Electron Transport Chain
1/23
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
24 Terms
What’s left over?
All of the carbon from glucose has been oxidized
We have been tracking carbon throughout
6 Carbon in Glucose
6 CO2 Molecules formed!
No carbon left to track!
Now we have to focus on the energy carriers!
Energy captured as potential chemical energy carried to ETC by NADH and FADH2
Recap
Glycolysis:
Oxidized glucose (C6H12O6) to 2 pyruvate (C3H3O3) molecules
Net 2 ATP and 2 NADH
These NADH are special and must be brought from cytosol into the mitochondria (more on this soon)
No Carbon lost yet!
Pyruvate Oxidation:
Moved pyruvate into the mitochondria, lost first CO2 (for each pyruvate!) and attached CoA group
Made 1 NADH for each acetyl-CoA formed (2 total)
Krebs Cycle:
Completely oxidize all remaining carbon from glucose to 4 CO2
Produce 3 NADH, 1 FADH2, and 1 ATP for each “turn” of cycle! (X2)
There is no remaining energy to be extracted from the carbon in glucose
It has all been oxidized to CO2
Energy has been trapped in ATP, transferred to high energy carriers (reduced), and lost as thermal energy
So far our cells have made a total of 4 ATP per molecule of glucose
Still not enough energy for us!
Need to get the energy stored in the high energy carriers!
We must now focus on the protons and electrons that have been transferred through chemical reactions
Electrons are stored in the bonds of high energy molecules
Protons attached (NAD+ reduced to NADH) and also have been accumulating in the matrix of mitochondria
Reduction of NAD+ and FAD to NADH and FADH2 (addition of protons (H+) and electrons)
Each time they are reduced, some protons accumulate within the matrix of the mitochondria
These accumulated H+ ions will be very important soon!
Types of Phosphorylation
1. Substrate Level Phosphorylation
Forms ATP directly
Add a Pi + ADP to make ATP
Enzyme catalyzed reactions
Yields less ATP during cellular respiration
Happens during Glycolysis and the Krebs Cycle
2. Oxidative Phosphorylation
Indirect ATP formation (involves intermediates)
Requires energy carriers (NADH or FADH2)
Series of redox reactions where oxygen is the final electron acceptor
Involves NADH and FADH2
Yields much more ATP during cellular respiration
Process begins with the synthesis of NADH and FADH2 during Glycolysis and the Krebs Cycle and is completed in the Electron Transport Chain
Electron Transport Chain
Occurs at the inner mitochondrial membrane
Sometimes referred to as the Cristae
All other parts of aerobic cellular respiration occur in matrix of mitochondria
The electron transport chain is the final destination of all oxygen that we breathe
Oxygen is carried all around the body, to every cell, with the task of being the final electron acceptor in the electron transport chain
Extracts the potential energy stored in high energy carriers (NADH and FADH2)
Used to synthesize ATP ... much more ATP!
Is made up of a series of parts that transfer electrons from NADH and FADH2 to the final electron acceptor oxygen!
