BIOL 201 - Cell Bio & Metabolism
1/220
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
221 Terms
First Law of Thermodynamics
in an isolated system, the total amount of energy remains constant over time
energy is neither created nor destroyed, just converted from one form to another
Second Law of Thermodynamics
an isolated system always tends towards disorder
metabolism
science of energy conversions
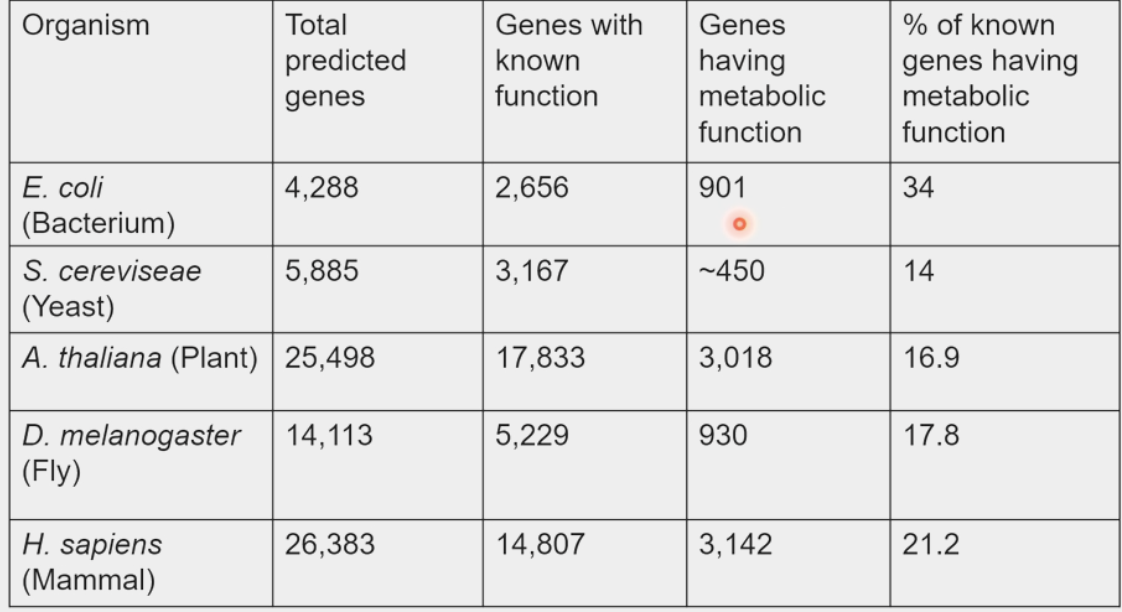
organisms devote 20% of genome to this
high-quality → low-quality energy
caveats of energy
organisms store energy during growth
organisms store energy temporarily during activity
inequality of energy
“Low quality’ = HEAT (random kinetic energy)
less order
“High quality” = Potential energy of mass on a pull
more ordered/structured
Joule’s Experiment
Organisms convert high-quality → low-quality energy
Gibbs free energy
available energy to an organism
H – “unavailable energy”
G = H – TS
Released by ATP hydrolysis
Cells capture it to create order
the change in this determines whether a process occurs spontaneously
Enthalpy
total energy in a system
H
Entropy
disorder in a given system
S
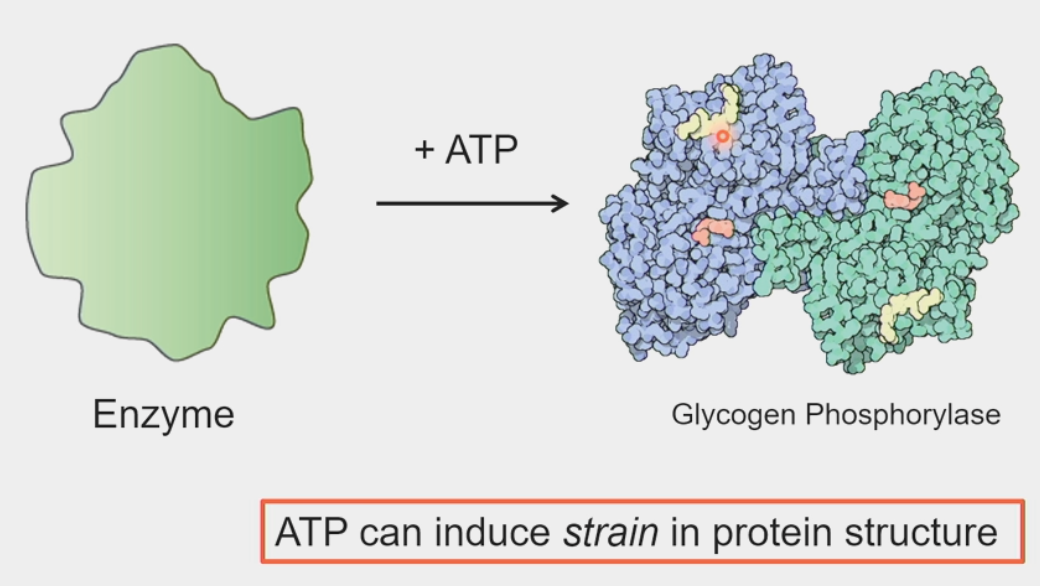
ATP binding
free energy can temporarily distort the 3D structure of proteins
induces strain
protein relaxes back into native configuration → chemical reaction forced to occur
change in free energy
determines whether a process occurs spontaneously
ΔG = Gfinal – Ginitial
ΔG = ΔH – TΔS
for “isolated system”, ΔH = 0
entropy always increases; ΔS > 0
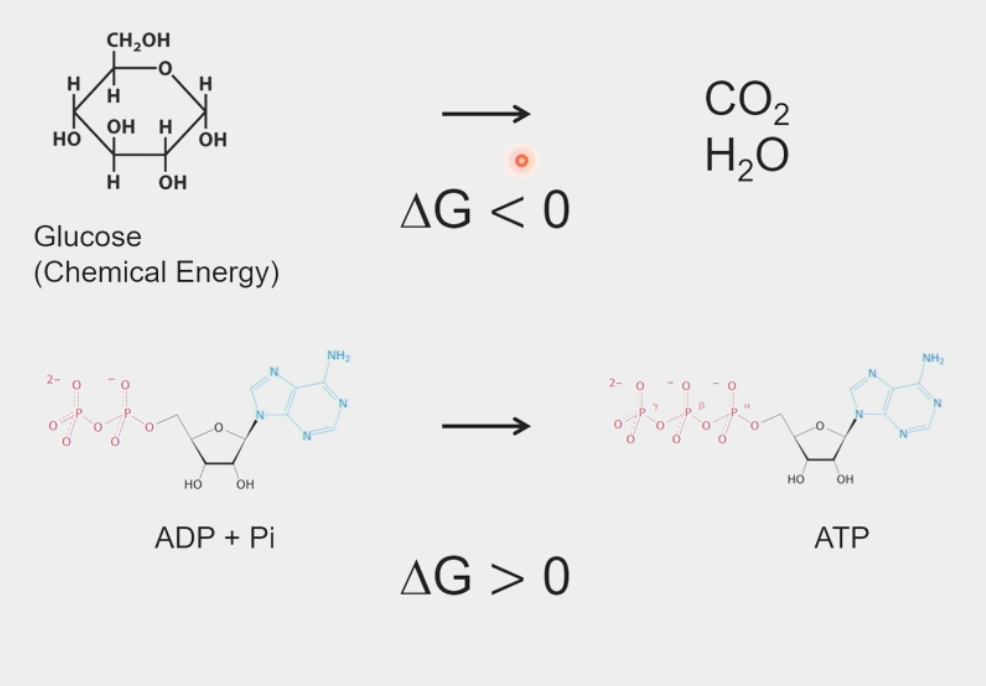
ΔG < 0 for a spontaneous process
values are standardized (ΔGº’)
T = 298K, P = 1atm, pH = 7.0, all concentrations = 1M
standard free energies are additive → “coupled” reactions
Redox reactions
Reduction = gaining an electron
Oxidation = losing an electron
provide a basis for energy transduction
fuels are oxidized by metabolic enzymes
NAD+
principal electron acceptor in metabolic redox reactions
GAPDH brings this into position to be reduced to NADH
may be useful in a wide range of therapies
ex: tuberculosis drug Isoniazid → active form binds NADH, inhibits cell wall synthesis enzyme
FAD used when available free energy can’t reduce this
requires ΔGº’ = 52.6 kcal/mol to capture e-
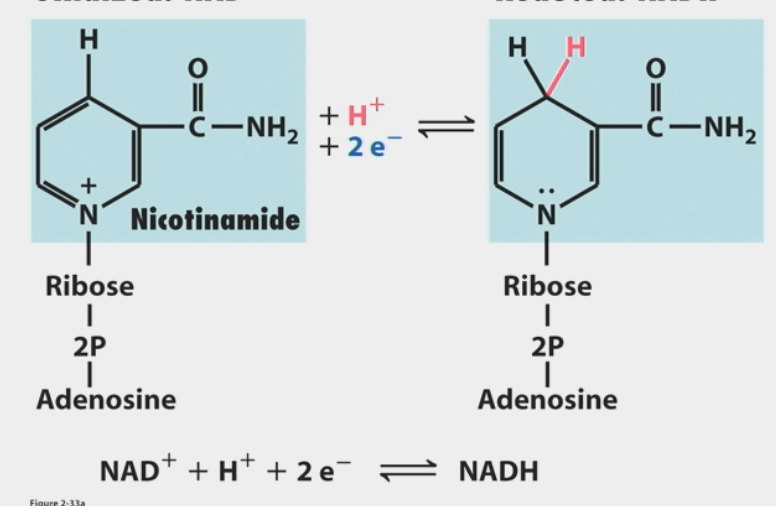
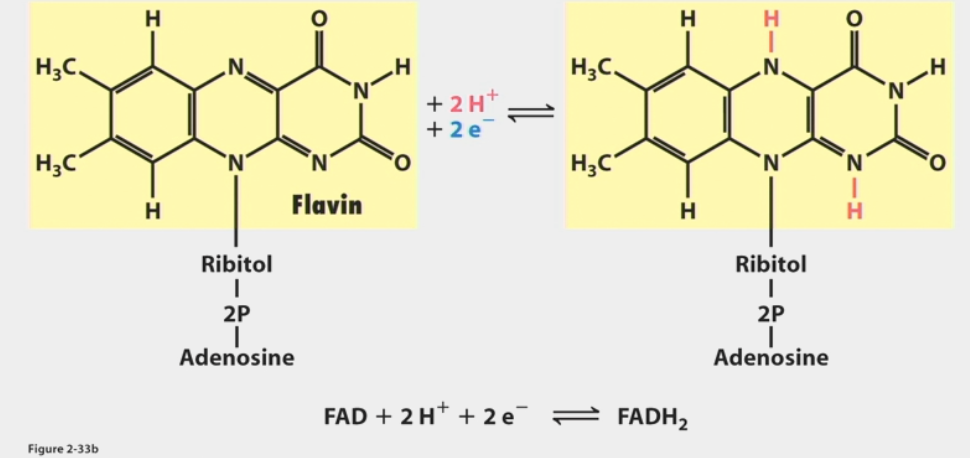
FAD
used when available free energy can’t reduce NAD+
less ΔGº’ required than NAD+ → NADH (43.4 kcal/mol vs. 52.6)
reduced to FADH2
ATP
“energy currency” of the cell (like a $20)
needs ΔGº’ = -7.3 kcal/mol to be hydrolyzed (20 kBT)
big enough to do something with but small enough to avoid too much waste
thermal energy
serves as baseline for cellular energy scales
〈E〉= 3/2kBT
kB = Boltzmann’s constant
kBT = 0.6 kcal/mol
average amount of energy an H2O molecule has when it collides
ATP hydrolysis ≈ 20 kBT (physiological conditions)
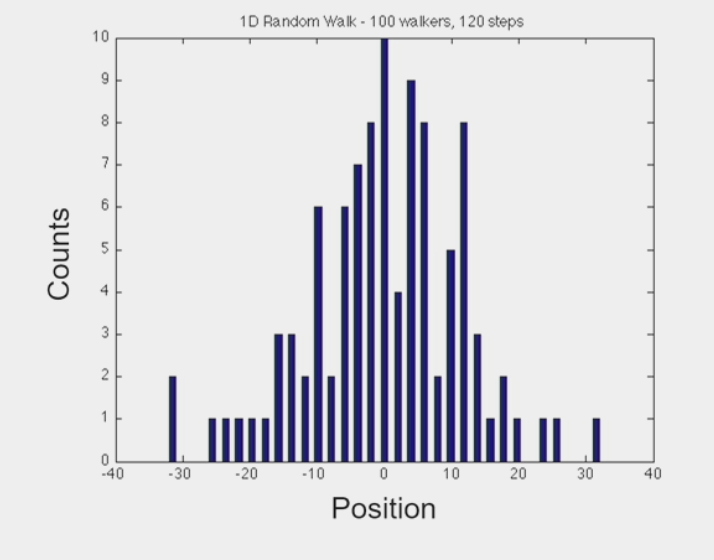
Brownian motion
the random motion of particles suspended in a medium
molecules continuously undergo small, random fluctuations
“random walk” → at any given moment, molecule shifts left OR right
as # “steps” increases, particles start to diffuse away from each other
molecules diffuse w/ characteristic diffusion coefficient (D)
smaller molecules = “faster” diffusion, larger D
Diffusion coefficient (D) describes spread of population of molecules
diffusion is mostly ACTIVE (a bit thermal, but it’s negligible in cells)
explores a lot of space, but on average gets nowhere
Mean Squared-Displacement: 〈x2〉= 2Dt
mechanical energy
energy that is possessed by an object due to its motion or due to its position
sum of kinetic and potential
cells and subcellular structures feel and produce mechanical forces
1 N (newton) = 1kg x 1 m/s2
cellular forces measured in pN (piconewton) to nN (nanonewton)
1 pN = 10-12 N
kBT = 4.1 pN nm
ATP hydrolysis ≈ 20 kBT ≈ 80 pN nm (a tiny amount of work)
electromagnetic energy
the various energies that travel as wavelengths through space at the speed of light
photons absorbed and emitted (electric + magnetic fields)
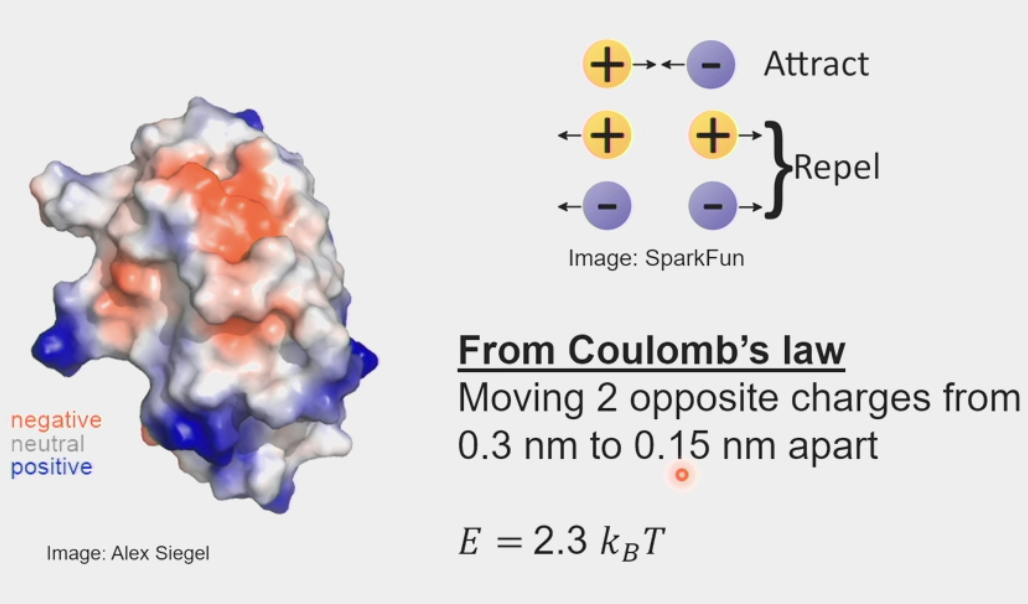
electrostatic potentials
surfaces of protons contain many charged residues
Coulomb’s Law: electrical force between two charged objects is directly proportional to the product of the quantity of charge on the objects
moving 2 opposite charges from 0.3 nm to 0.15 nm apart
E = 2.3 kBT (pretty small) → need a large surface area for 2 proteins to stick together
a little stronger than thermal energy
photons
elementary particle that is a quantum of the electromagnetic field
E (energy) = hν
h = Planck’s constant
ν = frequency in Hz
visible particle ≈ 2 eV = 80 kBT
kBT = 25 meV (milli-electric Volts)
absorbed to ultimately produce ATP in photosynthesis
breaking bonds
non-covalent bonds (ex. H-bonds)
2-12 kBT → varies because of random motion of thermal energy
electrostatic bonds
probability of breaking: P = e–E/kBT
ex. P = e-3 = 0.05 → 5% of breaking
1011 collisions per second
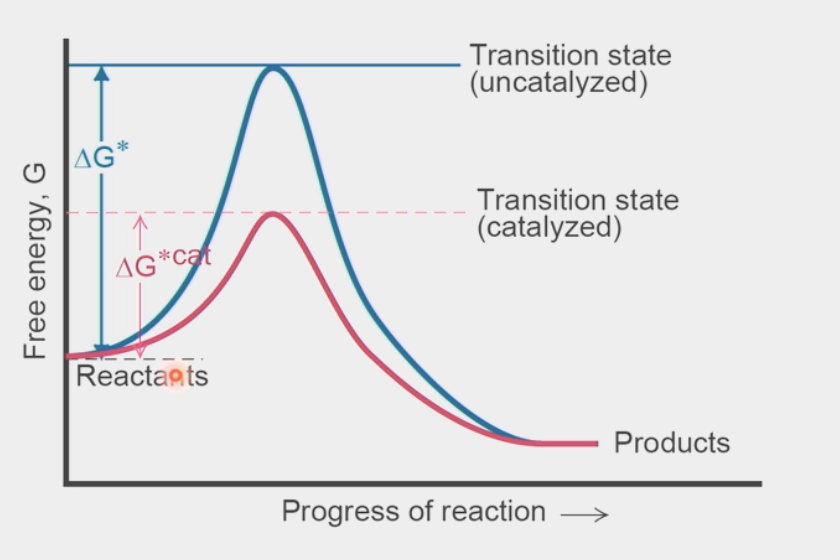
transition state
short-lived configuration of atoms at a local energy maximum (highest potential energy) in a reaction-energy diagram
catalysts lower this value → less ΔG required to get over this point
energy obtained from random collisions → if value is lower, then higher probability to get over it
products have LOWER ΔG than reactants (overall negative = spontaneous rxn)
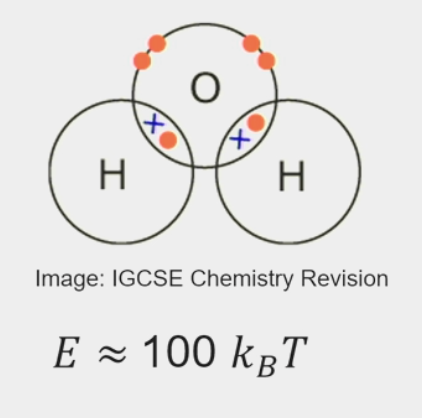
covalent bonds
chemical bond that involves the sharing of electrons to form electron pairs between atoms
very stable
E ≈ 100 kBT
probability of breaking: P = e-100 = 3.7×10-44
by itself, a collision will break this every 1024 years
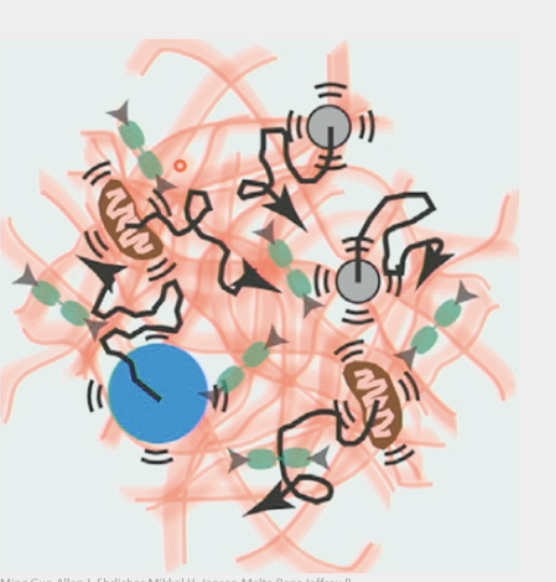
cytoplasm
an active material in the cell
far from equilibrium because of ATP hydrolysis
has different consistencies depending on the size of the particle in it
for ions: like water
for organelles/macromolecular complexes: like glass
carbon/ATP depletion causes transformation into glass consistency (molecules frozen)
very CROWDED
contains ions & H2O (0.1nm), sugars, amino acids, proteins & DNA & RNA(10-100 nm), organelles (1µm), etc.
