🏳️⚧️ Lecture 14: Regulation of CDK Activity and Transitions Between Stages of Cell Cycle
1/14
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
15 Terms
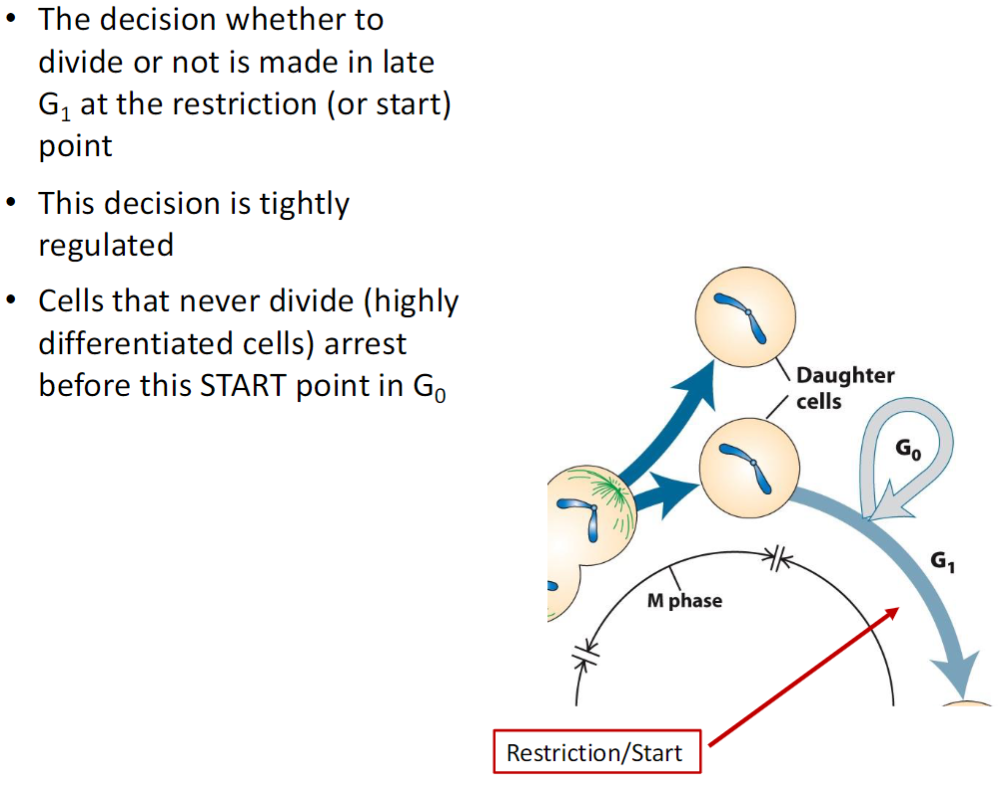
Cell Cycle – Restriction/Start Point
Decision to Divide
Occurs in late G1 at the restriction (or START) point
Determines whether the cell will continue through the cell cycle or pause
Regulation
Decision is tightly controlled by CDKs, cyclins, and other regulatory factors
G0 – Non-Dividing Cells
Cells that never divide (highly differentiated cells) arrest before the START point
Enter G0, a resting phase outside of the normal cell cycle
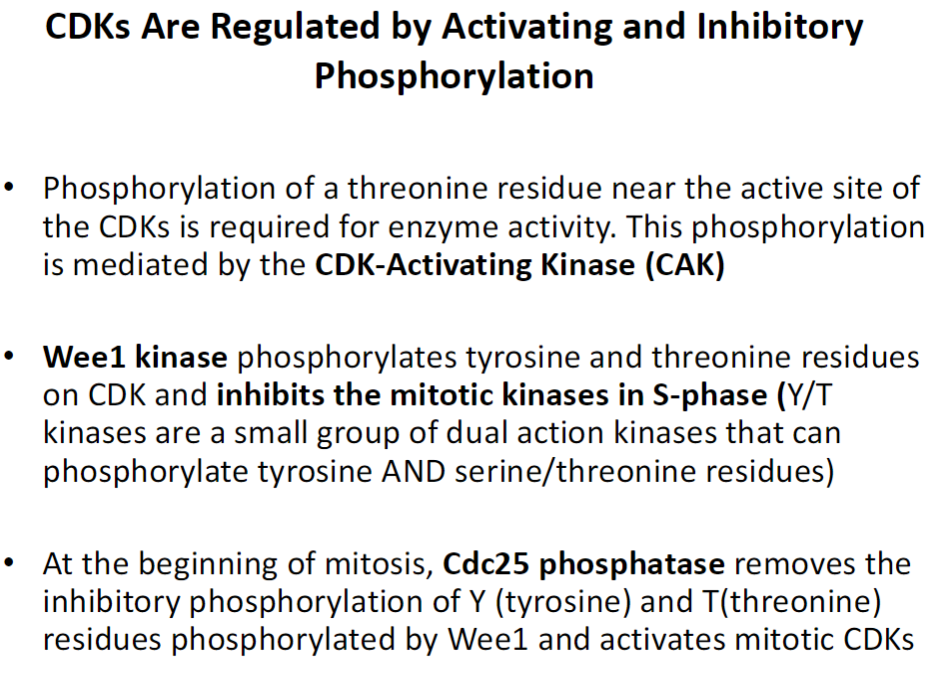
CDK Regulation – Phosphorylation
Activating Phosphorylation
Phosphorylation of a threonine (T) residue near the CDK active site is required for activity
Mediated by CDK-Activating Kinase (CAK)
Inhibitory Phosphorylation
Wee1 kinase phosphorylates tyrosine (Y) and threonine (T) residues on CDKs
Inhibits mitotic kinases during S-phase
