CH 13 DNA Replication and Recombination
1/35
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
36 Terms
Semiconservative model for DNA replication
Two DNA strands come apart
Each serves as a template for synthesis of new strands
= 2 newly made daughter strands
= 2 original template/parent strands

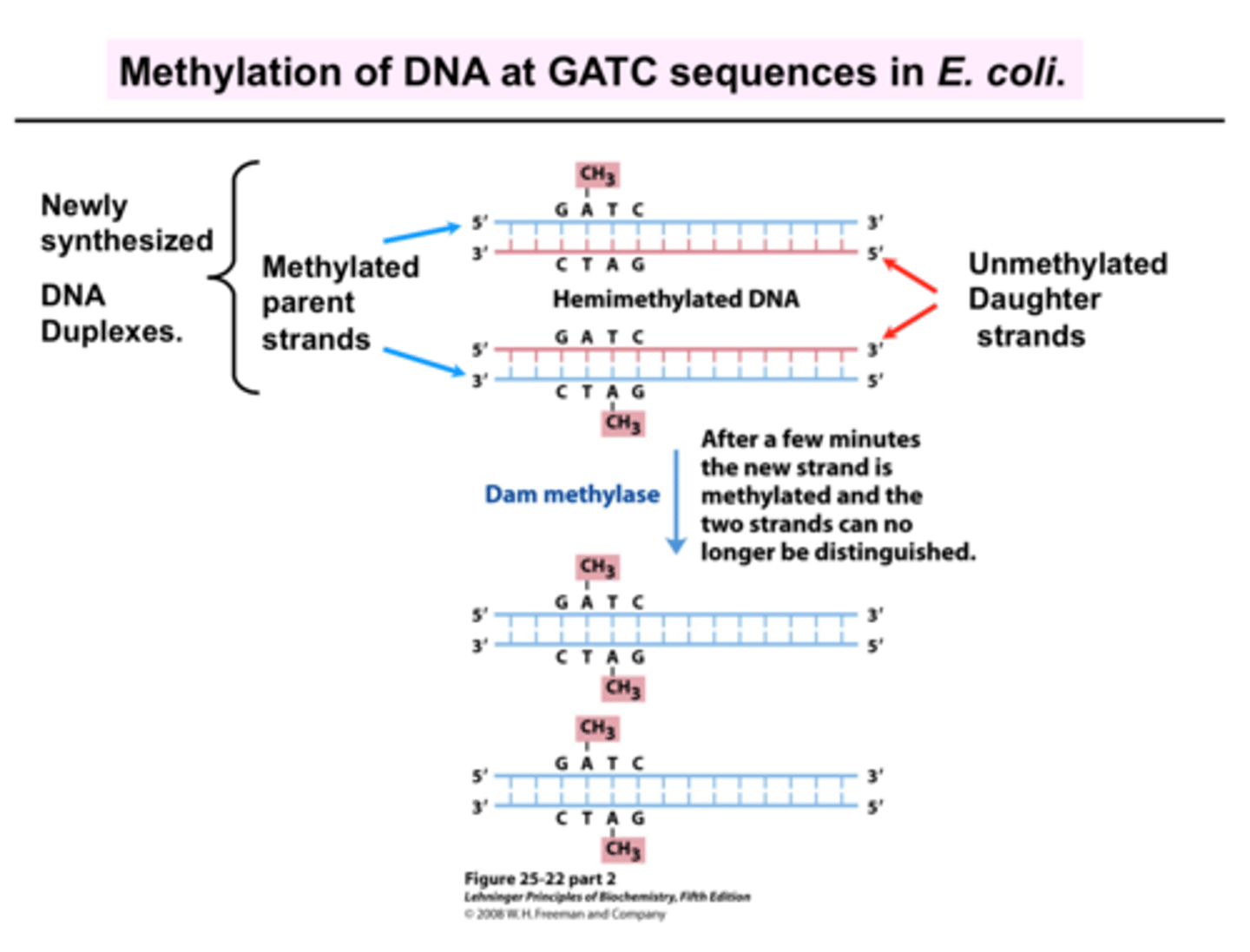
GATC methylation sites
regulate DNA replication
- Parent strands need to be methylated prior to replications (used to mark OG strand)
- New strands aren't methylated (until a bit a/f replication)
- Initiation of replication doesn't occur until DNA is methylated, preventing DNA replication from happening too quickly
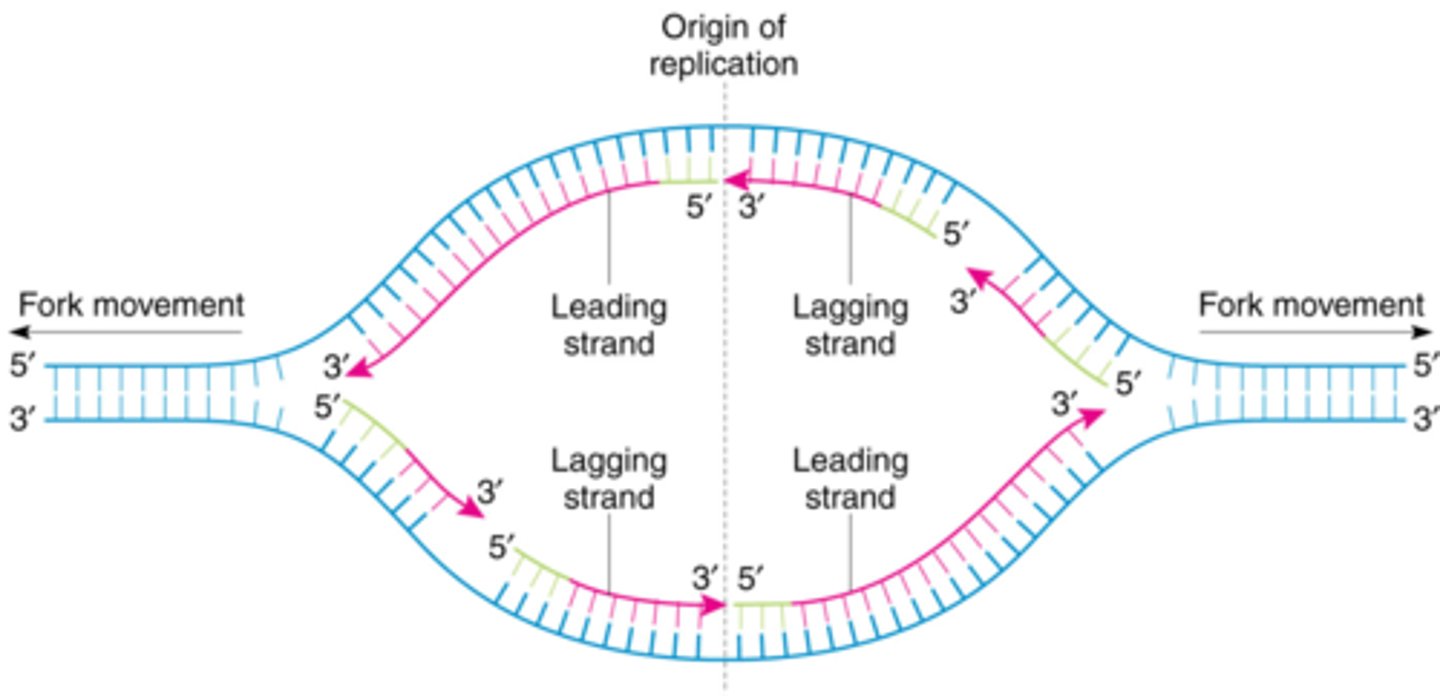
bidirectional replication
A type of DNA replication that occurs in two directions from the origin of replication.
