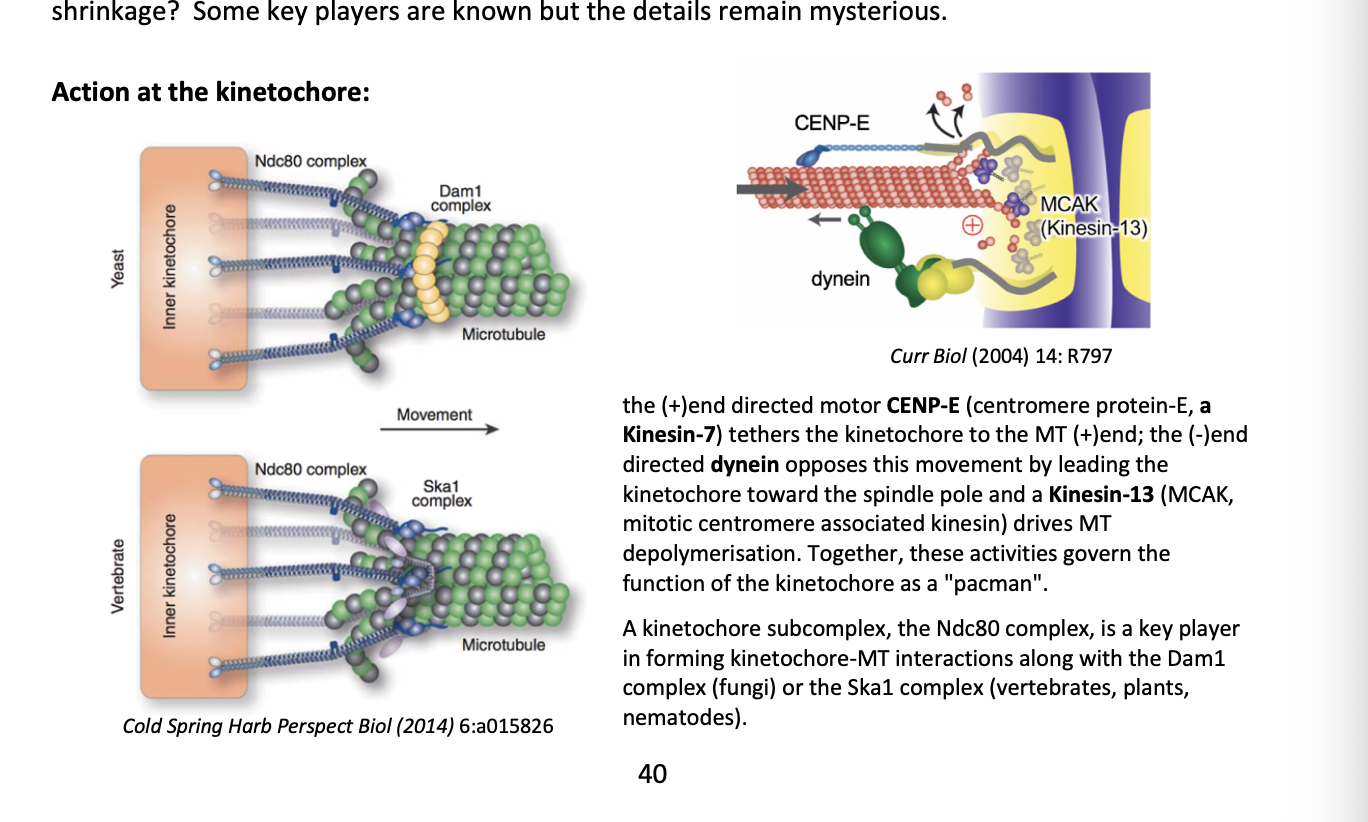
What is the Ndc80 complex
Kinetochore subcomplex
key player in forming kinetochore-MT interactions along
the Dam1 complex (fungi)
or
Ska1 complex (vertebrates, plants, nematodes)
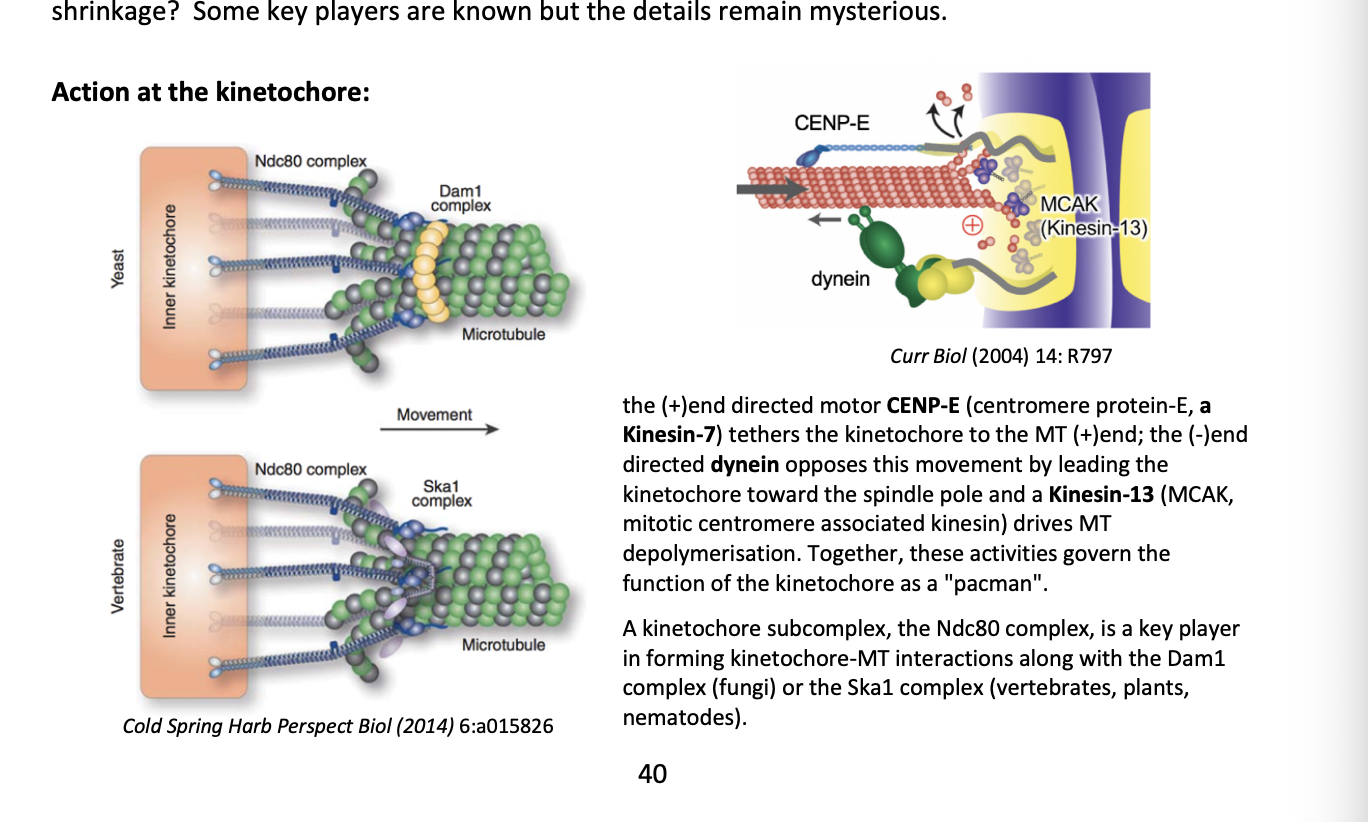
Anaphase B: what happens
spindle poles move further apart along with clustered chromosomes
pushed by bipolar (+) end directed Kinesin-5
causes the sliding of antiparallel MTs at the overlap zone
Dynein (-end directed motor)