Y/T kinases can phosphorylate both tyrosine and serine/threonine residues
Activation at Mitosis
Cdc25 phosphatase removes inhibitory Y/T phosphorylation added by Wee1
Activates mitotic CDKs to promote entry into mitosis

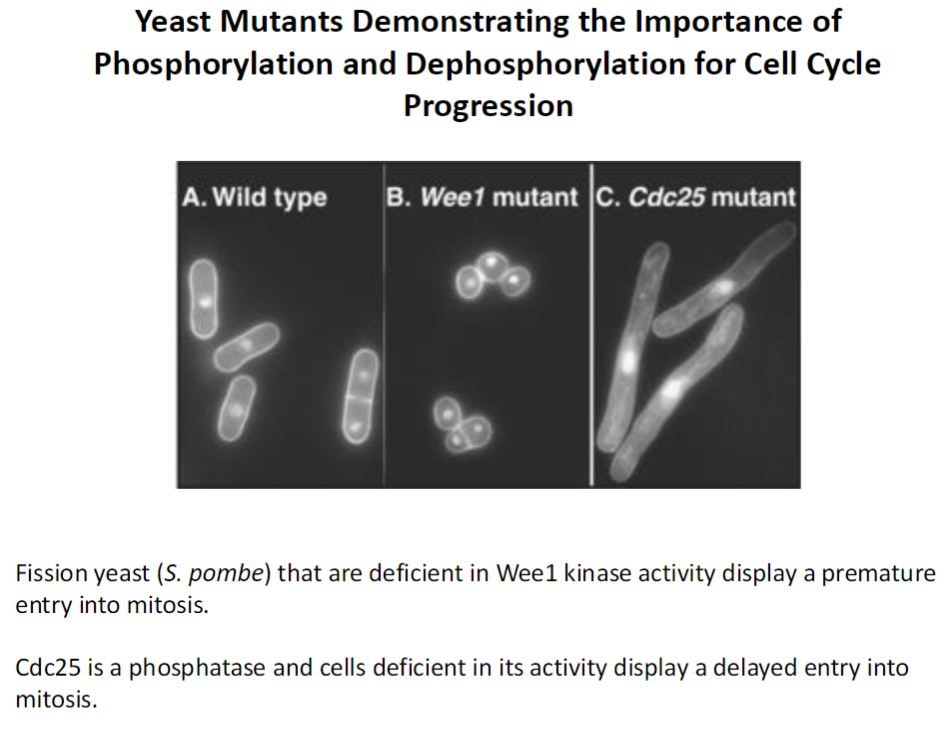
Yeast Mutants – CDK Phosphorylation
Wee1 Deficient Mutants
Fission yeast (S. pombe) lacking Wee1 kinase enter mitosis prematurely
Shows inhibitory phosphorylation by Wee1 is important to delay mitosis until the cell is ready
Cdc25 Deficient Mutants
Cells lacking Cdc25 phosphatase show delayed entry into mitosis
Shows removal of inhibitory phosphorylation is required to activate mitotic CDKs
Takeaway
Phosphorylation and dephosphorylation of CDKs are critical for proper cell cycle progression
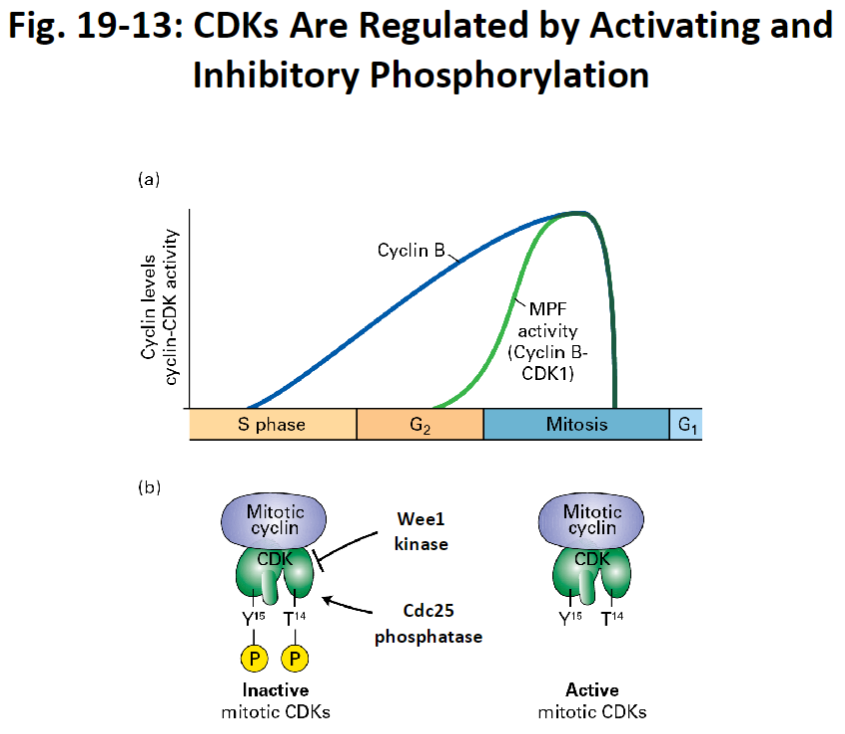
CDK Regulation – Activating and Inhibitory Phosphorylation
Wee1 Kinase
Adds inhibitory phosphate groups to CDKs
Prevents premature entry into mitosis
Cdc25 Phosphatase
Removes inhibitory phosphate groups from CDKs
Activates mitotic CDKs so the cell can enter mitosis
