🧬 Lecture A: Topics + Nucleus
1/18
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
19 Terms
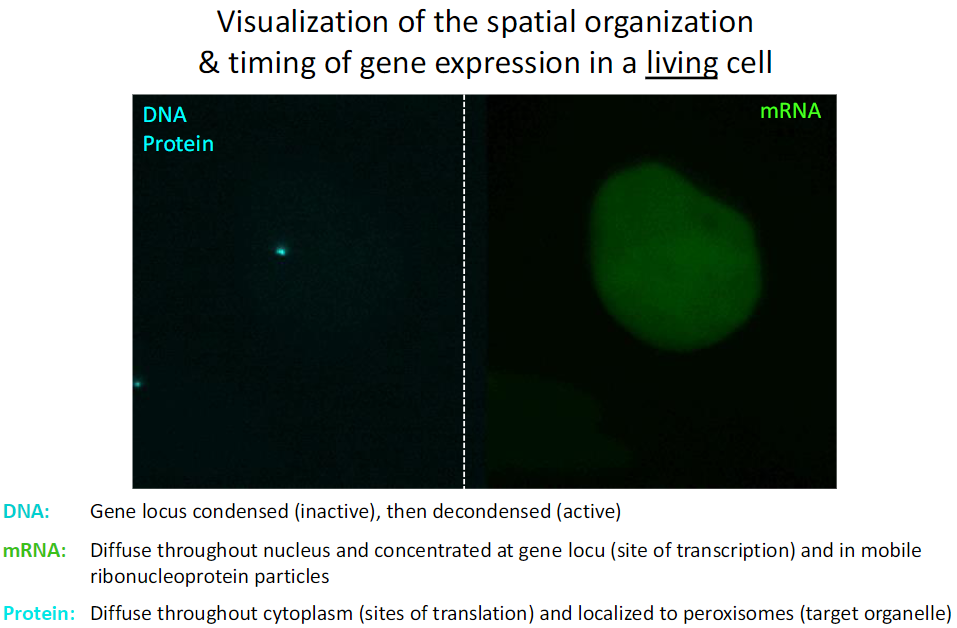
Spatial and Temporal Organization of Gene Expression
DNA
Gene locus condensed → inactive
Gene locus decondensed → active
mRNA
Diffuse throughout nucleus
Concentrated at gene loci (site of transcription)
Found in mobile ribonucleoprotein particles
Protein
Diffuse throughout cytoplasm (sites of translation)
Localized to peroxisomes (target organelle)
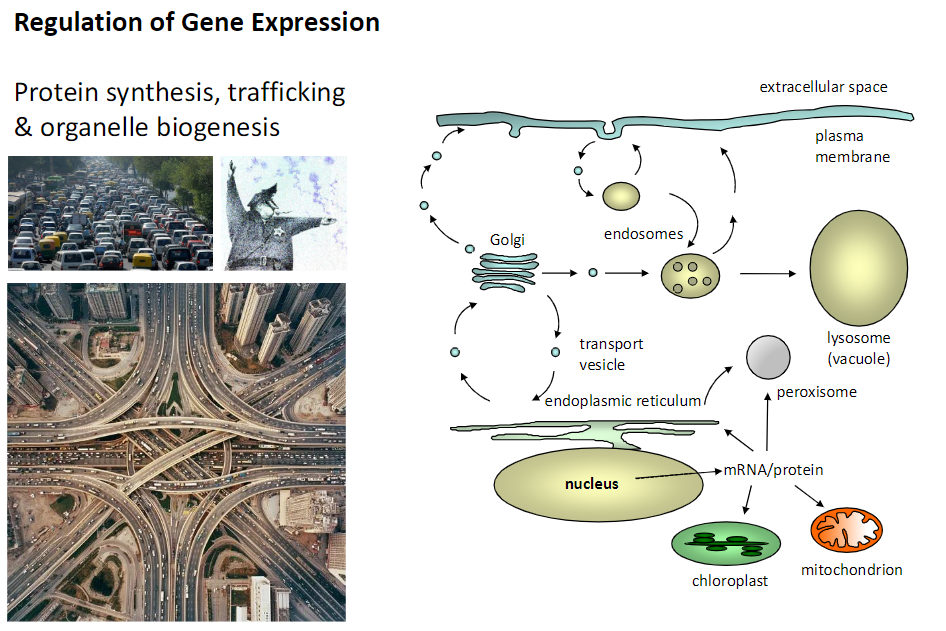
Protein Synthesis, Trafficking & Organelle Biogenesis
Nucleus
Site of mRNA transcription
Endoplasmic Reticulum (ER)
Site of protein synthesis and initial folding
Golgi
Modifies, sorts, and packages proteins for transport
Transport Vesicle
Carries proteins between organelles and to the plasma membrane
Lysosome (Vacuole)
Contains digestive enzymes for macromolecule breakdown
Mitochondrion / Chloroplast
Sites of energy production (ATP or photosynthesis)
Peroxisome
Site of oxidative reactions and specialized protein function
Endosomes
Involved in protein sorting and trafficking
Plasma Membrane / Extracellular Space
Final destination for secreted or membrane-bound proteins
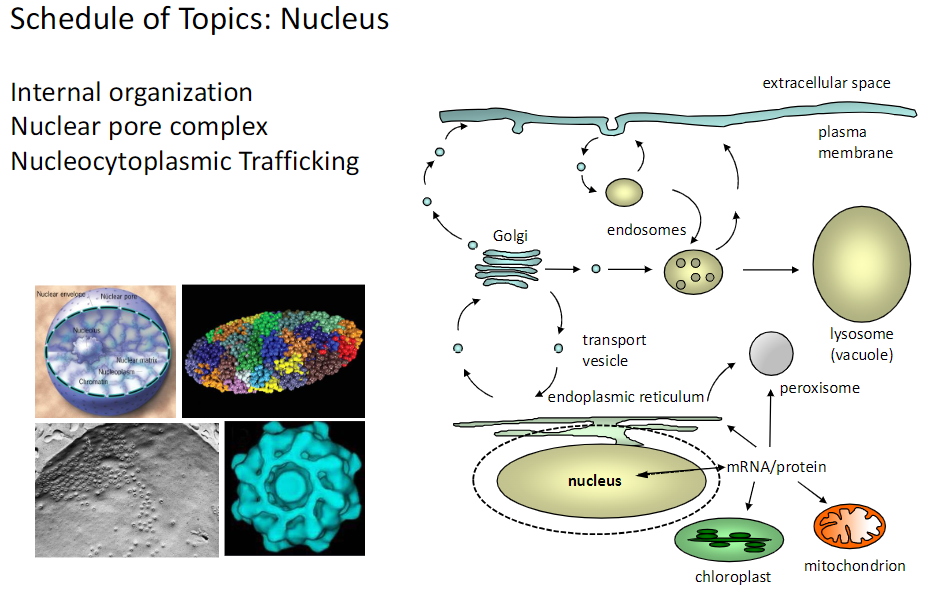