Involves 4 protein complexes and 2 mobile electron shuttles
Leads to the reduction of oxygen to water
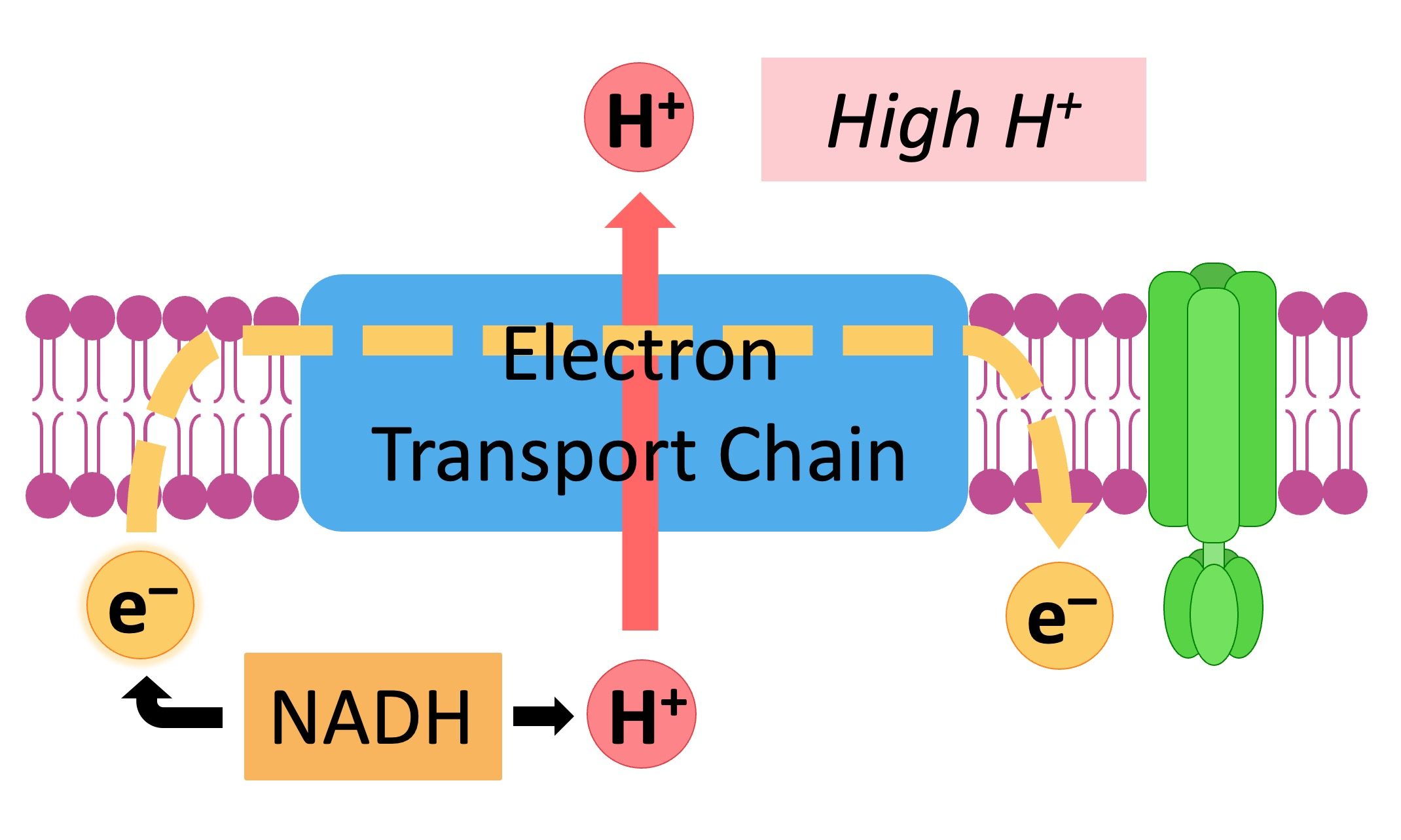
Electron/proteins
4 membrane proteins:
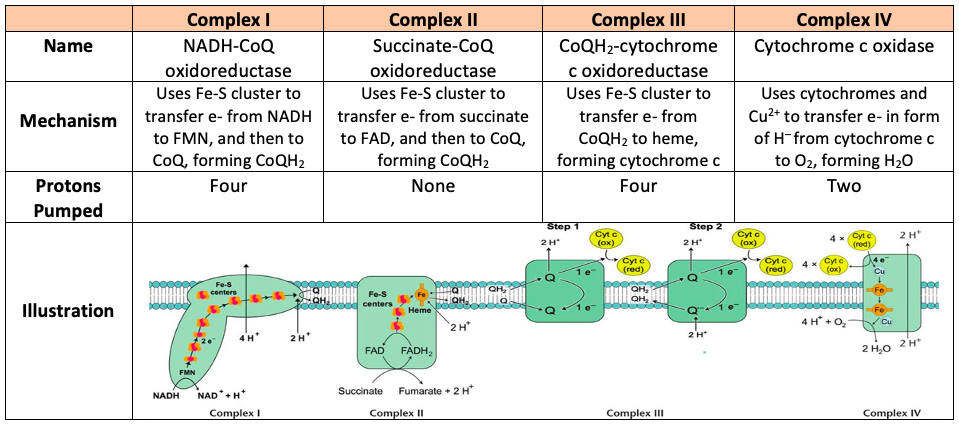
Complex I - NADH Dehydrogenase
Complex II - Succinate Dehydrogenase (Krebs)
Complex III - Cytochrome Complex
Complex IV - Cytochrome Oxidase
2 mobile electron shuttles:
Ubiquinone (UQ)
Also called coenzyme Q
Cytochrome C (cyt c)
Electron “chain” as each complex has increasing electronegativity and thus pass electrons off to next in line ending at oxygen
Oxygen is required for the electron transport chain to “start”
Before oxygen present, all carriers and high energy molecules are stable (full electron shells)
Oxygen interacts with Complex IV and removes a pair of electrons
Oxygen removes 2 electrons and also reacts with 2 H+ ions in the matrix to form a molecule of water
For each O2 we breathe → 4 electrons are pulled through the ETC and 2 water molecules are produced
This removal of electrons from complex IV causes a chain reaction where Complex IV cannot get its electrons back from oxygen (oxygen has higher electronegativity), so it takes electrons from Complex III (less electronegative) which then takes electrons from Complex I (less electronegative), which then takes electrons from NADH (less electronegative)