as packed as protein crystals (20-60% protein by weight)
rough-and-tumble place
proteins in constant motion → constantly smashing into each other
Brownian motion → “random walk”, 1-D: 50% step right, 50% step left
as # “steps” increases, particles start to diffuse away from each other
Diffusion coefficient (D) describes spread of population of molecules
cells have to fight to maintain spatial organization
collisions distort structure of individual proteins → conformational changes
large organelles/proteins move in place, tiny ions/particles basically move freely
viscous
inertia = resistance of an object to any change in its state of motion
viscosity = measure of a fluid’s resistance to flow
Reynold’s Number: Re = inertial forces/viscous forces = ρνL / µ = (density)(velocity)(Length) / (dynamic viscosity)
cells, organelles, proteins are in LOW Re environment (~10-4)
inertia is completely negligible here
elastic
elasticity = tendency of an object to return to its original shape after deformation
meshwork
long filamentous proteins → actin filaments
organelles, polymers, other structures define “pore size” → this is why larger macromolecular complexes are “trapped”/unable to move on their own
inertia
resistance of an object to any change in its state of motion
viscosity
measure of a fluid’s resistance to flow
Reynold’s number
dimensionless quantity that helps predict fluid flow patterns
Re = inertial forces/viscous forces = ρνL / µ = (density)(velocity)(Length) / (dynamic viscosity)
elasticity
tendency of an object to return to its original shape after deformation
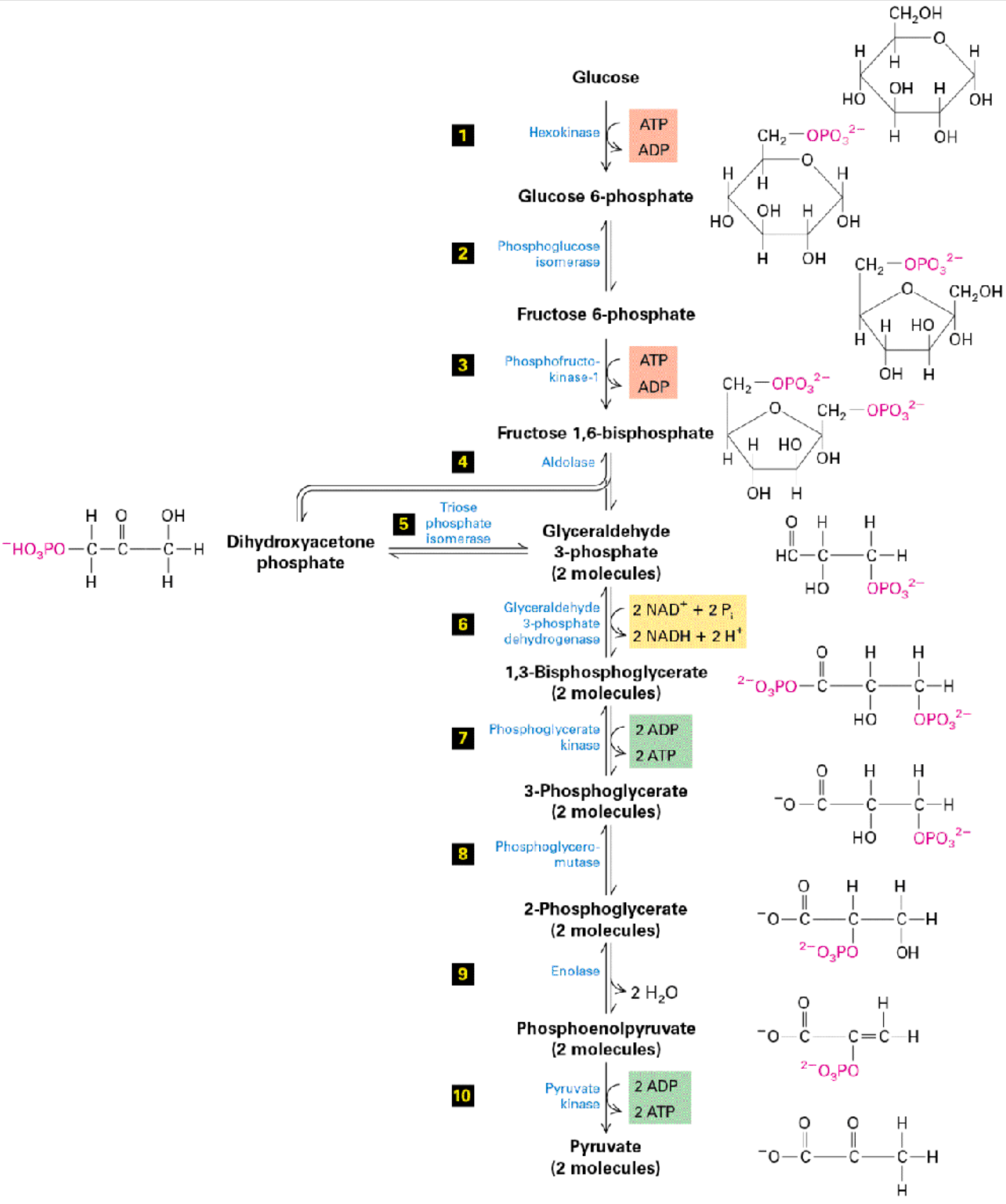
glycolysis
Stage 1 of metabolism
converts glucose (C6H12O6) into pyruvate
harvesting electrons
yields NET production of 2 ATP per 1 glucose
Phosphorylate glucose by adding Pi from ATP
Creates glucose 6-phosphate → goes on to form other metabolites
Catalyzed by hexokinase enzyme
Change glucose ring → fructose ring
hexameric → pentameric
catalyzed by phosphoglucose isomerase
fructose 6-phosphate → used for downstream processes like making glycolipids
Phosphorylate again w/ another ATP
produces fructose 1,6-bisphosphate
catalyzed by phosphofructokinase
Formation of dihydroxyacetone phosphate
catalyzed by aldolase
Split into 2 molecules
2 x glyceraldehyde 3-phosphate (G3P)
2 NAD+ used to make 2 NADH
Oxidizing G3P with glyceraldehyde 3-phosphate dehydrogenase
1,3-bisphosphoglycerate formed
Dephosphorylation
2 ADP → 2 ATP produced
3-phosphoglycerate formed by phosphoglycerate kinase
3-phosphoglycerate mutated → 2-phosphoglycerate
catalyzed by phosphoglycerol mutase
Condensation → H2O produced
done by enolase
Dephosphorylation
2 ADP → 2 ATP produced
pyruvate formed
catalyzed by pyruvate kinase
pump priming
generates useful metabolite
increases free energy of reactants
larger ΔG, smaller activation barrier to get over transition state
NOT catalysis
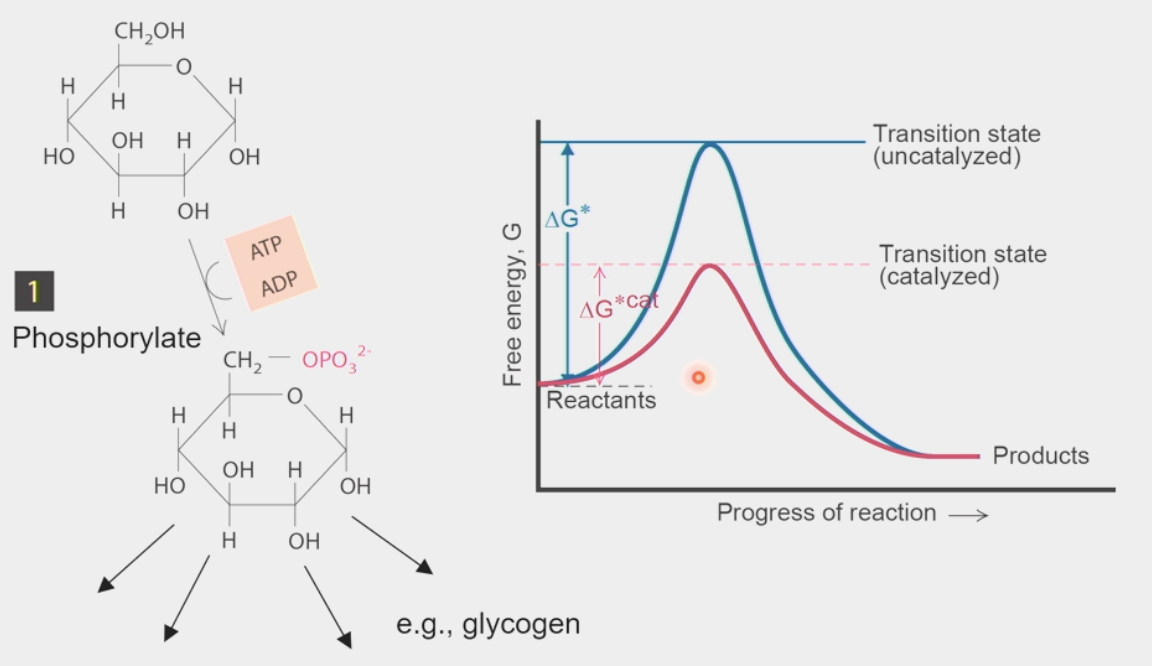
regulation of glycolysis
huge enzymes like “disassembly line”
cells must monitor glycolytic flux by conformational switches to enzymes modulated by allosteric activators and inhibitors
cancer cells → increased flux
mutations in glycolytic enzymes found in tumors → block product inhibition of enzyme
cause is DNA damage, NOT misregulation of glycolysis
steps w/ big –ΔG effectively irreversible
PFK1 → gatekeeper
also catalyzes step 3 (phosphorylation of fructose 6-phospate → fructose 1,6)
once this step is done, you can’t go back
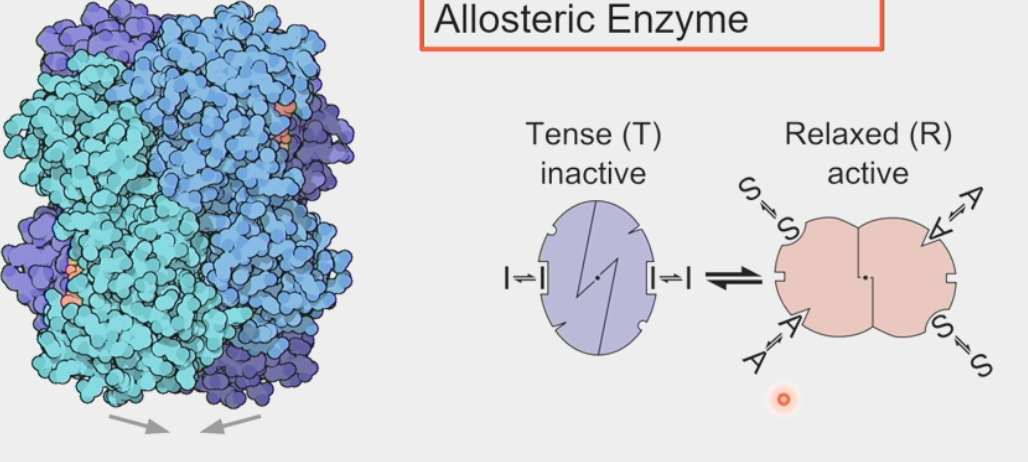
PFK1
phosphofructokinase → controls glycolytic flux
tetramer → 4 individual polypeptide chains linked together
switches between active and inactive states by conformational change
tense = inactive, relaxed = active
2 substrate binding sites open in active state
inhibitors at inactive state
allosteric enzyme
enables cells to have control over switching glycolysis on/off
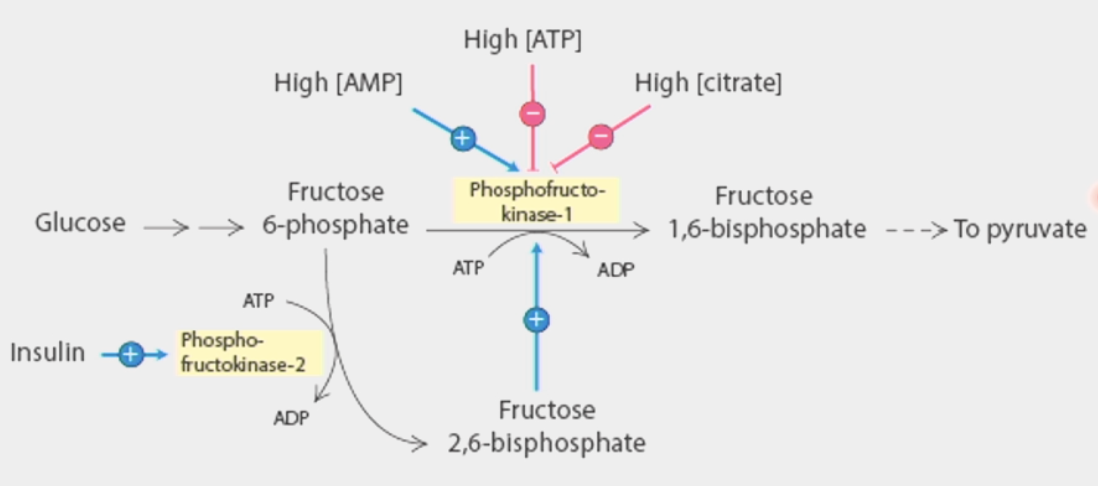
inhibited by its products → ATP and citrate
ATP also a substrate → as its concentration increases, it becomes an inhibitor
activated directly by AMP, indirectly by excess fructose-6-phosphate (starting material for step 3)
AMP = starting material for ATP (adenosine monophosphate)
PFK2 starts phosphorylating when there is excess → produces fructose 2,6-bisphosphate which acts as direct activator for PFK1
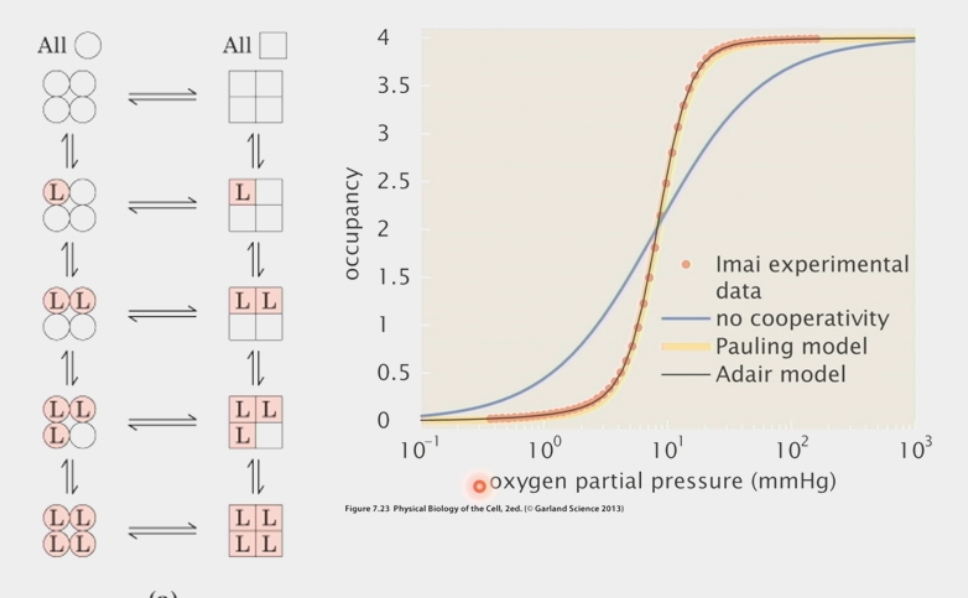
MWC model
explains allostery: when allosteric enzyme switches between active/inactive conformation, ALL 4 subunits must switch at once
one ligand binds in inactive state → rapid switch of molecule to active state
then all other ligands bind, enzyme performs rxn
as soon as one ligand is lost → rapid switch to inactive state
all other ligands dumped immediately
S-shaped sigmoid curve
used to find hemoglobin
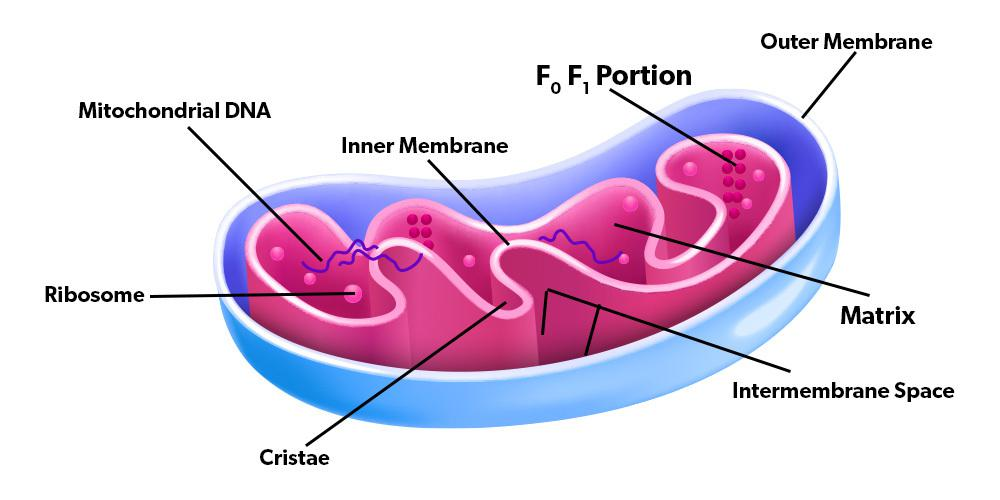
mitochondria
eukaryotic organelle
double membrane structure
form complex network of tubes
originated from bacteria → Endosymbiont hypothesis
aerobic respiration was the driver of this
evolutionary advantage
helped cells produce more power per gene
make their own ribosomes, segregate their own DNA (mtDNA), etc.
have their own genetic code & different codon usage
ex. UGA = Stop for standard code, Trp for mitochondria
determines protein length
function influences lifespan
ex. mice w/ homozygous mutation die very young
ex. mutations in mitochondrial proteins like ETC increased lifespan in C. elegans
mutations affect 1/5000 live births
can cause problems in all tissues/cell type of body
morphology varies
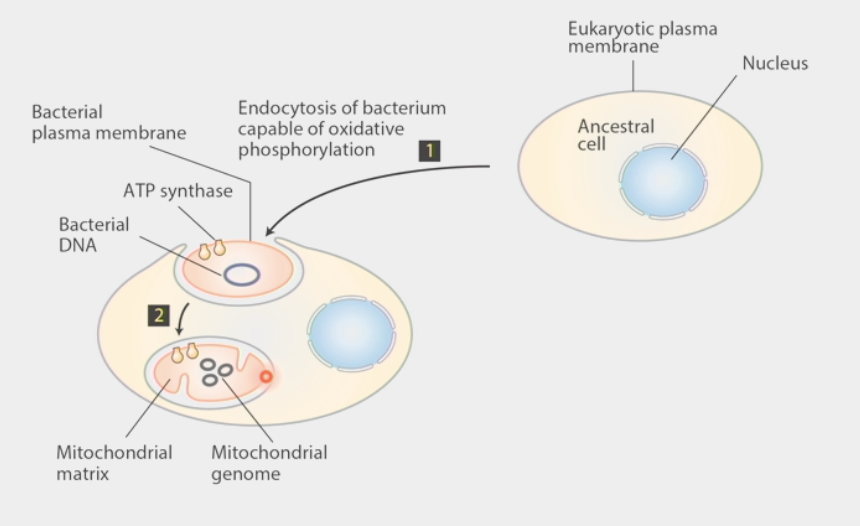
Endosymbiont hypothesis
ancestral cell w/ nucleus & eukaryotic plasma membrane
engulfed a bacteria capable of oxidative phosphorylation w/ ATP synthase, bacterial DNA, bacterial plasma membrane
bacteria became mitochondria w/ mitochondrial matrix and unique genome (mtDNA)
controversial hypothesis!!
