Biology 1A03 Theme 4
1/123
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
124 Terms
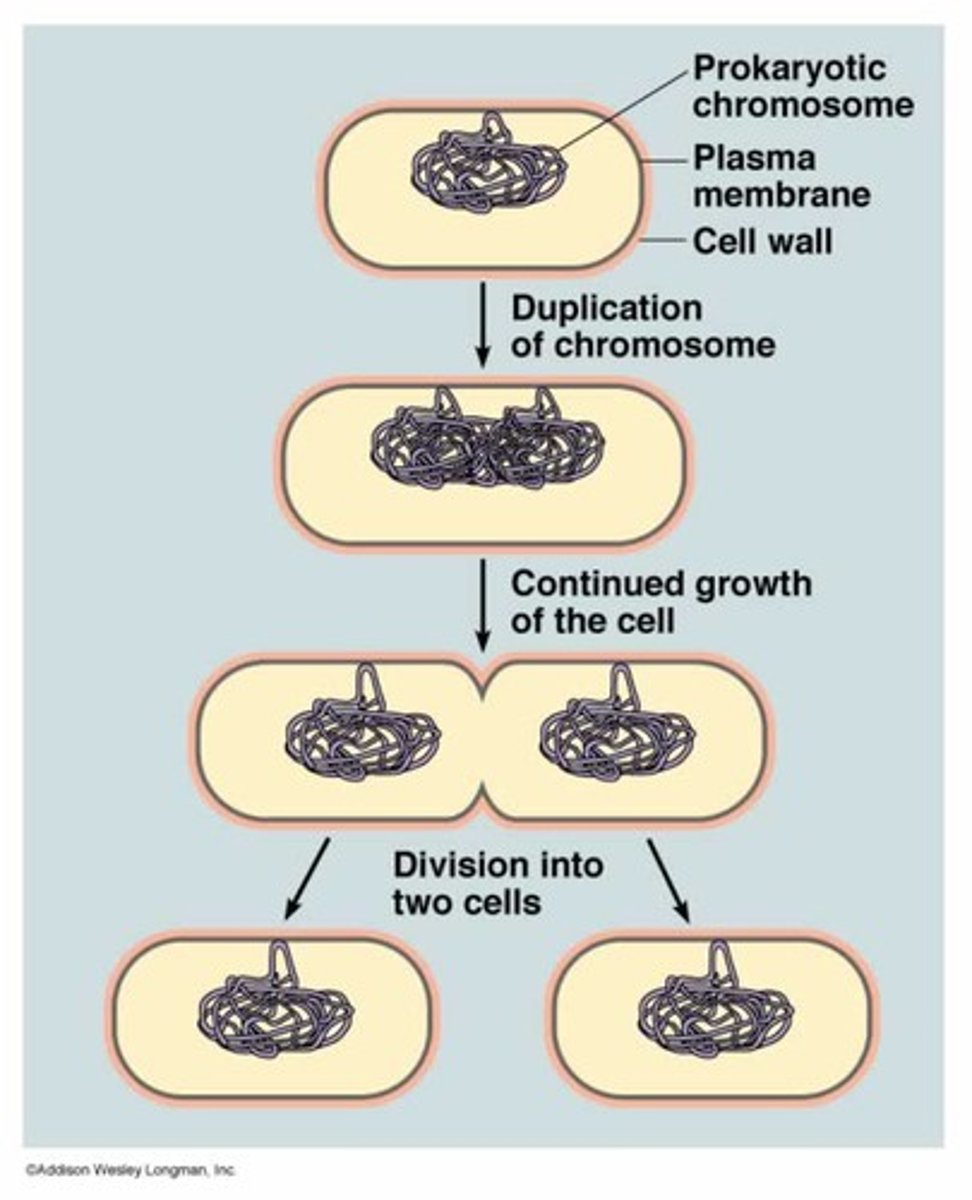
Binary Fission
a form of asexual division that makes two identical copies of the original organism. usually in prokaryote it is also reproduction
Cell divison
ability of preexisting cells to give rise to another cell
The process of binary fission
The cell elongates and new DNA is anchored to the plasma membrane
the bactera doubles in size
constriction occurs along the midpoint
new cell is formed with a new wall and plasma membrane
Stem cells
unspecialized cells that can reproduce indefinaetly under proper conditions and specialze into various cell types
Adult stem cells
stem cells that are not able to find rise to all cell types but are able to replace non reproducing specailzed cells
G0 phase
pause in the cell cycle to allow for growth
G1 phase
Prepare cell for DNA synthesis
S phase
DNA synthesis
G2 phase
Prepares cell for mitosis
Before mitosis what happens ?