Each of these complexes have an associated cofactor that can be in an oxidized or reduced state
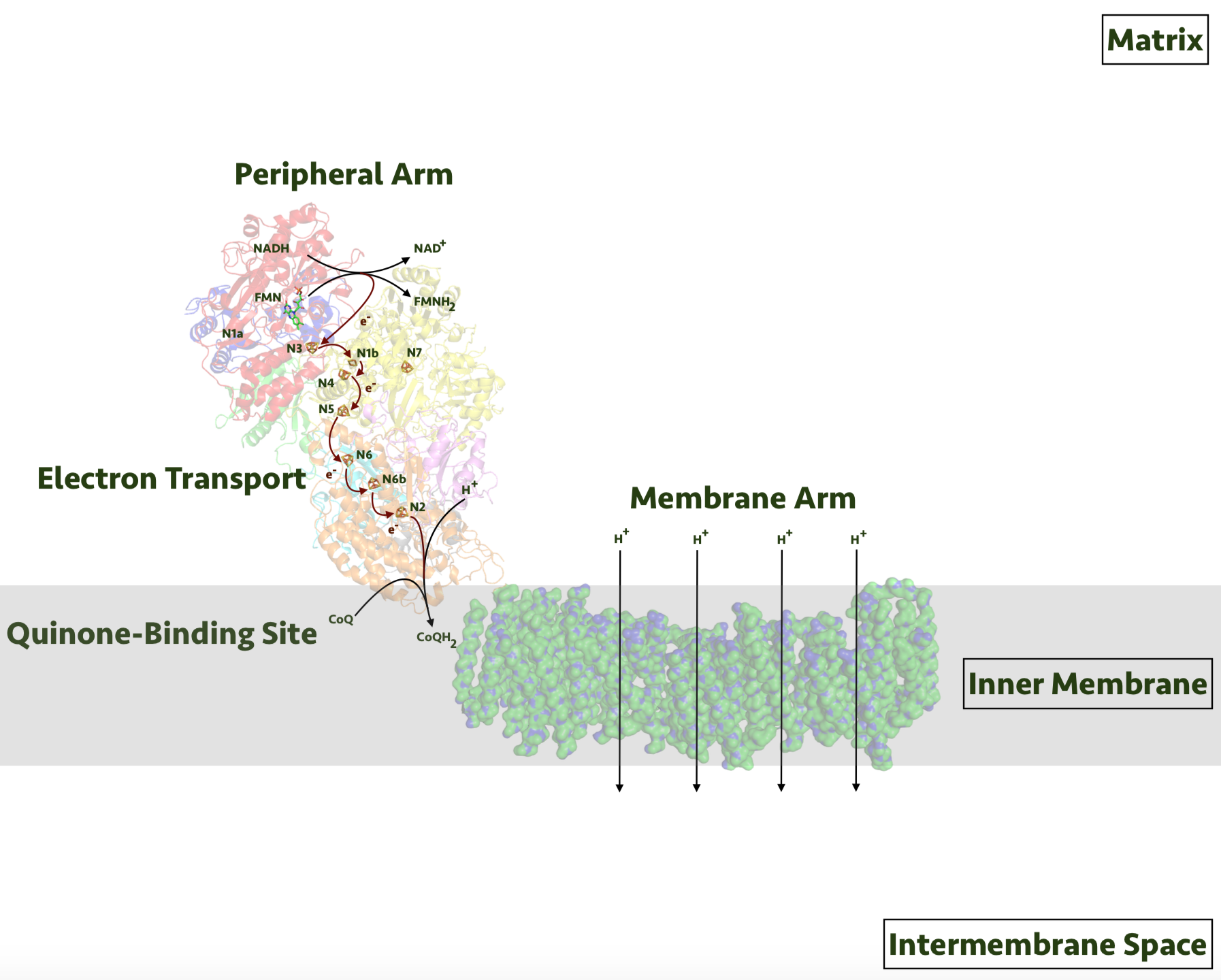
NADH Dehydrogenase (Complex I)
Lowest electronegativity in the Electron Transport Chain
Removes electrons from NADH
Oxidizes NADH to NAD+ and pumps protons (H+) ions into the intermembrane space from the mitochondrial matrix
This is against the protons electrochemical gradient - energy comes from the redox reaction as electrons move to more electronegative complexes
For each NADH, 4 H+ are pumped across membrane
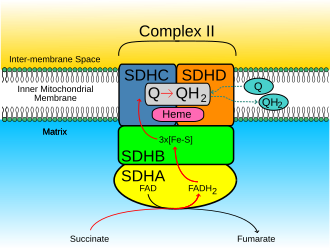
Succinate Dehydrogenase (Complex II)
Removes electrons from FADH2
Only enzyme that is involved in both the Krebs cycle and the electron transport chain
Peripheral protein that can associate with ubiquinone
Oxidize FADH2 to FAD+
No direct pumping of protons here (not a integral protein)’
Contributes to FADH2 generating less ATP than NADH
Cytochrome Complex (Complex III)
Removes electrons from Complex I (or electrons carried from Complex II)
Electrons move to Complex IV
Pumps 4 more H+ ions into intermembrane space
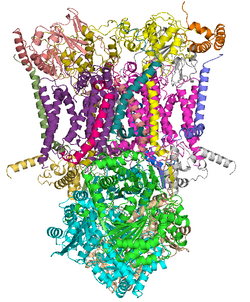