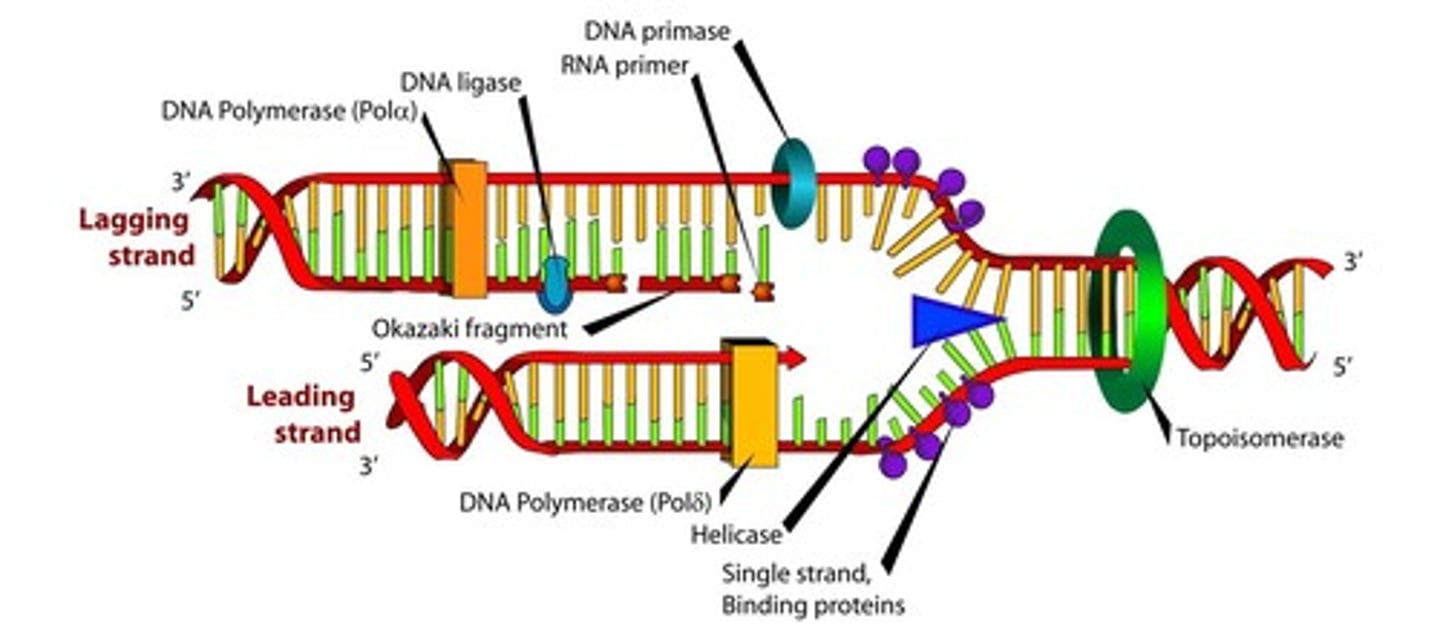
replication fork
- unwinding of helix (helicase)
-synthesis of RNA primers (RNA primase)
-Replication (DNA polymerase)
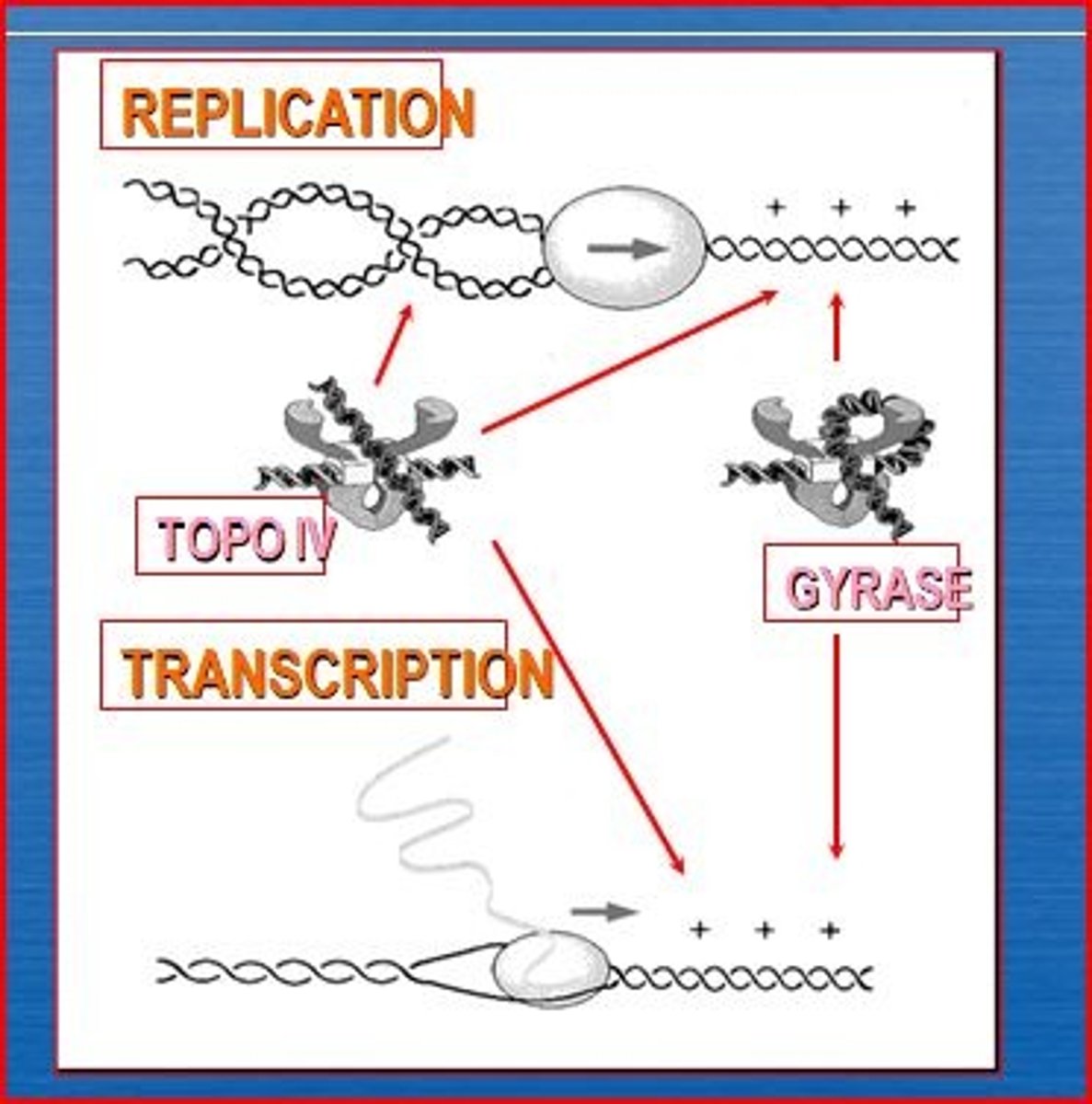
DNA helicase (Dna B)
An enzyme that unwinds the DNA double helix ahead of the replication fork.
DNA gyrase (topoisomerase II)
alleviates the torsional strain generated ahead of the replication fork by introducing negative supercoils.
RNA primers
Short RNA sequences that provide a starting point for DNA synthesis.
primase
An enzyme that synthesizes short RNA primers (~10-12 bp) to initiate DNA replication.
