Post Translational Modifications
1/39
Earn XP
Description and Tags
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
40 Terms
natural protein restriction
limited protein side chains …..but it can modify residues to create new functions
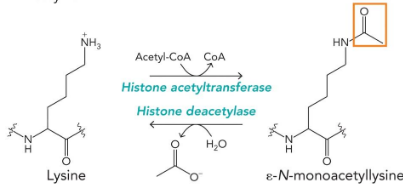
uncommon natural amino acids
modifications on amino acids incorporated into proteins, reversible modifications for regulation, intermediates in metabolism (urea cycle)
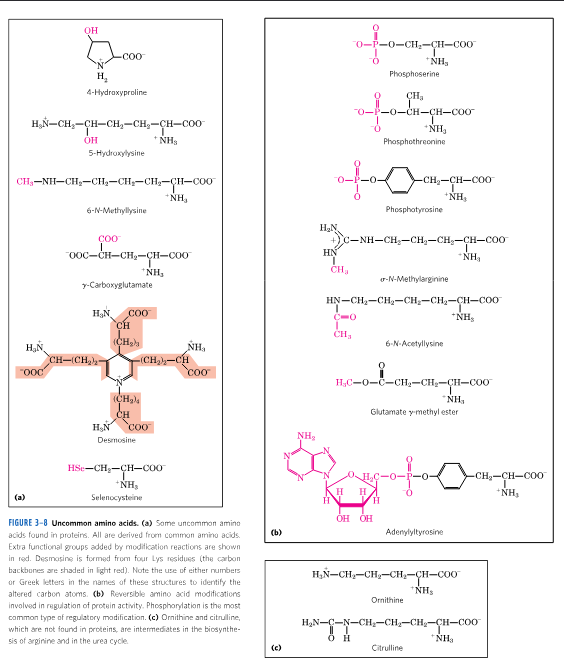
types of protein PTMs
phosphorylation
tyrosine sulfation
lysine acetylation
lysine ubiquitination
lysine methylation
arginine methylation
glycosylation
hydroxylation
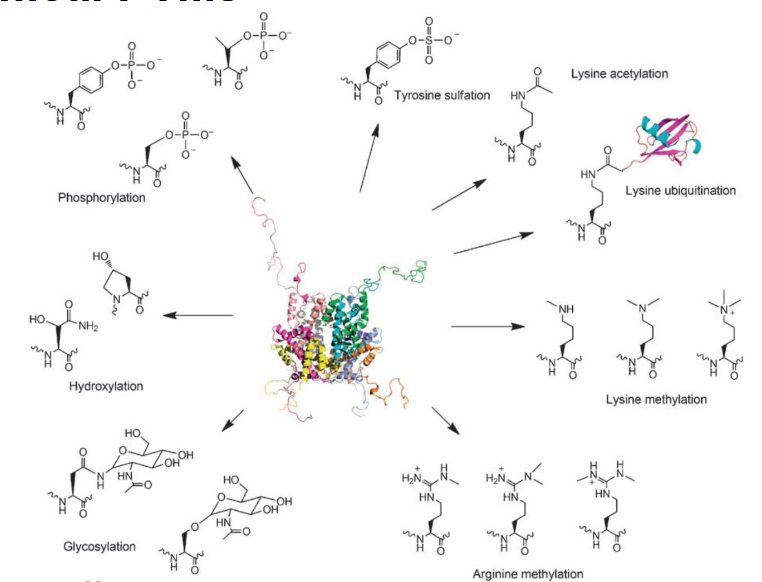
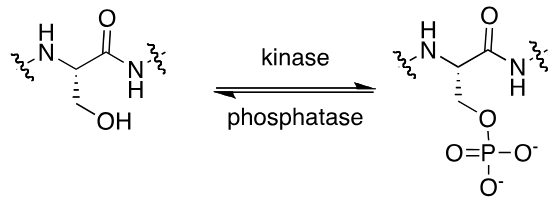
phosphorylation