alternative: aerobic proto-eukaryote enlarged & engulfed its respiratory surfaces (ATP synthase complexes)
central counter: host was already aerobic
DNA sequence analysis proved these wrong → chicken liver mitochondria closer to E.coli than bovine erythrocyte
mitochondria increased power per gene
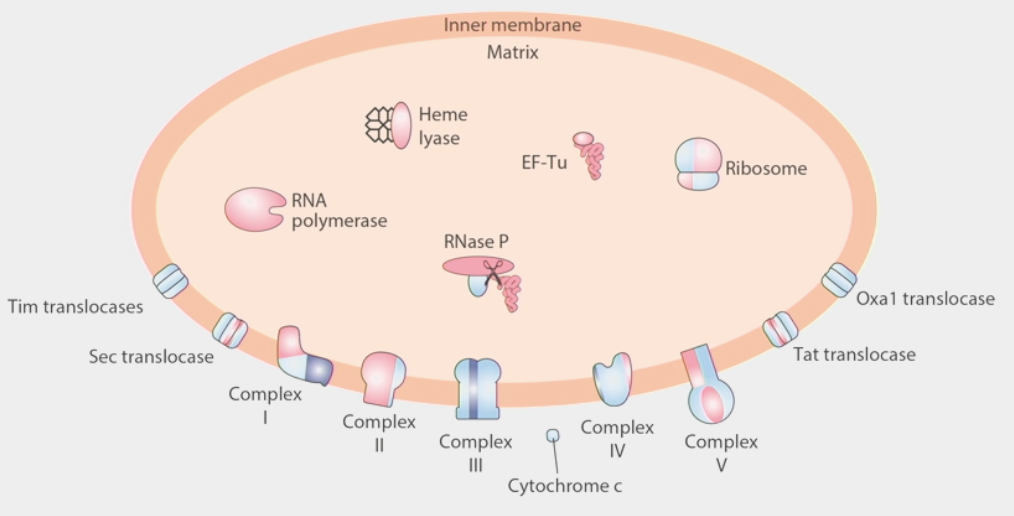
mitochondrial proteins
built from a combination of nuclear DNA and mtDNA
small % of mitochondrial proteome is built from the mitochondrial genome
ribosome: built from some proteins synthesized in cytoplasm and imported, combined w/ locally synthesized proteins
several translocases work as channels to bring cytosolic proteins into mitochondrial matrix
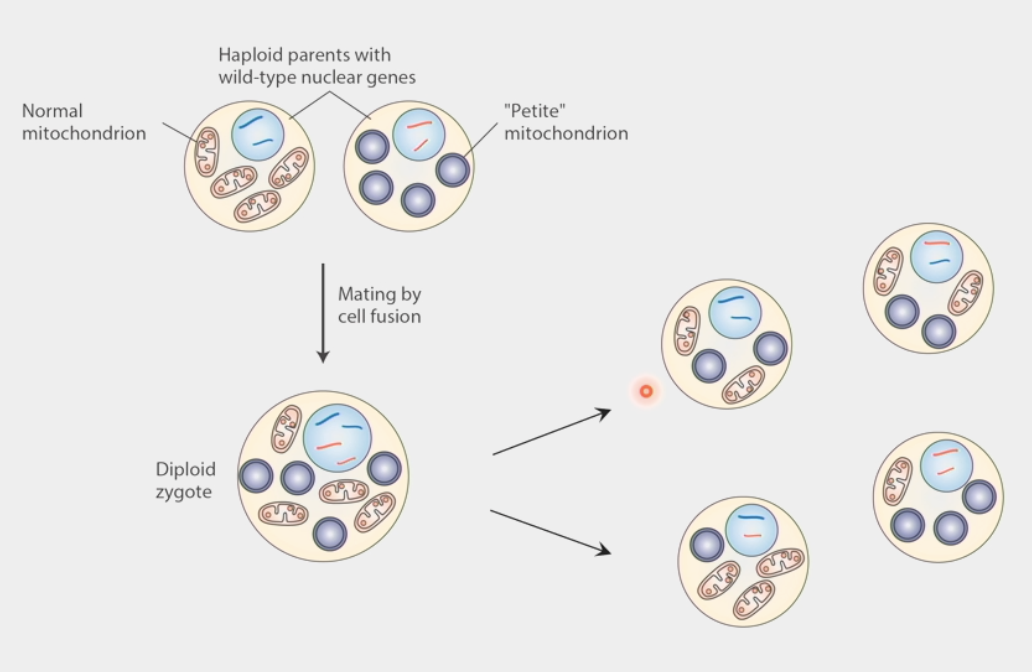
mtDNA inheritance
modeled from petite mutation in yeast (inhibits growth)
2 haploid (1n) parents w/ wild-type nuclear genes → one w/ normal mitochondria, one w/ petite
mating by cell fusion → diploid (2n) zygote
yeast sporulate → even segregation of nuclear DNA
mitochondrial segregation is RANDOM
petite cells occur by chance due to random segregation of mtDNA
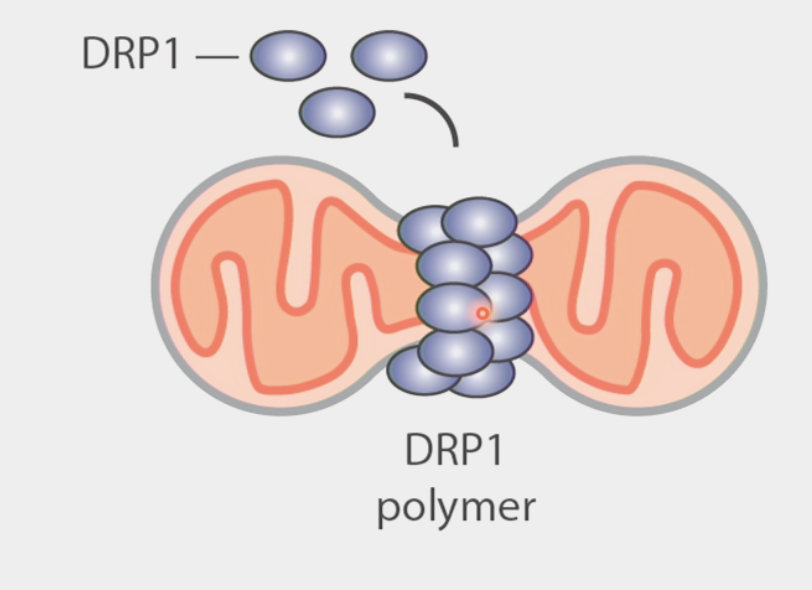
mitochondrial fission
the process by which mitochondria divide or segregate into two separate mitochondrial organelles
GTPase called DRP1 oligamerizes in ring shape on mitochondrial membrane and squeezes it apart
mutants have opposite phenotypes to fusion
continuous and ongoing at all times
promotes equal segregation of mitochondria into daughter cells
as cells enter mitosis, they fragment mitochondria into as many small pieces as possible
DRP1-/- cells segregate mitochondria asymmetrically
often occurs at point of Endoplasmic Reticulum-mitochondrial contact
triggers recruitment of DRP1
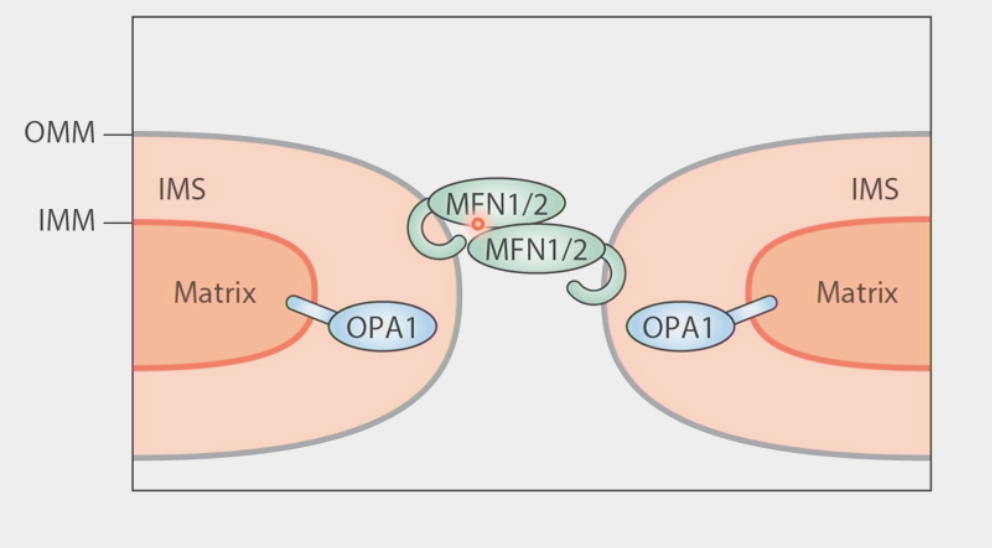
mitochondrial fusion
merging of two or more mitochondria within a cell to form a single compartment
2-step process because of the double membrane:
Mitofusins (MFN1/2) form trans-dimer extending from one mitochondria to another
conformational change in MFN1/2 brings outer membranes together
Inner membrane: 2 copies of OPA1 dimerize → undergo conformational change to smash inner membranes together
mutants have opposite phenotypes to fission
continuous and ongoing at all times
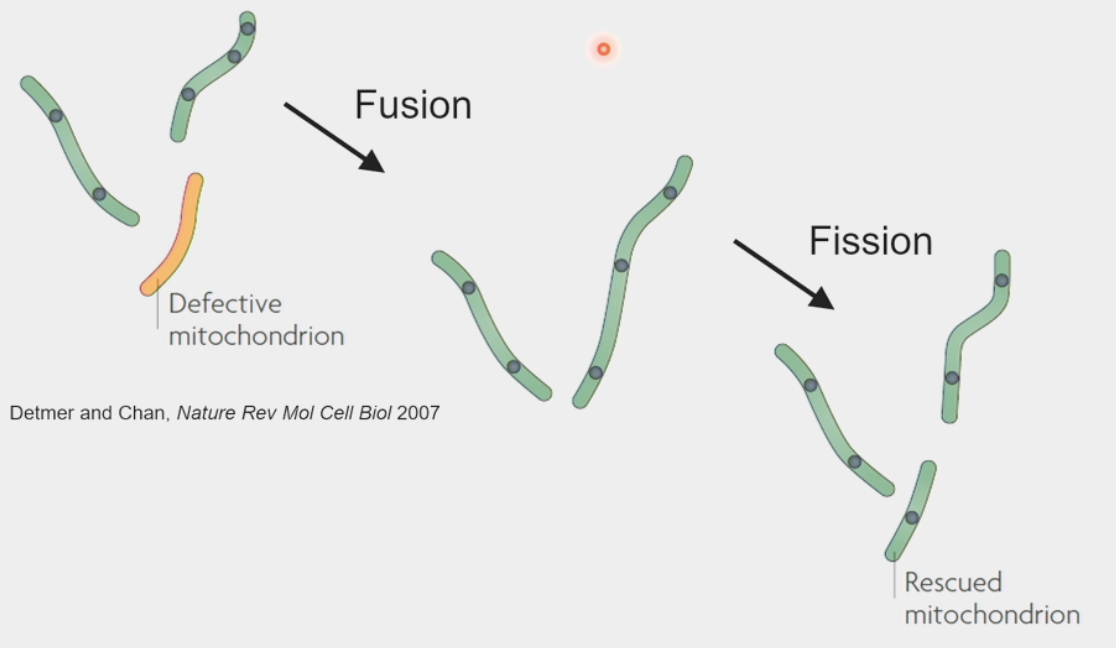
mitochondrial repair
damage accumulated to mtDNA and protein contents
reactive oxygen species → byproduct of metabolism, toxic waste!!
fission & fusion promote mixing of mitochondrial contents & enable “rescue”
defective mitochondrion fused w/ working mitochondrion → now able to produce proteins to rebuild machinery → undergoes fission again to create “rescued” mitochondrion w/ its own genome
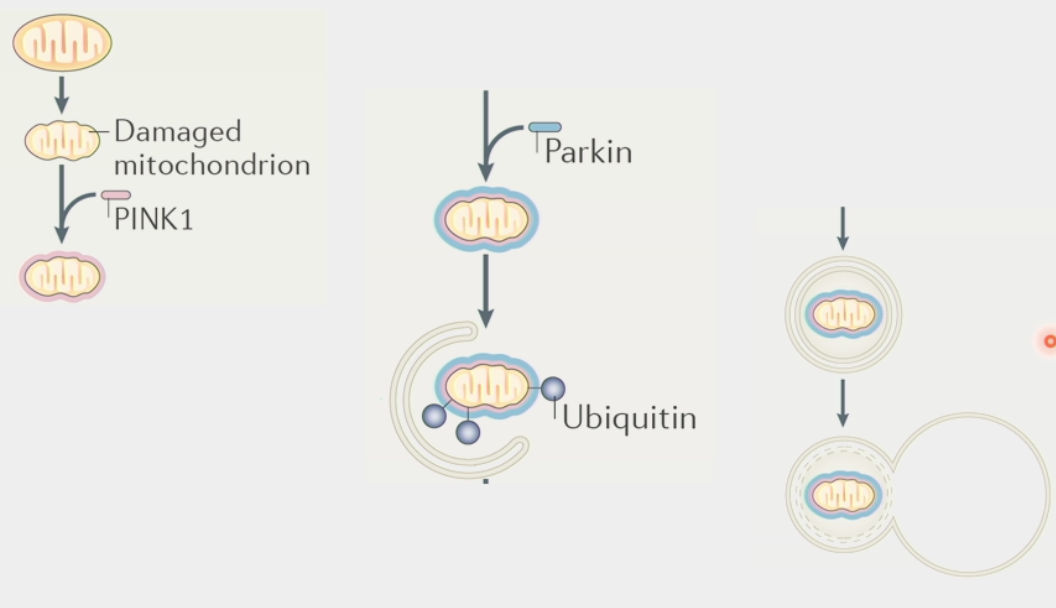
mitophagy
the process of cells degrading their mitochondria
clears out dysfunctional mitochondria and controls number
eaten by large multi-protein organelle autophagosome
spits out nucleic & amino acids for rebuilding
PINK1 → identifies/targets damaged mitochondria
Parkin → tags mitochondria for degradation
Ubiquitin ligase, recruits autophagosome
Autosomal Recessive Early-Onset Parkinson’s Disease linked to misregulation of this process
PINK1 & Parkin mutations - dysfunctional mitochondria not degraded
tolerated for decades, then triggers disease later in life
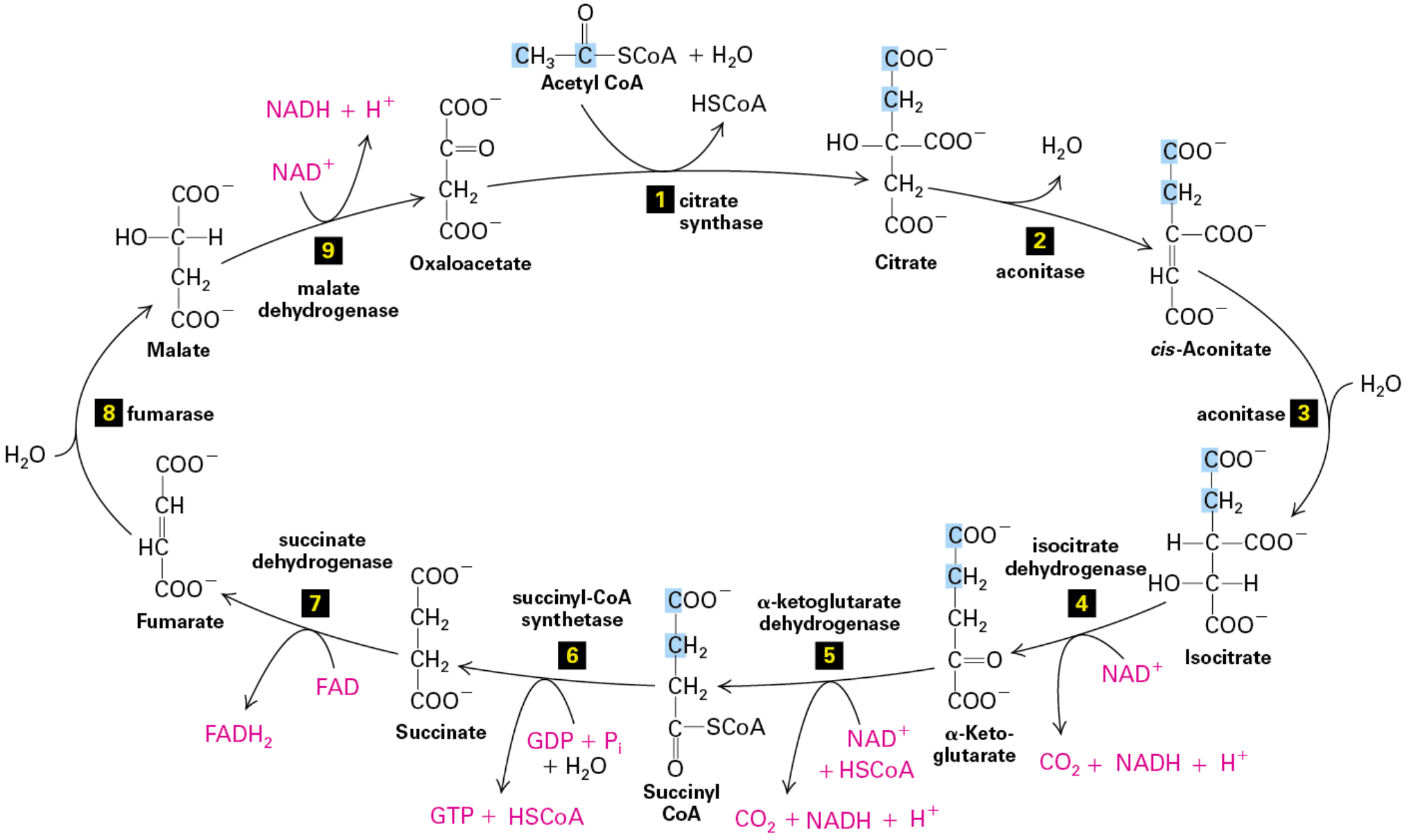
citric acid cycle
Stage II of metabolism
pyruvate enters cycle as C2 (2 carbon) acetyl group on acetyl-CoA
bound to CoA (Coenzyme A) in 3-step process
1. Decarboxylation
2. Oxidation (NAD+ → NADH)
3. Transfer of acetyl group
performed by huge pyruvate dehydrogenase complex
ΔGº’ = -80kcal/mol, basically irreversible rxn
Krebs measured O2 consumption of pigeon muscle w/ manometer (U-shaped pressure thingy)
addition of citrate increased rate of consumption
C2 acetyl group added to oxaloacetate (C4 carrier) by citrate synthase
citrate formed
oxaloacetate binding to citrate synthase creates binding site for Acetyl-CoA (INDUCED FIT)
Citrate converted → cis-Aconitate → isocitrate (less stable form)
done by aconitase
H2O out, then H2O in
C6 isocitrate converted → ⍺-ketoglutarate
done by isocitrate dehydrogenase
also converts NAD+ → CO2 + NADH + H+
⍺-ketoglutarate converted → C4 Succinyl-CoA
done by ⍺-ketoglutarate dehydrogenase (same steps as pyruvate dehydrogenase)
also converts NAD+ + HSCoA → CO2 + NADH + H+
Succinyl CoA converted → Succinate
done by succinyl-CoA synthetase
also converts GDP + Pi + H2O → GTP + HSCoA
Succinate converted → Fumarate
done by succinate dehydrogenase
also converts FAD → FADH2 (less energy to reduce than NAD)
Fumarate converted → Malate
done by fumarase
H2O comes in
Malate converted back to oxaloacetate
done by malate dehydrogenase
also converts NAD+ → NADH + H+
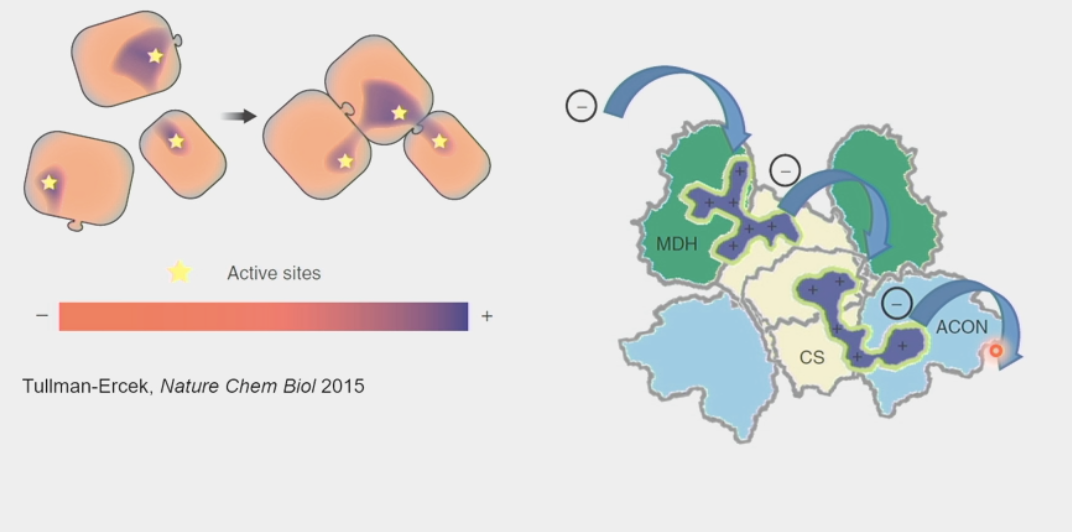
substrate channeling
enabled by large metabolic complexes
passing of the intermediary metabolic product of one enzyme directly to another enzyme or active site WITHOUT its release into solution
increase reaction rates and preserve orientation
ex. ETC
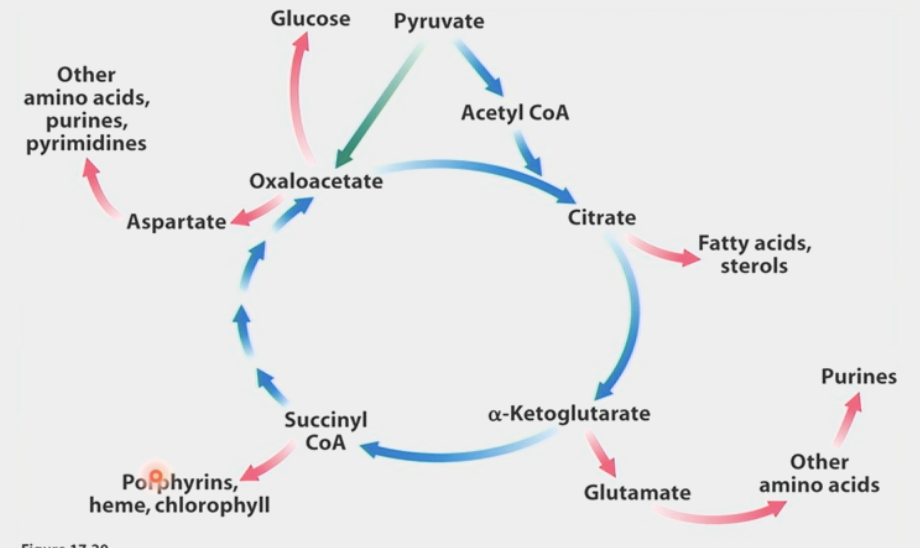
radioactive carbons
Carbon-14 allowed scientists to track fates of metabolites in citric acid cycle
learned that the 2C that enter cycle do NOT leave the first time around (from CO2 produced by steps 4 & 5)
carbons cleaved off from bottom to form CO2 → next carbons in molecule move down
symmetry of molecules makes succinate dehydrogenase unable to distinguish orientation after step 5
0% radioactivity 1st time → 50% radioactivity 2nd time, etc.