MPF (Maturation Promoting Factor)
Cyclin B-CDK complex
Cyclin B is made in advance but only acts during mitosis
Dephosphorylation by Cdc25 allows MPF to become active and trigger mitosis
G1–S Phase Transition *Only look at metazoans, but just know this entire thing was first discovered in yeast
Key Concept
Decision to enter S phase is tightly controlled in metazoan cells
Mechanisms were first discovered in yeast but are conserved across species
CDK-Cyclin Complexes
Drive the transition from G1 to S phase
Cyclin-CDK activity triggers DNA replication machinery
Regulation
Inhibitory proteins can bind CDKs to prevent premature S phase entry
Ubiquitin ligases degrade cyclins to reset the cycle
Activating kinases and phosphatases fine-tune CDK activity
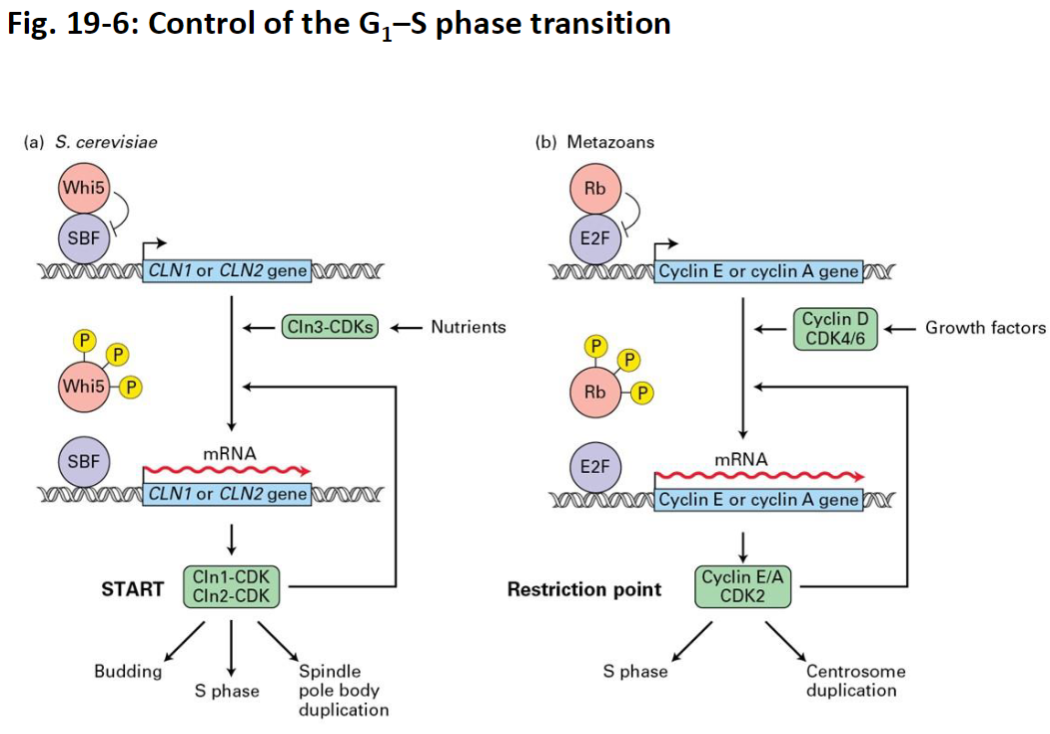
G1–S Phase Regulation – Mammals
Rb and E2F
In early G1, genes required for S phase (Cyclin E and A) are suppressed
Rb protein binds transcriptional activator E2F to block S-phase gene expression
Growth Factor Stimulation