modifies hydroxyl groups
enzymes: kinase/phosphatase
substrates: ATP/water
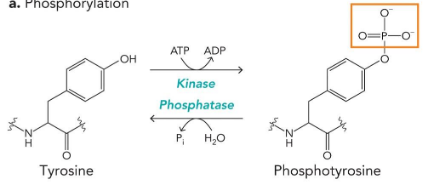
acetylation
lysine
enzymes: histone acetyltransferase/histone deacetylase
substrates: Acetyl-CoA/water
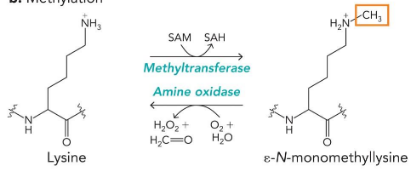
methylation
lysine, arginine, glutamate
enzymes: methyltransferase/amineoxidase
substrates: SAM/O2+water
often irreversible
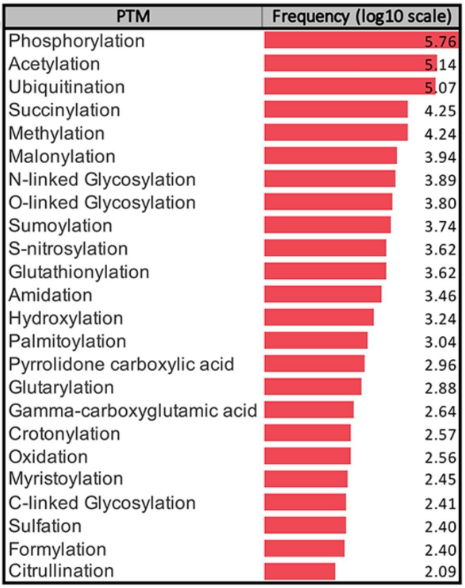
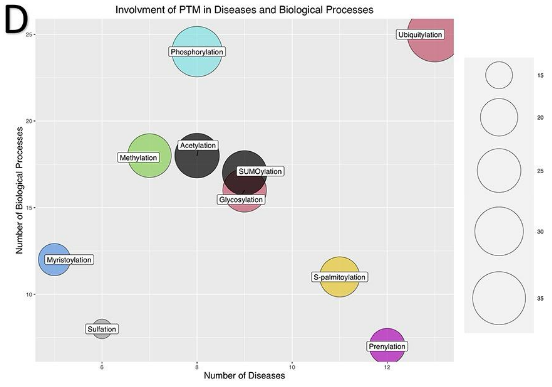
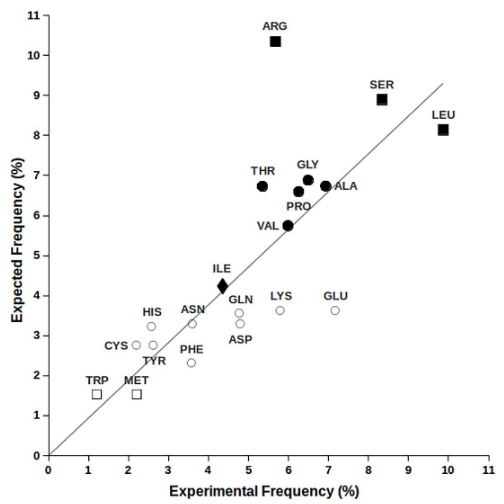
frequency of types of PTMs
Phosphorylation, acetylation, and ubiquitination account for ~90% of validated PTMs
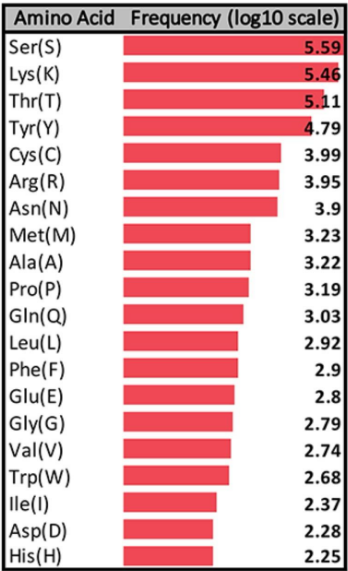
Frequency of SItes of PTMS
A wide variety, but Serine, Lysine, Threonine and Tyrosine dominate.