citric acid cycle enzymes
major oncogenes
mutations in isocitrate dehydrogenase produce “oncogenic metabolite”
produces 2-hydroxy-glutarate instead of ⍺-Ketoglutarate
inhibits histone demethylation → change in gene expression → cancer
TCA metabolites → source of biosynthetic precursors & cofactors
fatty acids, sterols, glutamate, other amino acids, purines, etc…
exit of intermediates from citric acid cycle
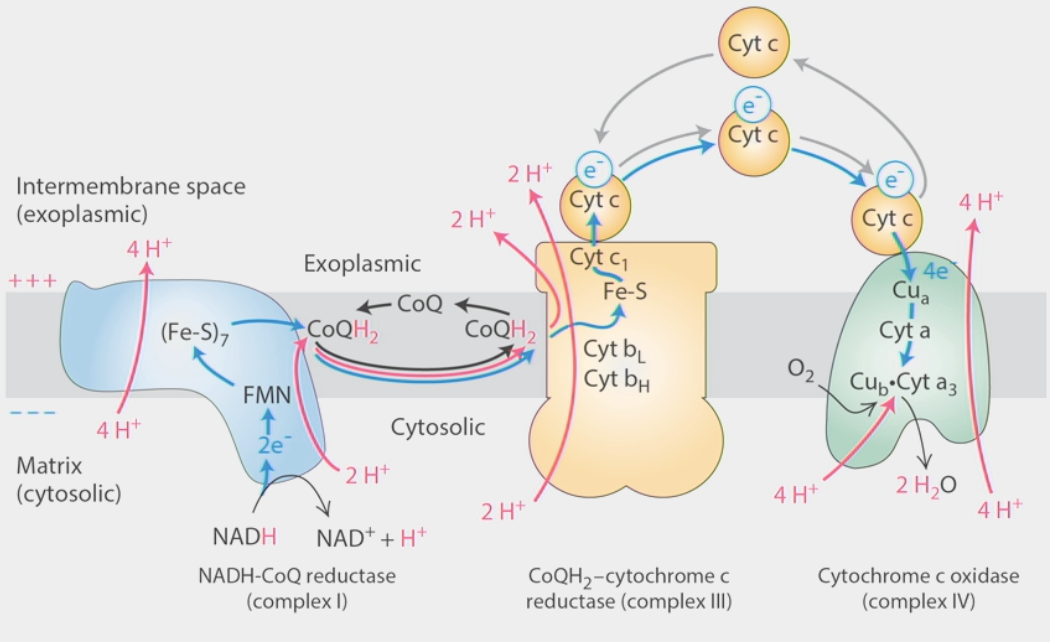
Electron Transport Chain
Stage III of metabolism
converts NADH reduction into a proton gradient
intermembrane space → positively charged
cytosolic matrix → negatively charged
iron clusters & other “prosthetic groups” pass e- downhill
decentralized electron clouds
Hemes, Fe-S clusters
Complex I: NADH-CoQ reductase
NADH → NAD+ + H+ sends 2 e- in
e- pass through Fe-S clusters (different e- comes out each time)
4H+ move against concentration gradient SEPARATE from e- transfer → transverse helix pump
e- passed to ubiquinone (CoQ) → lipid-like carrier
2H+ + 2e- + CoQ → CoQH2
CoQH2 transfers e- to complex III
Complex II: from citric acid cycle (succinate-CoQ reductase)
transfers e- to CoQ from succinate
NOT near other complexes!!!
Complex III: CoQH2-cytochrome c
e- transferred by CoQH2 to cytochrome C
2 e- hit Q0 site →1 goes up to Cyt c, other goes down, interacts w/ Qi site
Cyt c dissociates, e- released, new Cyt c binds to complex
unloaded CoQ leaves, new CoQH2 comes in → 2nd e- that goes down joins the 1st one in Qi site, makes another CoQH2 w/ 2H+
TOTAL: 4e- IN, 2H+ IN, 2CoQH2 IN → 2e- OUT (Cyt c), 1 CoQH2 OUT, 4H+ pumped ACROSS
Complex IV: Cytochrome c oxidase
passes e- from Cytochrome C → O2
e- pass down thru prosthetic groups (e- carriers), meet with O2 + 4H+ → 2H2O
1e- per Cyt C, 4 needed to produce 2 H2O out of 1 O2
4H+ also pumped across gradient
activity level measured w/ changes in pH
without O2 → TCA cycle & ETC STOP b/c NAD+ is depleted
ΔGº’ = 52.6 kcal/mol
electron carrier
iron clusters and other groups that pass electrons downhill
Hemes, Fe-S clusters (ex. in Complex I of ETC)
have decentralized electron clouds
reduction potential (readiness to gain e-) increases down the chain
ΔG decreases
electrons moved to higher reduction potential groups all the way to O2 in ETC
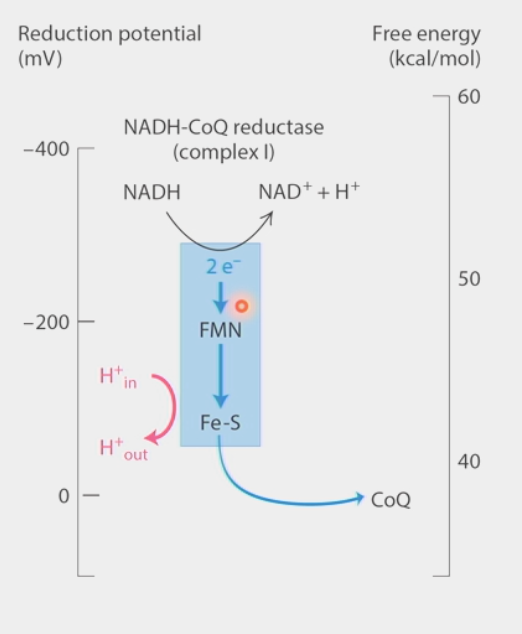
proton translocation
in Complex I of ETC
4 separate gated proton channels for 4H+ moving against concentration gradient
separate from electron transfer
channels have residues with regulated pKa
Ka = acid dissociation constant (affinity for protons)
pKa = -log10 Ka
lysine residues in Complex I bind and release protons in response to CoQ reduction
gate = 2 Lys connected to half-channels in membrane
transverse helix (t-helix) couples changes at CoQ site to conformational changes in pKa of Lys
sliding left: CoQ oxidized, H+ caught → right: CoQ reduced, H+ dropped → repeat
PUMP motion
300x a second!!
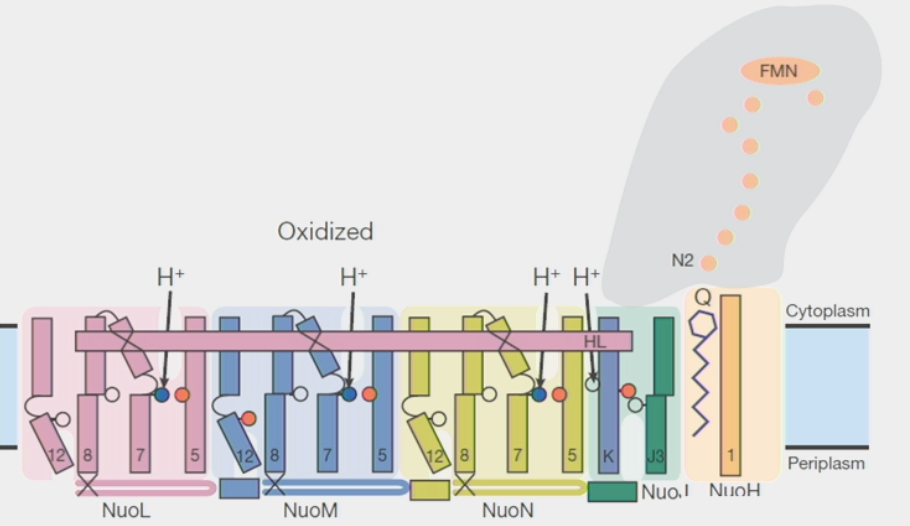
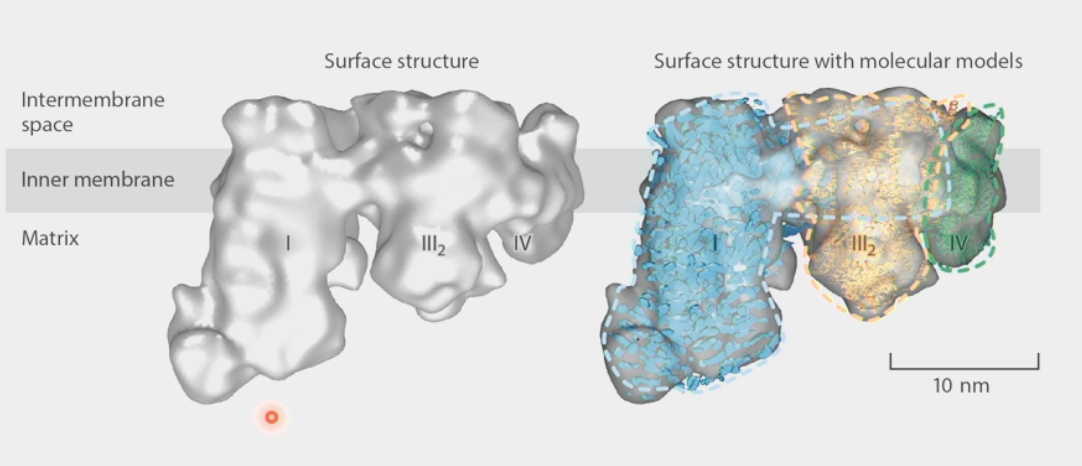
supramolecular assemblies
parts of the structure are held together by very strong interactions, but not necessarily by covalent bonds
ex. Electron Transport Chain → complexes are adjacent, molecules do not have to diffuse far
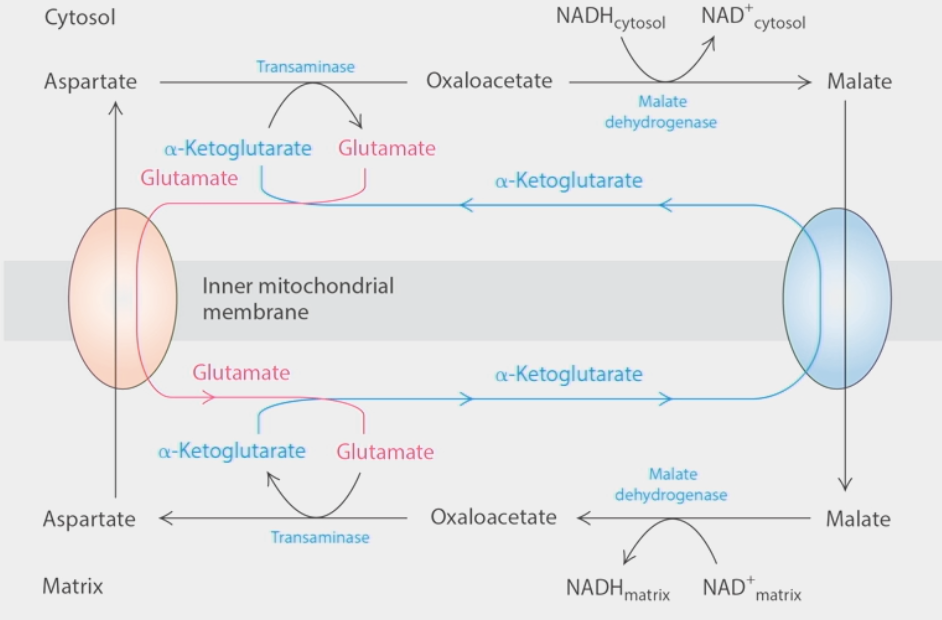
electron shuttle
how cells regenerate cytoplasmic NAD+ pools
2 transport molecules → transport intermediate metabolites (malate & aspartate) across inner mitochondrial membrane in opposite directions
malate: cytosol → matrix
aspartate: matrix→ cytosol
enzymes couple creation of malate from oxaloacetate to oxidation of NADH → NAD+
malate travels across membrane → is reduced back to oxaloacetate → NAD+ becomes NADH again
oxaloacetate created from aspartate by transaminase in cytosol, turned back into aspartate in matrix
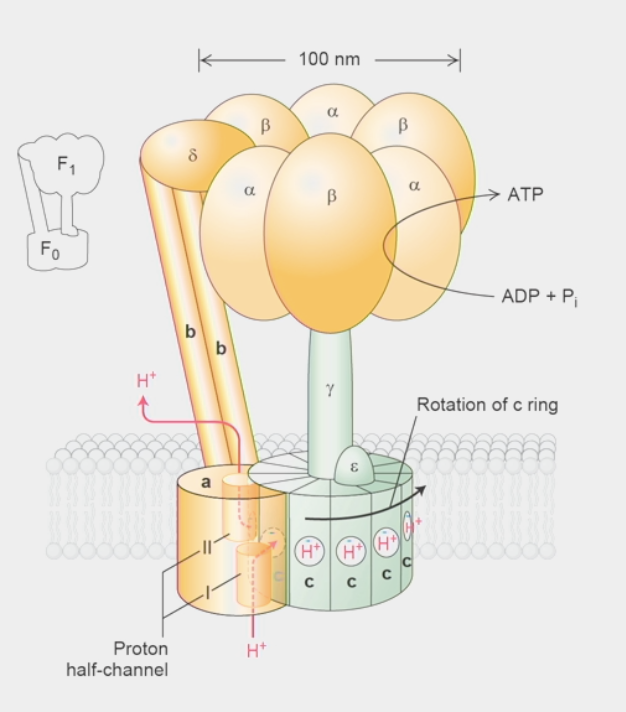
ATP synthase
molecule that harnesses proton-motive force from inner mitochondrial membrane to make ATP
H+ want to flow from high concentration → low concentration side: energy can be harvested!
Split into Fragment 0 & 1 (F0 & F1)
F0:
subunit a → proton half-channels I & II
H+ flow up through half-channels
c ring → made of 10-14 identical c subunits
H+ flow from half-channel I → bind to negatively-charged residue in c subunit
ROTATES 360º until H+ reaches half-channel II → flows up & out
rotation drives enzymatic activity in F1 → ATP synthesis
stator → made of 2 b subunits & δ subunit
holds ⍺ & β subunits in the head stationary while c ring rotates
F1:
𝛾 subunit → non-symmetric shape, rotates!!
sticks up from c ring to head group
rotation physically pushes β subunits thru 3-stage cycle of reaction where they change conformations
130 rotations per second = 1000 H+ = 300 ATP per synthase per second!
β subunits → 3 configurations
O config = OPEN → ADP + Pi can pop IN and OUT
L config = LOOSE → ADP + Pi trapped but NON-reactive
T config = TIGHT → ADP + Pi converted to ATP (REVERSIBLE)
12 H+ needed for 360º rotation = 3 ATP
ancient innovation → same structure found in bacteria, yeast, chloroplasts, etc.
can run backwards to generate proton gradient FROM ATP
ΔGº’ = 3 × 7.3 kcal/mol (50% efficient from ETC)
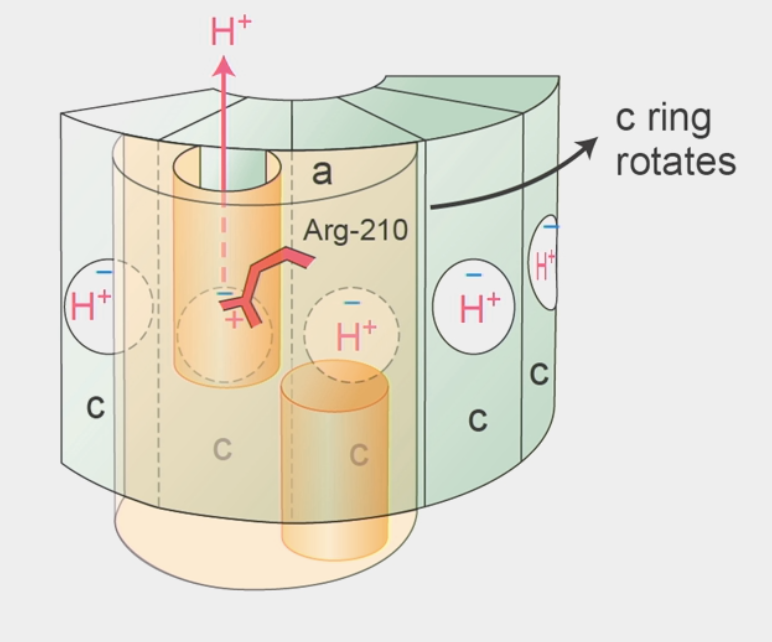
c ring rotation
a subunit has Arg-210 residue (like Lys, can be protonated/deprotonated)
carries positive charge → interacts w/ negative residue in a c subunit
proton comes up through half-channel I → binds w/ same negative residue and displaces Arg-210 to next c subunit
Arg-210 undergoes conformational change → kicks out fully rotated H+ in next subunit to bind w/ negative residue
H+ leaves out of half-channel II
c ring rotates always in ONE direction!!
Arg-210 back in original position → REPEAT process
“Brownian Ratchet” → rotation driven by HEAT/random thermal collisions
must be “rectified” to avoid breaking 2nd Law by H+ concentrations on sides of membrane
Arg-210 = pawl of ratchet
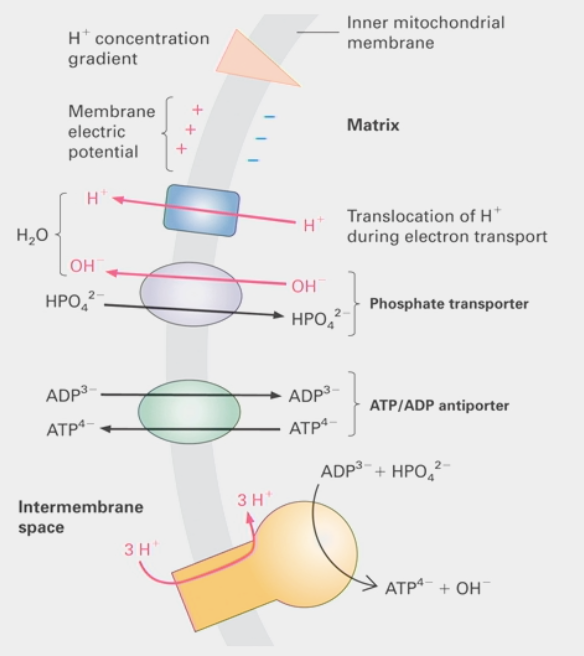
ATP/ADP antiporter
integral membrane protein that uses secondary active transport
uses energetically favorable movement of one molecule down its electrochemical gradient
how ATP escapes mitochondrial matrix
lets ATP out while letting ADP in
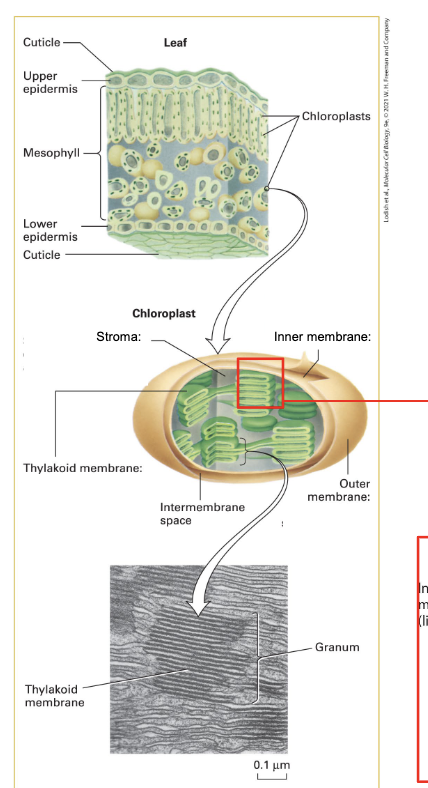
chloroplast
double membrane organelle
outer membrane = highly permeable to small molecules
inner membrane = less permeable, requires transporters
intermembrane space = stroma (like cytoplasm)
stroma contains thylakoids (membranous structure) → form stacks called grana
enclose distinct space called lumen
photoelectric effect
emission of electrons from a material caused by electromagnetic radiation
ex. ejection of electrons from a metal when light falls on it
photosynthesis takes advantage of this!