Growth factors trigger signal transduction pathways
Stimulate expression of Cyclin D
Cyclin D/CDK Activity
Cyclin D/CDK4 and Cyclin D/CDK6 phosphorylate Rb
Phosphorylation causes Rb to dissociate from E2F
E2F Activation
Free E2F stimulates transcription of Cyclins E and A
Cyclin E-CDK2 further phosphorylates Rb
Also activates S-phase promoting factors to drive DNA replication
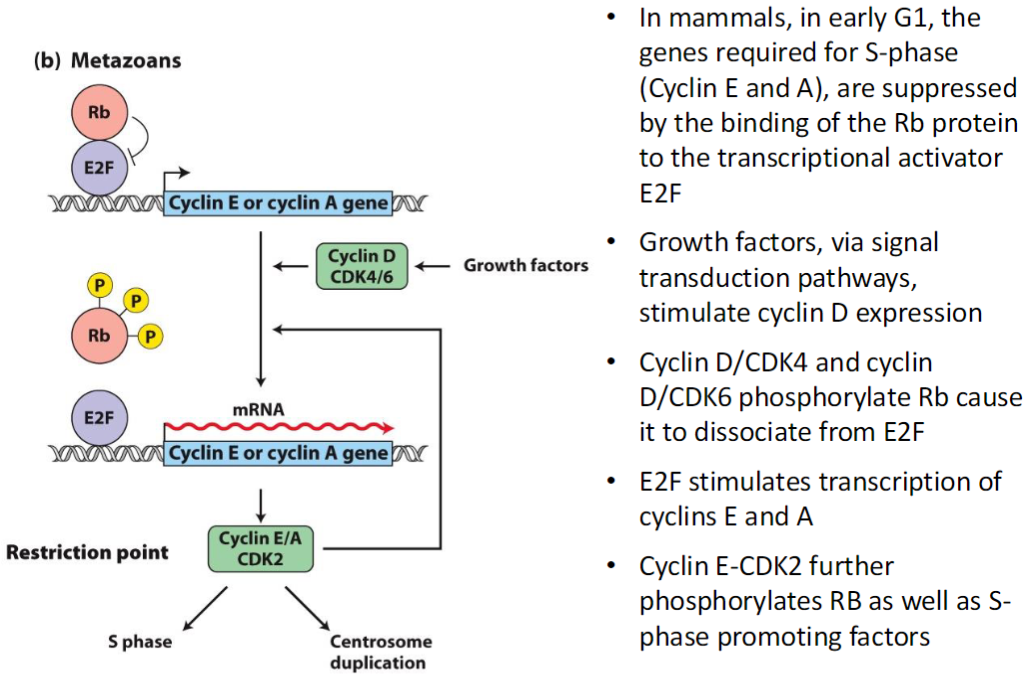
S-Phase Entry – Yeast
Sic1 Inhibition
Sic1 protein inhibits S-phase CDKs when unphosphorylated
Prevents premature entry into S phase
Phosphorylation of Sic1
Sic1 is phosphorylated at 6 sites
Multiple phosphorylations make Sic1 a very effective target
Proteolysis and Activation
Phosphorylated Sic1 is recognized by SCF ubiquitin ligase
Sic1 is degraded by the proteasome
Degradation releases S-phase CDKs
S-phase cyclins and CDKs become active
Cell can enter S phase and begin DNA replication
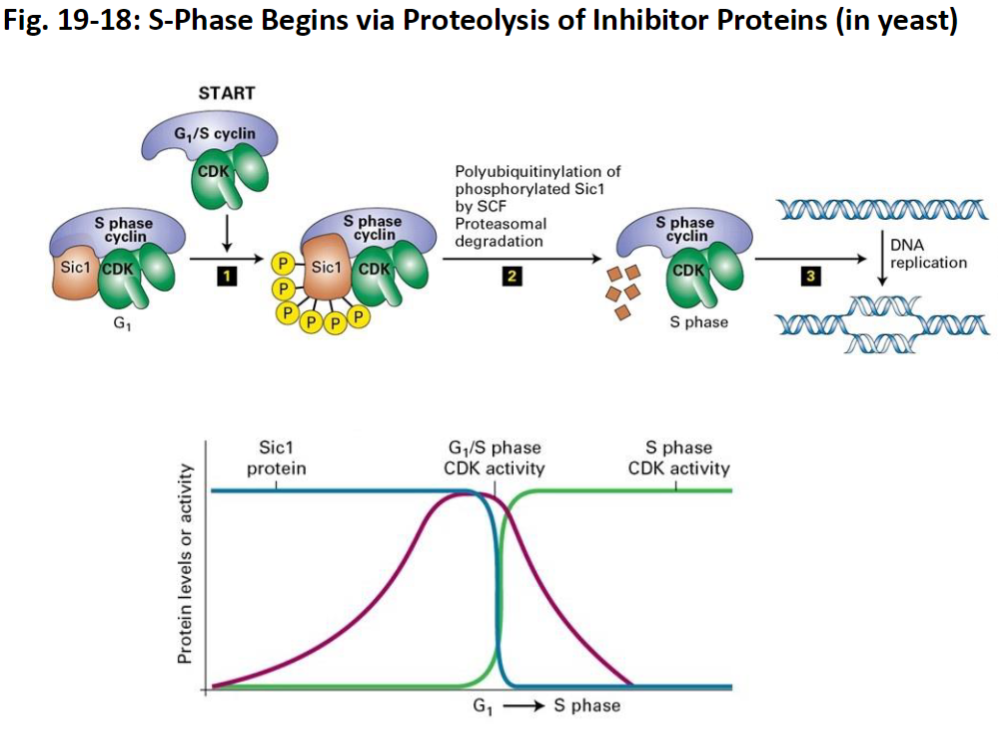
Fig. 19-19: Molecular Mechanisms Governing the Initiation of DNA Replication
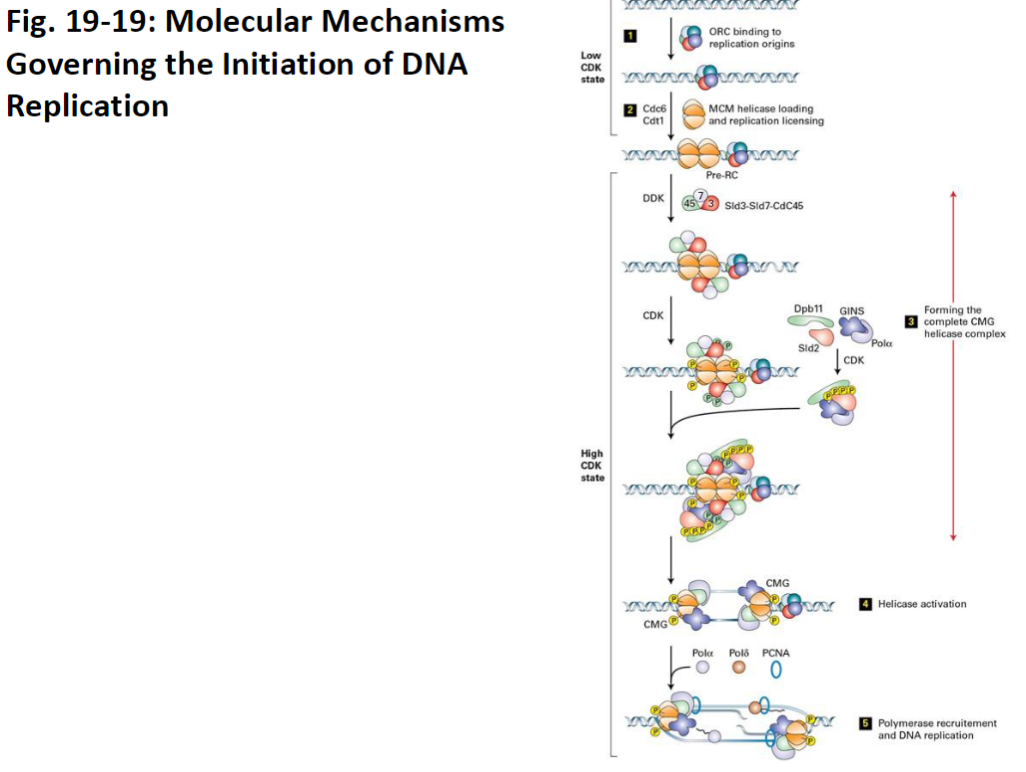