leading strand
The strand of DNA that is synthesized continuously in the same direction as the replication fork (one primer needed)
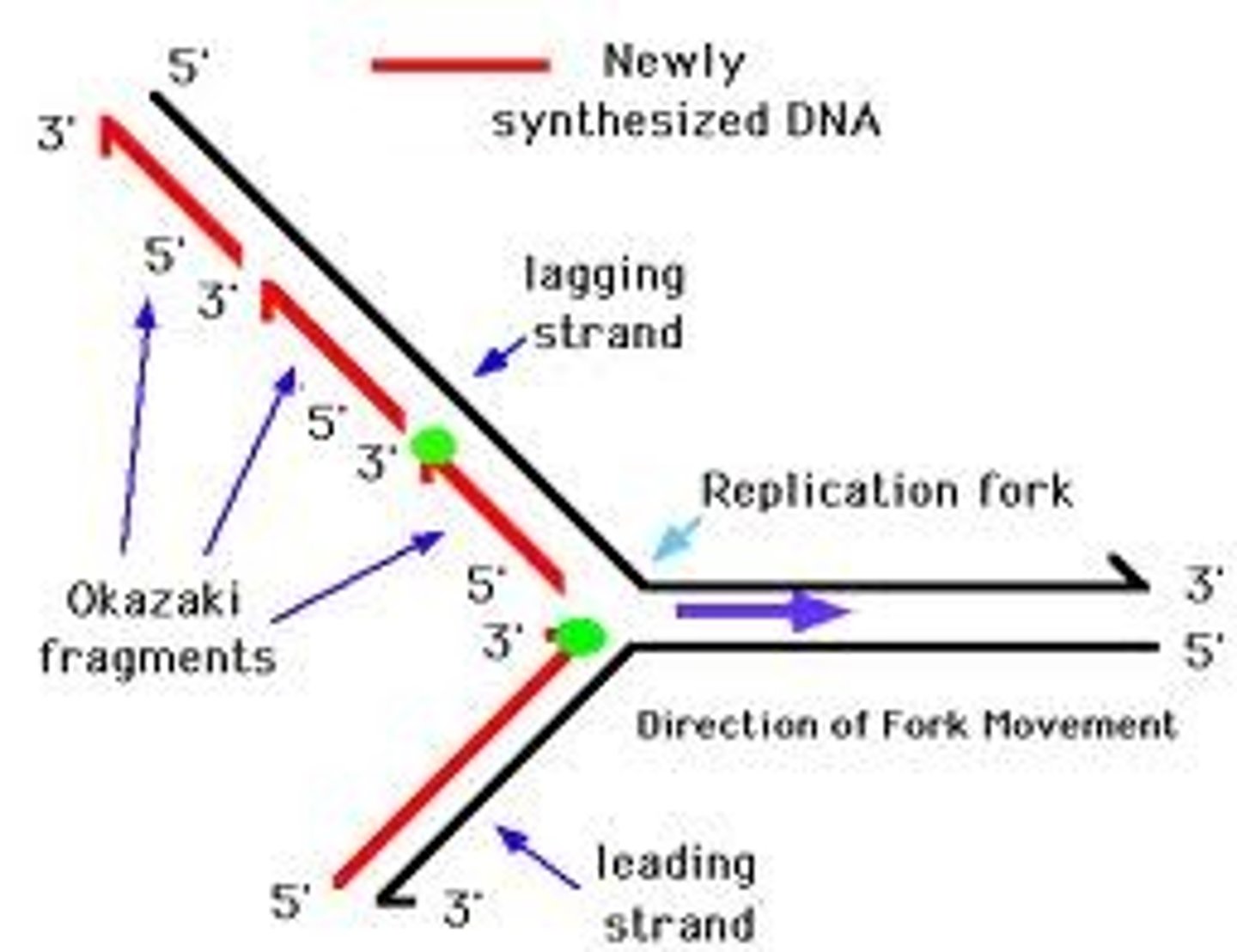
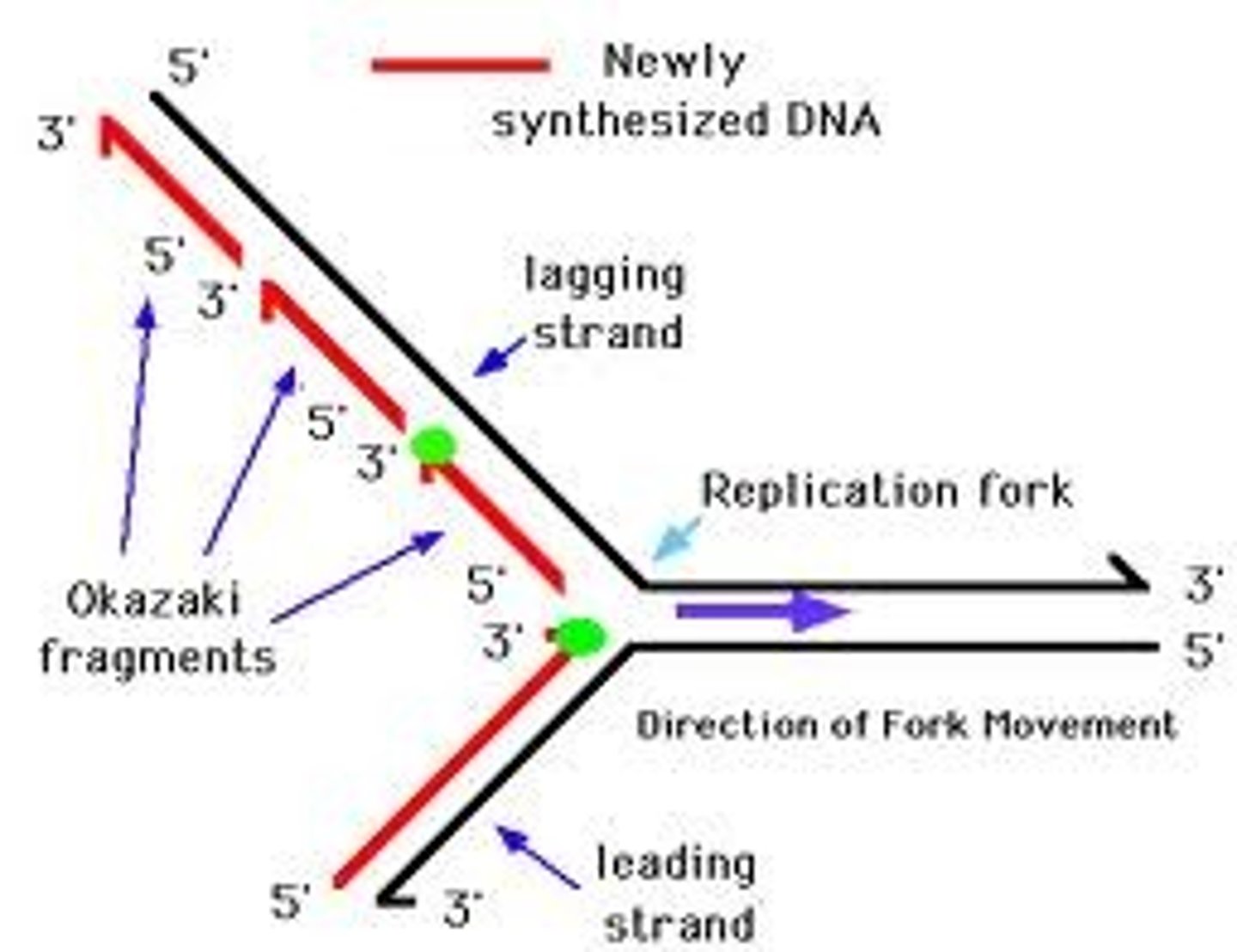
lagging strand
The strand of DNA that is synthesized discontinuously in short segments opposite to the direction of the replication fork (multiple primers)
DNA polymerase
An enzyme that synthesizes new DNA strands by adding nucleotides to a pre-existing chain.
- some have repair function
Okazaki fragments
Short segments of DNA synthesized on the lagging strand during DNA replication.
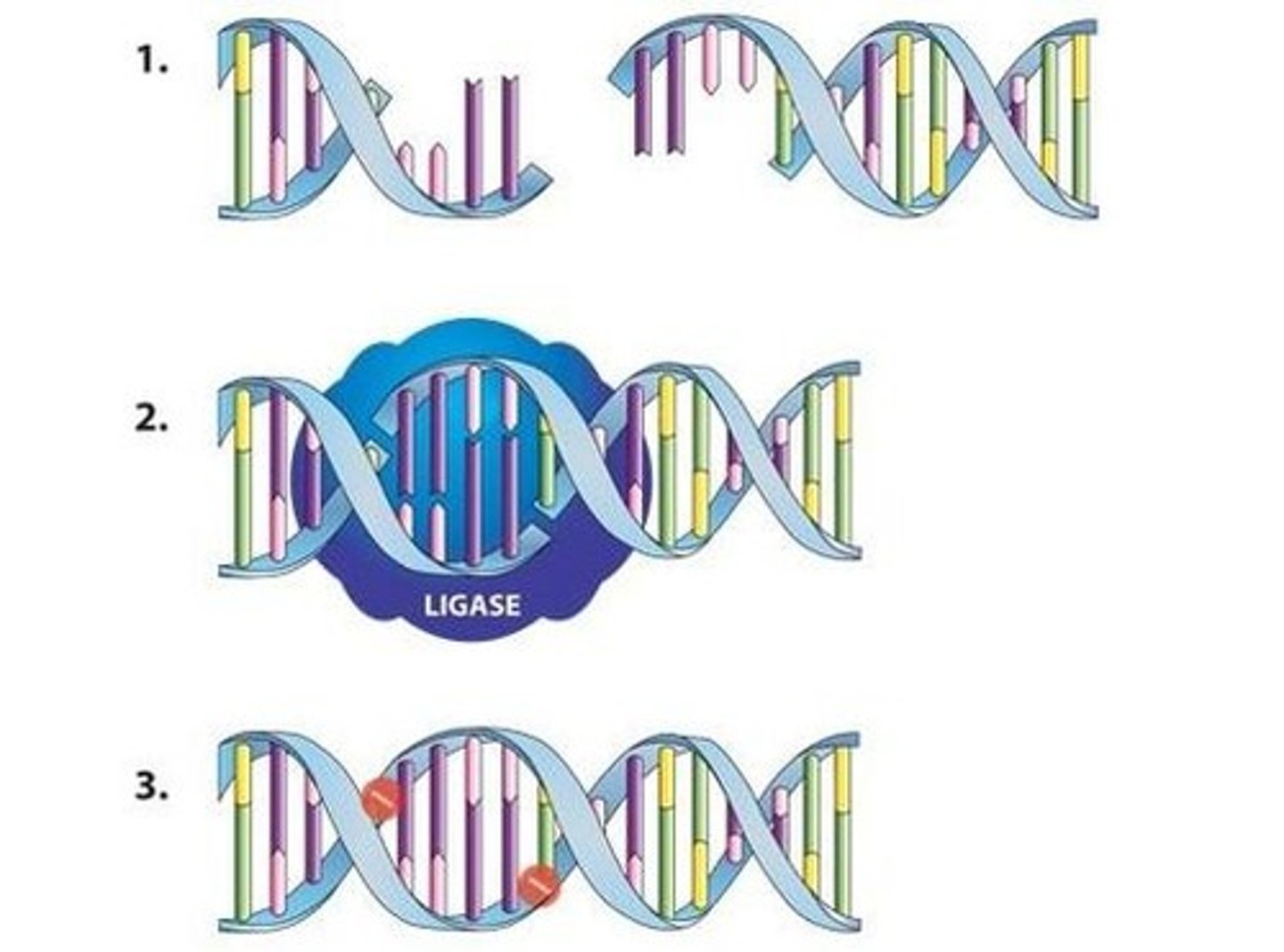
DNA ligase
An enzyme that joins Okazaki fragments together to form a continuous DNA strand.