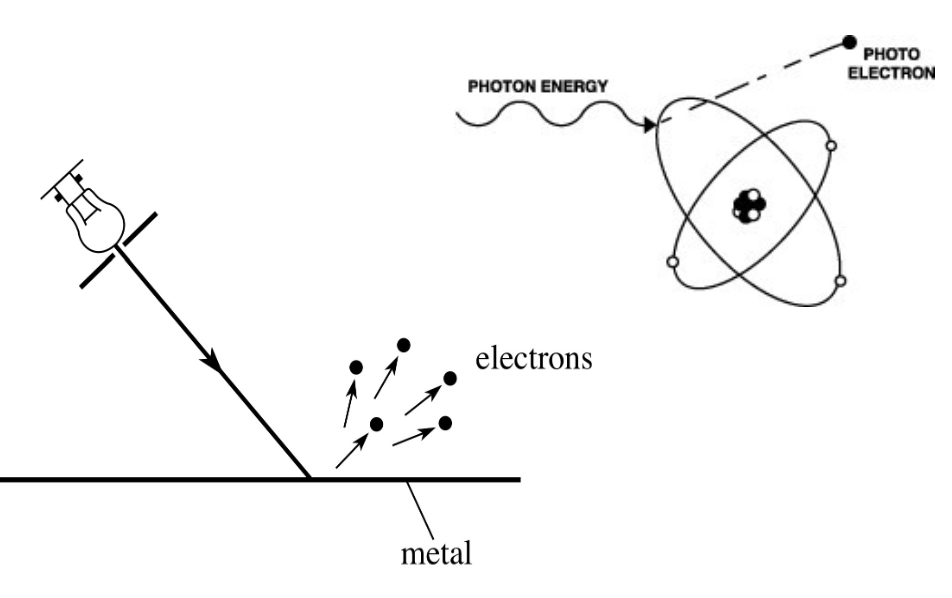
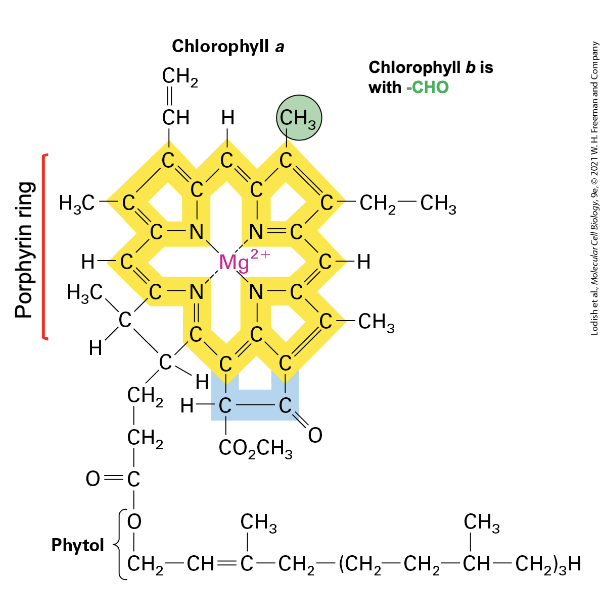
chlorophyll
photosynthetic pigment that absorbs light energy
has porphyrin ring structure → central ion is Mg2+, has hydrophobic tail
extensive system of conjugated double bonds around Mg2+ where e- move
when light (photon) strikes it → excites e- → they delocalize over bonds surrounding Mg2+
excited state is unstable
excitation energy can be lost in 3 ways:
as heat + fluorescence
transferred to neighboring pigments through resonance energy transfer
eject and transfer e- w/ high energy to a nearby e- acceptor → ground state achieved by acquisition of low energy e- from nearby e- donor
look green because mainly blue and red wavelengths of visible light are absorbed while green is reflected
largely found in thylakoid membrane in photosystems
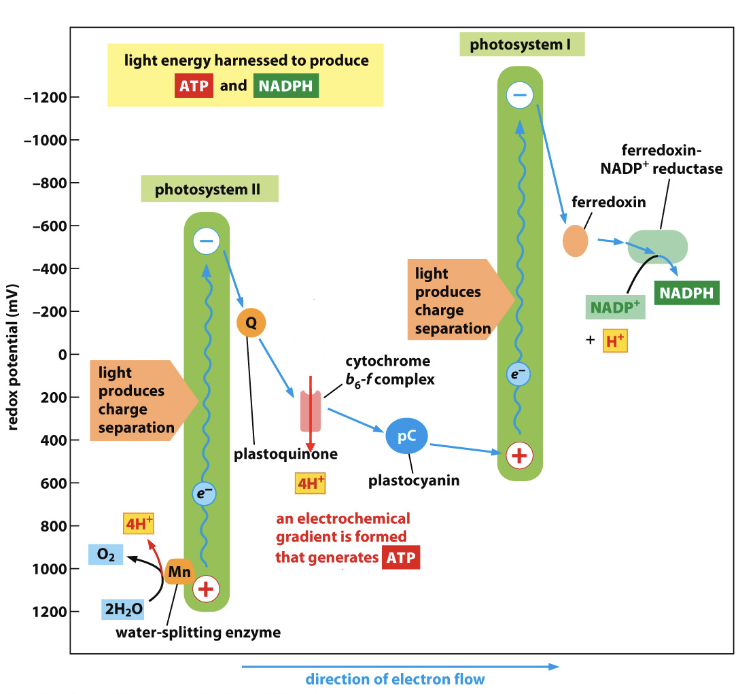
photosystems
where light reactions occur during photosynthesis
2 of them are coupled in thylakoid membrane → light harvested twice
Z scheme of redox potential
evolved from cyanobacteria (produced first O2 in atmosphere = Great Oxidation Event) → algae → plants
has a Light Harvesting Complex (LHC) → consists of hundreds of bridging chlorophylls
has a reaction center w/ special pair of chlorophylls
light energy captured by chlorophylls in LHC → funneled to special pair in reaction center
capture light of different wavelengths → absorbed energy transmits one-by-one thru resonance energy transfer
energy excites e- → ejects from special chlorophyll in reaction center to e- carrier
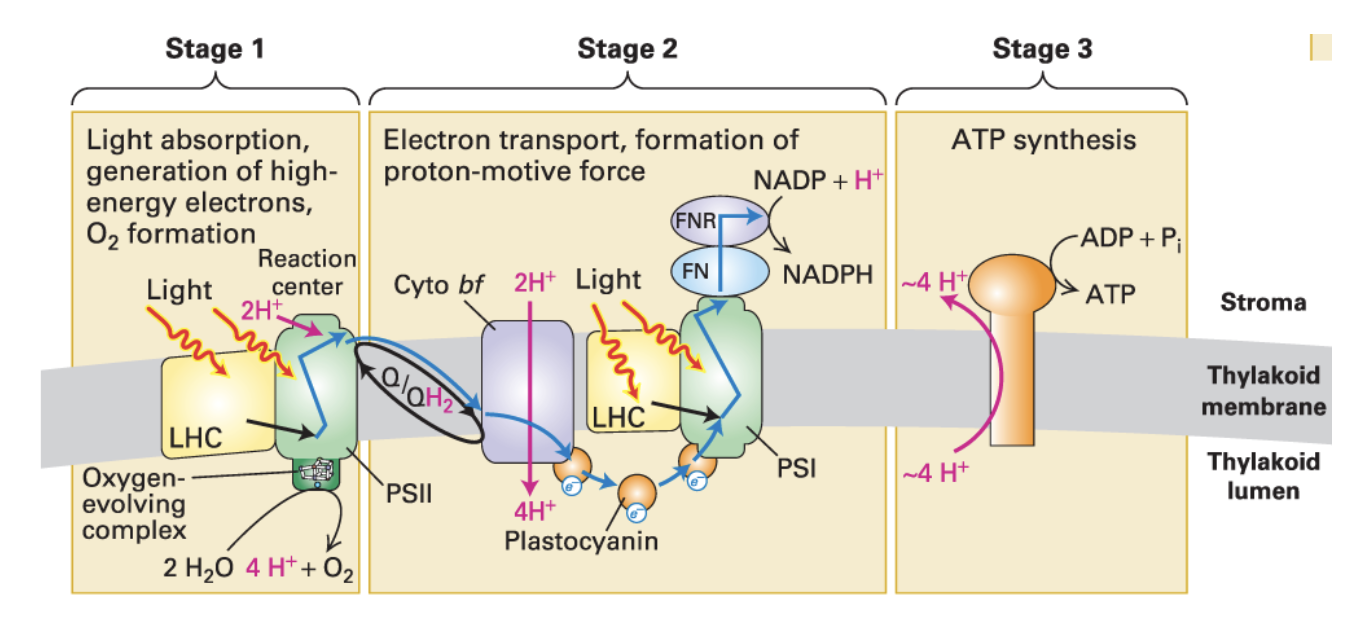
light reactions
3 major complexes: PSII, Cyto bf, PSI
5 electron carriers: H2O, plastoquinone (Q), plastocyanin (pC), ferredoxin (Fn,) NADPH'
2 key enzymes: Water-splitting complex (O2-evolving), ferredoxin-NADP+ reductase (FNR)
Stage 1:
Light absorption, generation of high-energy e-, O2 formation
Stage 2:
During e- transport, H+ are pumped into thylakoid membrane → proton-motive force formed
Stage 3:
ATP synthesis
Photosystem II
discovered second, but comes first
light energy funneled from LHCs → reaction center ejects excited electron from P680 chlorophyll → transferred to Q
Q picks up 2H+ and is reduced to QH2
ionized P680 extracts 4e- from water-splitting complex, one at a time from 2H2O → oxidized to form O2 → released into lumen → atmosphere (all O2 has been generated this way)
4H+ also released to lumen → contribute to proton gradient across thylakoid membrane → produces ATP
NET: 2H2O → 4H+ + 4e- + O2
P680 = ONLY molecule on Earth that can extract e- from H2O
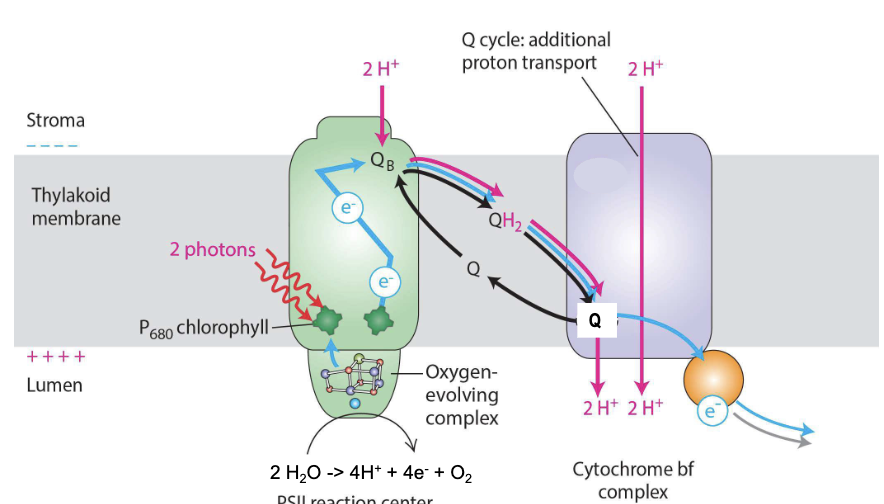
Cytochrome bf complex
functional equivalent of cytochrome c reductase (Complex III) in mitochondria
Reduced QH2 passes its 2e- to plastocyanin (pC) in 2 steps (Q cycle)
increases efficiency of proton pumping from stroma to lumen for ATP synthesis
NET: 2QH2 + 2pCoxidized + 2H+stroma → QH2 + Q + 2pCreduced + 4H+lumen
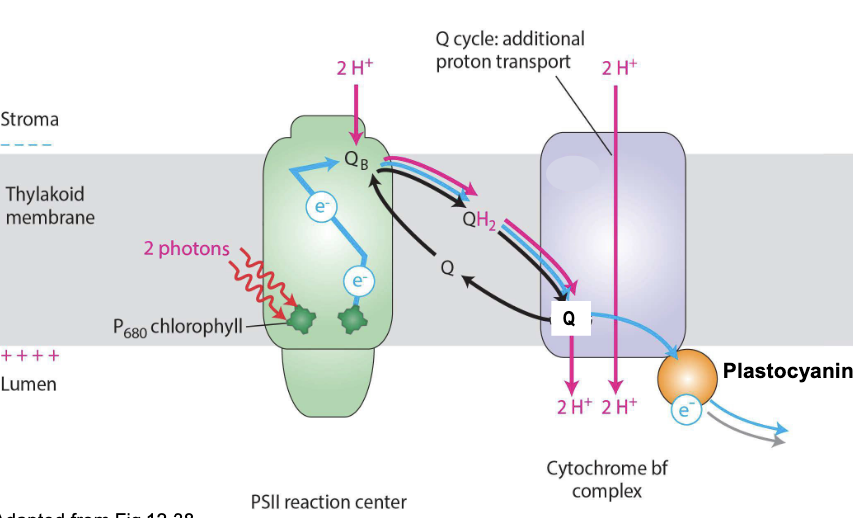
Photosystem I
discovered first, comes last
pC passes e- to P700 (special pair chlorophyll) → energy of e- boosted by light energy (photons)
boosted e- move within reaction center → ferredoxin (Fn)
Fn transfers e- w/ high energy → NADP+, which picks H+ to form NADPH
requires action of ferredoxin-NADP+ reductase (FNR)
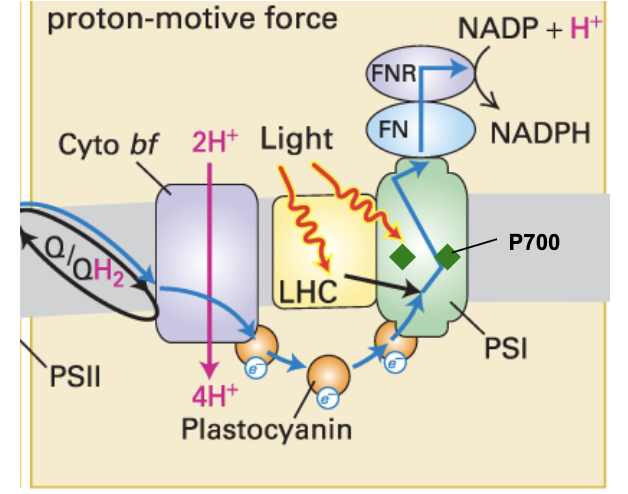
cyanobacteria
PSII with water-splitting complex + PSI
extract and transfer e- from H2O to make ATP & NADPH for CO2 fixation into sugar
produced the oxygen that fundamentally transformed Earth's atmosphere → Great Oxidation Event ~ 2.0-2.5 bya
then transferred by endosymbiosis → algae → plants
3 key innovations:
P680 chlorophyll in PSII → when oxidized has greater affinity for e-
Special H2O splitting complex in PSII
2 light-harvesting photosystems in tandem
RuBisCO
constitutes >50% of total chloroplast proteins → most abundant on earth!!
8 large and 8 small subunits
active in presence of CO2, Mg2+, light
when light intensity increases, Rca (Rubisco activase) removes inhibitor → promotes conformational change in ATP-dependent manner to activate
action of Rca is under regulation of Thioredoxin (Tx) in light-dependent manner
catalyzes CO2 fixation: CO2 (1C) + RuBP (5C) = Intermediate (6C) → 2x 3-Phosphoglycerate (3C)
happens in chloroplast stroma
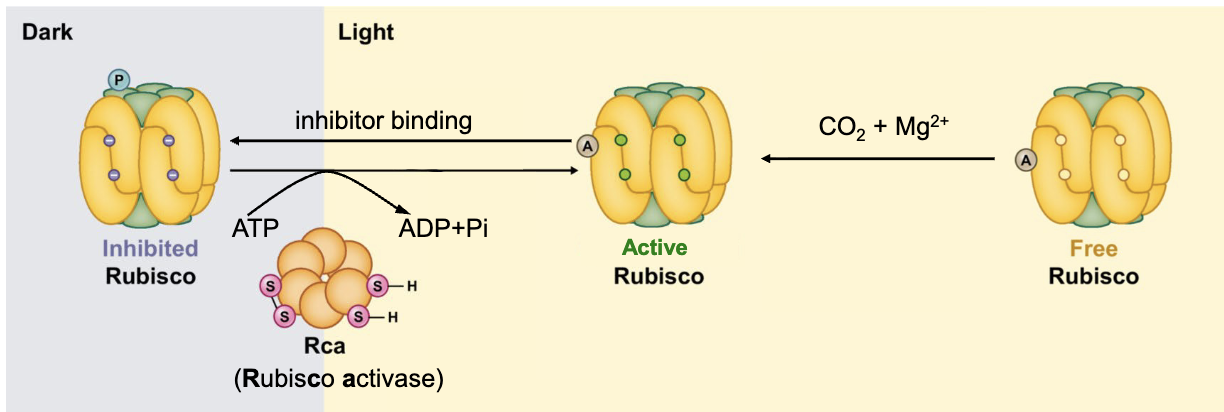
Calvin Cycle
fixes CO2 → glyceraldehyde-3-phosphate (G3P)
6 CO2 → 12 PGA (3C) → 12 ATP used to make 12 1,3-bisphosphoglycerate (3C) → 12 NADPH used to make 12 G3P (3C)
2 G3P used to produce sucrose
10 G3P recycled → interact w/ 7 enzymes → 6 Ribulose 3-phosphate (5C) → 6 ATP used to convert to 6 RuBP (5C) → REPEAT!
18 ATP TOTAL, 12 NADPH TOTAL NEEDED
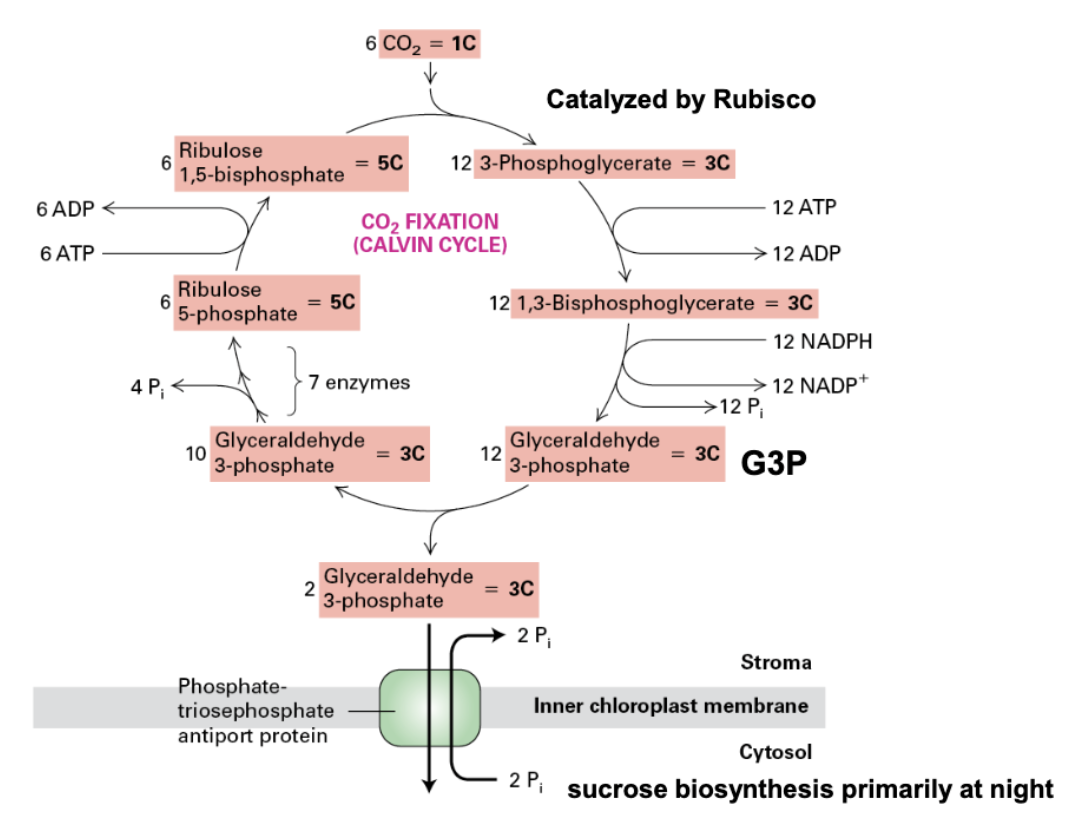
sucrose production
converted from G3P primary at night (nocturnally)
basically the reverse of glycolysis
2 G3P (3C) → 1 Fructose 1,6-bisphosphate (6C) x2
1 of of the fructose molecules stays as is
other fructose → glucose 1-phosphate (6C) → UDP-glucose (UTP needed)
UDP-glucose + Fructose 6-phosphate = 1 sucrose 6-phosphate (12C), UDP OUT → Sucrose (12C)
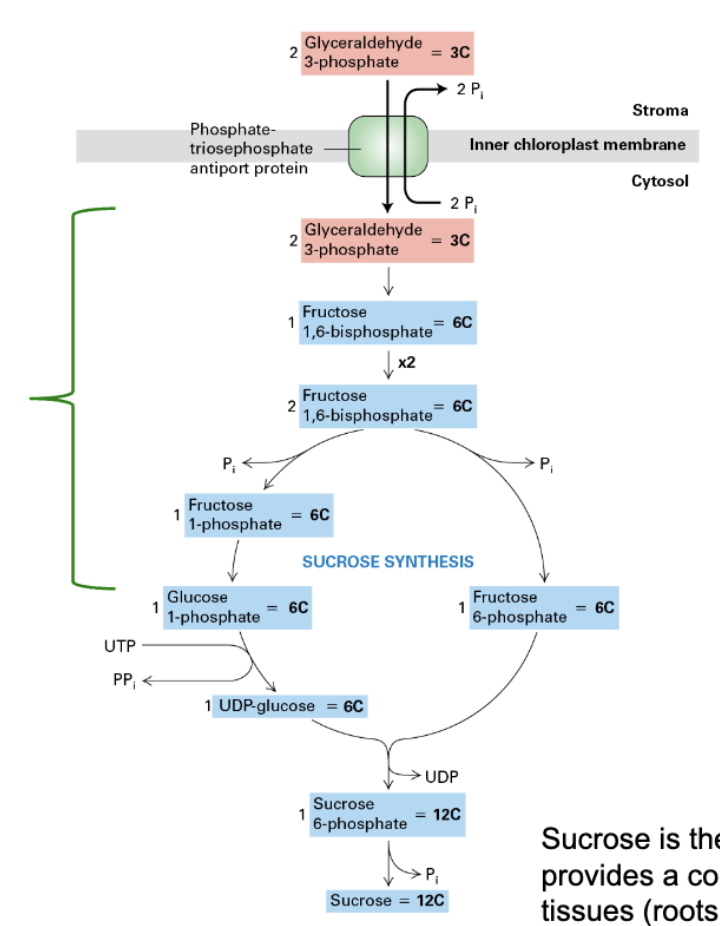
sucrose
main sugar transported within plants
provides constant energy supply to non-photosynthetic tissues (roots, fruits, etc.)