DNA sequences are replicated and the newly synthezied molecules are assocaited with histones and other chromosomal proteins for tight comactoion
Centromere
fully repilicated and highly compacted that phat the paired centromeres appear fused
Sister chromatids
chromsome that is duplicated into identical copies
23 distinct chromosome pairs
46 chromosomes...22 homologous pairs and 2 sex chromsomes
M phase
mitosis and cytokinesis occurs
Prophase
Each chromosome will appear as identical sister chromatids that are joined at their centromeres
• Centrosomes (duplicated cellular microtubule organizing centers) begin to radiate long microtubules, forming a mitotic spindle (crucial for separating the chromosomes into the two daughter cells)
• Centrosomes become positioned at opposite poles of the cell
mitotic spindle
long microtubules that help with mitosis
Prometaphase
breaking down of the nuclear envelope
kinetochore microtubules radiate from centrosome and attach to the kinetochore regions to pull chromosome to poles of the cell during mitosis
polar microtubules radiate from centrosome and interact with each other to push the poles of cell away from eachother during mitosis
kinetochores
specialized protein structures that associate with one of the sister chromatids on either side of centromere
kinetochore region
areas on the chromosome that kinetochore proteins attach to to pull chromosome
polar microtubules
they raadiate from centrosome to help push poles of cell away from eachother
Metaphase
alignment of chromosome at the centre of the cell in the metaphase plate
metaphase plate
center of the cell where chromosomes align
Anaphase
Kinetochore microtubules begin to shorten
sister chromatids seperate into individual chromosomes that are pulled towards the opposite spindle poles of the cell
Polar microtubules push against eachother and help elongage the cell
Equal segregation of chromosomes at the two ends of the dividing cell
Cytokinesis
Division of cytoplasm and therefore of the cell
cytokinesis in animal cells
• Animal cells: begins with the formation of a contractile ring made up of motor proteins that contract bundles of actin fibers along the midline of the cell
• Cleavage furrow separate the cell into two distinct and separate daughter cells
cytokinesis in plant cells
• Plant cells: lay down a newly developed cell wall along a cell plate region in the middle of the dividing cell (complete once the forming cell wall fuses with the original)
Cyclin protein
Mitosis promoting factor combined with a cyclin dependant kinase (CDK) that control the progression of the cell cycle
Cyclin dependant kinase (CDK)
with the cyclin protein form the CDK that control the cell cycle progression
G1/S cyclin -CDK complex
needed for the transition from G1 to S phase and helps prepare cell for DNA replication by increasing expression of histones and unwinding DNA
S cyclin CDK complex
helps initiate DNA synthesis
M cyclin CDK complex
Initiates process of mitosis
Checkpoints
points in cell division that can pause cell division untill the preparation for the next stage is completed and serve as an opportunity for damage to be repaired
DNA damage checkpoint
located at the end of the G1 phase and only allows undamaged DNA to enter the S phase for replication
How does the DNA damage checkpoint work?
kinases phophorylate p53 that can inhibit the cell cycle when turned on
p53
protein that can inhibit the cell cycle when turned on (usually present at low levels but upon phosphorylation can accumulate and act as a transcription factor that turns on genes that inhibit the cell cycle
CDK inhibitor
p53 leads to the production of a CDK inhibitor that can bind to and block the activity of the G1-S cyclin CDK complex and stop the cell cycle in G1
DNA replication Checkpoint
At the end of G2 pase and only allows progression when ALL DNA is replicated
Spindle assembly checkpoint
before anaphase and during mitosis
cell only completes mitosis
cell
Regualatory proteins can monitor the degree to which the sister chromatids are attached to the microtubules and unattached kinetochores create a "wait" signal that recruits assembly checkpoints that are activated by a lack of tension in the centromere area
seperase
enzyme that splits sister chromatids
Semiconservative model
model of DNA replication that says DNA replication consists of the parental DNA splitting and forming two new DNA double helices each with 1 parental and one daughter DNA strand
Conservative model
two daughter strands
dispersive model
fragments from the parental and daughter strands
proof of the semiconservative model
DNA was added to a medium of 15N after being grown in 14N and the result was DNA with one 15N containing strand and one 14N containing strand
Where does DNA replication begin?