Bacteria DNA Pols in Replication
- DNA pol 1 and III = normal replication (synthesizes strands)
- DNA pol II, IV and V = DNA repair and replication of damaged DNA
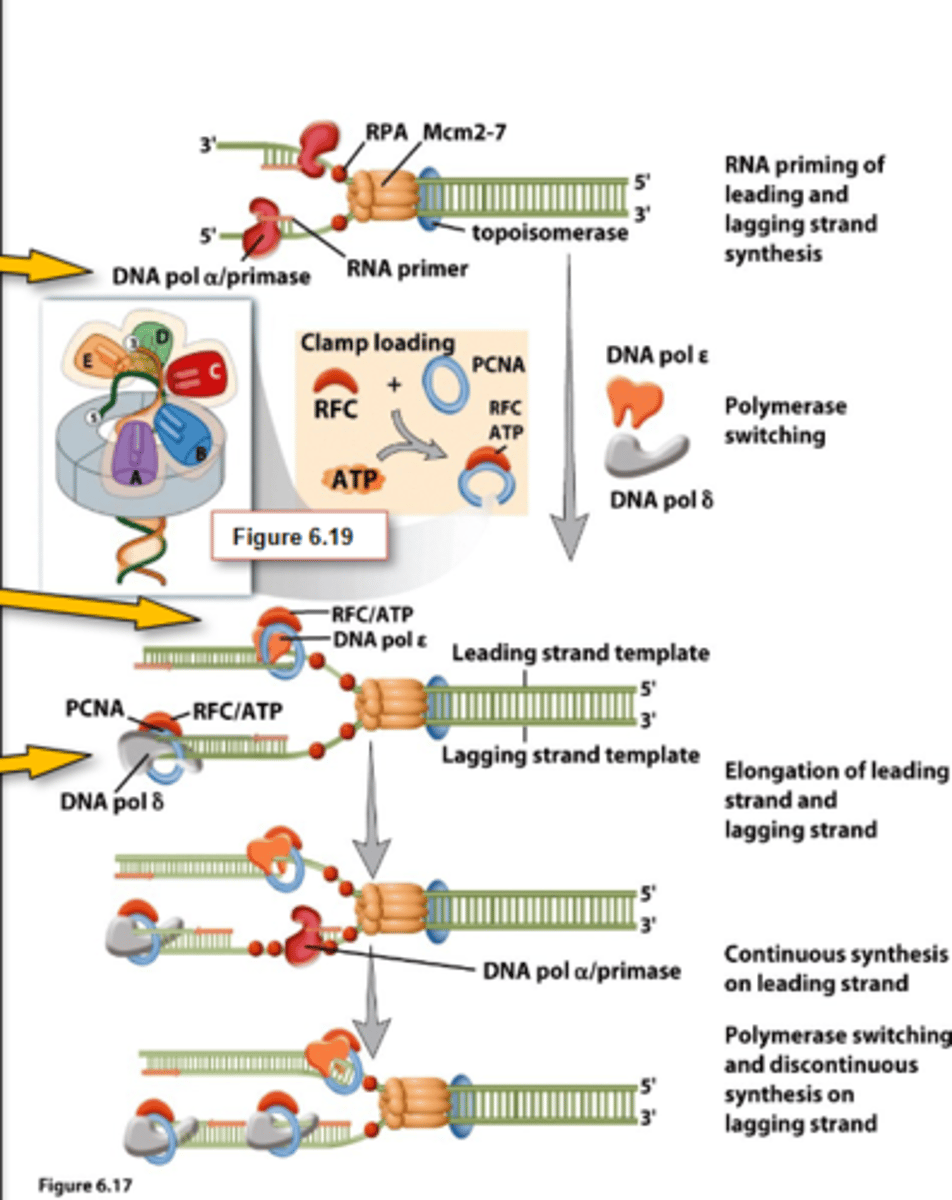
Eukaryotic DNA Pols in Replication
-Alpha (α), delta (δ), epsilon (ε) = Nuclear DNA
~Translesional & nuclear DNA
-Alpha (α): associates with primase to synthesize and RNA primer following by DNA nucleotides
-Epsilon (ε): synthesizes leading strand
-Delta (δ): synthesizes lagging strand
~ gamma (γ) = Mitochondrial DNA
Processive enzyme
An enzyme that catalyzes consecutive reactions without releasing its substrate.
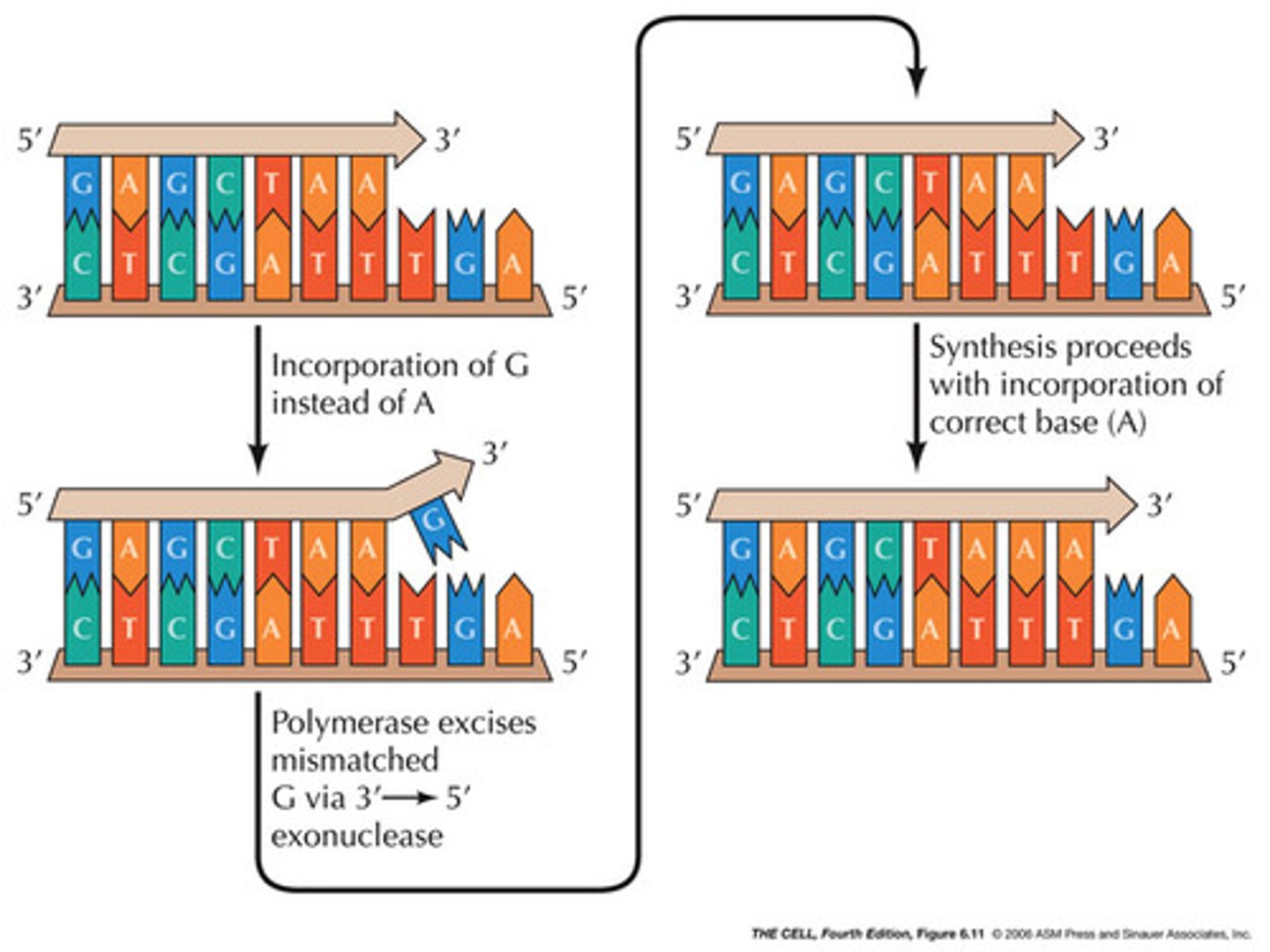
proofreading
The process by which DNA polymerases correct errors in DNA synthesis.