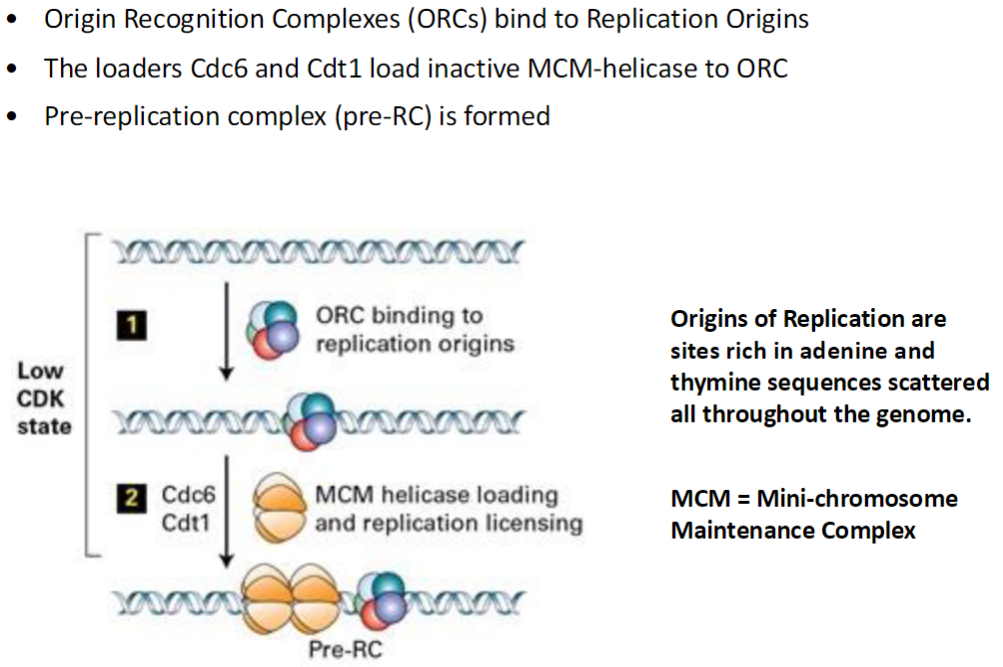
DNA Replication – Pre-Replication Complex
Origin Recognition Complexes (ORCs)
Bind to replication origins (specific DNA sites rich in adenine and thymine)
Mark where DNA replication will start
Loading of MCM Helicase
Cdc6 and Cdt1 act as loaders
Place inactive MCM helicase onto ORC
Pre-Replication Complex (pre-RC)
Formation of pre-RC occurs once MCM is loaded
Prepares DNA for replication in S phase
MCM (Mini-Chromosome Maintenance) Complex
Helicase that unwinds DNA during replication
Essential for replication fork progression
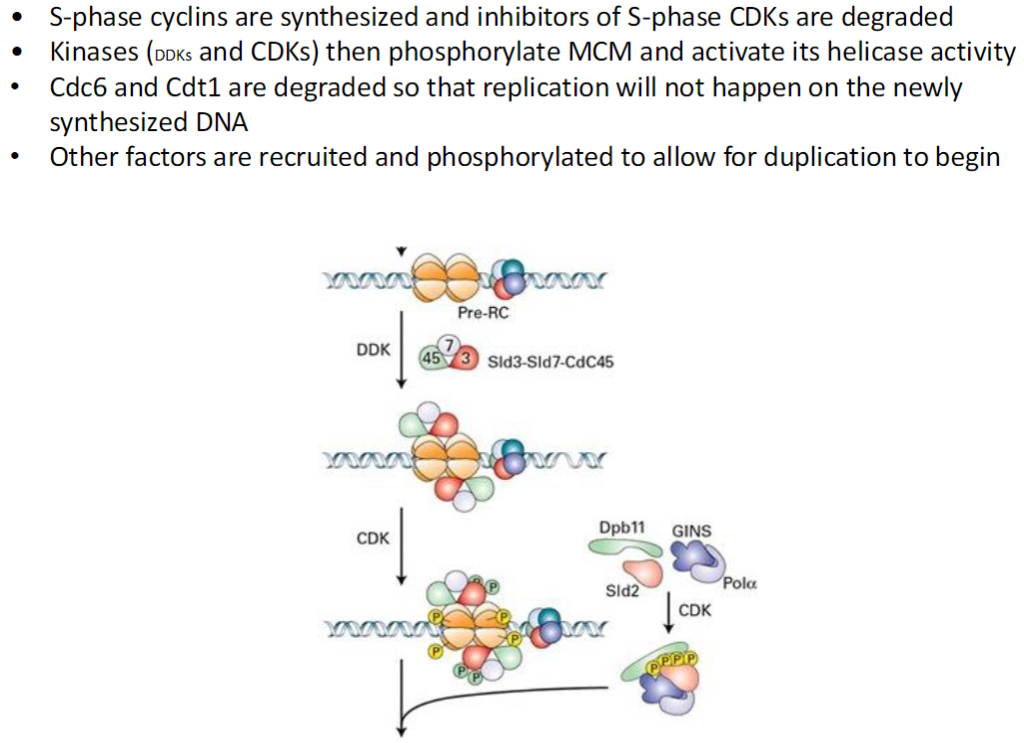
S-Phase Initiation – Activation of Replication
S-Phase Cyclins and CDK Inhibitors
S-phase cyclins are synthesized
Inhibitors of S-phase CDKs are degraded
This allows CDKs to become active
Activation of MCM Helicase
Kinases such as DDKs and CDKs phosphorylate MCM
Phosphorylation activates the helicase activity for DNA unwinding
Prevention of Re-Replication
Cdc6 and Cdt1 are degraded after loading MCM