Methionine modifications
f-Met
Phenylalanine modification
amine removal/alteration
Frequency distribution og sites and types of PTMs
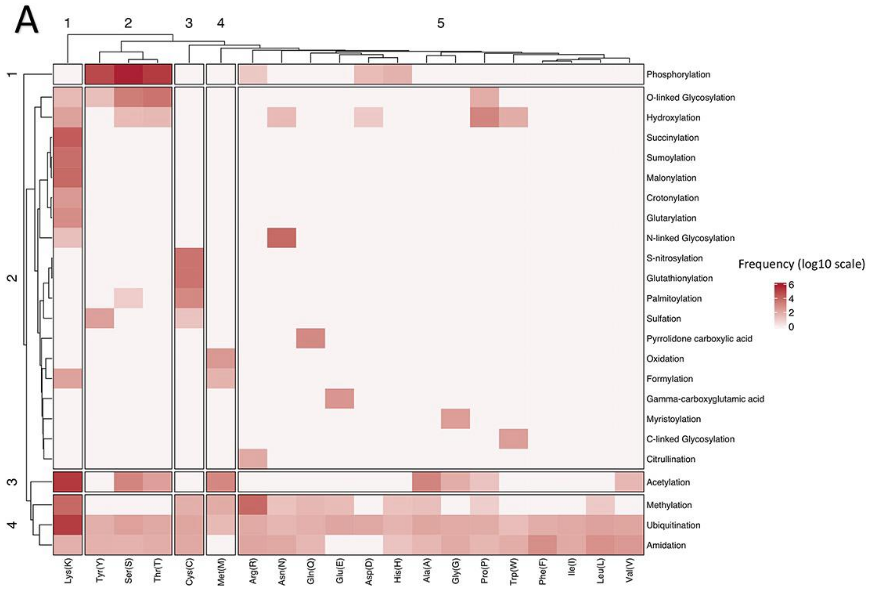
involvement of PTM in disease and biological processes
PTM compartmentalisation
proteasome + ubiquitination
phosphorylation + cell signalling
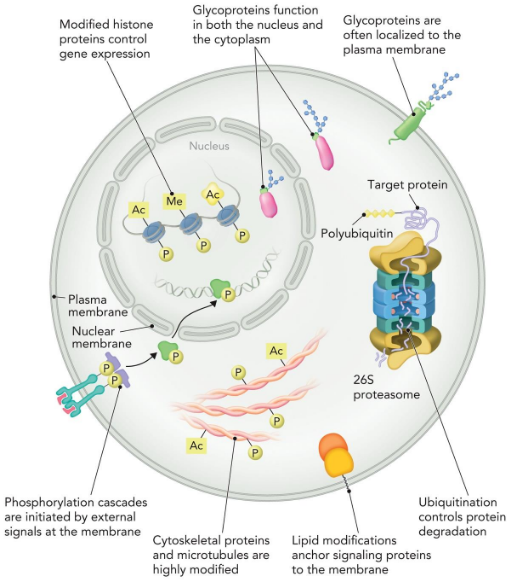
Molecular consequences of post-translational modifications
• enzyme function and assembly
• protein lifespan
• protein–protein interactions
• cell–cell and cell–matrix interactions
• molecular trafficking
• receptor activation
• protein solubility
• protein folding
• protein localization
protein kinase
enzyme that modifies serine, threonine, tyrosine and sometimes histidine on proteins by chemically adding phosphate groups to them
functional change of the target protein by changing enzyme activity, (decrease or increase in rate of achemical reaction), cellular location, or association with other proteins.