anchored at the cell cortex helps to pull the poles apart
Integrity of the spindle midzone depends on additional MAPs
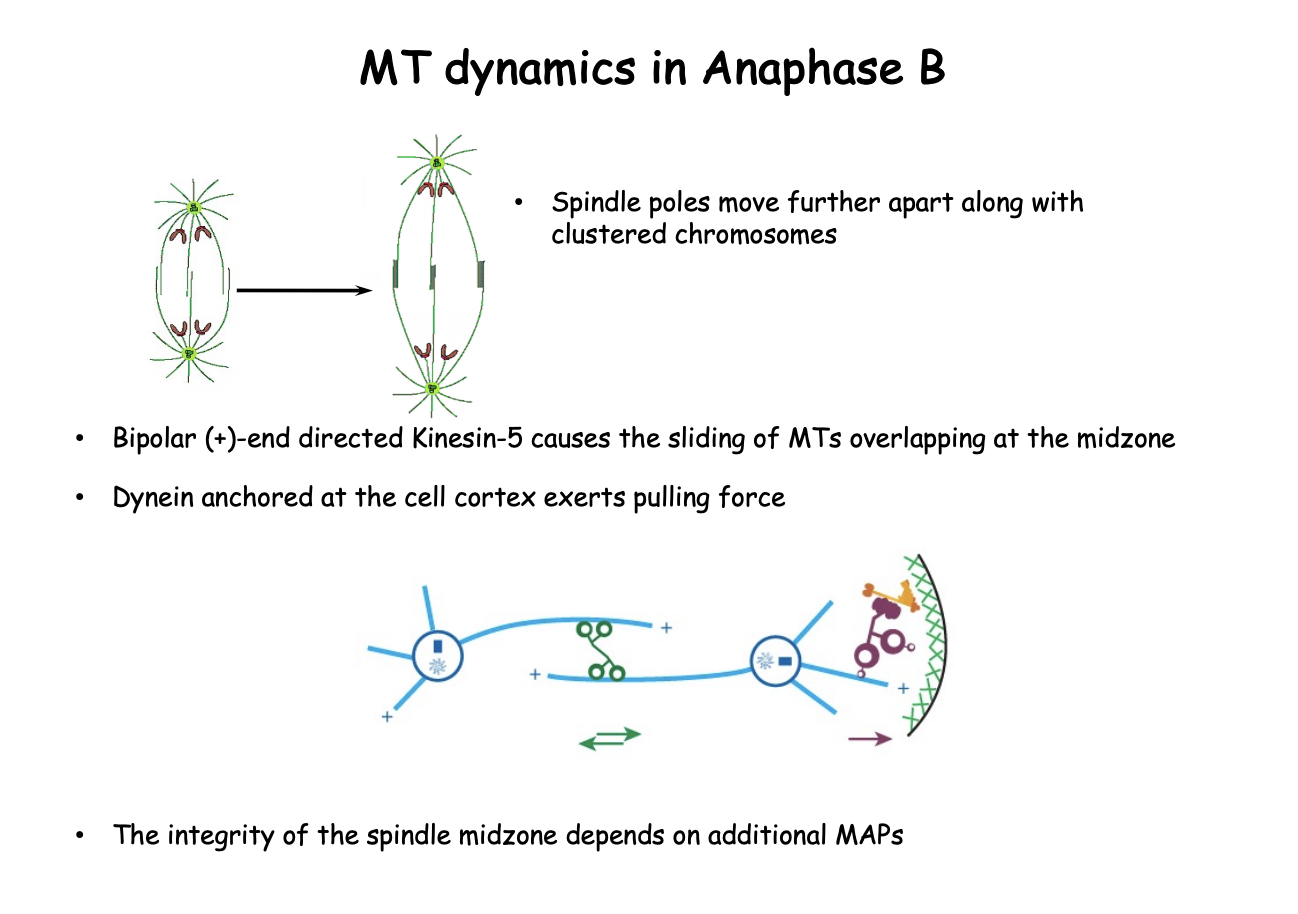
Mictrotubules during anaphase, telophase and cytokinesis
the spindle gives way to a postmitotic bridge (D)
connects the separating daughter cells (central spindle)
these remnants form a very dense structure→ midbody during cytokinesis
plays a key role in abscission
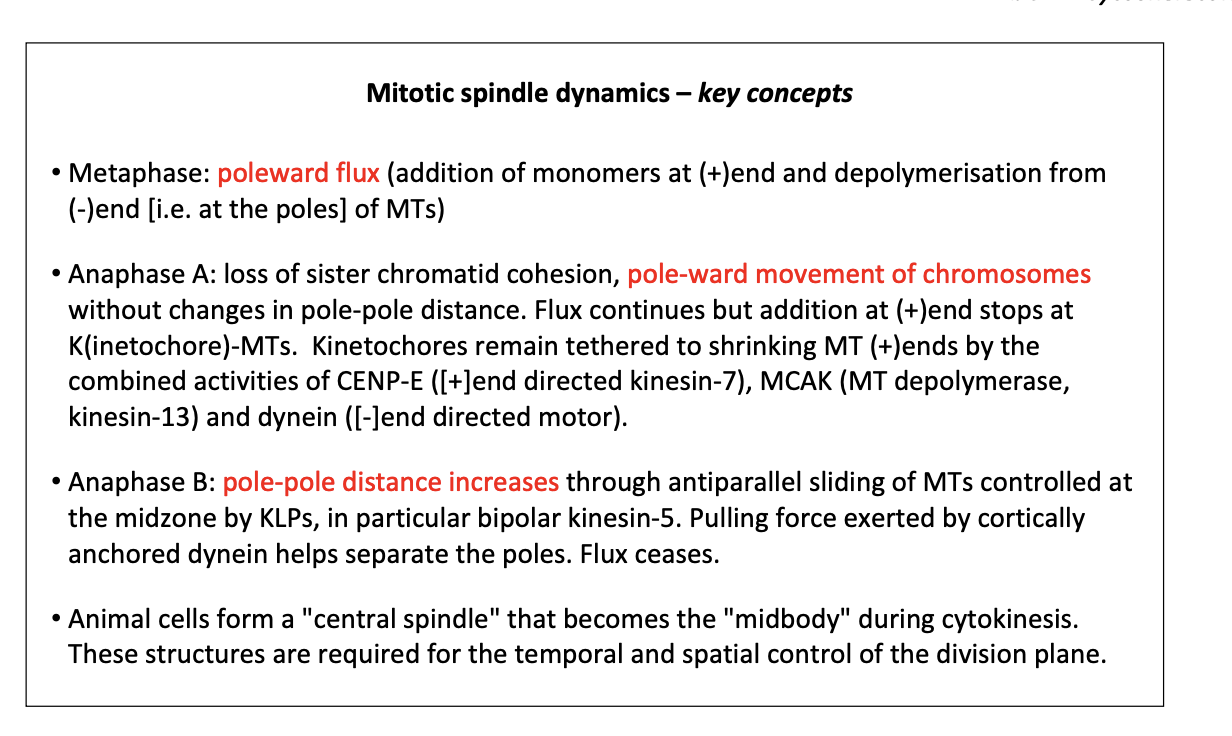