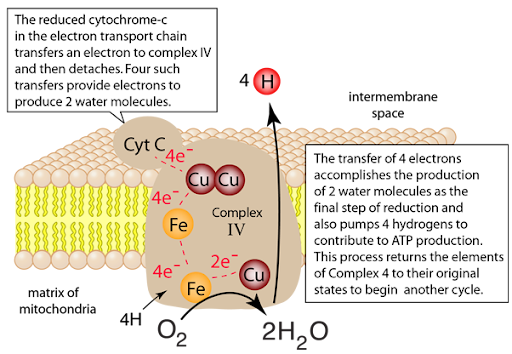
Cytochrome Oxidase (Complex IV)
Highest electronegativity in the Electron Transport Chain
Aside from oxygen
Oxidized by oxygen (loses electrons)
Site of water formation
Pumps more protons into the intermembrane space (2H+)
Further increases the electrochemical gradient being built
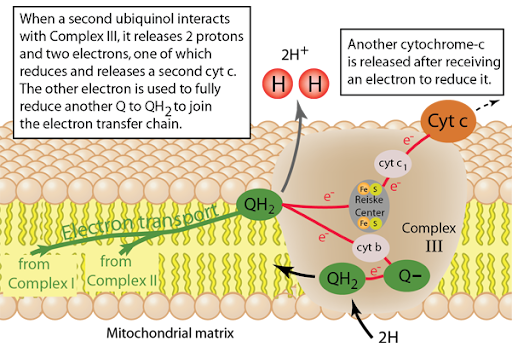
Electron Shuttles
Carry electrons from one protein complex to the next
1. Ubiquinone
Carries electrons from Complex I and Complex II and transfers them to Complex III
Hydrophobic molecule - found in the core of the membrane (among the fatty acid tails)
Carries and releases protons into intermembrane space
2. Cytochrome c
Carries electrons from Complex III and transfers them to Complex IV
Found on the intermembrane space side (facing the matrix) of the membrane (hydrophilic)
Cyanide Poisoning
Cyanide blocks cytochrome c oxidase so that the electrons cannot be transferred to oxygen
Irreversible, non-competitive inhibitor
Results in cellular respiration stopping
No more energy is made
Amygdalin is a common compound found in seeds of apples, almonds, apricots, cherries, and peaches along with many species from the family of roses
Hydrolysis of amygdalin gives rise to hydrogen cyanide