S phase of the cell cycle at specific regions along DNA
Origins of Replication
Where DNA replication begins
What direction is DNA copied and elongated?
copied 3' to 5 and elongates 5' to 3'
Replication forms
region of DNA where it is being unfound
RNA primer
5-10 nucleotides long and is required since the DNA polymerase can only begin to elongate from an existing peice
DNA polymerase
enzyme that synthesizes the replciated DNA stran from primers that anneal to template strand
Leading strand
requires only one primer and is continually replicated
lagging strand
has discontinuous replication forks containing okazaki fragments that needs a separete primer for each fragment
What replaces the RNA primers then?
another DNA polymerase comes into to repalce them
okazaki fragments
fragments of dna on the lagging strand
DNA Helicase
enzyme that can unwind the DNA double helix by breaking the hydrogen bonds between base pairs
Single stranded binding proteins
bind to and stabelize each parental strands untill elongation can begin so the strands don't rejoin
Topoisomerases
able to bind upstream of replication form to minimize torsional strain brought about from the unwinding
RNA primase
synthezies the short RNA primers that are needed to begin replication
DNA polymerase III
does most of the elongation work in prokaryotes
DNA polymerase I
responsible for removing RNA primers after DNA replication and replacing them with DNA nucleotides
DNA ligase
joins the 3' end of a fragment to adjacent DNA nucleotide by catalyzing the phosphodiester bond which brings the okazaki fragments together
Proof Reading DNA
during replication errors can occur but DNA polymerase proof reads each nucleotide but some errors can even avoid that so enzymes also help with correction
Why do the ends become shorter in linear Eukaryotic DNA
the DNA becomes shorter and shorter because once RNA primers removed, the DNA polymerase can not add nucleotides onto it
Telomeres
contain six nucleotide sequences between hundreds to thousands of times leading to many tandem repeats of GT that serve as a buffer zone so coding regions are protected
Telomerase
specific type of reverse transcriptase that is able to synthesize DNA from an RNA template and plays a key role in aging and cancer progression
PCR
Polymerase chain reaction a means of amlifying or replicating DNA in a tube
How is PCR set up
The sample DNA is put into a tube with essential ions and salts and dna primers, dNTPs and DNA polymerase (taq polymerase) and the tubes are places in a thermocylcer
dNTP
deoxyribonucleotides
Taq Polymerase
heat tolerant DNA polymerase
Denaturation
heating to break hydrogen bonds
Annealing
Cooling to allow primers to anneal
Extension
DNA polymerase extends and polymerizes the daughter strands using four dNTPs starting from primers and extending 5' to 3'
Thermocylcer
machine that goes from heating and cooling cycles to facilitate replication
Gel Electrophoresis
seperated molecules based on size . used to seperate DNA fragments from other sources to visualize DNA molecules and since DNA is negatively charged it is attracted to the positive end of the gel
Small molecules in gel
they travel faster and longer
large molecules in gel
slower and shorter distance
Sandard Ladder
the DNA is compared to this which has all the DNA sizes
DNA sequencing
developed by Fredrick Sanger that could sequence small fragments
Sangar Sequencing Technique
Dideoxy chain termination method, and required all key components of DNA replication Dideoxynucleotides (ddNTPs) are missing the -OH group at the 3' position and would not allow for further elongation of a growing DNA strand sing the -OH group is required for the attachment of the next nucleotide