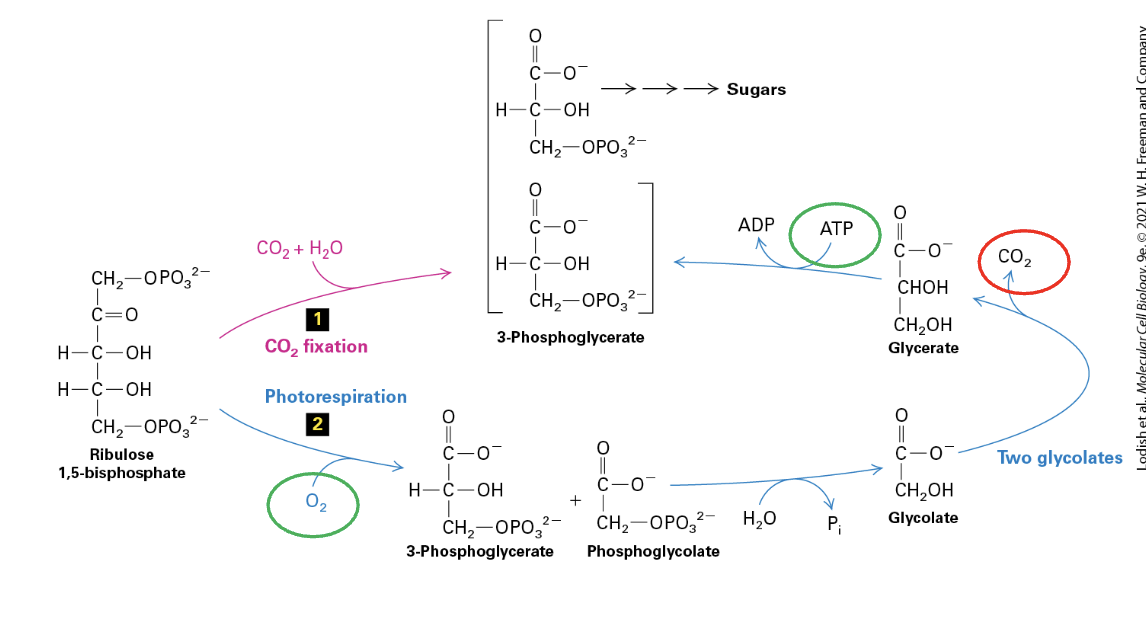
photorespiration
catalyzed by RuBisCO (also an oxygenase) when CO2 is low
O2 binds to RuBP → PGA + Phosphoglycolate + H2O → Glycolate x2 → CO2 OUT, Glycerate produced → ATP used up to convert back to PGA
produces toxic 2-phosphoglycolate that needs to be converted back to PGA (a COSTLY process)
C3 plants
major plants that fix CO2 in all mesophyll cells of leaves
no special features to combat photorespiration
not as efficient at fixation
ex. rice
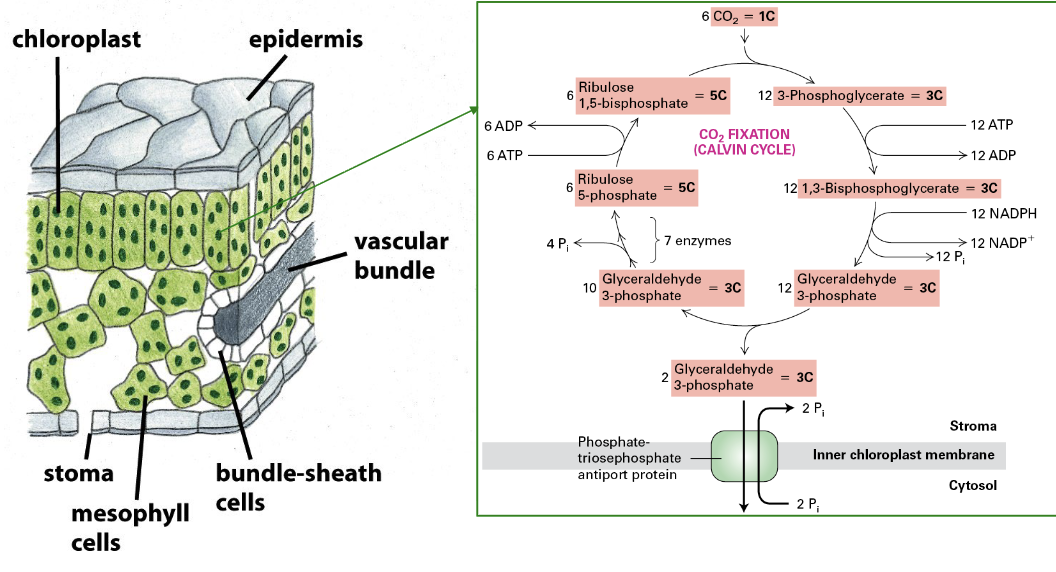
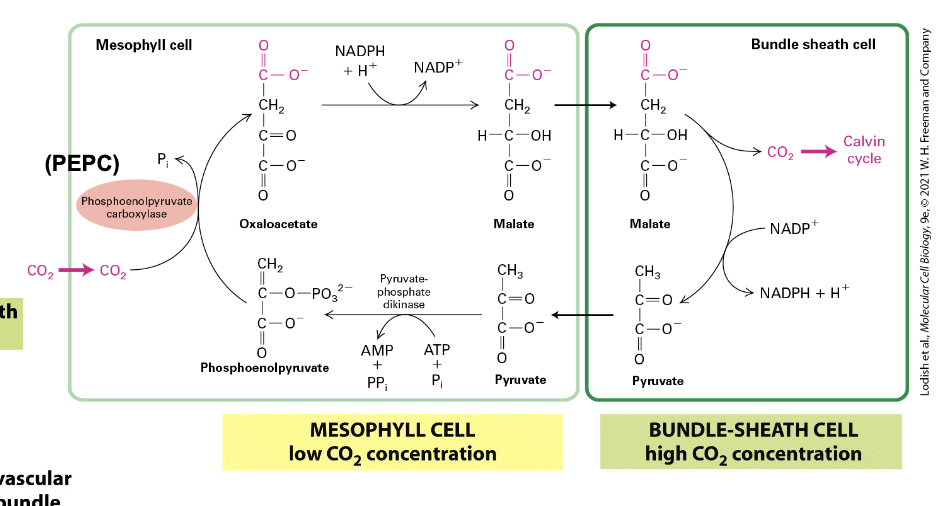
C4 plants
CO2 fixation is compartmentalized in these plants
ex. maize
enhances fixation and reduces photorespiration
PEP carboxylase takes up CO2 → low concentration in mesophyll
used to convert to oxaloacetate → NADPH needed → Malate → bundle-sheath cells
bundle-sheath cells are deeper in leaves → high CO2 concentration → Calvin Cycle
Malate → NADP+ used → Pyruvate → Mesophyll → Phosphoenolpyruvate → + CO2 → Oxaloacetate → REPEAT
projects trying to change C3 crop composition to this to be more efficient
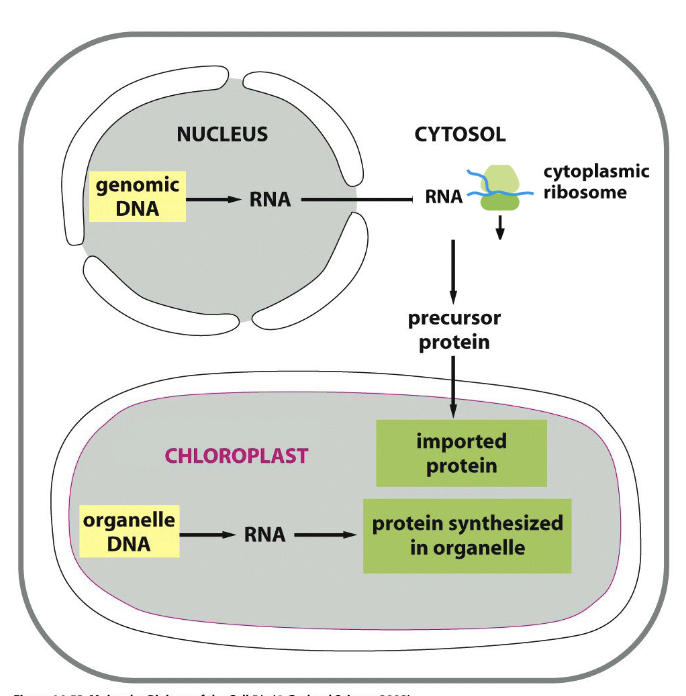
chloroplast genetic systems
most prokaryotic genes were transferred → nucleus during evolution
2 separate systems to make functional chloroplast
~4500 genes → nuclear genome → transcribed in nucleus → translated by cytoplasmic ribosomes → imported in chloroplasts
~90 protein coding genes still maintained in chloroplasts → transcribed & translated WITHIN organelle
>300 chloroplast genomes sequenced
all contain ~45x rRNAs + tRNAs and ~90x protein-coding genes → encode proteins found in PSI, PSII, Cyto bf, ATP synthase, RuBisCO, proteins involved in gene transcription & protein translation
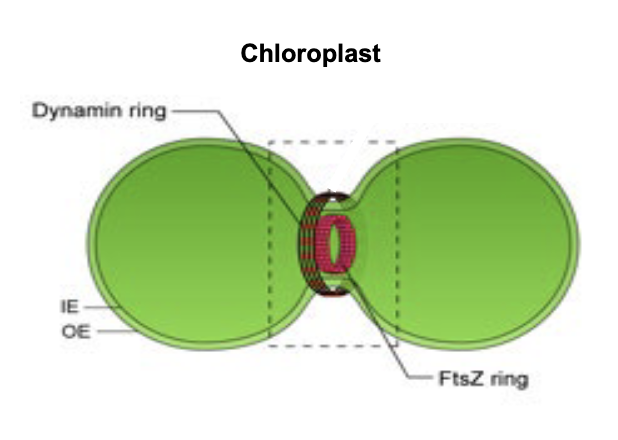
chloroplast fission
FtsZ: tubulin-like protein
self assembles into dynamic ring of protofilaments (Z-ring) beneath inner membrane of chloroplasts
Z-ring acts as scaffold for recruitment of other cell division proteins → generates contractile force → membrane constriction → division
Dynamin: GTPase protein responsible for endocytosis in the eukaryotic cell
comes from cytosol → forms dynamin ring for outer membrane fission
chloroplast fusion unclear → linked w/ tubular channels called stromules
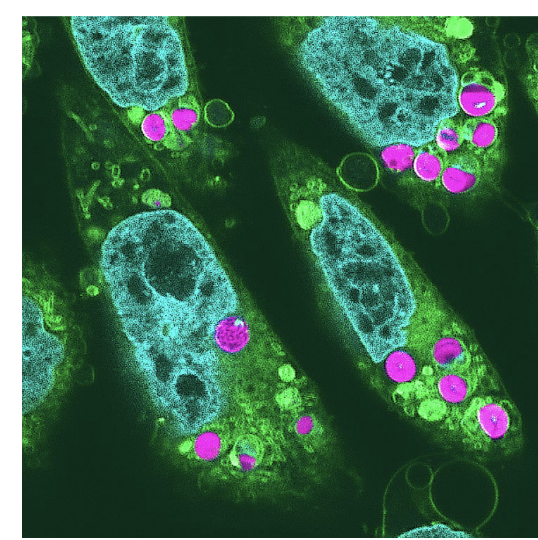
animals that photosynthesize
Elysia timida (sea slug) → has chloroplasts on its back
Scientists created plant-animal hybrids (in hamster cells) w/ chloroplasts
maintained electron transport activity in cultured cells for 2 days after incorporation
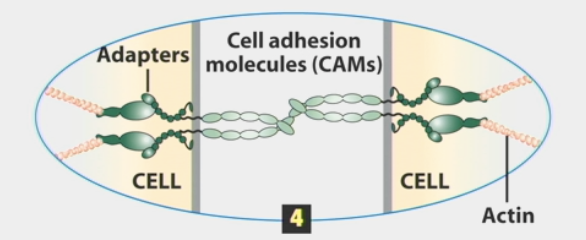
cell-cell adhesion
physical connection
molecules bind to one another and to intracellular proteins
mediated through membrane proteins called cell adhesion molecules (CAMs)
strengthened by adding up many weak interactions
cell alters adhesion by deciding how many interactions are on its surface
sorts out cells of different types into clusters
H.V. Wilson sponge experiments: mixing cells of two different species makes them grow separately, adhere only to cells of their own species
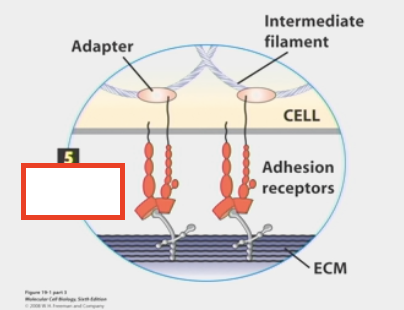
cell-matrix adhesion
cells sticking to a non-membrane surface (the ECM)
adhesion receptors or CAMs stick to ECM and get information from it
ECM released by other cells to send info
cell adhesion molecules
membrane proteins that mediate cell adhesions
cells express a wide, diverse range
many classes (ex. Cadherins, Integrins, Claudin, etc.) that each perform a specific function
each class has many individual molecules that share similar structures
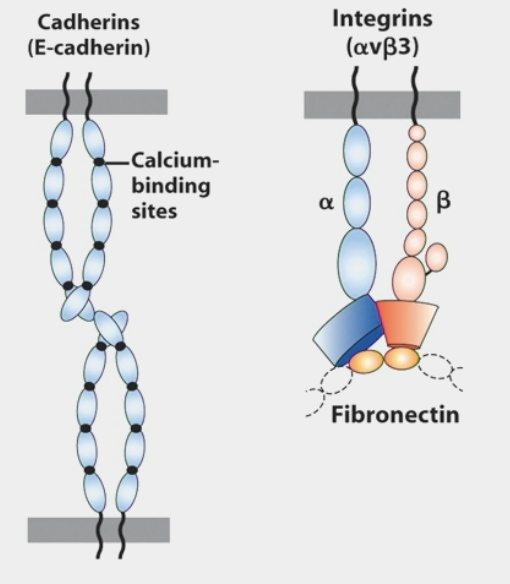
Ex. Cadherins
homotypic interactions: extracellular domain binds to similar extracellular domain of another cell
calcium binding sites
Ex. Integrins
can function as both cell-cell and cell-matrix molecules
homotypic adhesion
adhesive interactions between cells of the same type
heterotypic adhesion
adhesive interactions between cells of different types
trans interactions
intercellular/adhesive
CAMs on one cell bind to the CAMs on an adjacent cell
