M3: Nuclear Biology
1/43
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
44 Terms
What is the proteome of an organelle?
the complete and distinct ‘collection’ of proteins that originate from ribosomal synthesis (primarily in the cytosol and ER) of an organelle
What is required for function of distinct proteins?
distinct proteins must be targeted from their site of synthesis to their correct organelle location for function
What does targeting proteins require?
Targeting of proteins requires specific “address labels”, and these are specific amino acid sequences
can be uni-directional or bi-directional
What are the different modes of transport used?
gated, transmembrane and vesicular
What is gated transport?
bi-directional, folded proteins, into and out of nuclear pores to cytosol
What is transmembrane transport?
unfolded proteins
What is vesicular transport?
budding of vesicles, trafficked to another organelle which fuses and releases into organelle
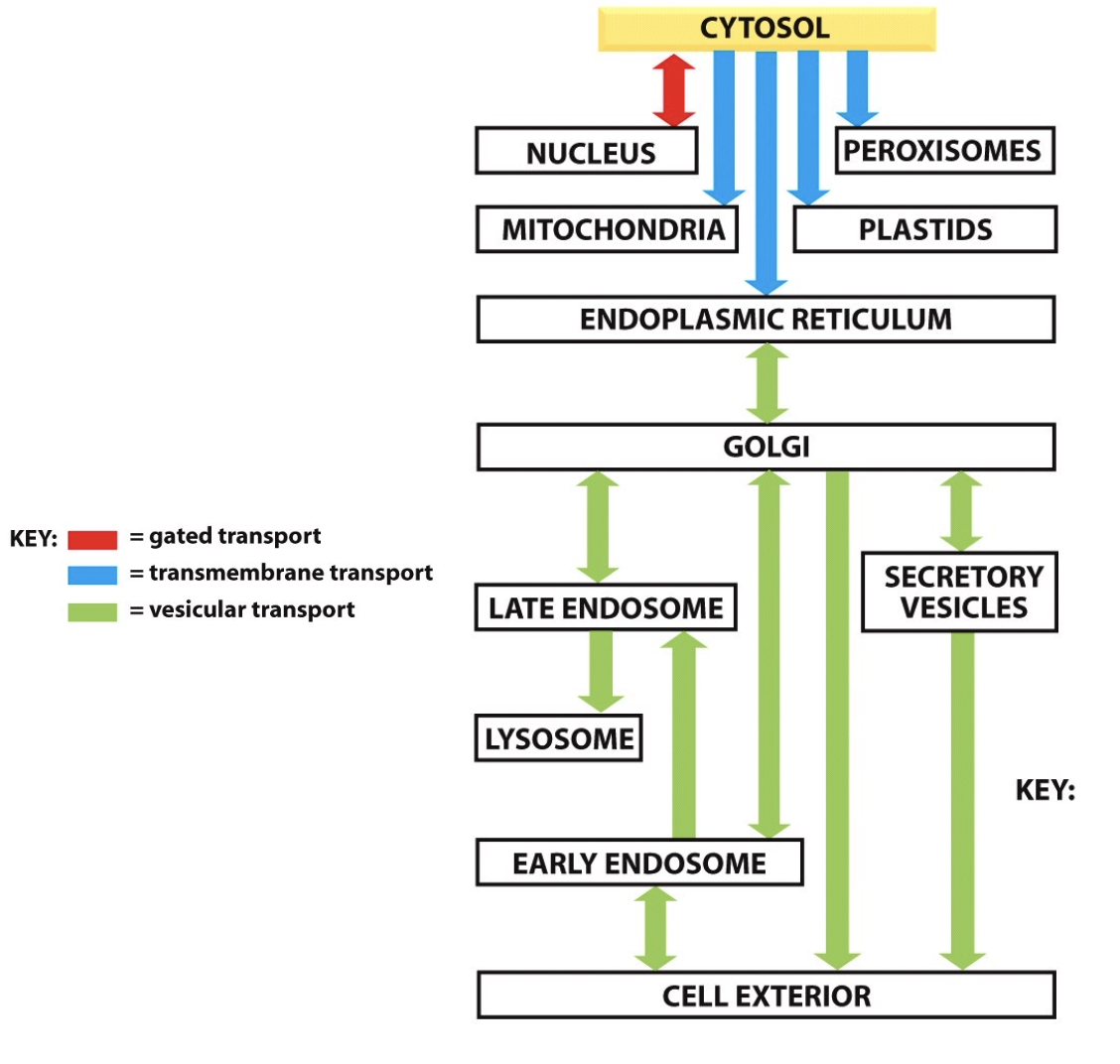
What is the hierarchy of intracellular protein trafficking?
general = cytosol→ER→golgi→vesicle→exterior
What is fluorescence microscopy and examples?