o Leads to a series of interrupted daughter strands, each terminating DNA replication at the site where the ddNTP is incorporated
Shot Gun method
Could sequence larger phases
breaks the entire genome into different sized peices and proceeds with three phases
Shot gun phase 1
random sequencing of the DNA in each fragment
Shot gun phase 2
identifies the regions of overlap between the fragments and assembles the continous sequence of nucleotides in the DNA molecule that make up the chromosome
the overlapping regions helped to identify the order of the seqeuence
Shot gun Phase 3
annotating the sequences to best identify the regions of genomic DNA that encodes genes (regulatory and non-coding). usually done by a computer
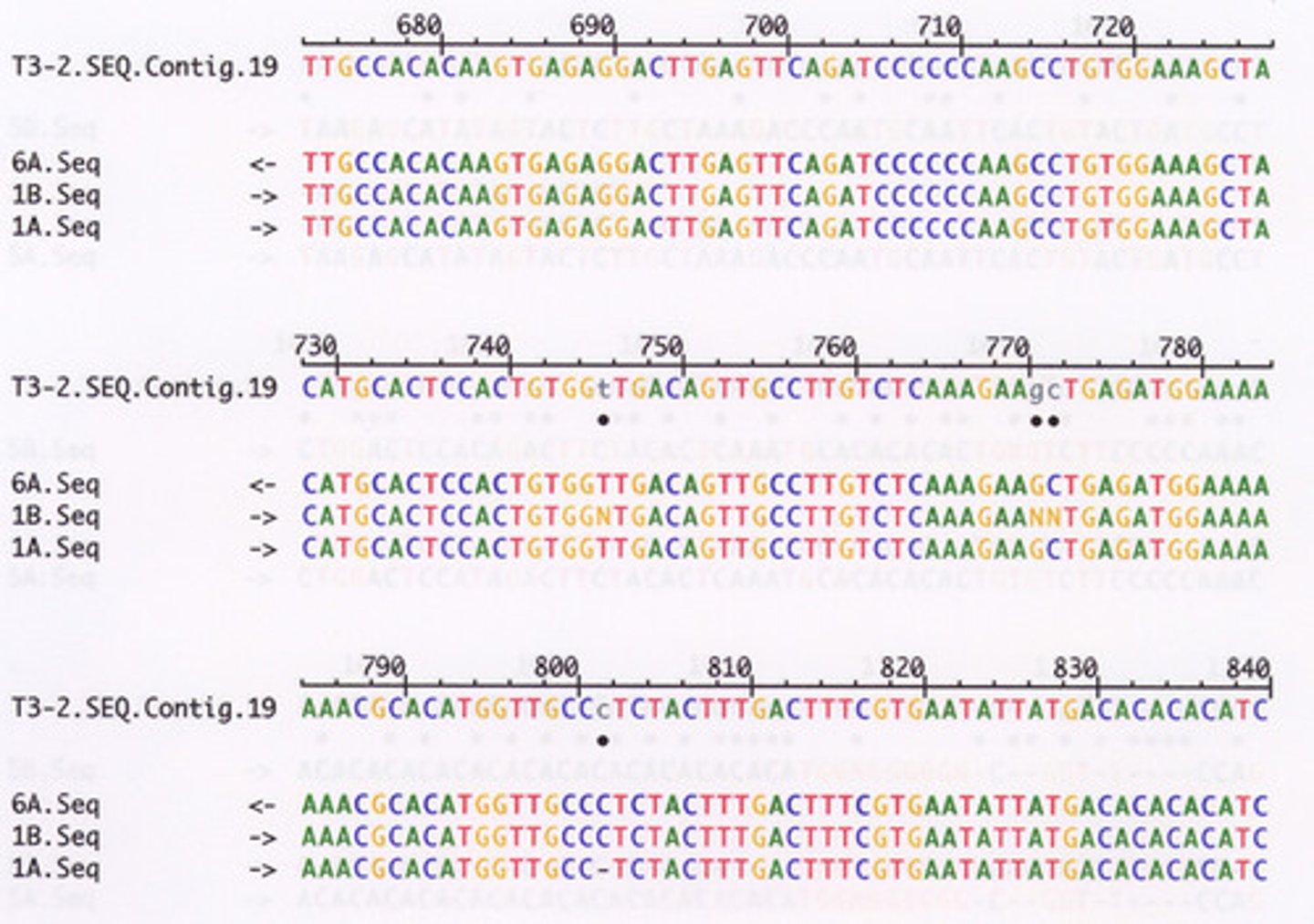
contigs
the overlapping regions of the DNA
DNA mutations
errors in the DNA that can be corrected but sometimes cannot be thus allow the mutations to propogate
What creates DNA mutations?
environmental factors, spontaneous muations, errors during replication, etc.
Somatic mutation
occur in non germline cells and cannot be inherited
Germline cells
the cells that come together to produce offspring (egg and sperm)
germline mutation
occur in germline cells and can be inherited
What happens to a mutation in a non dividing or G0 phase cell?
the effect of the mutation is negligble
How can we prove that mutations are random?
Joshua and Esther Lederberg in 1952 showed that mutations such as those of antibiotic resistance in bacteria are random and not directed
• Stamping a plate with a cloth and then stamping a new plate to transfer bacteria is referred to as replica plating, and preserves the relative arrangement of colonies on the new plate relative to the first agar (non-selective supplemented nutrients) plate
Replica Plating
Stamping a plate with a cloth and then stamping a new plate to transfer bacteria is referred to as replica plating and it preserves the arrangement of te colonies on a new plate
Joshua and Esther Lederberg Experiment
showed that mutations are random through replica plating
○ Whether the mutation occurred in the dish with penicillin or when it was without penicillin
- They took bacteria from the same colony from the non - selective agar and cultured them in a medium with penicillin
○ Led to a culture of bure antibiotic resistant bacteria
- Since the colonies were never exposed to penicillin before isolation, it can be concluded that the mutation was already existing prior to extraction, thus proving that mutations are random.
Mutagens
things that cause mutations such as radiation or chemicals
DNA ligase
enzyme to repair breakage in the DNA backbone
Mismatch Repair
Mismatching of a single nucleotide during replication can be corrected by the DNA polymerase and other proofreading proteins
What happens when DNA base pairs are mismatched?
they create a kink in the DNA that is recognized by proteins that scan the DNA for damage
What is the sign of a mismatch in DNA?
A kink in the molecule
How are mismatches fixed?
the strang with the wrong base is cleaved by nuclease (dna cutting enzyme_a distance away from the mismatch and another enzyme removes the nucleotides including the mismatched one
other enzymes such as DNA polymerase and DNA ligase intorduce synthesis to close the gap to form an intact DNA
nuclease
enzyme that cuts DNA
Base excision Repair
occurs when there is a Uracil that might end up in DNA