30% of all proteins may be modified by kinase activity
human genome codes for 518 protein kinases, about 2% of all genes
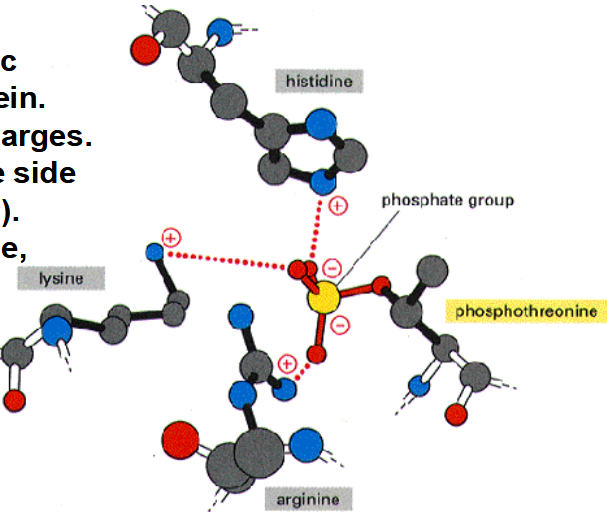
Why phosphorylation?
1. Causes allosteric changes in protein.
2. Two negative charges.
3. Attracts positive side chains (Lys, Arg).
4. Occurs on Serine, threonine, and tyrosine.
Why methylation?
no change in charge for Arg/Lys
change in sterics, H-bonding
alter protein-protein interactions
Histone methylation of lysines by histone methyltransferases can reorganize chromatin, change gene expression
Why acetylation?
Removes a charge
Hydroxylation
proline
enzymes: prolyl hydoxylase
substrate: O2 + alphaketoglutarate + Vit C → CO2 + succinate
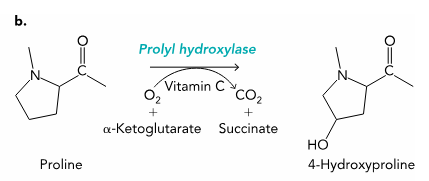
Why hydroxylation?
collagen
How common an amino acid is hydroxyproline?
very, collagen is a big part of cell structure in animals; Hyp is more common that seven coding amino acids (~4% of all proteins)
Is there an aminoacyl-tRNA synthetase for this amino acid?
No, proline is added then oxidised
Cysteine modifications
disulfide crosslinks - extracellular environment is generally oxidizing
Nitrosylation - signaling through a diffusable gas
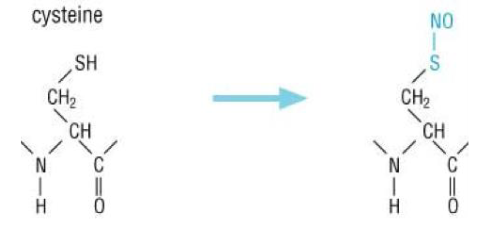
Nitrosylation
cysteine
enzyme:
substrate: NO

Destination of proteins
dictated by PTMs
proteins synthesized initially by free ribosomes are destined for the nucleus, mitochondria, peroxisomes, and chloroplasts.
Proteins synthesized on the ER membrane are destined for the plasma membrane or for various membrane compartments in the cytosol. ER-synthesized proteins are often posttranslationally modified with covalently attached lipids and carbohydrates.