Internal Organization – Nuclear Pore Complex & Trafficking
Nuclear Pore Complex
Gateway for molecules between nucleus and cytoplasm
Nucleocytoplasmic Trafficking
Movement of mRNA, proteins, and other macromolecules through the nuclear pore
Key Point
The nuclear pore complex controls transport to maintain proper cellular function
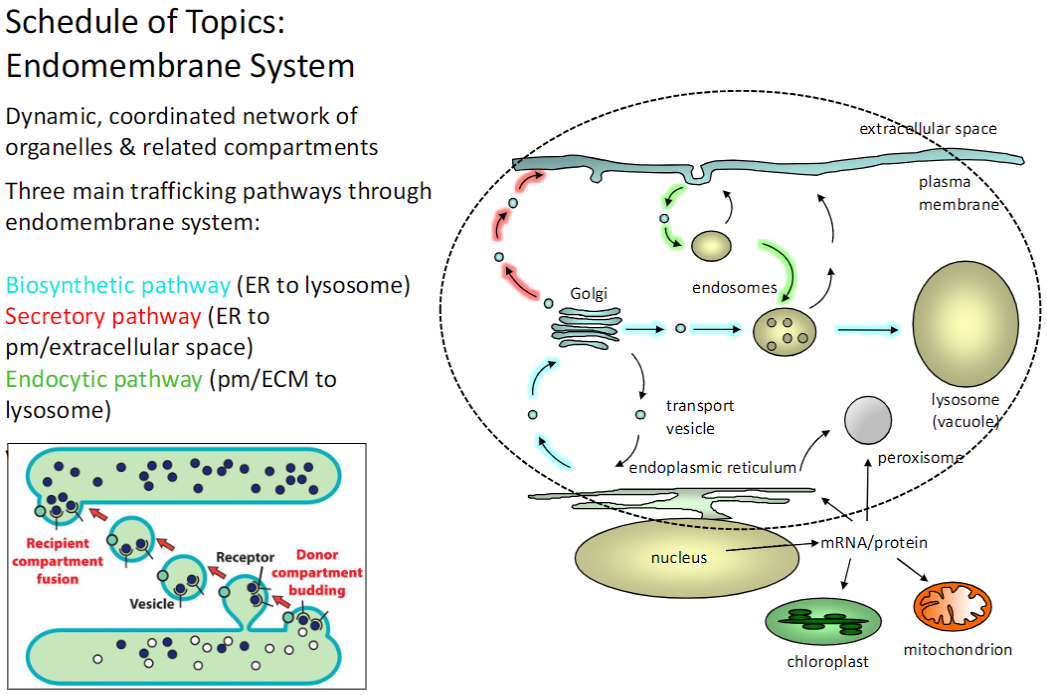
Endomembrane System – Overview
Definition
Dynamic, coordinated network of organelles and related compartments
Trafficking Pathways
Biosynthetic Pathway
From ER → lysosome
Secretory Pathway
From ER → plasma membrane / extracellular space
Endocytic Pathway
From plasma membrane / ECM → lysosome
Key Point
The endomembrane system organizes protein and membrane trafficking through biosynthetic, secretory, and endocytic pathways
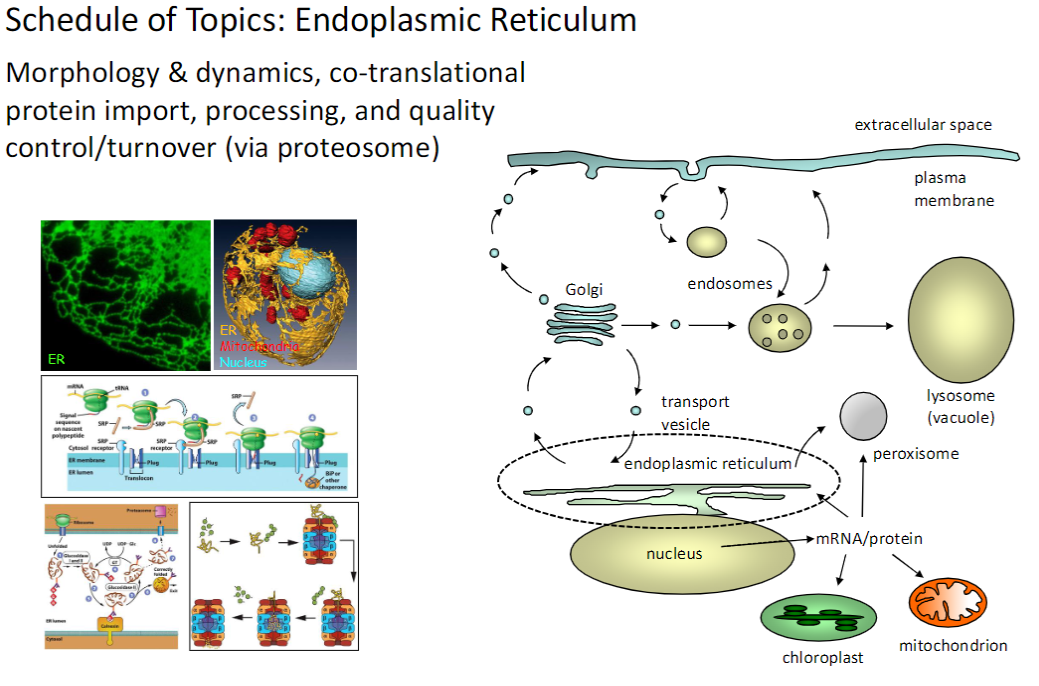
Protein Processing & Quality Control
Morphology & Dynamics
Structure and organization of organelles involved in protein synthesis and trafficking
Co-Translational Protein Import
Proteins are imported into organelles while being synthesized
Processing
Proteins undergo folding, modification, and sorting
Quality Control / Turnover
Misfolded or damaged proteins are degraded by the proteasome
Key Point
Cells maintain protein function through co-translational import, processing, and proteasome-mediated turnover