Mitotic spindle dynamics key concepts

Cytokinesis and Cell separation: why is coordination between the mitotic apparatus and division plane needed
ensure cytokinesis end stages DO NOT occur before chromosome separation has been completed
prevents unevent chromosome distribution or damage
how is cell division plane arranged in respect to spindle axis
Orthogonal
But are all cell dvisions symmetrical?
No
some are asymmetric
E.g for cell diversity generation in development
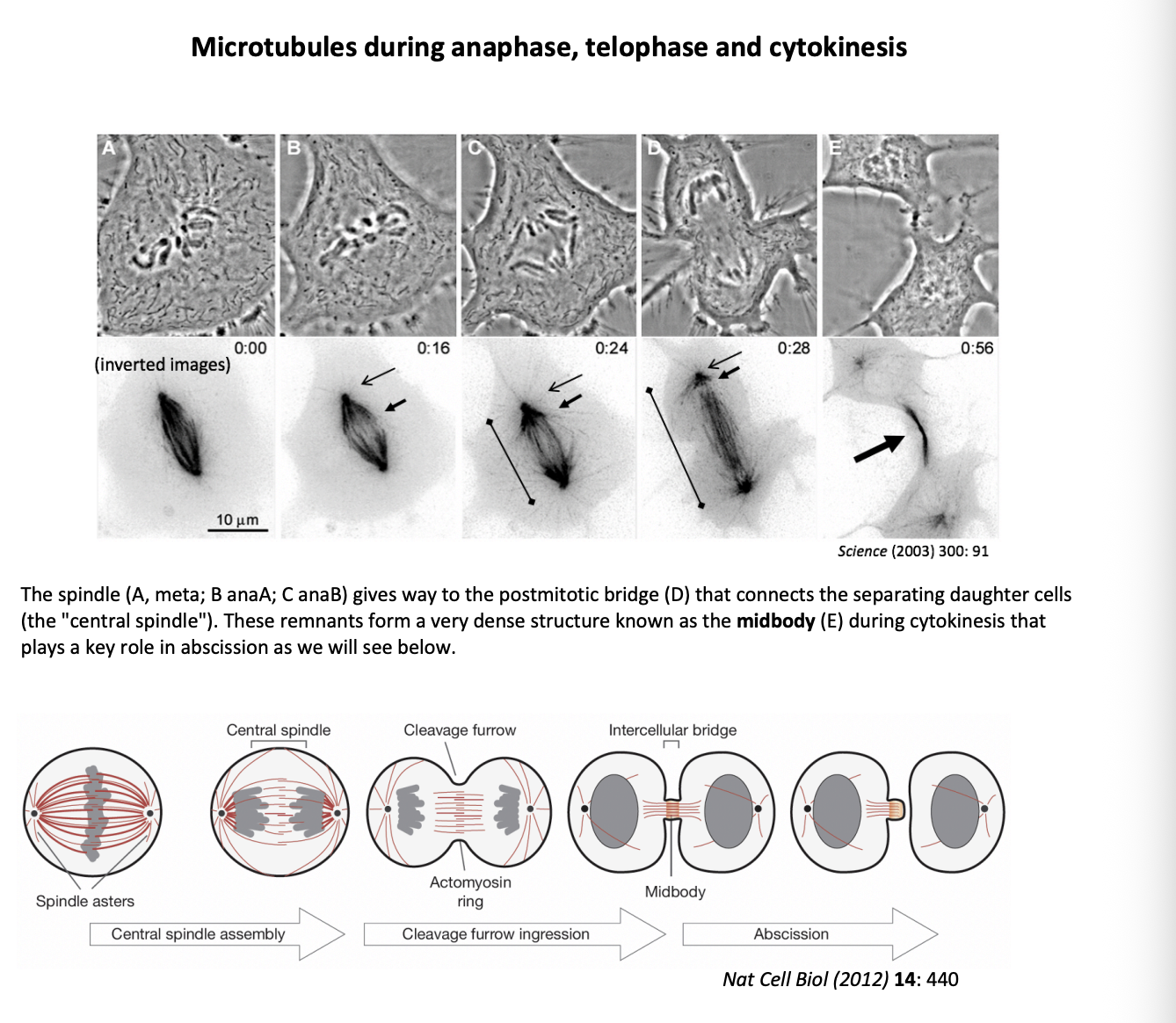
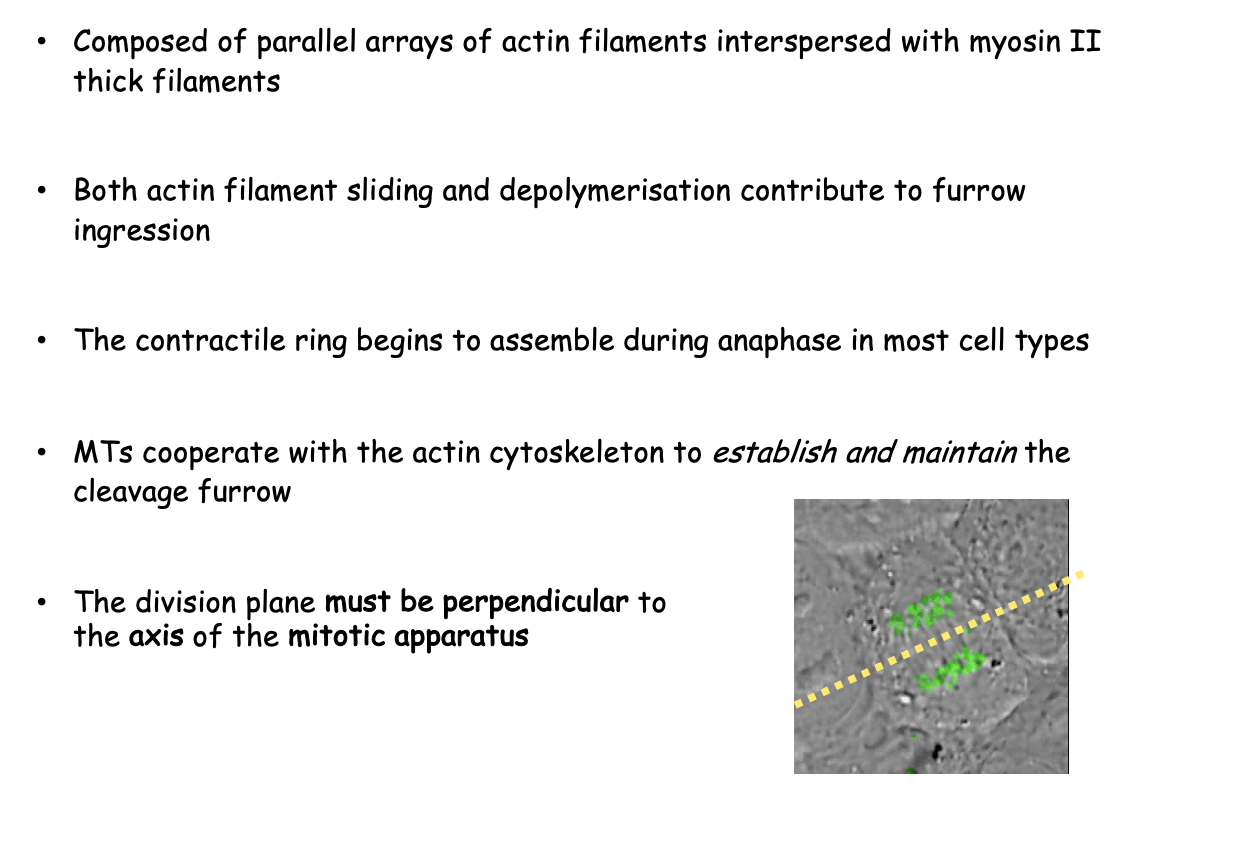
In animal cells, what is the position of the cleavage plant specified by
Mitotic apparatus
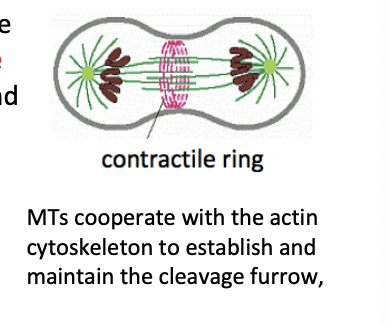
What does cytokinesis involve
formation of a furrow that encircles the cell
deformation of the plasma membrane
insertion of new membrane components
force of furrow ingression is provided by contractile ring
composed of arrays of F-actin interspersed with myosin II thick filaments