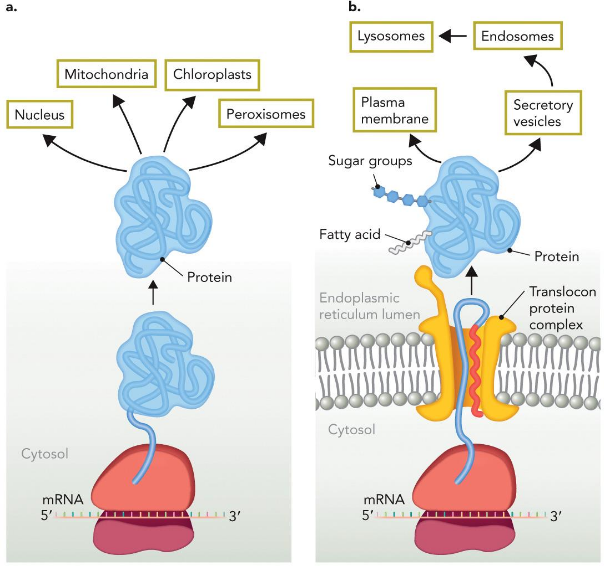
Lipid Modifications
anchor proteins on membranes
Prenylation
makes thioether
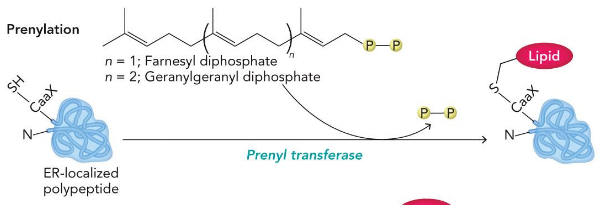
myristoylation
make amide
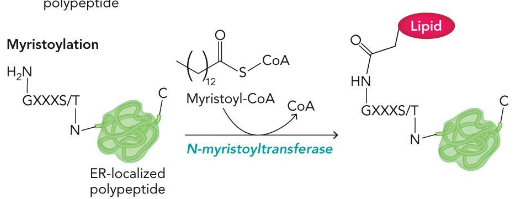
palmitoylation
make thioester
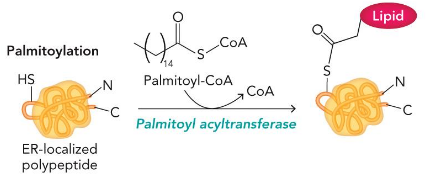
Glycosylation
sugars on proteins
Why glycosylation?
increases solubility - sugars = constrained water
influences structure and molecular interactions
sialic acid
N-linked sugars
attached to Asn
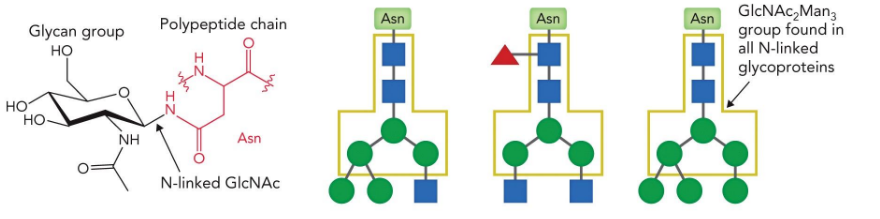
O-linked sugars
attached to Ser/Thr
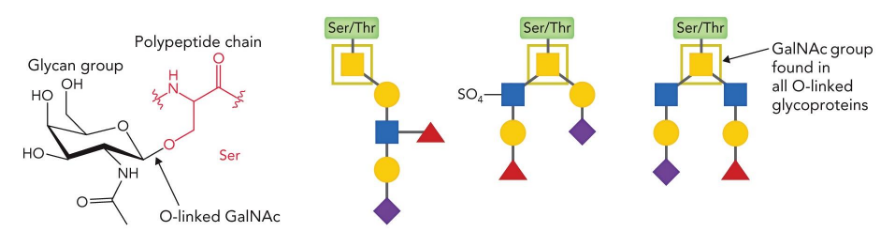
CD2
cell adhesion receptor expressed by human T-cells and natural killer cells. - Presentation of extracellular binding domain dependent on glycosylation/lipid composition. N-linked
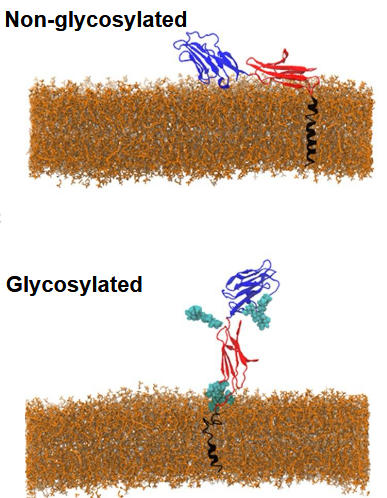
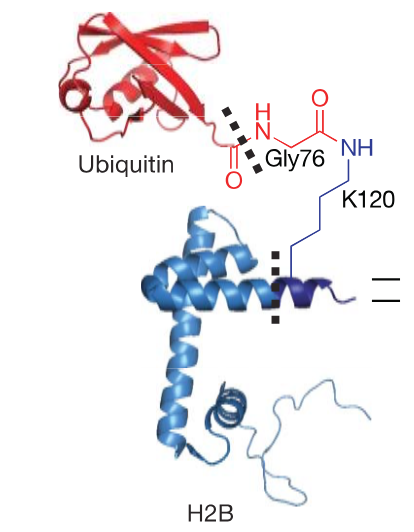
Why can’t we just express this protein?