-uses 3' to 5' exonuclease
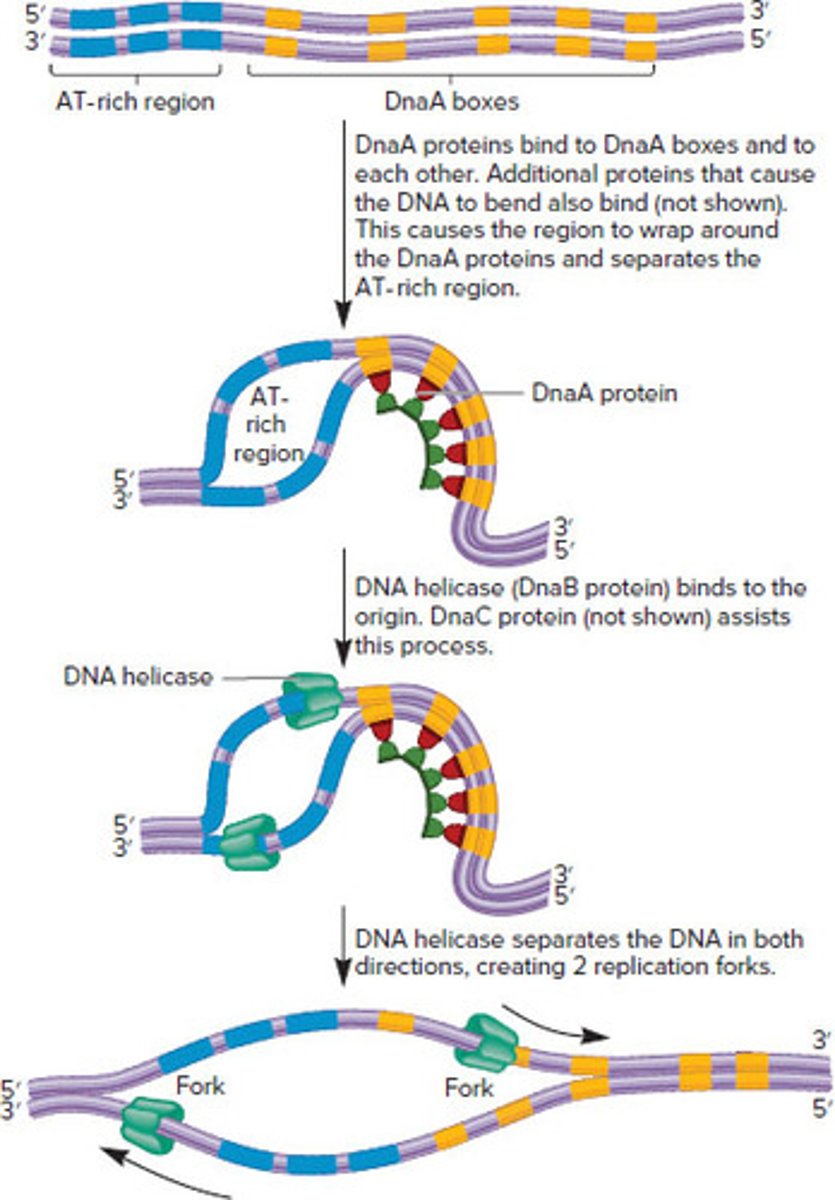
Replication initiation in Bacteria
1. DnaA proteins: bind to DnaA box sequences in oriC, and additional DnaA proteins bind to each other
2. Proteins cause the region to bend around the DnaA proteins and separates to AT-rich region
(ATs are easier to separate because of two hydrogen bonds instead of three)
3. DNA Helicase (DnaB protein) and DnaC unwind DNA. Separating the strands in both directions beyond the origin
(Use ATP for energy to drive strands separation)
4. DNA helicases travel along DNA 5' to 3' & two replication forks proceed outward from oriC (bidirectional replication)
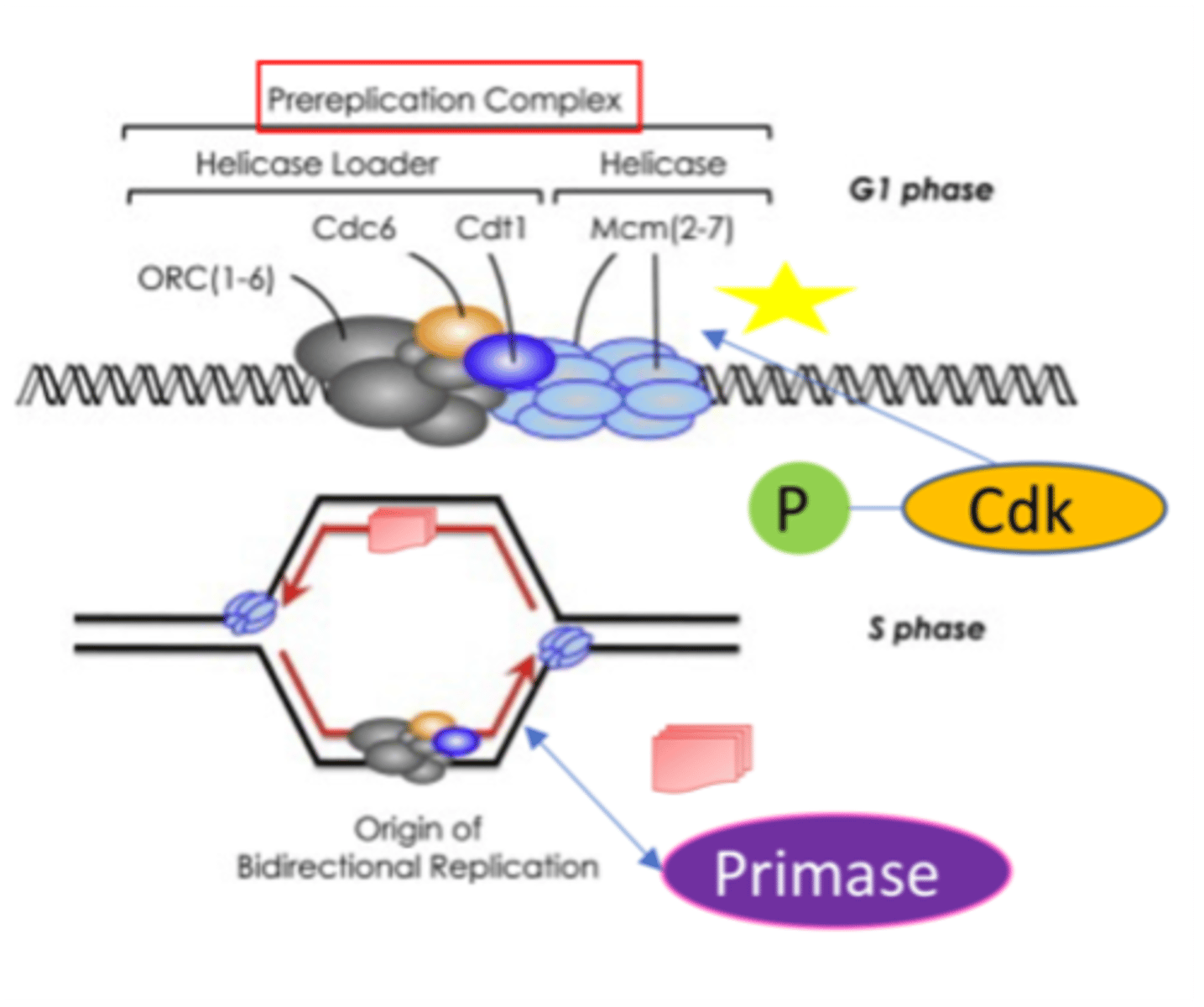
Replication initiation in Eukaryotes
1. Prereplication complex (preRC): assembles to begin replication
- Includes origin recognition complex (ORC)
- ORC = six proteins & acts as initiator of DNA replication
2. ORC promotes the binging of Cdc6l Cdt1, and MCM helicase completing process of DNA replication licensing
3. DNA replication can then proceed