Ensures replication occurs only once per cell cycle
Recruitment of Additional Factors
Other proteins are recruited and phosphorylated
This prepares the replication machinery to begin DNA duplication
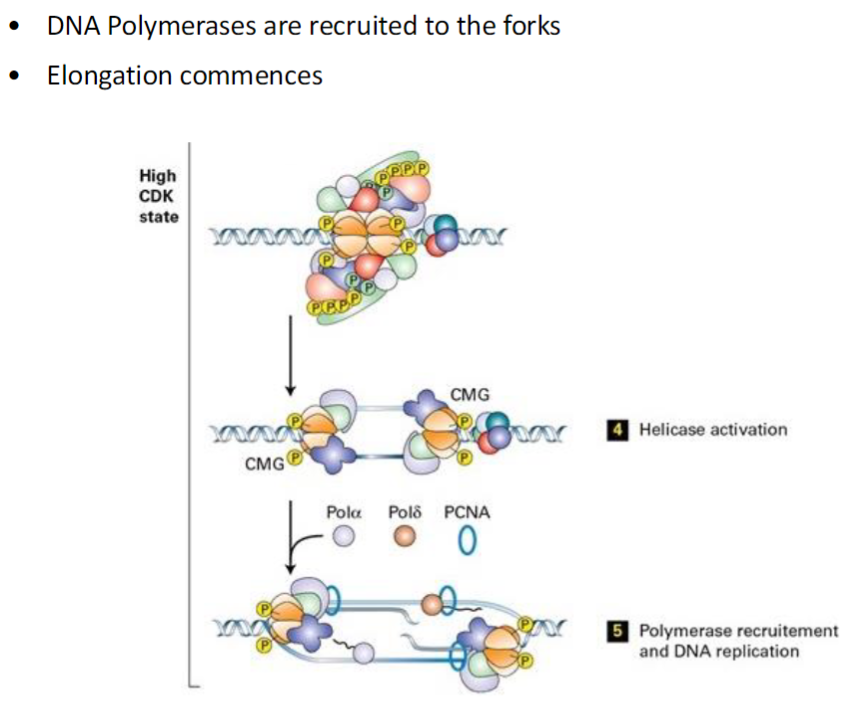
DNA Replication – Elongation
Polymerase Recruitment
DNA polymerases are recruited to the replication forks
These enzymes synthesize new DNA strands complementary to the template
Elongation
DNA synthesis commences
Replication proceeds bidirectionally from the origin
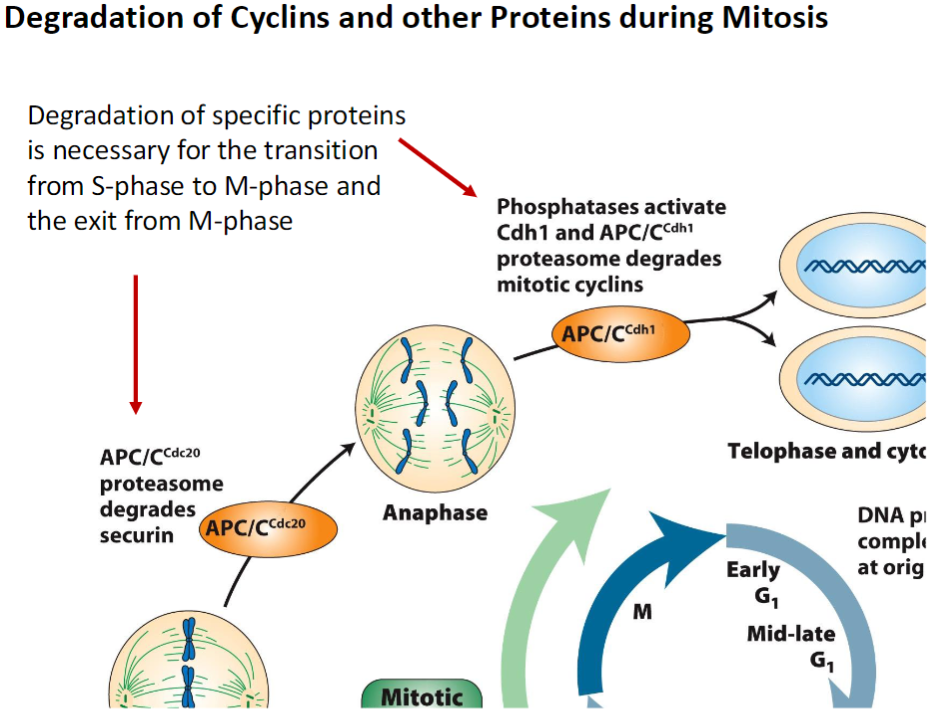
Protein Degradation During Mitosis
Purpose
Degradation of specific proteins is necessary for cell cycle progression
Ensures proper transition from S-phase to M-phase and exit from M-phase
Mechanism
Proteins are tagged for destruction, often by ubiquitin ligases
Targeted degradation allows regulated activation and inactivation of cyclins and other factors
Cell Cycle Surveillance Mechanisms
Checkpoints
Ensure next stage of cell cycle does not start before preceding stage is complete
Checkpoint Pathways