usually combined with cis interactions
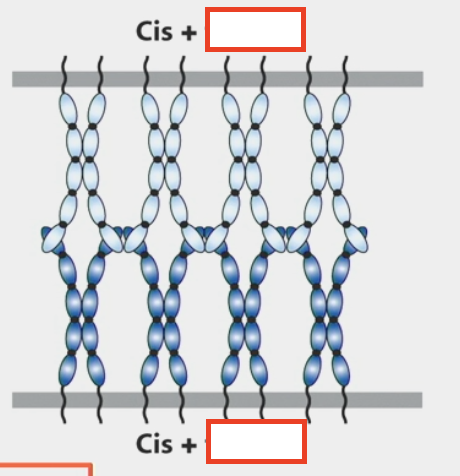
cis interactions
lateral/in the same cell
monomeric CAMs on one cell bind to one or more CAMs in the same cell’s plasma membrane
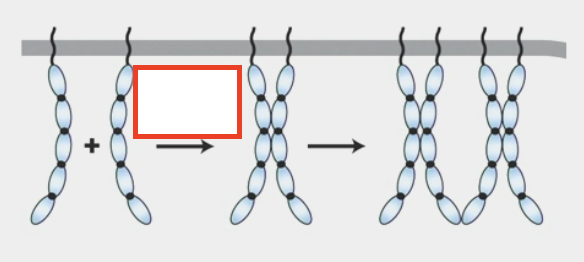
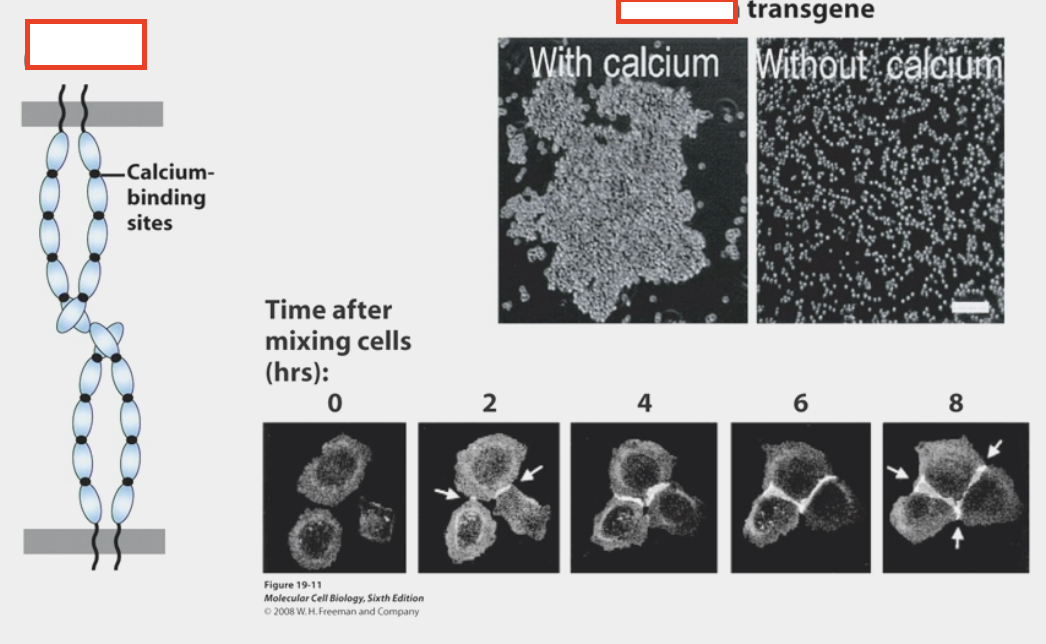
cadherins
calcium-dependent cell-cell adhesion molecules
removing calcium → cells no longer adhere
connect to cytoskeleton via adapter proteins that bind to intracellular domain or to other proteins (ex. Catenins, Vinculin, VASP, ZO1) → eventually bind to actin
adapter proteins
physically link one protein to another protein by binding to both of them
directly / indirectly (via additional adapters) connect cell-adhesion molecules or adhesion receptors to elements of the cytoskeleton OR to intracellular signaling proteins
extracellular matrix
a complex combination of proteins and polysaccharides that is secreted and assembled by cells into a network in which the components bind to one another
often involved in holding cells and tissues together
mostly made up of Type IV collagen fibrils, but also Laminin, Entactin, Perlecan
Connected to intermediate filament cytoskeleton via Laminin
Collagen acts as intermediary between connected cells
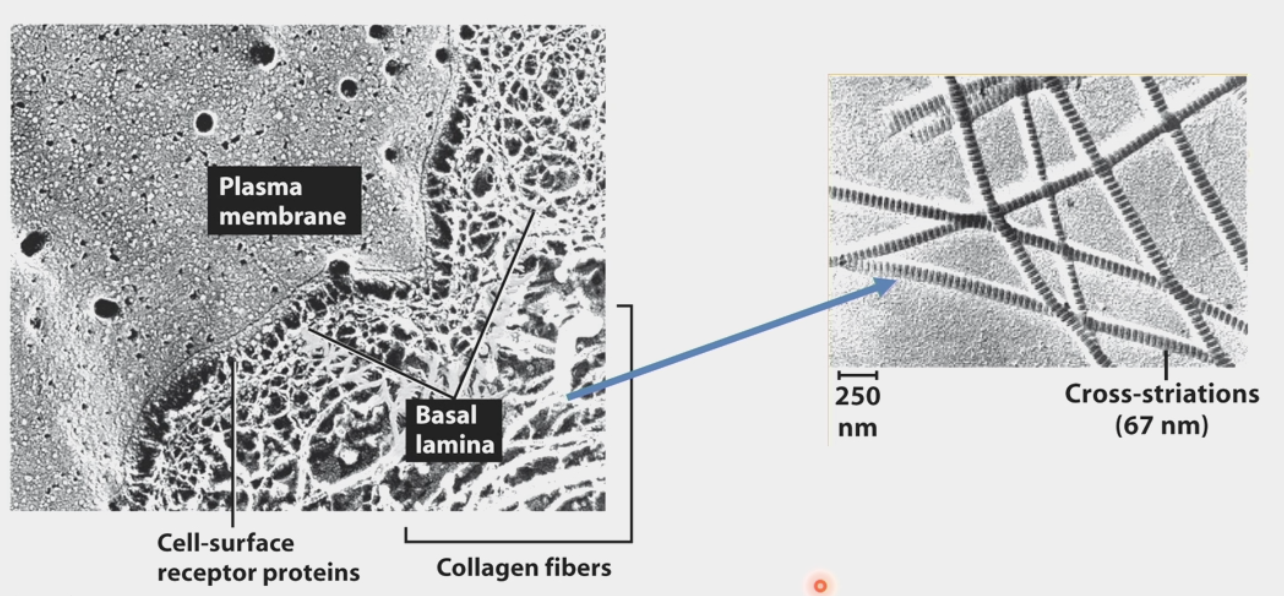
collagen
makes up the ECM (type IV)
Type I, II, III, IV
25% of protein mass of human body
triple-helix structure like a rope
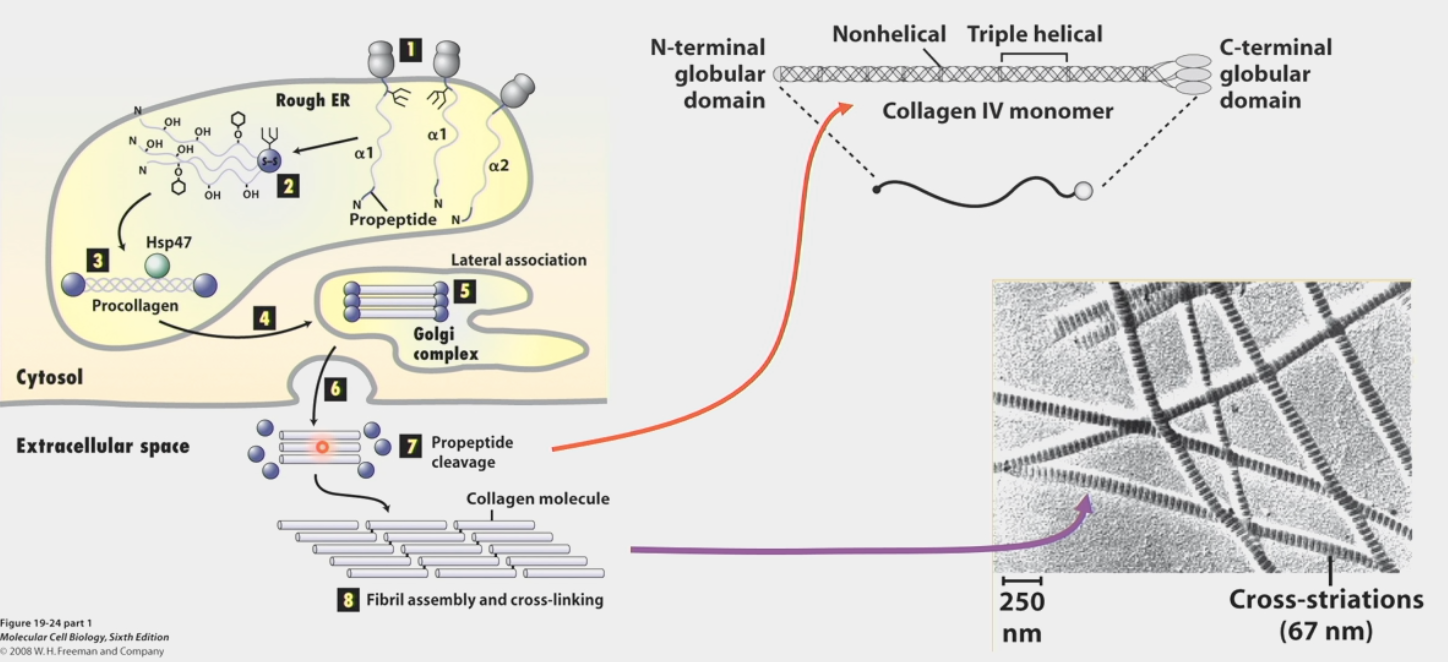
built from propeptides that are cleaved and cross-linked
Protein peptides form from Rough ER
Processed in various ways
3 wind around each other to form procollagen
Transport to Golgi complex
Processing in Golgi; lateral association
Export out of cell
Propeptide cleavage
Higher order fibril self-assembly and cross-linking
sends important information to the cell that it responds to, changes shape or direction of migration
scurvy (vitamin C deficit) defect results indirectly from lack of this
Vitamin C is necessary cofactor for an enzyme that hydroxylates propeptides (strong interactions → higher annealing temp)
heterogeneous polymers
characteristic of Collagen IV networks
globular domains at the ends of triple helices bind to other collagen molecules in specific ways
C-terminal end forms dimer w/ C-terminal end of another collagen
N-terminal end forms tetramer w/ 3 other N-terminal ends
combine in endless series to get network of collagen that attracts other ECM components
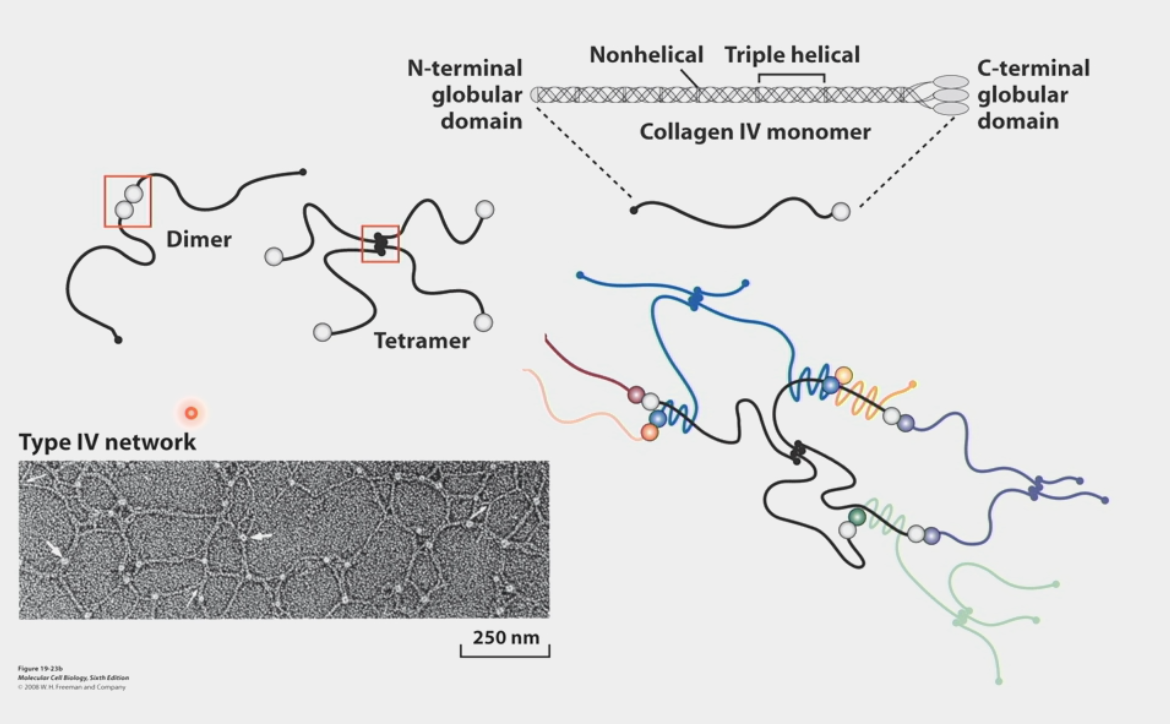
laminin
multi-adhesive ECM protein
triple-helix and cross structure
has different globular domains from various peptides that can either attach to each other heterogeneously like Type IV collagen or attach to other trans-membrane proteins (integrins, collagen, other cellular receptors)
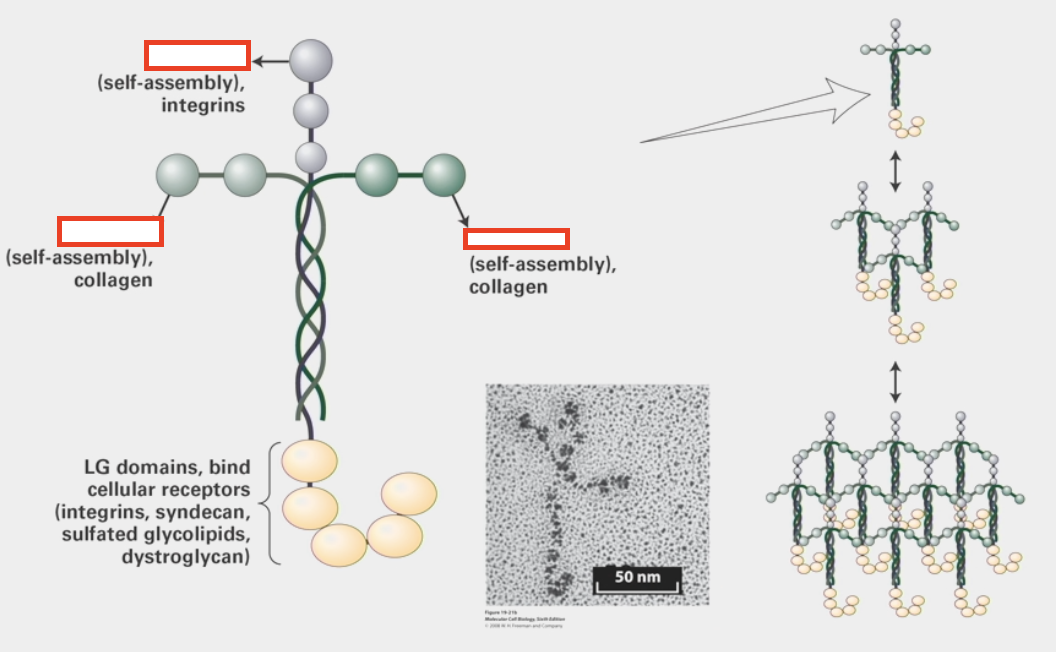
integrins
cell-matrix adhesion molecules
straighten upon activation
cell controls ability of laminin binding to them
Talin signal binds to intracellular domain, Kindlin binds to Talin
Activates extended conformation change that is able to bind to laminin
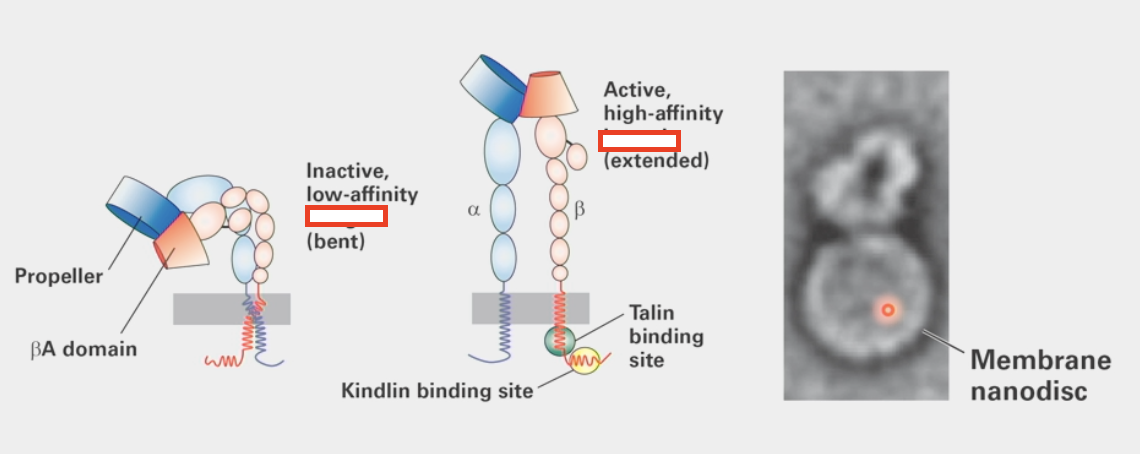
desmosome
a trans-membrane structure by which two adjacent cells are attached
formed from protein plaques in the cell membranes linked by intermediate filament cytoskeleton
forms trans-cellular network of “cables”
disruption compromises epithelial tissue integrity
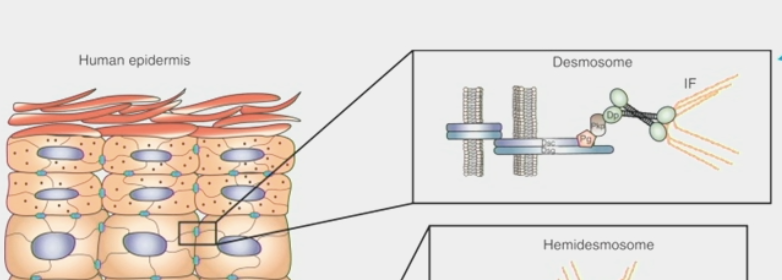
focal adhesion complexes
integrins cluster into these rather than being uniformly spread
act like the “feet” of the cell → important for cell movement
attach to actin filaments (dynamic force-generating cytoskeleton)
connect to signaling pathways (intracellular) through intermediate proteins that interact w/ other receptors receiving non-adhesion info
have a layered structure
membrane-apposed integrin signalling layer w/ integrin cytoplasmic tails, adhesion kinase, paxillin
intermediate force-transduction layer w/ talin and vinculin
uppermost actin-regulatory layer w/ zyxin, vasodilator-stimulated phosphoprotein, a-actinin
integrin/ECM layer? actin layer?