- PreRC converted to an active replication site by phosphorylation
- DNA replication proceeds bidirectionally from origin
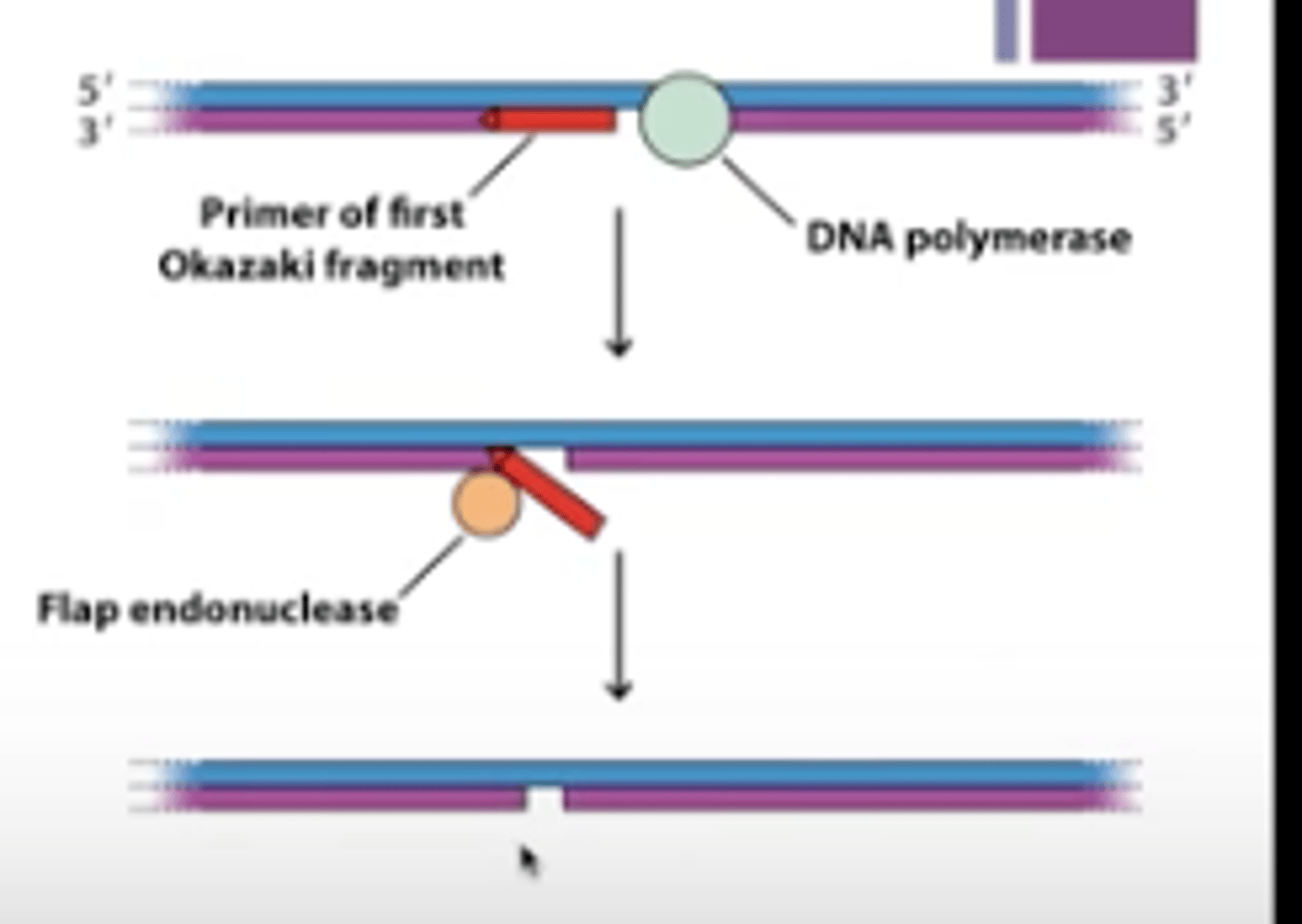
flap endonuclease
removes primers in eukaryotes at end of DNA replication
- Removes small RNA flaps generated by DNA pol delta
DNA pol delta elongates Okazaki fragments until it runs into RNA primer, pushing it up to create flap, then flap endonuclease removes flap and this repeats until RNA primer is completely removed
telomeres
-At end of both ends of linear eukaryotic chromosomes
-Special proteins bind to sequences
(too short for primer to bind final bit of DNA on lagging strand)
senescence
The process of aging at the cellular level, often associated with telomere shortening.
-cells lose ability to divide
telomerase
An enzyme that extends telomeres, preventing their shortening during DNA replication.
- maintain length of telomere by adding DNA to end
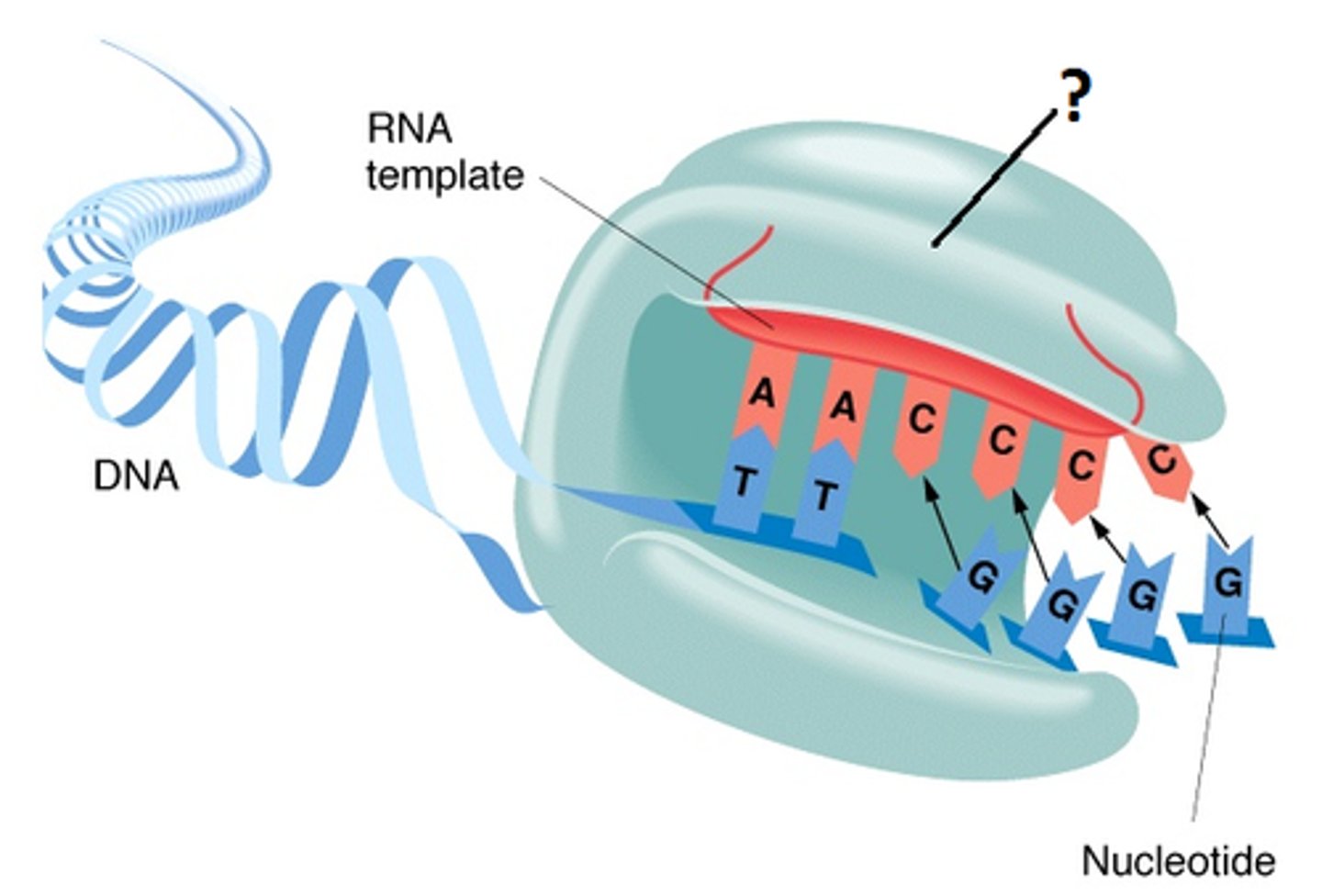
how does telomerase prevent telomere shortening?