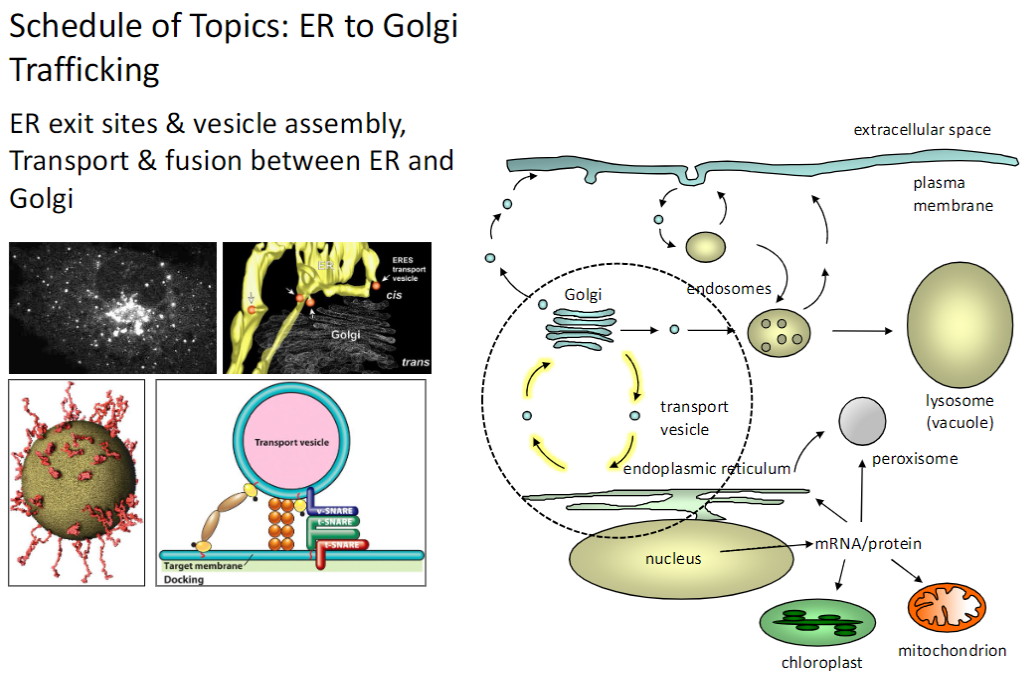
ER to Golgi Transport
ER to Golgi Transport
ER Exit Sites & Vesicle Assembly
Regions of the ER where proteins and lipids are packaged into transport vesicles
Transport & Fusion
Vesicles move from ER → Golgi and fuse with Golgi membranes for further processing
Key Point
ER exit sites and vesicle transport are essential for protein sorting and trafficking between organelles
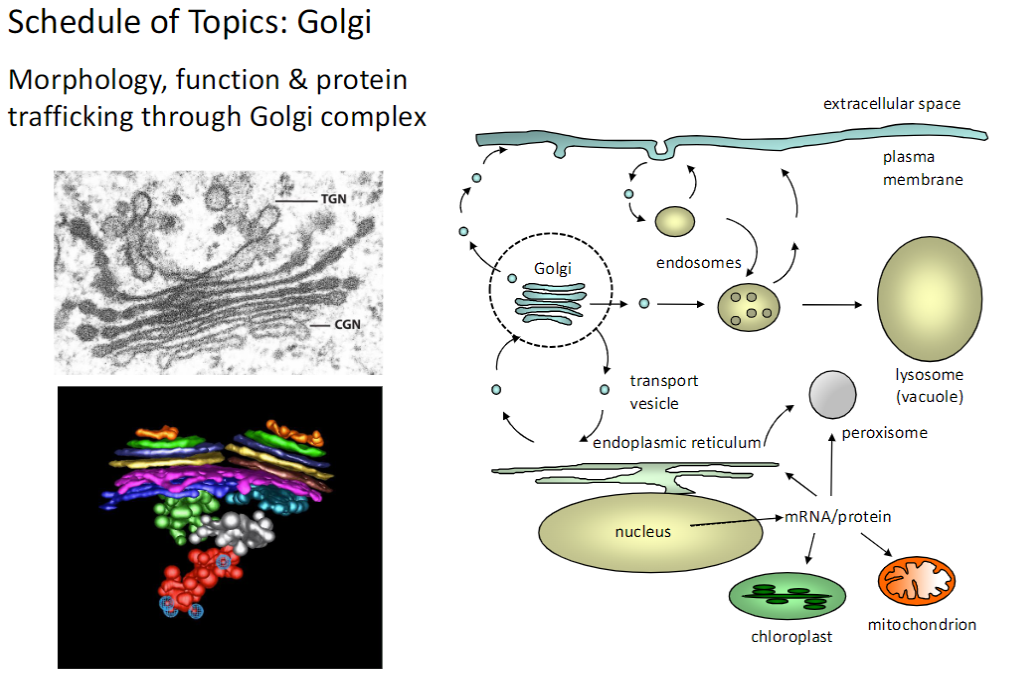
Golgi Complex – Structure, Function & Trafficking
Morphology
Stacked cisternae with distinct cis, medial, and trans regions
Function
Modifies, sorts, and packages proteins for delivery to organelles, plasma membrane, or extracellular space
Protein Trafficking
Proteins arrive from ER, undergo processing, and are shipped to their final destinations
Key Point
The Golgi complex is central for protein maturation, sorting, and trafficking
Lysosomes, Endosomes & Clathrin-Coated Vesicles
Lysosomes (Vacuoles)
Contain digestive enzymes for breaking down macromolecules
Endosomes
Involved in sorting and trafficking of internalized material
Clathrin-Coated Transport Vesicles
Vesicles with a clathrin protein coat that mediates transport between organelles and from plasma membrane → endosomes