protein added to side chain of Lys
removal of signal sequences
type of PTM
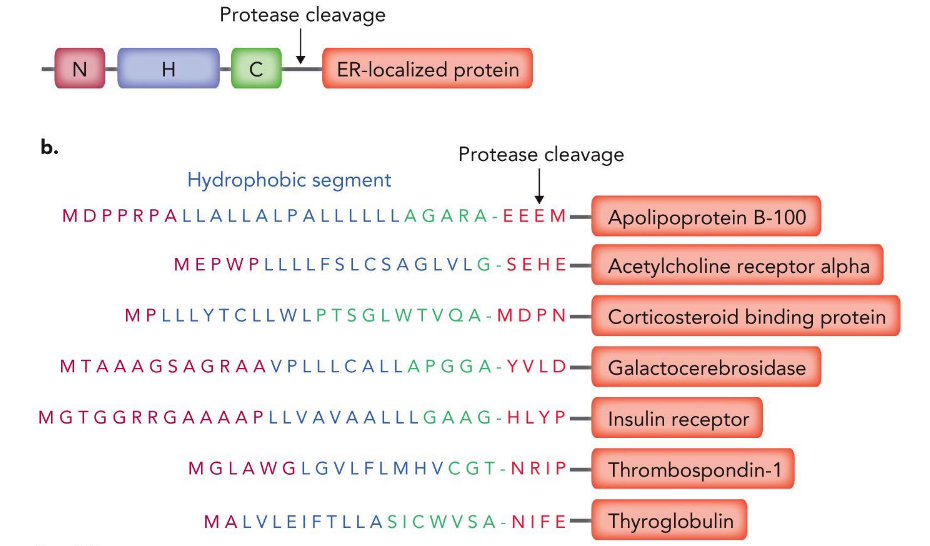
Why are proteins are eventually targeted for degradation in the cell?
Prevents build-up of abnormal or unwanted proteins
t1/2 of eukaryotic proteins vary from 30 sec to many days
Some proteins, like hemoglobin, last the lifetime of the cell (~110 days) for erythrocytes (red blood cells)
Some proteins are rapidly degraded because of incorrectly inserted amino acids or damage (perhaps due to chemical modification)
Enzymes involved in metabolism are often turned over rapidly (type of regulation)
Proteins that are unfolded or aggregated (that pose a risk to the cell)
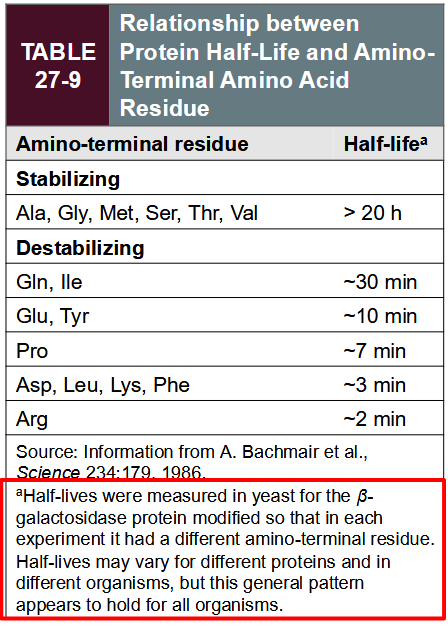
protein half-life prediction
N-terminal rule is important for predicting the half-life of a given protein
First residue after removal of Met is important for proteolytic processing
small, uncharged = stabilising
large, charged = destabilising

Why are there ~600 E3 ligases but only one E1?
Specificity for the target protein is provided by E3, not E1