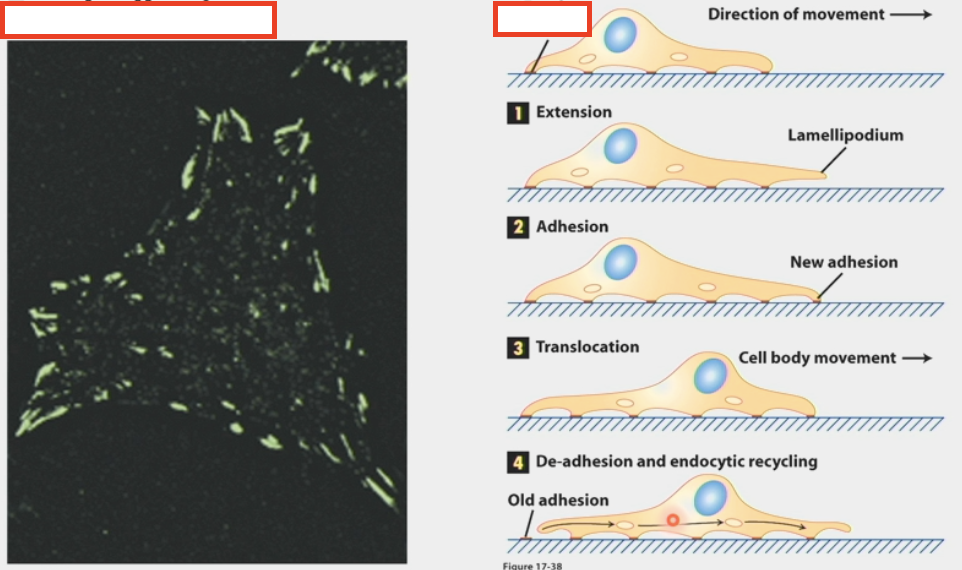
endocrine signaling
type of chemical signaling between cells
endocrine glands secrete hormones that affect distant target cells through blood/circulatory system
ex. epinephrine (adrenaline) secreted by the adrenal glands, insulin secreted by pancreas
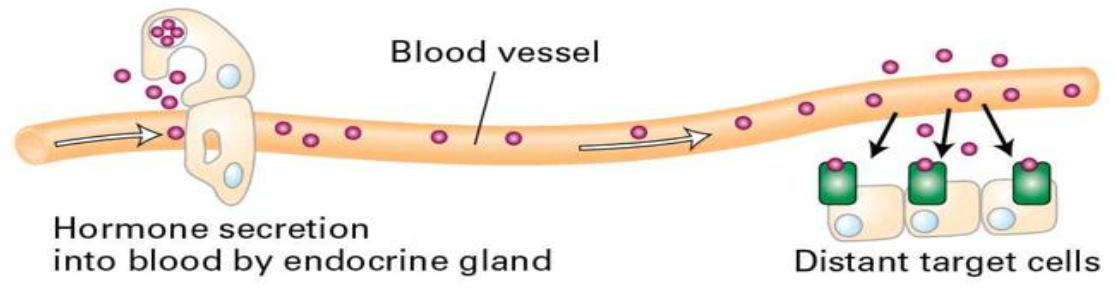
chemical signalling
type of communication between cells
signals are received by cell surface receptors
usually chemicals that are soluble so they can be transported through blood
receptors help signals cross lipid bilayer and transduce to interior of cell
signals change internal state of cell:
modification of cellular metabolism, function, movement
OR modification of gene expression, development
cellular response can be short-term OR long-term
short-term = rapidly executed, rapidly inactivated, NOT “all or nothing”
long-term = time frame is less critical, “decision” is more critical, developmental, signals can determine cell fate
property of ligand binding to receptors according to chemical equilibrium
competitive balance between stimulation and inhibition
ligand binding
binding to receptors according to chemical equilibrium
R + L ←→ RL
R = receptor, L = ligand, RL = ligand bound to receptor
reversible chemical rxn
Kd = ([R][L]) / [RL]
Kd = concentration of unbound [L] at which half of the total molecules of R are associated with L
as ligand concentration increases, there is a higher fraction of surface receptors with a bound ligand
but the physiological response of cells is often more sensitive than anticipated
signalling pathways = series of REVERSIBLE chem rxns → binding drives rxn towards HIGH PKA activity
[GPCR] + [hormone] ←→ kbind[GPCR][hormone] / krelease[GPCR:hormone] ←→ [GPCR:hormone]
![<ul><li><p>binding to receptors according to chemical equilibrium</p></li><li><p>R + L ←→ RL</p><ul><li><p>R = receptor, L = ligand, RL = ligand bound to receptor</p></li><li><p>reversible chemical rxn</p></li></ul></li><li><p>K<sub>d</sub> = ([R][L]) / [RL]</p><ul><li><p>K<sub>d</sub> = concentration of unbound [L] at which <strong>half</strong> of the total molecules of R are associated with L</p></li></ul></li><li><p>as ligand concentration increases, there is a higher fraction of surface receptors with a bound ligand</p><ul><li><p>but the physiological response of cells is often <strong>more sensitive</strong> than anticipated</p></li></ul></li><li><p><strong>signalling pathways</strong> = series of <strong>REVERSIBLE</strong> chem rxns → binding drives rxn towards <strong>HIGH</strong> PKA activity</p><ul><li><p>[GPCR] + [hormone] ←→ k<sub>bind</sub>[GPCR][hormone] / k<sub>release</sub>[GPCR:hormone] ←→ [GPCR:hormone]</p></li></ul></li></ul><p></p>](https://assets.knowt.com/user-attachments/8c9fec2a-3c7e-408e-9b12-1b9ffb729902.png)
adrenaline
hormone that triggers short-term responses
in cardiac muscle: increases contraction
in liver: converts glycogen to glucose for release into bloodstream, inhibits glycogen synthesis
in skeletal muscle: converts glycogen to glucose
used as vasoconstrictive medicine (EpiPen)
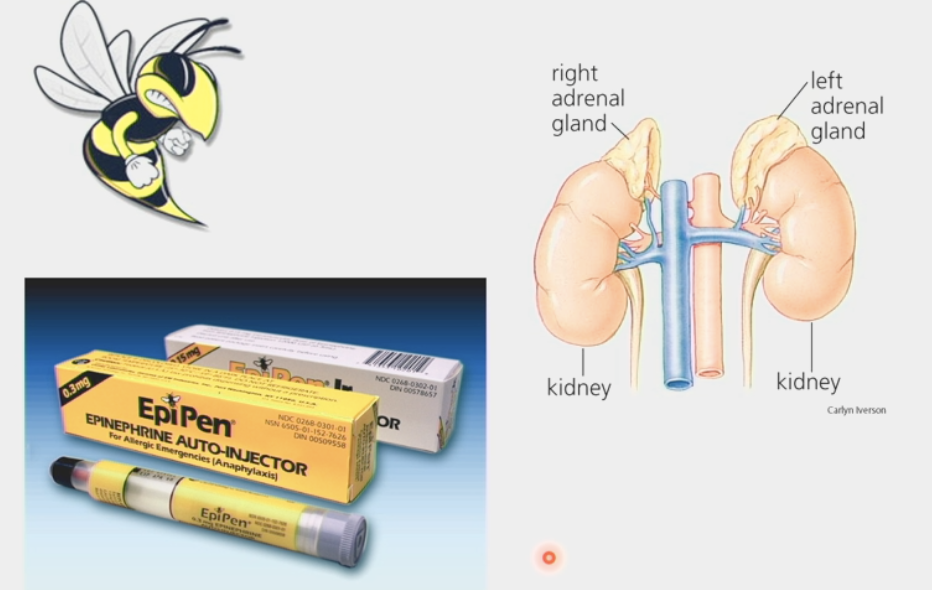
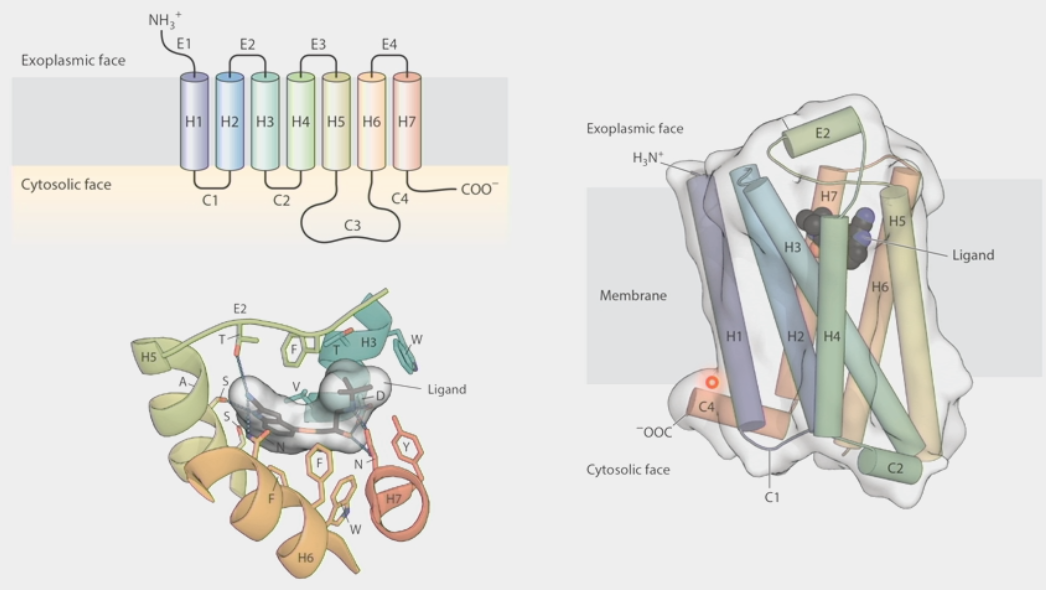
G-protein coupled receptors
large family of receptors that respond to many hormone signals
have 7 trans-membrane domains
loops on exoplasmic face and cytosolic face are specific for a particular ligand to bind on
ligand binding changes overall conformation of receptor → changes ability to bind other proteins
hormone binding recruits the G-protein
binding of hormone induces conformational change in receptor
activated receptor binds to G⍺ subunit
Binding of GTP to G⍺ triggers dissociation of G⍺ both from receptor and from Gβ𝛾
Hormone dissociates from receptor; G⍺ binds to effector molecule in GTP-bound state & activates it
Hydrolysis of GTP → GDP causes G⍺ to dissociate from effector and reassociate with Gβ𝛾
not only for hormones!
subfamily that has different specificities for odorants (smells)
have own specialized Golf G-protein
~400 genes in humans, ~1200 in mice, dogs, etc.
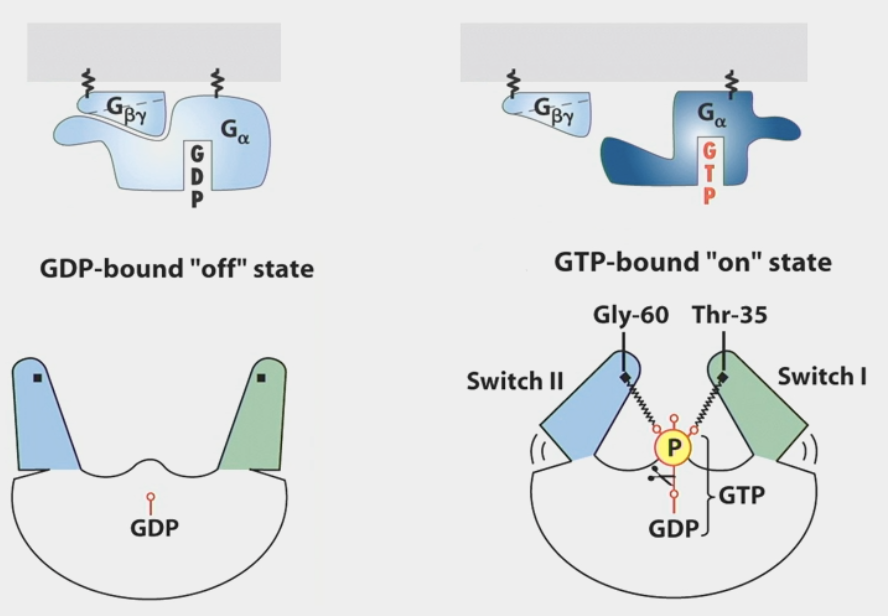
G-proteins
trimeric GTPases that transduce hormone signals
⍺, β, 𝛾 subunits tethered to membrane through attached lipid
GDP-bound “off” state: subunits are close together
GTP-bound “on” state: ⍺ subunit dissociates from other subunits
automatic off switch → subunits have enzymatic activity to convert GTP back to GDP
dissociate within seconds of ligand binding
β𝛾 subunits can also activate effector molecules (ex. K+ channel)
some can inhibit effector molecules while others activate them (competitive balance)
ex. stimulatory hormones epinephrine, glucagon, ACTH v.s. inhibitory hormones PGE1 & adenosine
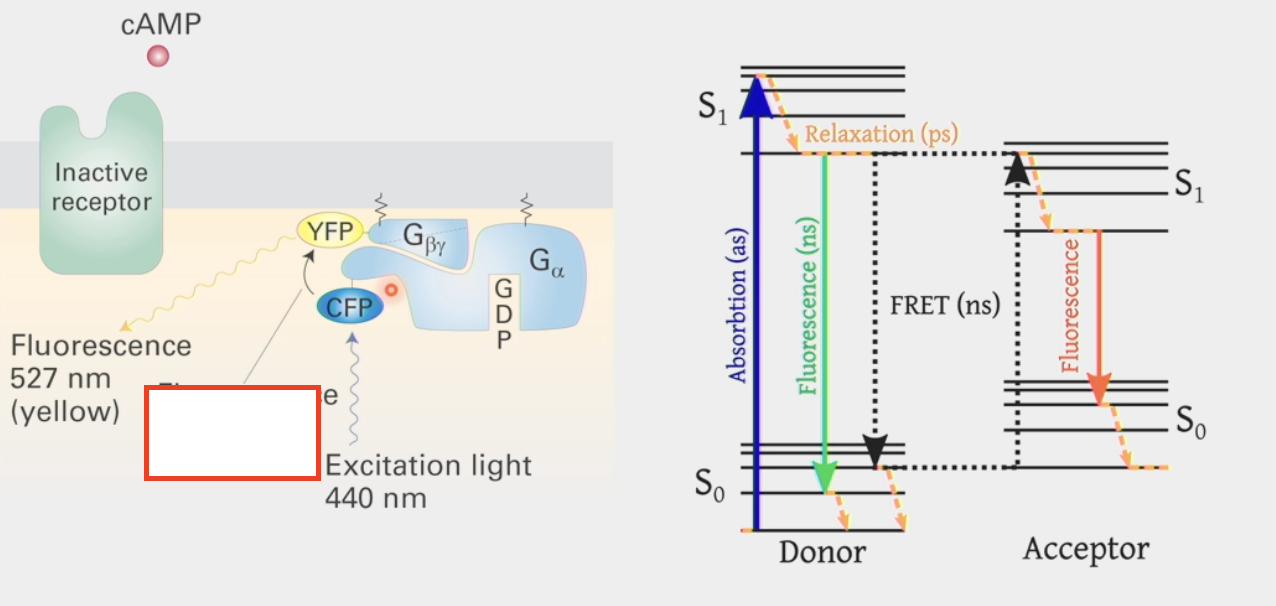
Fluorescence resonance energy transfer
biophysical technique to determine the proximity of two molecules
attach fluorescent molecules w/ different absorption & emission spectra to the two proteins to see if they are touching
shine excitation light on one fluorescent protein and see if it reflects light back or transfers to another fluorophore with an absorption spectrum that is equal to the 1st fluorophore’s emission spectrum (ex. CFP and YFP)
second fluorophore will emit light in a different emission spectrum if in close proximity to 1st fluorophore
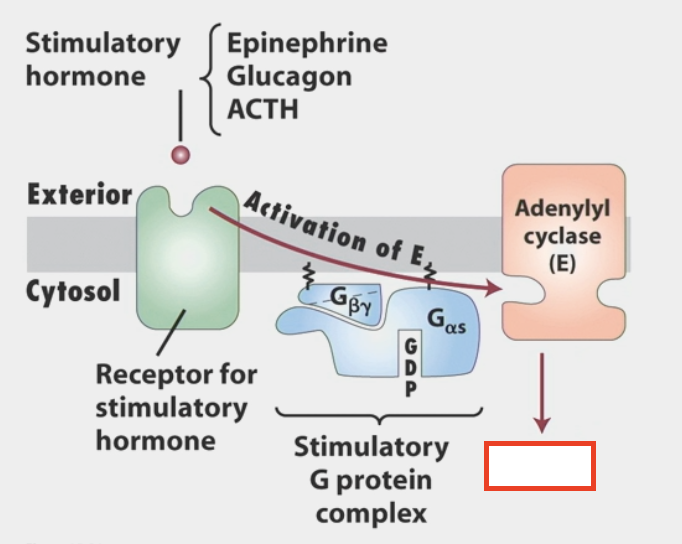
cAMP
made by adenylyl cyclase (common effector of activated G-proteins)
stimulatory hormone can be epinephrine, glucagon, ACTH
second messenger
activates Protein Kinase A (PKA)
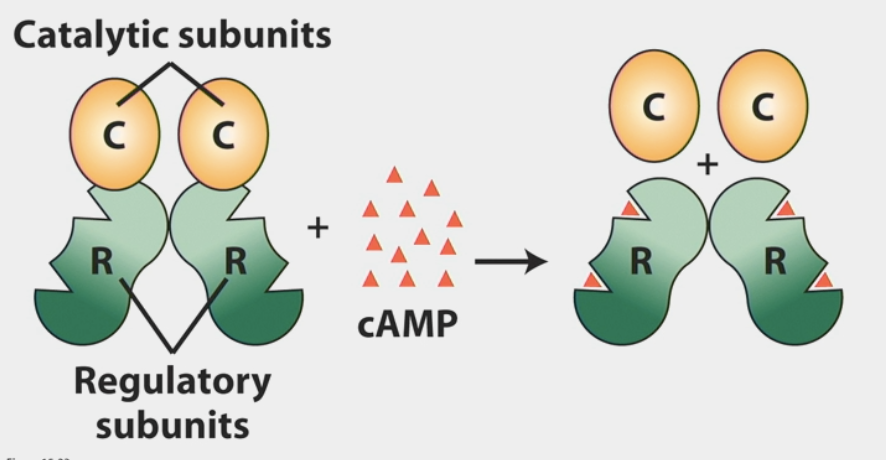
Protein Kinase A
activated by cAMP
a protein that regulates activity of other proteins by covalently adding phosphate groups to them
consists of 4 subunits (2 regulatory, 2 catalytic)
when activated → catalytic sites begin phosphorylating other proteins
directly controls molecules of glycogen metabolism
stimulation of glycogen breakdown
inhibition of glycogen synthesis
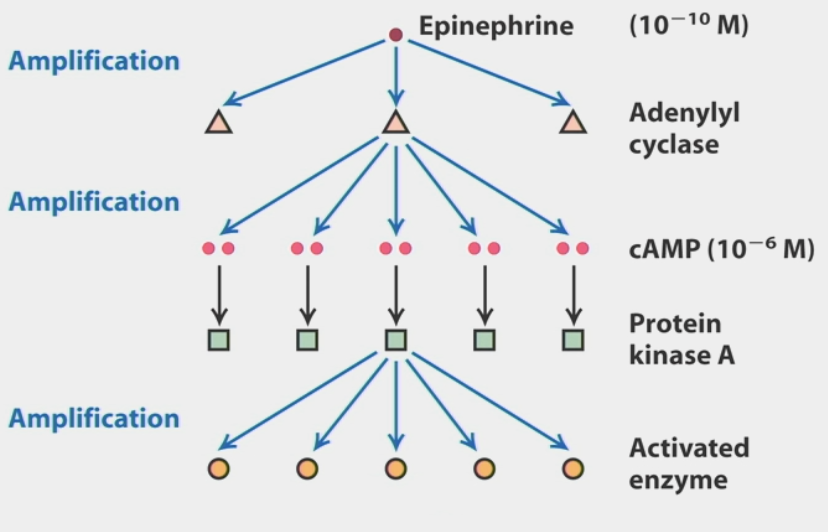
signal amplification
due to multi-step activation
ex. single epinephrine signal (10-10 M) → many adenylyl cyclases → cAMP (10-6 M) → PKA → many activated enzymes → more product
response of cells is more sensitive than anticipated!!
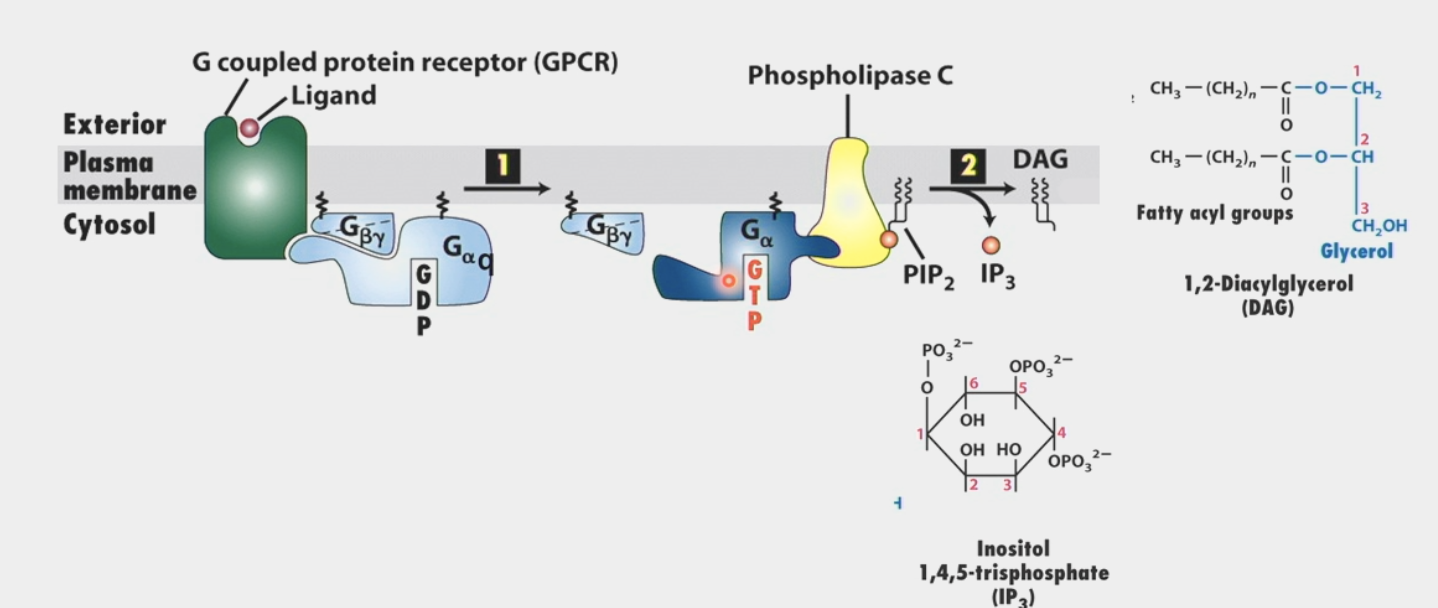
second messengers
common feature of signaling pathways
intracellular signaling molecules released by the cell in response to exposure to extracellular signaling molecules
different effector molecules will release different ones
ex. IP3 and DAG are produced by stimulation of Phospholipase C (effector)
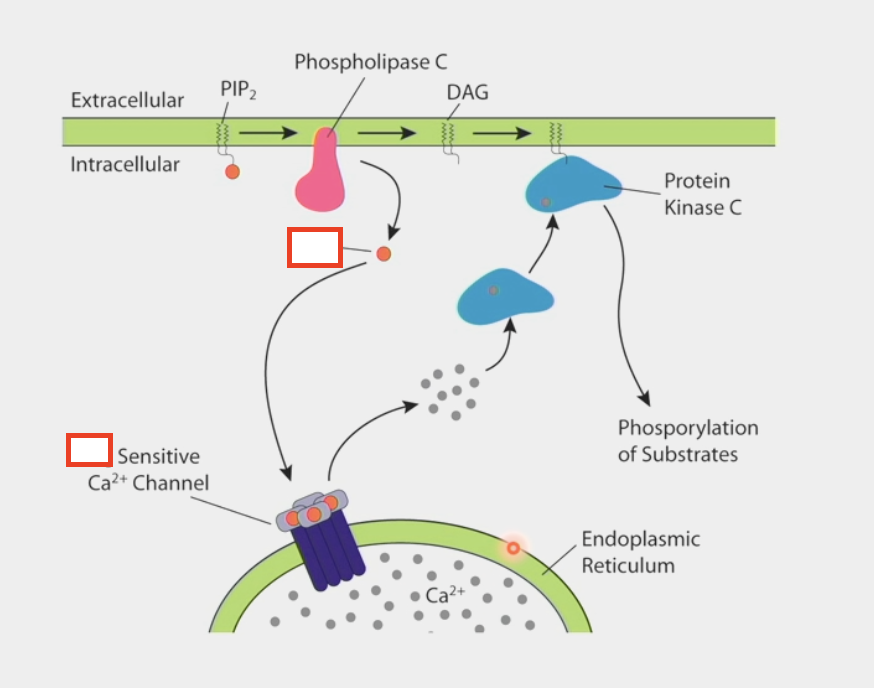
IP3
second messenger
binds to Ca2+ channel in the ER
allows calcium to be released from internal stores → cytoplasm
activates Protein Kinase C
binds to other second messenger DAG → phosphorylation of substrates
nuclear hormone receptors
intracellular receptors for steroid hormones
hormones can diffuse through lipid bilayer membrane into the cell
control gene expression and other processes