Key Point
These compartments and vesicles coordinate protein and membrane trafficking and degradation within the cell
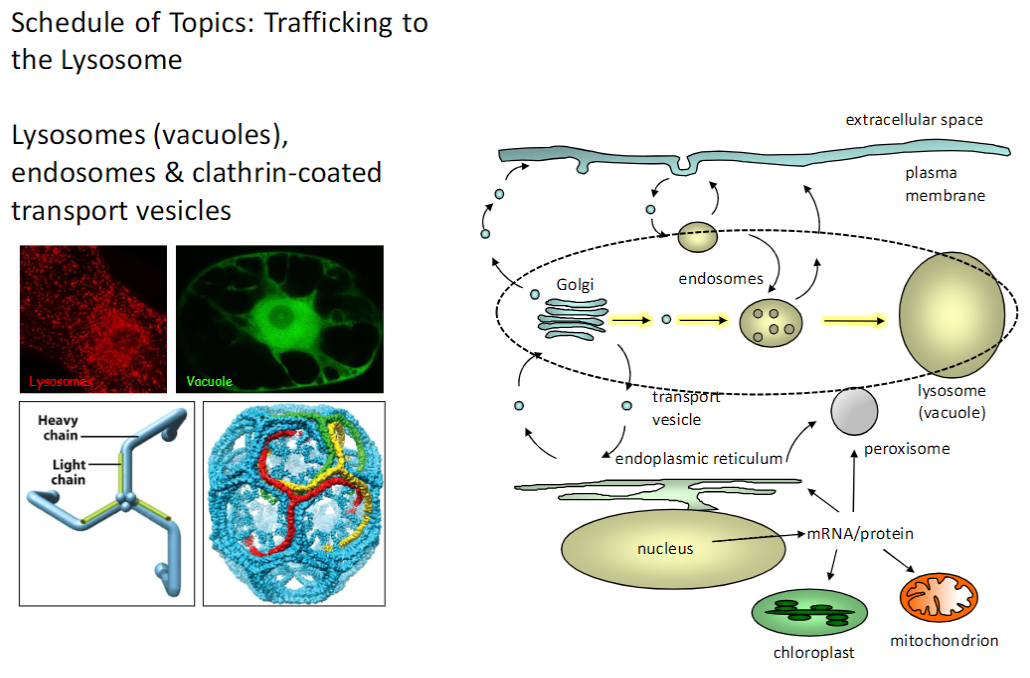
Cellular Transport – Exocytosis & Endocytosis
Exocytosis
Vesicles fuse with the plasma membrane to release proteins or other molecules outside the cell
Receptor-Mediated Endocytosis
Specific molecules are internalized via receptors on the plasma membrane
Endosomes
Sort and direct internalized molecules to their proper cellular destinations
Phagocytosis
Engulfment of large particles or pathogens into phagosomes for degradation
Key Point
Cells use exocytosis, endocytosis, and phagocytosis to control intake, processing, and release of materials
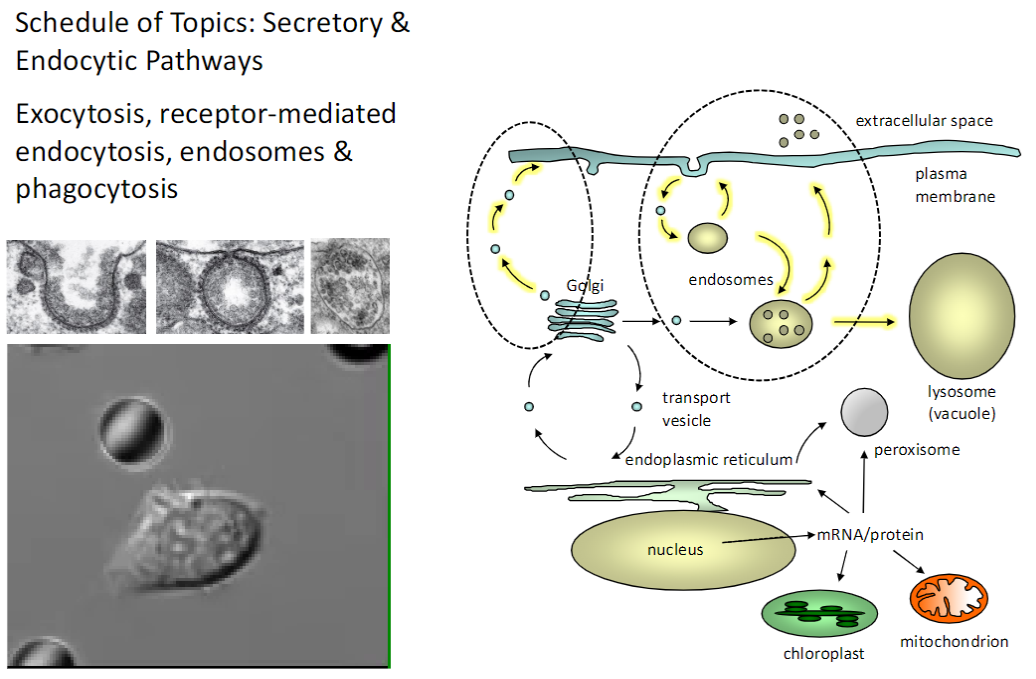
Mitochondria & Chloroplasts – Structure, Dynamics & Protein Import
Morphology
Mitochondria: double membrane, inner membrane folded into cristae
Chloroplasts: double membrane, internal thylakoid membranes
Dynamics
Fission, fusion, and movement within the cell to meet energy needs
Protein Import
Proteins are synthesized in the cytoplasm and imported into organelles via specific translocases
Key Point
Mitochondria and chloroplasts rely on imported proteins and dynamic morphology for proper energy production and function