Comprised of event sensors (detect problems), signaling pathways (relay information), and effectors (halt cell cycle and activate repair)
Functions
Arrest cell cycle progression in response to DNA damage or improper spindle assembly
Activate repair pathways to maintain genomic integrity

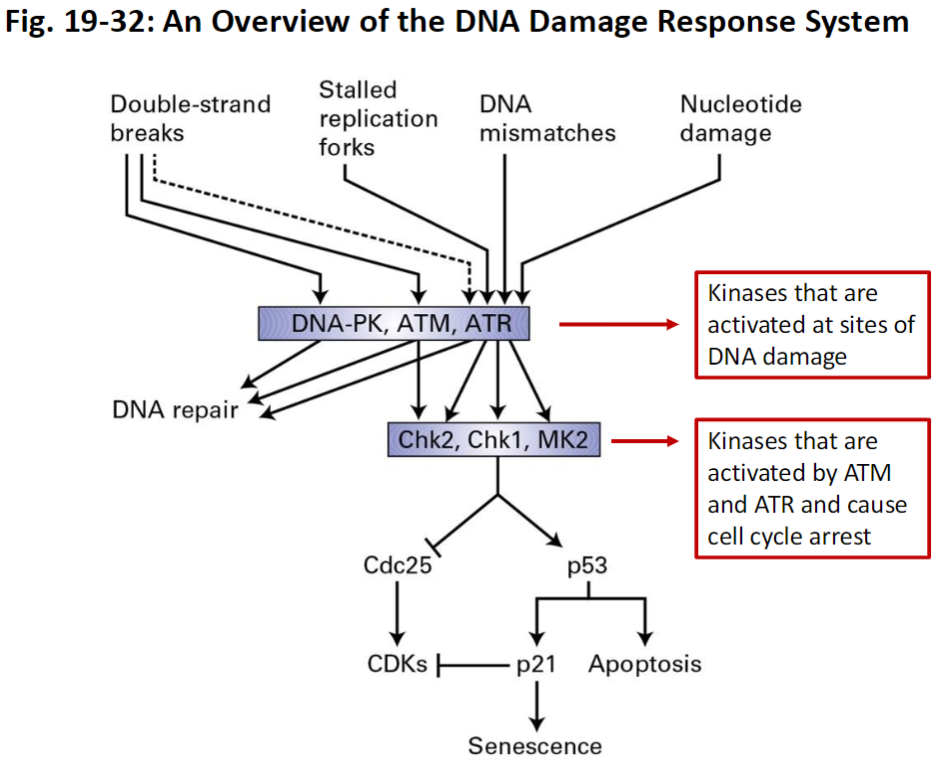
DNA Damage Response System
DNA Damage Sensors
Detect double strand breaks, stalled replication forks, DNA mismatches, or nucleotide errors
Most mutations occur in non-coding DNA and are harmless
Mutations in coding DNA can disrupt proteins and lead to cancer
Key Kinases
ATM – activated by double strand breaks
ATR – activated by stalled replication forks and other DNA stress
DNA-PK – activated by double strand breaks
Downstream Kinases
Chk1, Chk2, MK2 – activated by ATM/ATR
Cause cell cycle arrest
Inhibit Cdc25 phosphatase
Slow CDK activity to prevent mitosis until DNA is repaired
p53 Pathway
Activated by DNA damage
Can trigger apoptosis in irreparable cells
Activates p21 which inhibits CDKs to halt cell cycle progression