Evidence that myosin powers cytokinesis
EXP: Inject anti-myosin antibodies (into the tight cell of the 2 cell embryo)
RESULT: cytokinesis fails
Special cytokinesis events in specific examples
Early embryoninc division in Drosophila
cytokinesis is suppressed
nuclei divide in common cytoplasm
Oogenic cyst of Drosophila
Cytokinesis is incomplate
cells remain connected by cytoplasmic bridges
Budding and fission yeast
cytokinesis results from activity of an actomyosin contractile ring (similar to animals)
COMBINED with the deposition of a septum
Furrow ingression is powered by a contractile ring

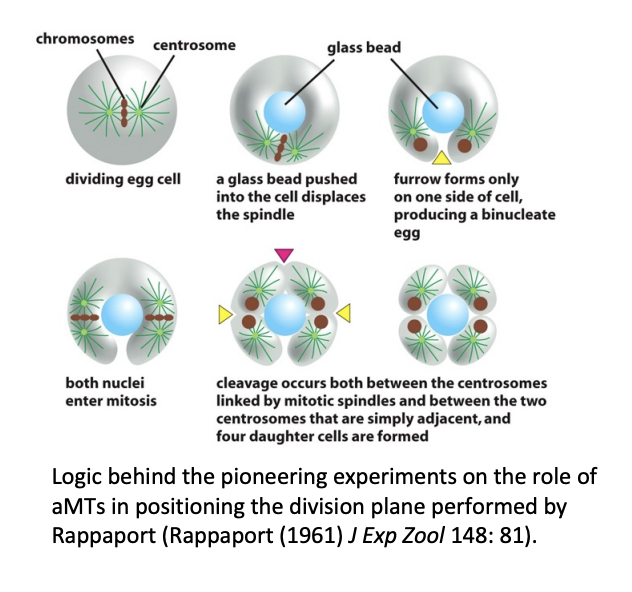
Influence of mitotic apparatus on the position of the division plane in animals: aMTs or central spindle?
aMTs may be sufficient to localise the division plane in large embryonic cells
e.g Rappaport 1961
BUT in other experiments: when glass barrier was placed to interfere with aMTs, cytokinesis could still occur