Dose is the poison as always
Recall the entire Electron Transport Chain and draw a Diagram
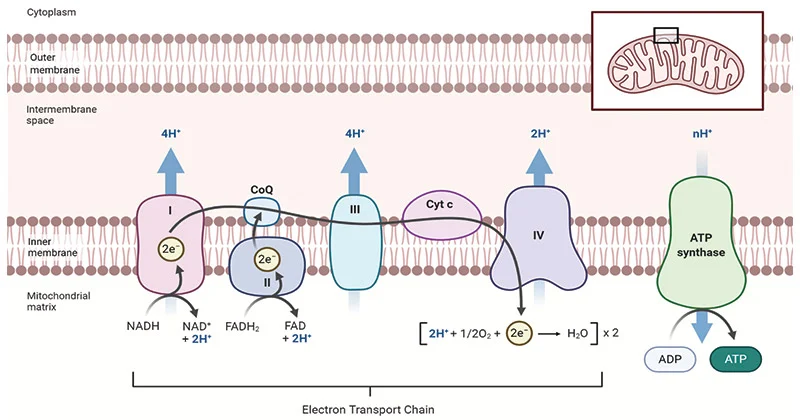
Chemiosmosis
Throughout the ETC, protons have been accumulating in the intermembrane space after having been pumped through the membrane at each of the protein complexes (using the energy from the redox reactions)
This has created a large electrochemical gradient (proton gradient)
This electrochemical gradient means that protons want to move back into the matrix of the mitochondria
Remember diffusion!
This is harnessed as the “Proton Motive Force”
Utilized at ATP Synthase
This is not part of the ETC but is located close by
Used to make ATP!
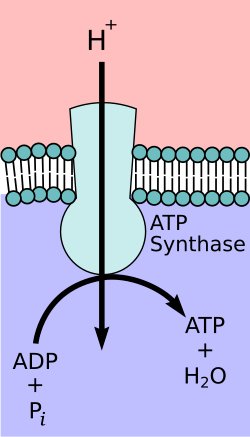
ATP Synthesis
Chemiosmosis - synthesis of ATP using the energy of the electrochemical gradient and ATP synthase enzyme
Synthesizes ATP via oxidative phosphorylation
Utilizes the energy source of the proton gradient (proton motive force)
Acts like a water mill - protons moving “downstream” create energy to turn a wheel
Protons move through a channel protein in the membrane and pass into the matrix of the mitochondria (down their electrochemical gradient)
This in turn generates power that allows for ATP synthesis!