- Contains protein (TERT) and RNA (TERC) = RNA is complementary to DNA sequences found in the telomeric repeat
- Allows telomeres to bind to 3' overhang & add DNA lost on telomere
Homologous recombination
process where DNA segments that are similar or identical to each other break and region
- crossing over
-sister chromatid exchange
sister chromatid exchange (SCE)
crossing over that occurs between sister chromatids
- Sister chromatids are genetically identical; therefore, SCE does not produce new combination of alleles
genetic recombination
The process of exchanging genetic material between different organisms or between different chromosomes.
-crossing over
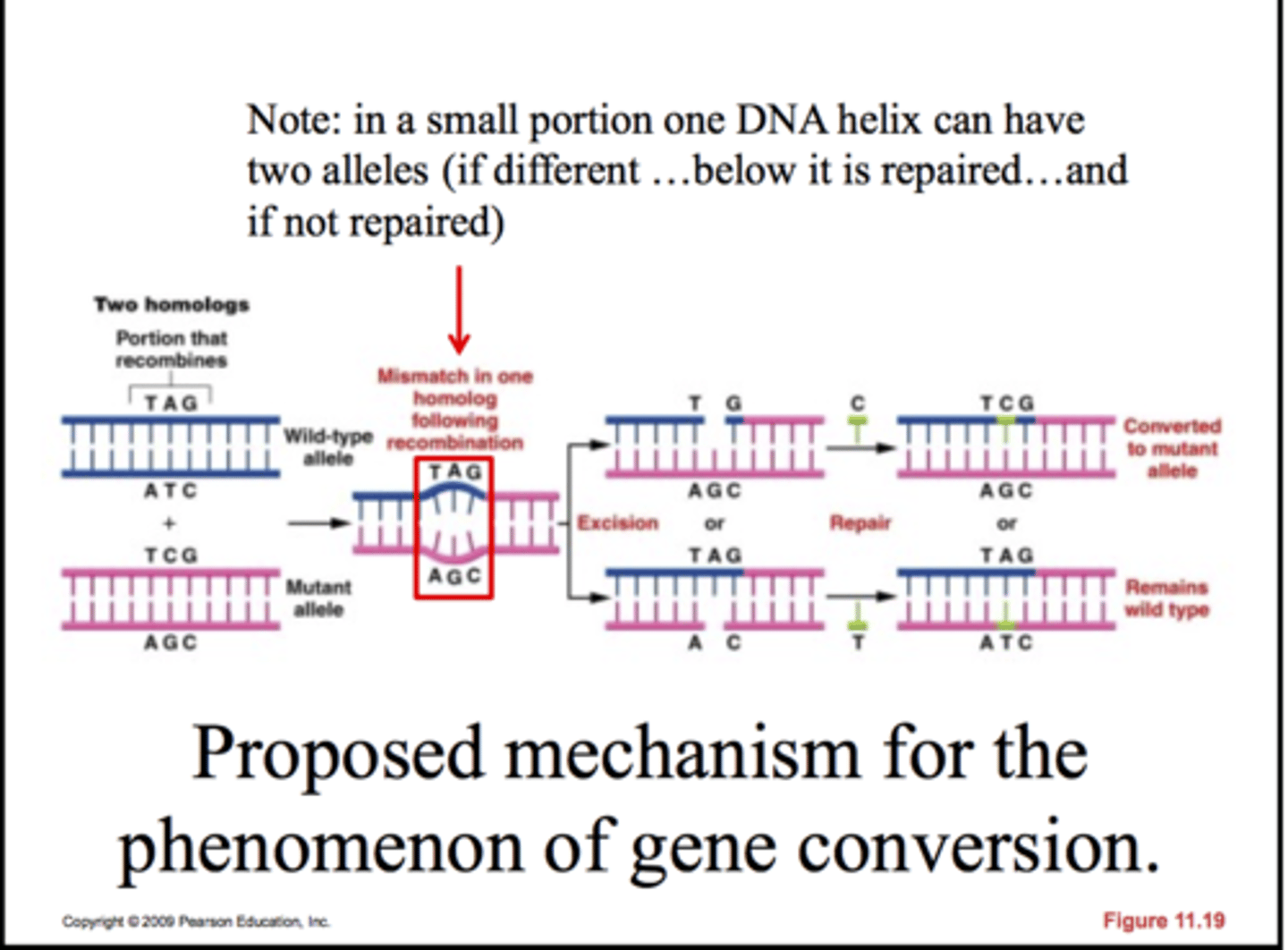
gene conversion
A process that results in the non-reciprocal (not between homologous parts) transfer of genetic information.
-one allele is converted to allele on the homologous chromosome
- two ways: DNA mismatching repair & DNA gap repair synthesis
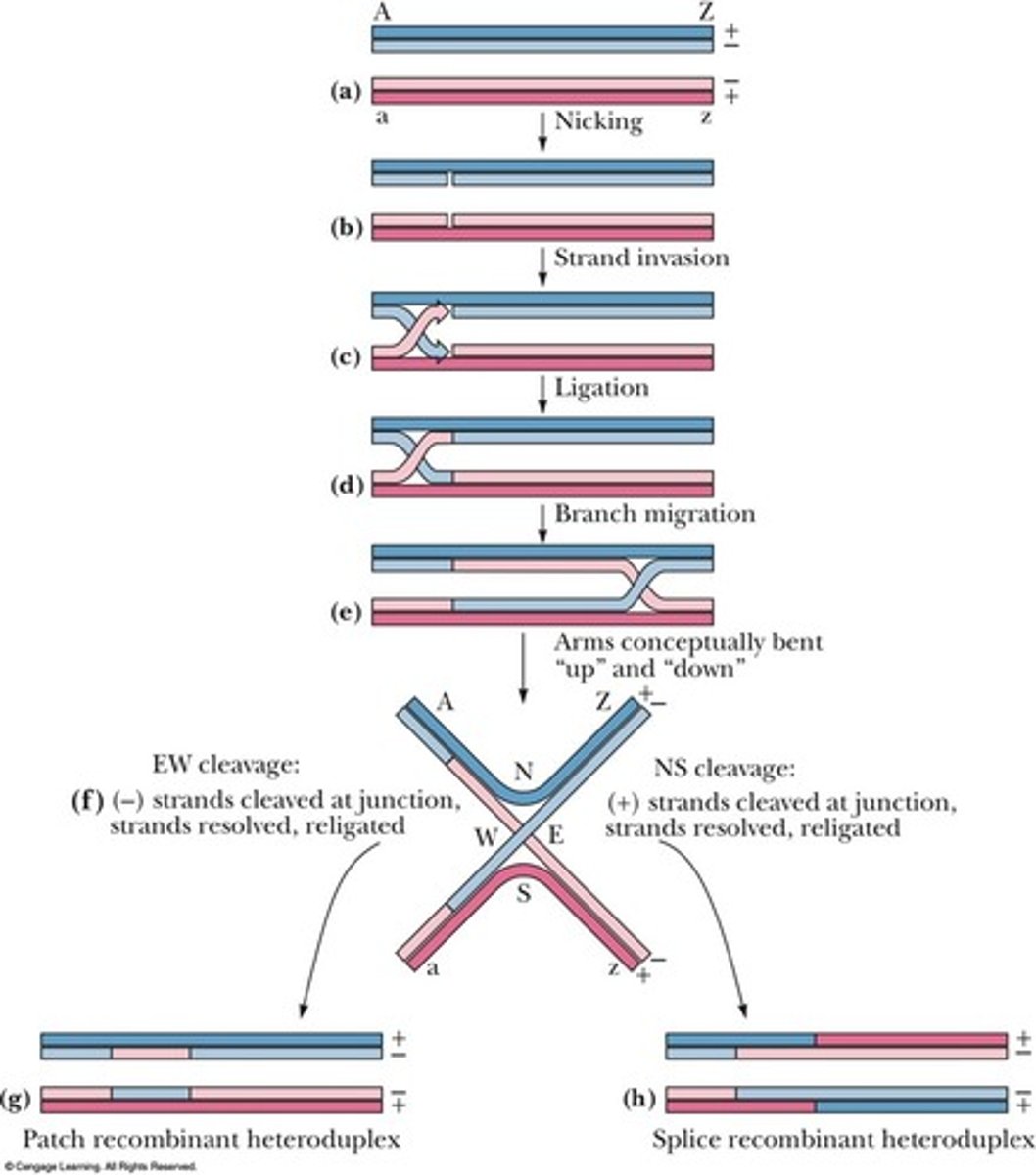
Holliday model
A model that describes the mechanism of homologous recombination.