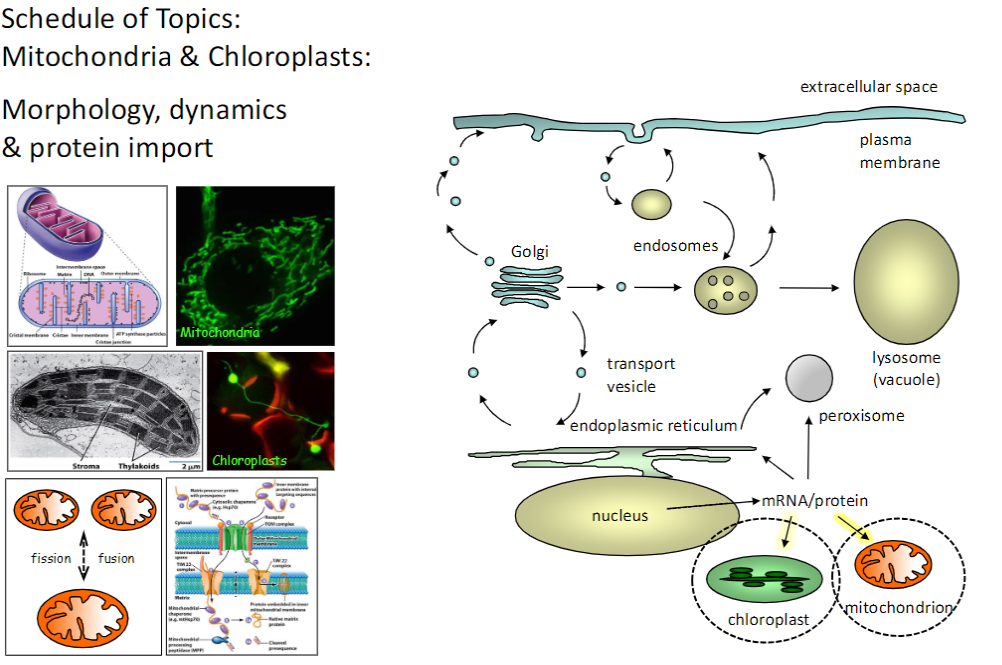
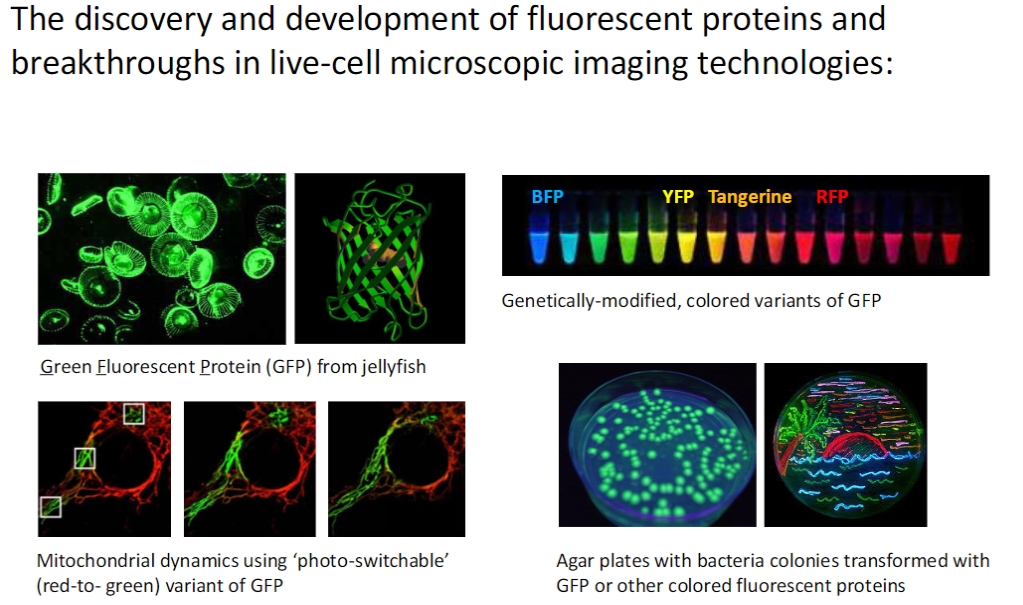
Fluorescent Proteins & Live-Cell Imaging
Discovery & Development
Green Fluorescent Protein (GFP) from jellyfish
Creation of genetically modified colored variants of GFP
Applications
Bacteria colonies on agar plates expressing GFP or other fluorescent proteins
Study of mitochondrial dynamics using photo-switchable GFP (red ↔ green)
Variants
BFP – Blue Fluorescent Protein
YFP – Yellow Fluorescent Protein
Tangerine – Orange variant
RFP – Red Fluorescent Protein
Key Point
Fluorescent proteins allow visualization of dynamic processes in live cells using advanced microscopy
Fluorescent Proteins in Living Cells & Organisms
Transformed Organisms
Bacteria, Yeast, Worm, Fly, Plant, Fish, Cat, Mouse, Human, Primate
Purpose
Visualize cellular and developmental processes in living cells and organisms
Key Point
GFP and its variants can be used across a wide range of species to study dynamic biological processes in real time
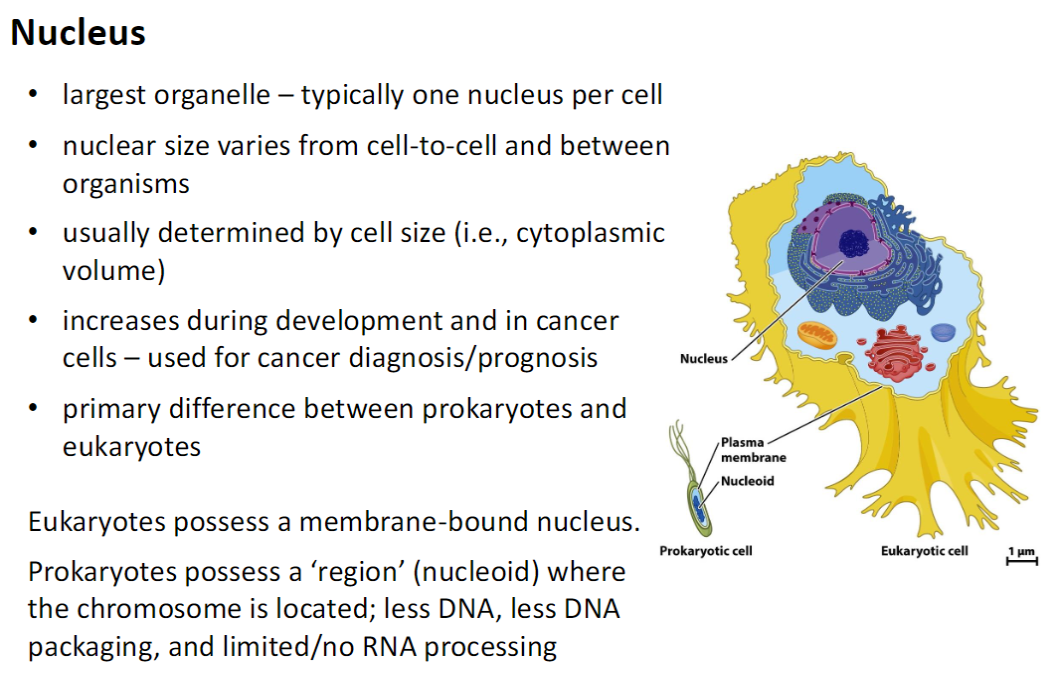
Nucleus – Structure & Function
Size & Number
Largest organelle, usually one nucleus per cell
Size varies between cells and organisms
Generally correlates with cell (cytoplasmic) volume