Approximately 1 ATP is generate per 4 electrons
NADH pumps 10 electrons total ~ 2.5 (3) ATP per NADH
FADH2 pumps 6 electrons ~ 1.5 (2) ATP per FADH2
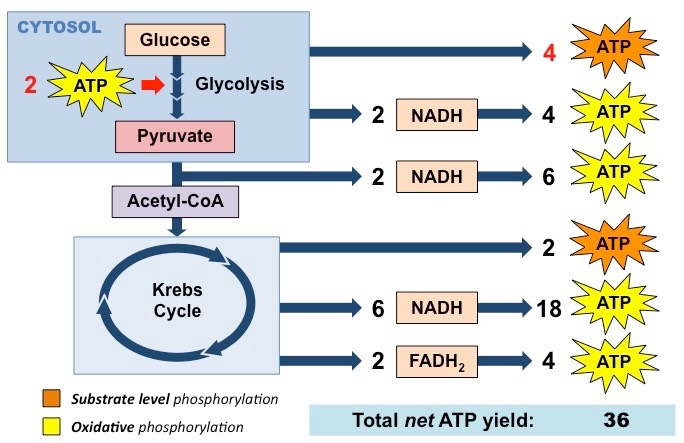
Total ATP from Cellular Respiration
2 ATP and 2 NADH from Glycolysis (these NADH are special and lose some energy entering the mitochondria - info on the next slide on shuttles)
2 NADH from Pyruvate Oxidation
2 ATP, 6 NADH and 2 FADH2 from Krebs Cycle
Electron Transport Chain
2 NADH from Glycolysis - 4 ATP
2 NADH from Pyruvate Oxidation - 6 ATP
6 NADH and 2 FADH2 from Krebs Cycle - (3*6 + 2*2) - 22 ATP
~28-34 ATP made by Oxidative Phosphorylation and 4 ATP made by Substrate Level Phosphorylation
32-38 Total ATP made during Cellular Respiration*
This is a theoretical max and is likely a slight overestimate due to inefficiency in the cell, differences in cell type, and some proton “leak” causing differences in ATP yield per NADH and FADH2
Efficiency:
Glucose = 686 kcal/mol
ATP = 7.3 kcal/mol (38 = 277.4 kcal/mol)
277.4 kcal/mol / 686 kcal/mol = 0.404 X 100%
40.4%
NADH Shuttles
Electrons from NADH made during glycolysis must enter into the mitochondria. Mitochondrial membrane is impermeable to NADH
Glycerophosphate Shuttle
NADH transfers electrons (reduces) DHAP to G3P
G3P enters the mitochondria and is oxidized back to DHAP
This transfers electrons to form FADH2
Therefore the electrons from these NADH produced during glycolysis only yield the same amount of ATP as FADH2
This shuttle is commonly found in brain and muscle tissues
High energy demand = need energy fast
Malate-Aspartate Shuttle
NADH transfers electrons (reduces) oxaloacetate to malate in the cytoplasm
Malate enters the mitochondria and is oxidized back to oxaloacetate
This regenerates NADH inside of the matrix
Therefore electrons from these NADH produced during glycolysis yield the same amount of ATP as normal
Predominantly in heart, liver, kidneys, etc.
Uncoupling
Electron Transport and ATP synthesis can be “uncoupled”
This is done via uncoupling proteins (alternate entry point other than ATP synthase)
Proton motive force no longer used to generate ATP
Common in brown adipose tissue
Instead of the energy being used to make ATP, it is all released as thermal energy
This is a key feature of body temperature regulation in hibernating animals, certain birds, and young animals
If you consume 4 molecules of glucose, how much ATP is made via substrate level phosphorylation? How much is made via oxidative phosphorylation? Show your work.
Glycolysis: Net 2 ATP
Krebs cycle: 2 turns per glucose → 2 ATP (as GTP). Total SLP per glucose = 4 ATP
SLP total = 16 ATP
2. ATP from oxidative phosphorylation (OxPhos). This comes from NADH and FADH₂ generated earlier.
Glycolysis: 2 NADH
Pyruvate oxidation: 2 NADH
Krebs cycle: 6 NADH + 2 FADH₂
Total per glucose:
10 NADH
2 FADH₂
1 NADH = 3 ATP
1 FADH₂ = 2 ATP
10 NADH × 3 = 30 ATP
2 FADH₂ × 2 = 4 ATP
OxPhos per glucose = 30 + 4 = 34 ATP
34 × 4 = 136 ATP
Grand Total:16 + 136 = 152 ATP
Explain how the proton electrochemical gradient is “built up” and how this is then used to generate ATP. Be sure to use scientific terminology.
The proton electrochemical gradient is central to ATP production via oxidative phosphorylation, and it is established and used through a process called chemiosmosis.