expressing fluorescent molecules in cells and exciting them at appropriate wavelengths to see how much/localisation
fluorescent dyes/probes
fluorescent dyes attached to antibodies
fluorescent proteins
fluorescent dyes/probes
non-protein molecules that absorb light and re-emit it at a longer wavelength
often used in the fluorescent labelling of biomolecules and can be smaller or more photostable than fluorescent proteins but cannot be genetically encoded
JC-1 (Mitochondria) and ER-Tracker Green (ER)
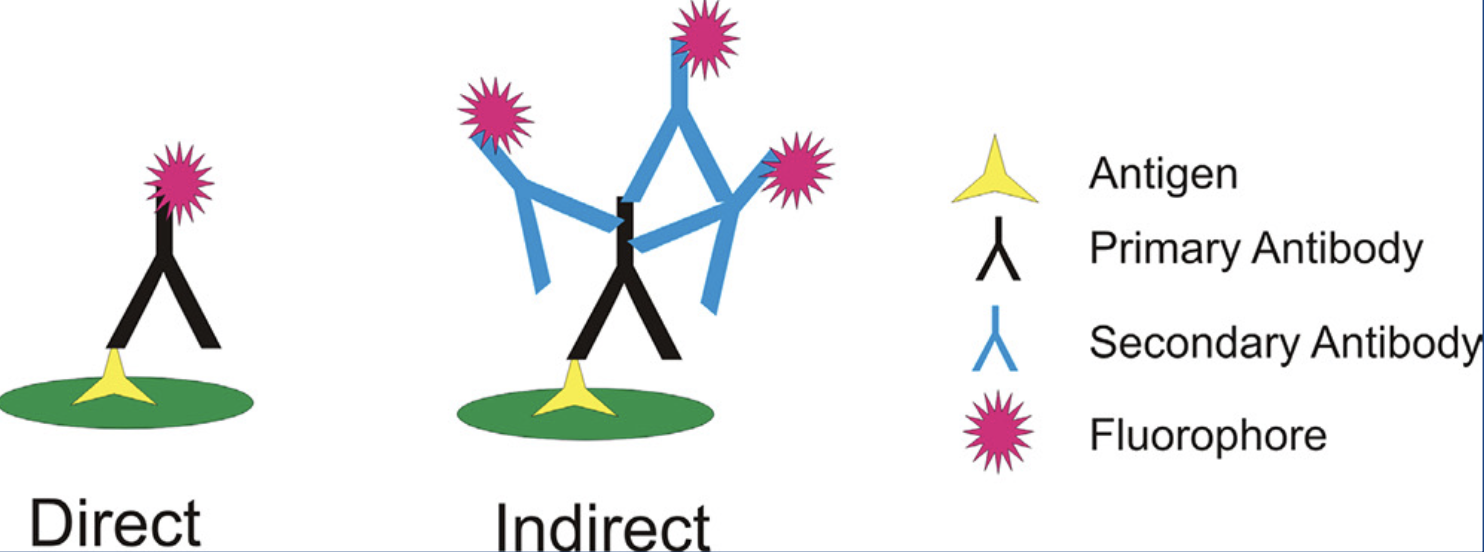
Immunofluorescence Assay – Direct versus Indirect
direct IFA: a single antibody directed against the target of interest. The primary antibody is directly conjugated to a fluorophore.
indirect IFA: two antibodies. The primary antibody is unconjugated and a fluorophore-conjugated secondary antibody directed against the primary antibody is used for detection.
more common
Fluorescent Proteins
GFP in 1992
has been engineered to produce a vast number of variously coloured mutants, fusion proteins, and biosensors that are broadly referred to as fluorescent proteins
Fluorescent Proteins applications
Fusion tagging: most common, GFP can be fused to the N- or C-terminus of a protein, which allows the scientist to visualise when and where the gene is expressed.
Forster resonance energy transfer (FRET): decipher how close two proteins are
Biosensors: measure different properties in cells
Split EGFP:
GFP:
beta-barrel protein green fluorescent protein
Split EGFP:
seperate last strand of beta barrel (strand 11) from rest of GFP → not fluorescent
attach strand 11 to a protein, strands 1-10 at a particular compartment of a cell
if protein goes to compartment, strands 1-10 and 11 will reform and fluoresce
eukaryotic nucleus is…
surrounded by nuclear membrane that provides compartmentalisation to the process that occur in the nucleus
must be crossed by RNA species to allow translation on cytosolic ribosomes (RNA export)
RNA export:
i) Cytosolic proteins must re-enter the nucleus to perform nuclear functions (e.g. transcription factors, DNA polymerase) (PROTEIN IMPORT)
ii) Proteins once in the nucleus can also exit the nucleus (PROTEIN EXPORT)
nuclear membrane = nuclear envelope
A double membrane envelope surrounds the nucleus.
Membranes such as these prevent movement of large or hydrophilic molecules.
Movement across the nuclear envelope therefore requires some controlled gateway and mechanism for crossing the gateway.
Requirements for traversing the nuclear membrane
1: A gateway (regulated/controlled) that allows the correctly addressed protein into the nucleus and other correctly addressed proteins out of the nucleus.
Proteins are FOLDED
2: A mechanism to address protein to the nucleus (import = into nucleus)
3: A mechanism to send proteins back to the cytosol once their nuclear function is
complete (export = from nucleus)
NPC
nuclear pore complex
maintain integrity of nucleus by preventing macromolecules to freely difuse in and out of nucleus (transport FOLDED CARGO)
proteins smaller than 40kDA can diffuse through NPC ie. small proteins may enter via diffusion; proteins larger need specialised transport factors therefore need RECOGNITION
What defines the mode of transport across NPC?
size of protein/molecule for transport - gated diffusion barrier
size of human NPC
1000 proteins (nucleoproteins)
total molecular mass of 110 Mda (Mega dalton)
aka. massive
NUPs
nucleoporins = proteins of the Nuclear Pore Complex
around 30 of them, many repeats to make up NPC
regulation of NPC proteins
FG repeats - phenylalynine glycine repeats
create regions of disorder in proteins
ordered folded domain allows them to be anchored to NPC