Increases during development and in cancer cells – useful for diagnosis and prognosis
Prokaryotes vs Eukaryotes
Eukaryotes: have a membrane-bound nucleus
Prokaryotes: have a nucleoid – less DNA, minimal DNA packaging, limited/no RNA processing
Key Point
The nucleus stores genetic material and distinguishes eukaryotic from prokaryotic cells
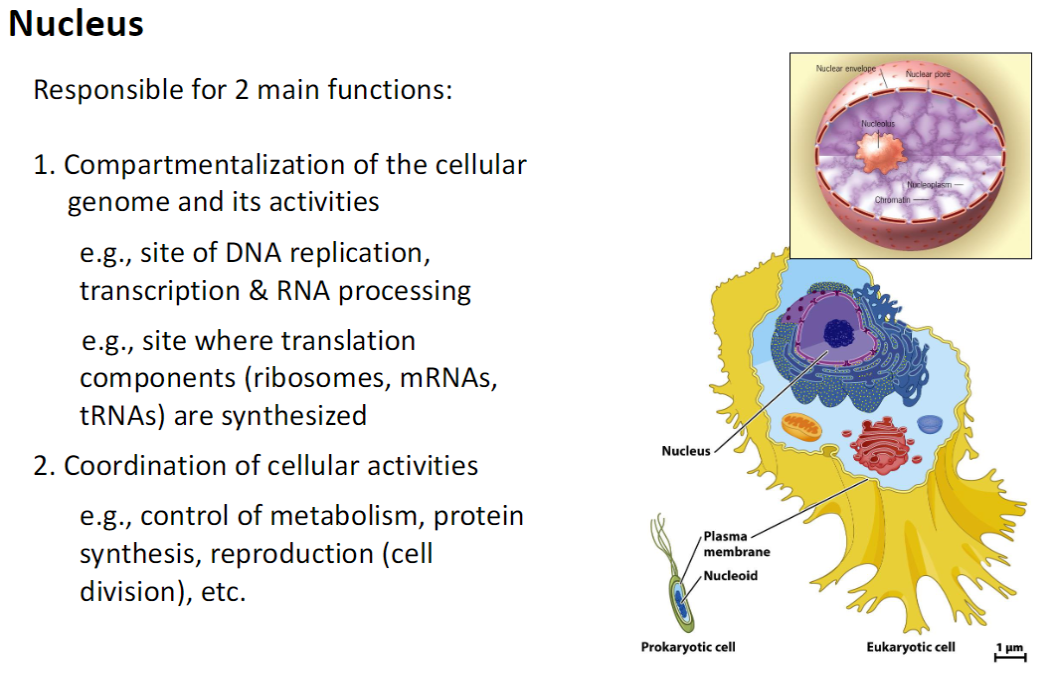
Nucleus – Main Functions
1. Compartmentalization
Houses the cellular genome and its activities
Site of DNA replication, transcription, and RNA processing
Synthesis of translation components: ribosomes, mRNAs, tRNAs
2. Coordination of Cellular Activities
Controls metabolism, protein synthesis, and cell division
Key Point
The nucleus organizes genetic material and coordinates essential cellular functions
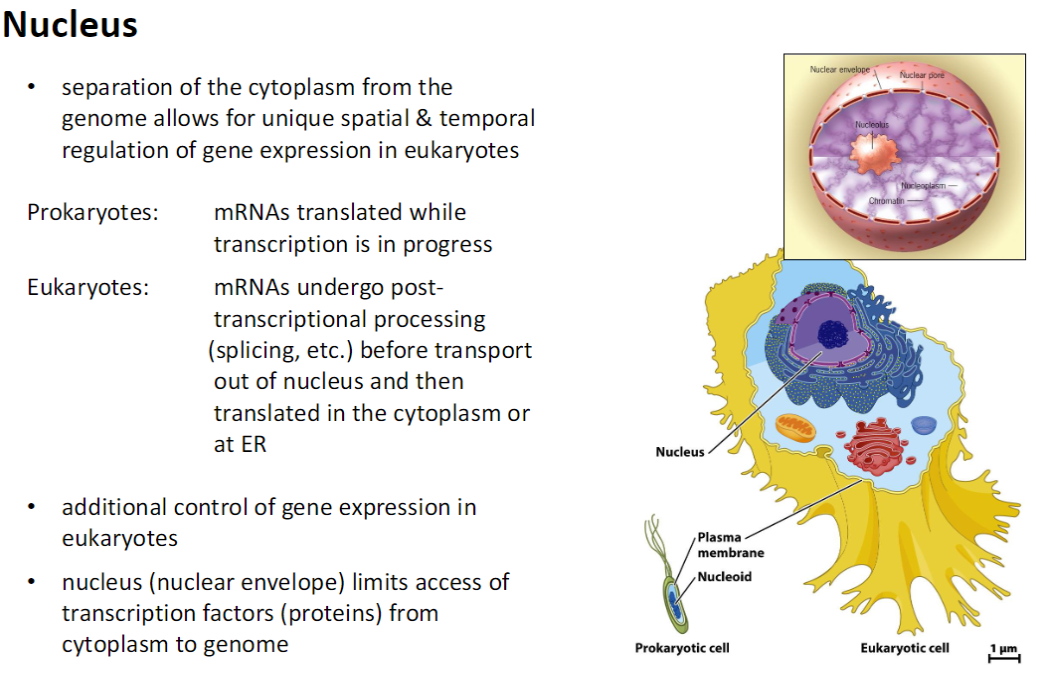
Nucleus – Gene Expression Regulation
Spatial & Temporal Separation
Cytoplasm separated from genome enables controlled gene expression in eukaryotes
Prokaryotes vs Eukaryotes
Prokaryotes: mRNAs are translated during transcription
Eukaryotes: mRNAs undergo post-transcriptional processing (splicing, etc.) before being transported out of the nucleus for translation in the cytoplasm or at the ER
Additional Control
Nuclear envelope restricts access of transcription factors from cytoplasm to DNA