This occurs in the inner mitochondrial membrane via the electron transport chain (ETC).
a. Electron transport
High-energy electrons from NADH and FADH₂ are passed through a series of protein complexes:
Complex I, II, III, and IV
These electrons ultimately reduce oxygen to water at Complex IV.
b. Proton pumping
As electrons move through the ETC:
Complexes I, III, and IV use the released energy to actively transport H⁺ (protons) from the mitochondrial matrix to the intermembrane space.
This creates two gradients:
c. The electrochemical gradient (proton motive force)
Also called the proton motive force, it has two components:
Chemical gradient (ΔpH)
Higher [H⁺] in the intermembrane space than in the matrix
Electrical gradient (ΔΨ)
Intermembrane space becomes more positively charged relative to the matrix
Together, this stores potential energy across the membrane.
2. How the gradient is used to generate ATP
Protons flow back into the matrix through the enzyme ATP synthase.
a. Facilitated diffusion of protons
H⁺ cannot freely cross the membrane
They move down their electrochemical gradient through ATP synthase
b. Mechanical rotation and catalysis
Proton flow drives rotation of the F₀ subunit
This induces conformational changes in the F₁ subunit
c. ATP formation
These conformational changes allow:
ADP + Pi → ATP
This mechanism is described by the binding change model:
ADP and Pi bind
ATP is synthesized
ATP is released
(a) Do the electrons in NADH have the most or the least free energy in the electron transport chain?
(b) The electrons in NADH form bonds as they move through the electron transport chain.
Do these bond formations use or release energy?
a. The electrons in NADH have the most free energy when they enter the electron transport chain.
Why?
NADH is a high-energy electron carrier.
It donates electrons to Complex I at the start of the chain.
These electrons are at a high energy state (high free energy).
What happens next?
As electrons move through the ETC:
They pass through a series of carriers with increasing electronegativity
They lose free energy step by step
This released energy is used to:
Pump protons (H⁺)
Build the proton motive force
Final state
At the end of the chain, electrons are transferred to oxygen (O₂), forming water
At this point, they have very low free energy
b. The bond formations release energy.
As electrons from NADH move through the electron transport chain, they pass to molecules with progressively higher affinity for electrons. Each transfer involves forming new, more stable bonds (for example, when electrons ultimately help form water).
Because these new bonds are lower in energy (more stable) than the previous ones, the excess energy is released. This released energy is then used to pump protons across the inner mitochondrial membrane, creating a gradient that drives ATP production.
Which stages of aerobic cellular respiration occur in the mitochondria, and which stages do not?
Occur in the mitochondria
Pyruvate oxidation (link reaction) – happens in the mitochondrial matrix
Citric acid cycle (Krebs cycle) – also in the mitochondrial matrix
Electron transport chain and oxidative phosphorylation – on the inner mitochondrial membrane
Do NOT occur in the mitochondria
Glycolysis – takes place in the cytoplasm (cytosol)
What important molecule is needed for oxidative phosphorylation but not needed for substrate-level phosphorylation?
The key molecule is oxygen (O₂).
Oxidative phosphorylation requires oxygen because it acts as the final electron acceptor in the electron transport chain. Without oxygen, the chain backs up and ATP production stops.
Substrate-level phosphorylation does not require oxygen. It generates ATP directly by transferring a phosphate group from a substrate to ADP (as in glycolysis and the Krebs cycle).
What is the primary function of the proton-motive force?
The primary function of the proton-motive force is to drive the production of ATP.
As protons (H⁺) are pumped across the inner mitochondrial membrane during the electron transport chain, they create an electrochemical gradient (the proton-motive force). This stored energy is then used when protons flow back into the mitochondrial matrix through ATP synthase.
This flow powers ATP synthase to convert ADP + Pi → ATP.
Give an example of how uncoupling is used by organisms to increase survival.
example is heat production in mammals, especially in cold environments.
In brown adipose tissue (brown fat), organisms use an uncoupling protein called UCP1. Instead of allowing the proton-motive force to drive ATP production, UCP1 lets protons flow back into the mitochondrial matrix without making ATP. This process is called uncoupling.
Because the energy from the proton gradient isn’t used to make ATP, it is released as heat instead.
Why this increases survival:
Helps maintain body temperature in cold conditions
Especially important for newborns and hibernating animals, which can’t generate enough heat by shivering