1. 2 helixes are nicked identical places and one stranded segment of each invades other
2. Segment when connected to opposite helix create heteroduplexes, create larger hetero segments by migrating down branch
3. Intangled DNA is resoved by another nick in both helixes allowing the two changed helixes to separate (2 ways)
- Horizontal cut = 2 heteroduplexes & no recombinants
- Vertical cut (involving a 180 degree rotation) creates a 4-way intersection that when cut vertically = 2 heteroduplexes & recombinant
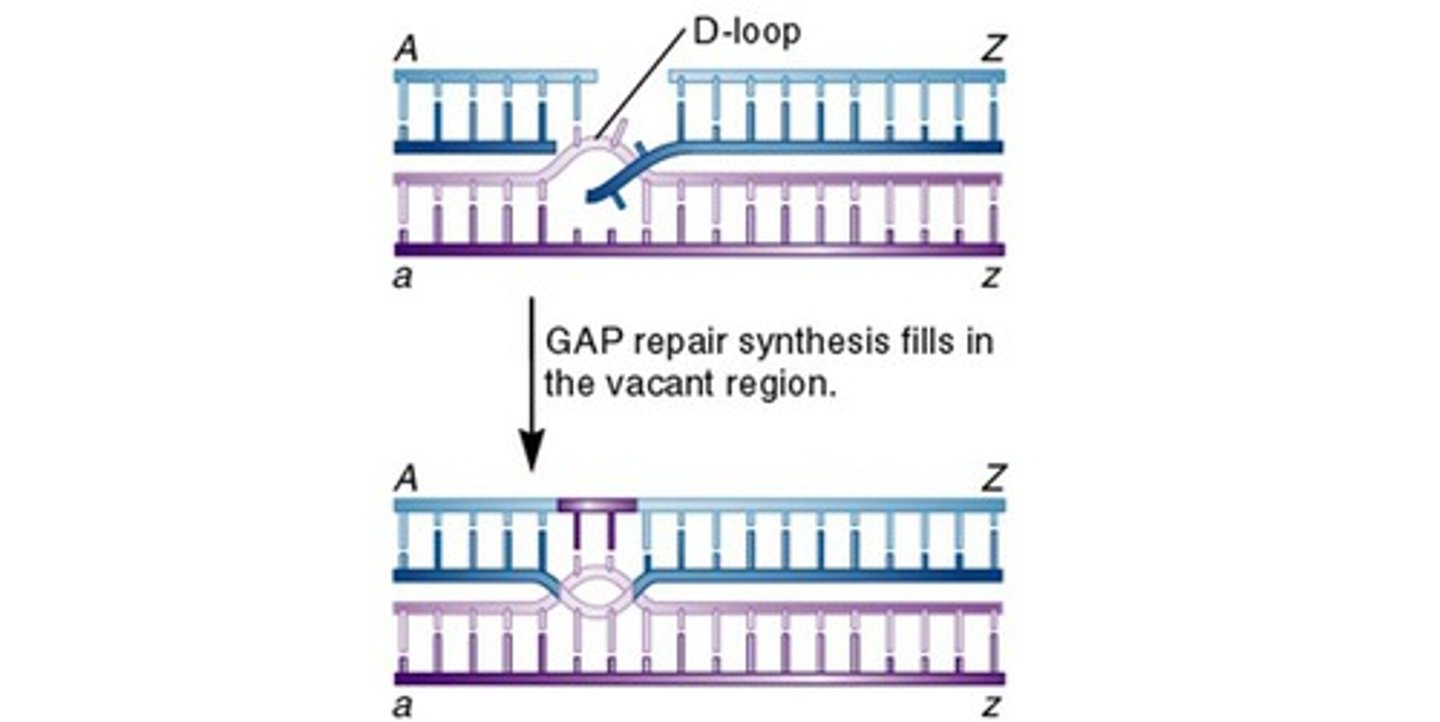
double-strand break model
Like Holliday bit more likely-> only 1 DNA helix (duplex) is nicked (double strand break)
- Both strands of DNA duplex breaks and those strands invade another duplex (forms D loop on invaded duplex)
- Gap repair synthesis fills in all other parts of structures (each has mix of the other)
- Can produce recombinant or nonrecombinant chromosomes depending on branch migration
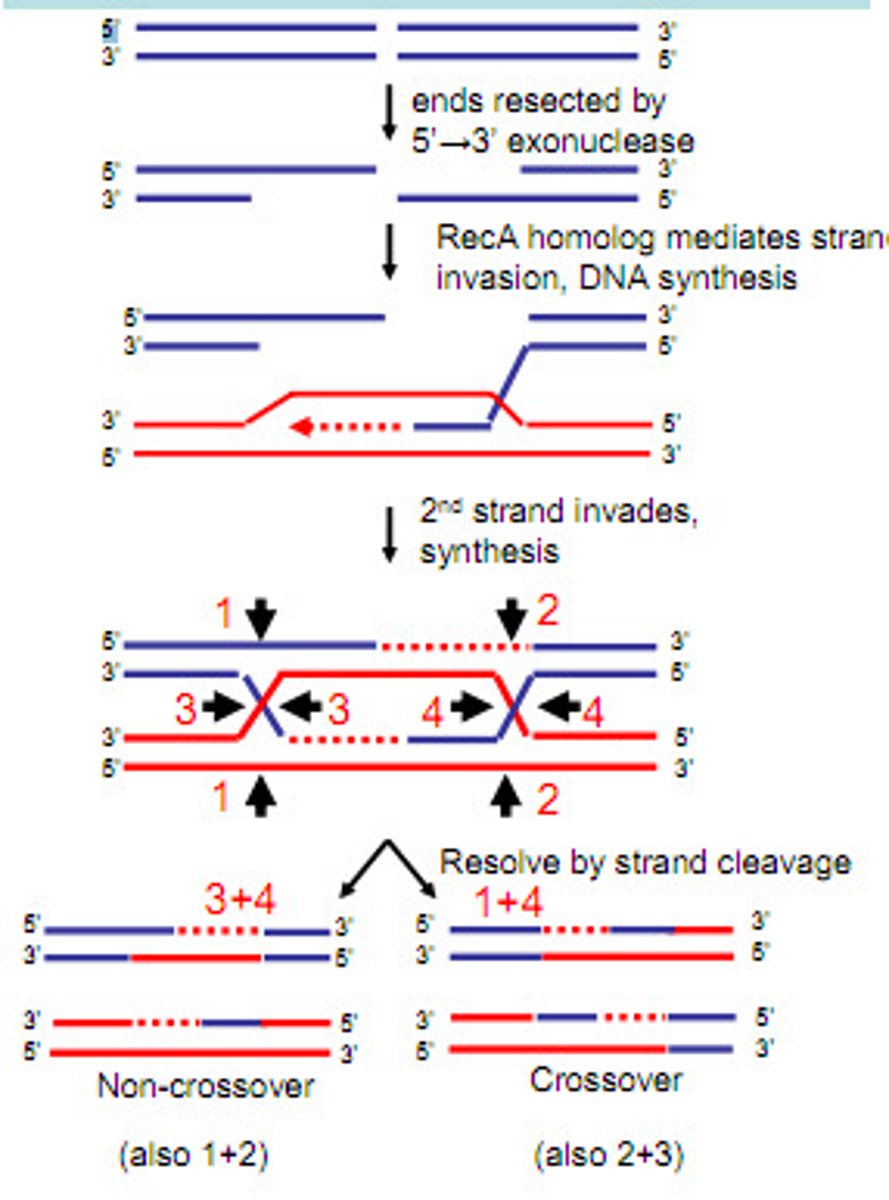
DNA gap repair synthesis
The process of filling in gaps in DNA during replication or repair.
Single Stranded Binding Protein
Binds to single strand DNA and prevents from re-forming a double stranded structure until replication is complete.
DNA pol III
Synthesizes DNA using RNA primers in leading and lagging strands
DNA pol I
Removes RNA primers, fills in gaps with DNA
Dna A
Binds to DnaA boxes within the origin to initiate DNA replication
Dna C
Aids DnaA in the recruitment of DNA helicase to the origin