Overall Function
Facilitates DNA repair, prevents damaged cells from dividing, and removes potentially tumorigenic cells
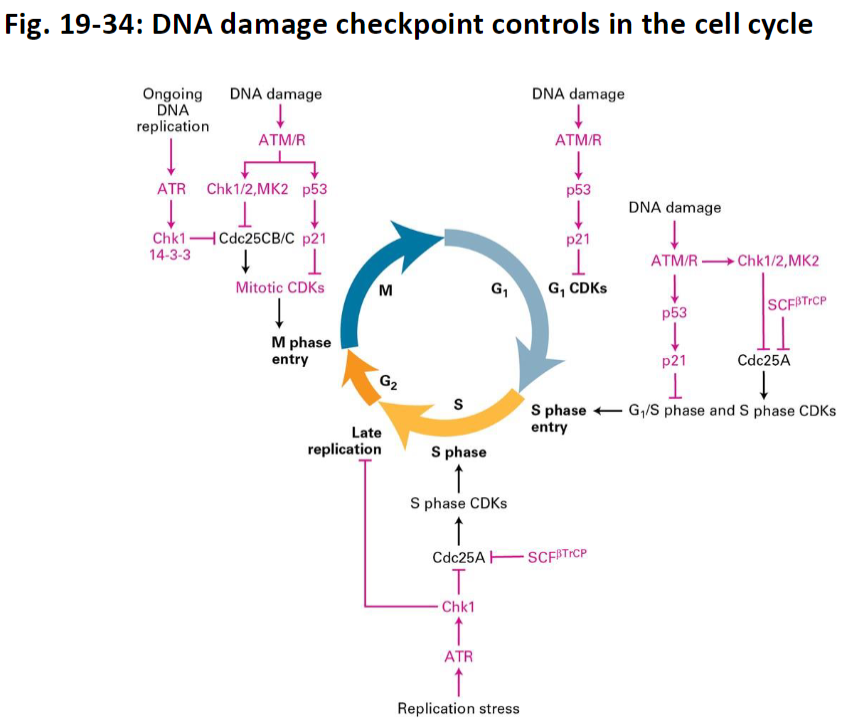
DNA Damage Checkpoint Controls
Checkpoint Timing
Kinases act at multiple points in the cell cycle:
G1 – before S-phase entry
S – during DNA replication
G2/M – entry into mitosis
G1 Checkpoint
ATM/ATR detect DNA damage
Inhibit G1 CDKs
Prevent entry into S-phase if damage is present
S-Phase Checkpoint
Replication stress or stalled forks activate ATR and Chk1
Inhibit Cdc25 phosphatase
Regulate S-phase CDKs to pause DNA synthesis until problems are resolved
G2/M (Mitotic) Checkpoint
Excess DNA damage or incomplete replication activates ATM, ATR, Chk1/2, MK2
Inhibit Cdc25 and other activators of mitotic CDKs
Prevent entry into mitosis until replication is complete and DNA is repaired
Overall Function
Ensures that each phase of the cell cycle only proceeds when prior events are complete
Prevents propagation of damaged DNA and reduces risk of cancer