Provides extra regulation of gene expression in eukaryotes
Key Point
Separation of the nucleus allows more precise temporal and spatial control over gene expression
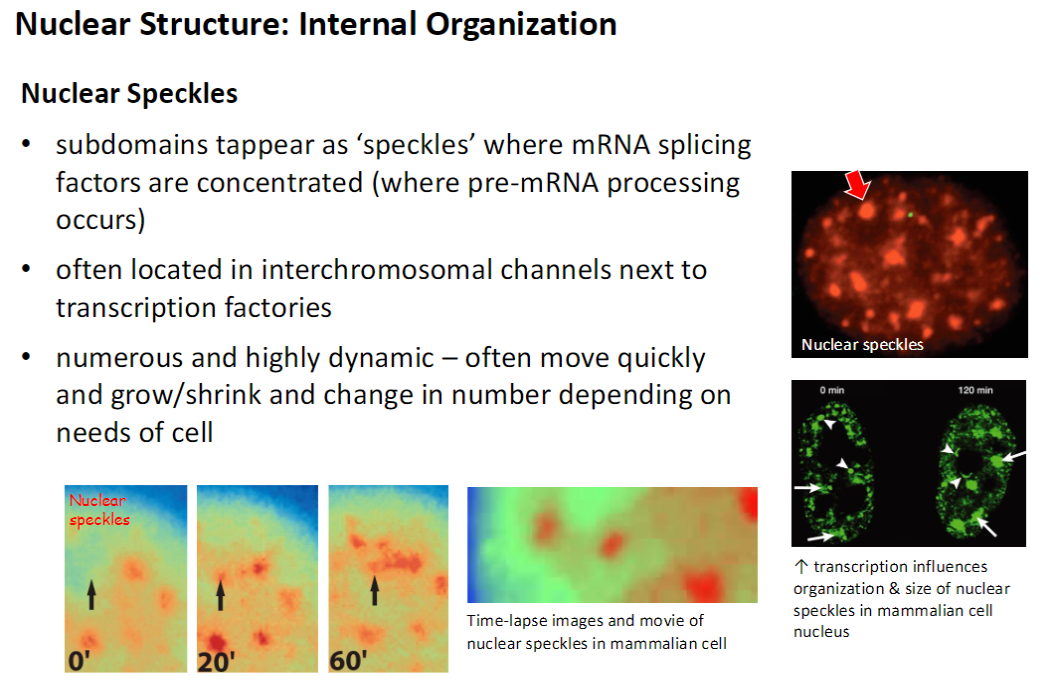
Nucleus – Nucleoplasm & Nucleolus
Nucleoplasm
Fluid-filled interior of the nucleus, highly organized
Contains >30 specialized subdomains for specific functions
Subdomains are not membrane-bound
Nucleolus
Most conspicuous nuclear subdomain, dense and granular
Size and number (1–5) depend on cell metabolic activity – more active cells have larger and more nucleoli
Function: produces ribosomes
Site of rDNA transcription, rRNA processing, and initial ribosomal subunit assembly
Final ribosome assembly occurs in the cytoplasm
Key Point
The nucleoplasm organizes nuclear functions, while the nucleolus specializes in ribosome production
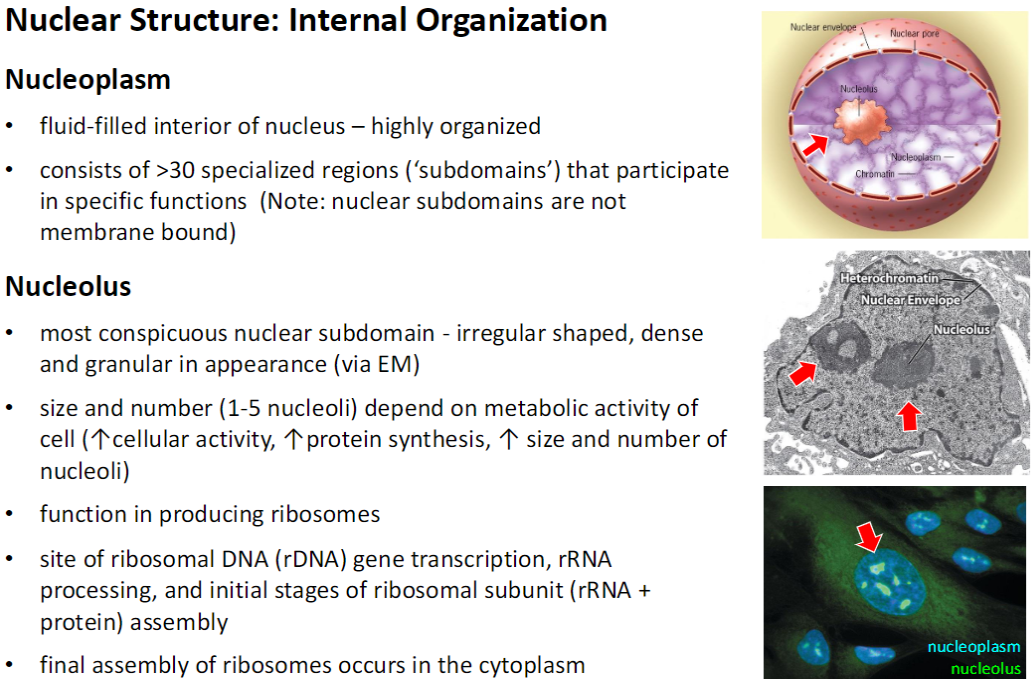
Nucleus – Chromosome Organization
Chromosomal Subdomains
Chromosomes are organized into discrete subdomains during interphase
Gene location often correlates with activity
Euchromatin (actively transcribed genes) found at periphery of subdomains
Inter-Chromosomal Channels
Regions between subdomains that prevent unwanted DNA-DNA or DNA-protein interactions
Transcription Factories
Regions where genes with similar functions cluster to share regulatory elements
Key Point
Nuclear organization supports efficient and regulated gene expression
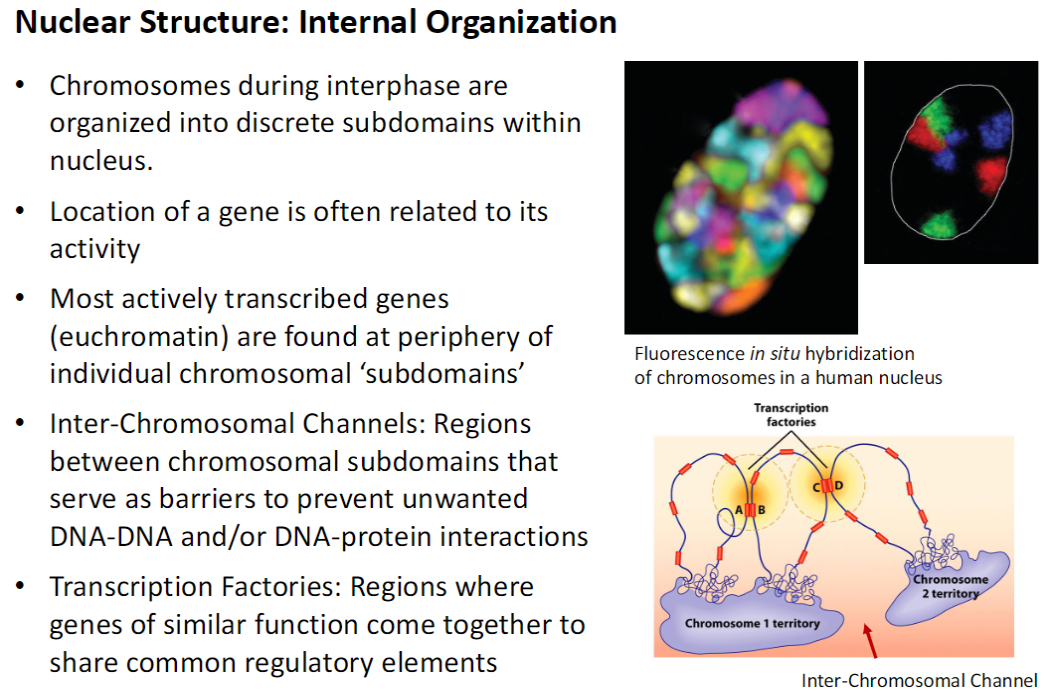
Nucleus – Nuclear Speckles
Definition & Function
Subdomains appearing as ‘speckles’ where mRNA splicing factors are concentrated
Site of pre-mRNA processing
Location
Often found in interchromosomal channels near transcription factories
Dynamics
Numerous and highly dynamic – move, grow/shrink, and change number based on cellular needs
Increased transcription affects organization and size of speckles in mammalian cells
Key Point
Nuclear speckles organize splicing machinery and adapt dynamically to support gene expression
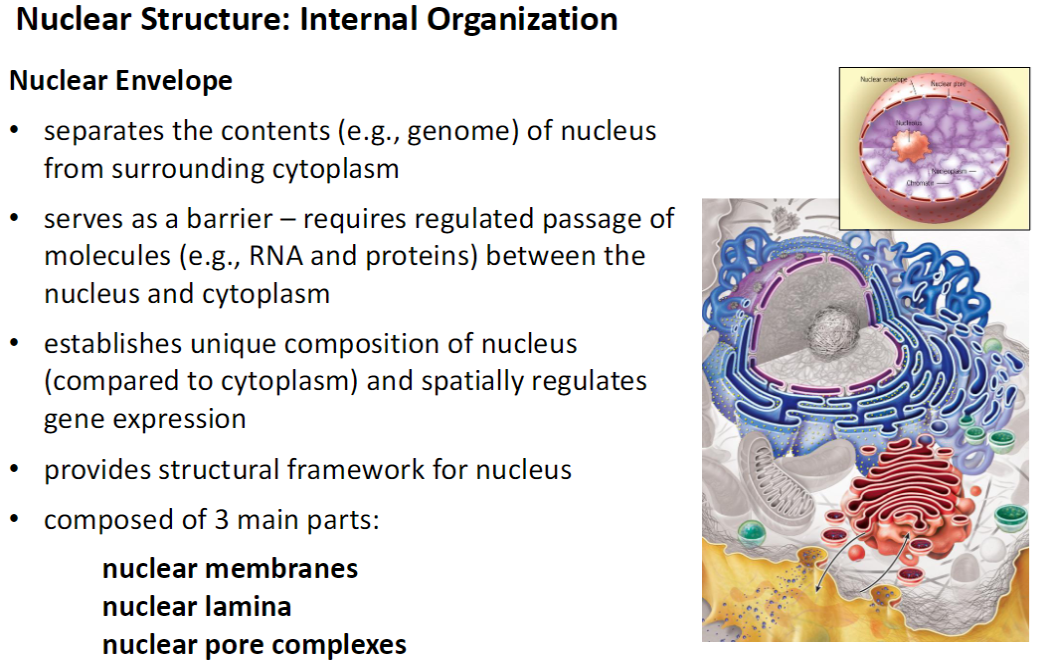
Nucleus – Nuclear Envelope
Definition & Function
Separates nuclear contents (e.g., genome) from cytoplasm
Acts as a barrier, allowing regulated passage of molecules like RNA and proteins
Establishes a unique nuclear composition and spatially regulates gene expression
Provides structural framework for the nucleus
Composition
Nuclear membranes
Nuclear lamina
Nuclear pore complexes
Key Point
The nuclear envelope maintains nuclear compartmentalization and controls molecule transport