Grade 12 ms aziz biochem flashcards
1/42
Earn XP
Name | Mastery | Learn | Test | Matching | Spaced |
|---|
No study sessions yet.
43 Terms
Chemical Bond
A chemical bond is the force that holds atoms together in molecules or compounds. It forms due to the interaction of electrons between atoms to achieve greater stability, often by fulfilling the octet rule (having a full valence shell, usually eight electrons).
Types of Chemical Bonds:
Ionic Bond: Formed when electrons are transferred from one atom to another, creating charged ions (e.g., NaCl).
Covalent Bond: Formed when two atoms share electrons (e.g., H₂O, CO₂).
Metallic Bond: Involves a "sea of electrons" shared among metal atoms, contributing to properties like conductivity and malleability (e.g., copper, gold).
Hydrogen Bond: A weak bond formed between a hydrogen atom in one molecule and an electronegative atom (N, O, or F) in another (e.g., DNA base pairing).
Mass number
The mass number (A) of an atom is the sum of protons and neutrons in its nucleus. It determines the isotope of an element.
A=Number of protons+Number of neutronsA = \text{Number of protons} + \text{Number of neutrons}A=Number of protons+Number of neutrons
Example:
Carbon-12 (12C^{12}C12C) has 6 protons and 6 neutrons, so its mass number is 12.
Carbon-14 (14C^{14}C14C) has 6 protons and 8 neutrons, so its mass number is 14.
Importance:
Determines the identity of isotopes.
Affects atomic mass but not chemical properties.
3. Atomic Number
The atomic number (Z) of an element is the number of protons in its nucleus. It uniquely identifies an element and determines its position in the periodic table.
Z=Number of protonsZ = \text{Number of protons}Z=Number of protons
Example:
Carbon has an atomic number of 6 (always 6 protons).
Oxygen has an atomic number of 8 (always 8 protons).
Key Differences from Mass Number:
Atomic number (Z) only counts protons, while mass number (A) includes both protons and neutrons.
The atomic number is always fixed for an element, but the mass number varies with isotopes.
Isotope
An isotope is an atom of the same element with the same atomic number (same number of protons) but a different mass number (different number of neutrons).
Example:
Carbon-12 (12C^{12}C12C) → 6 protons, 6 neutrons (stable).
Carbon-14 (14C^{14}C14C) → 6 protons, 8 neutrons (radioactive).
Key Difference from Radioisotope:
Isotopes can be stable or unstable, but radioisotopes are always unstable and emit radiation.
Medical & Industrial Purposes:
Carbon-13 is used in medical breath tests to diagnose infections.
Oxygen-18 is used in tracer studies to analyze biological processes.
Deuterium (Hydrogen-2) is used in nuclear fusion research and pharmaceutical development.
Radioisotope
A radioisotope is an unstable isotope that undergoes radioactive decay, emitting alpha, beta, or gamma radiation to become stable.
Example:
Carbon-14 (14C^{14}C14C) decays into nitrogen-14.
Iodine-131 (131I^{131}I131I) is used in thyroid treatment.
Key Difference from Isotope:
All radioisotopes are isotopes, but not all isotopes are radioactive.
Radioisotopes decay over time, releasing radiation, while stable isotopes remain unchanged.
Medical & Industrial Purposes:
Medical Imaging: Technetium-99m for bone and heart scans.
Cancer Treatment: Cobalt-60 emits gamma rays to kill tumors.
Thyroid Therapy: Iodine-131 treats hyperthyroidism and thyroid cancer.
Carbon Dating: Carbon-14 measures the age of fossils.
Industrial Use: Americium-241 is used in smoke detectors, and radiotracers track fluid movement in pipes.
Intramolecular Bonds
Intramolecular bonds are strong chemical bonds that hold atoms together within a molecule or compound. They determine the chemical structure and identity of a substance.
Types of Intramolecular Bonds:
Ionic Bonds
Formed between a metal and a nonmetal through the transfer of electrons.
One atom loses electrons (forming a positive ion, cation) while another gains electrons (forming a negative ion, anion).
Example: NaCl (Sodium chloride) – Sodium (Na) donates an electron to Chlorine (Cl), forming Na⁺ and Cl⁻, which attract each other.
Covalent Bonds
Formed when two nonmetals share electrons to achieve stability.
Can be polar (unequal sharing of electrons, e.g., H₂O) or non-polar (equal sharing, e.g., O₂).
Example: H₂O (Water) – Oxygen shares electrons with two Hydrogen atoms.
Metallic Bonds
Found in metals, where valence electrons are delocalized, forming a "sea of electrons" around positively charged metal ions.
This allows metals to be malleable, ductile, and conduct electricity.
Example: Copper (Cu) – Free-moving electrons enable conductivity.
Intermolecular Bonds
Intermolecular bonds are weaker forces of attraction between molecules. They influence physical properties like boiling point, melting point, and solubility.
Types of Intermolecular Bonds:
Hydrogen Bonds
A strong type of dipole-dipole interaction that occurs when hydrogen (H) is bonded to a highly electronegative atom (N, O, or F).
Responsible for water's high boiling point and DNA base pairing.
Example: H₂O – Each water molecule forms hydrogen bonds with others.
Dipole-Dipole Forces
Occur between polar molecules where partial positive and negative charges attract.
Stronger than London dispersion forces but weaker than hydrogen bonds.
Example: HCl – The partial positive H is attracted to the partial negative Cl in another molecule.
London Dispersion Forces (Van der Waals Forces)
Weak forces present in all molecules, but most significant in non-polar molecules.
Caused by temporary dipoles due to electron movement.
Example: Argon (Ar) – The weak attractions allow noble gases to exist as liquids at low temperatures.
Hydrogen Bonding
Definition:
A hydrogen bond is a type of intermolecular force that occurs when a hydrogen atom (H) is covalently bonded to a highly electronegative atom (Nitrogen (N), Oxygen (O), or Fluorine (F)) and experiences an attraction to a lone pair of electrons on another electronegative atom in a nearby molecule.
How It Forms:
Covalent Bond Formation: A hydrogen atom forms a covalent bond with an electronegative atom like N, O, or F. This creates a partial positive charge (δ⁺) on H and a partial negative charge (δ⁻) on the electronegative atom due to unequal electron sharing.
Attraction to Another Molecule: The partially positive hydrogen in one molecule is attracted to a lone pair of electrons on a highly electronegative atom (N, O, or F) in a neighboring molecule.
Formation of a Hydrogen Bond: This attraction forms a weak, but important, hydrogen bond, stabilizing the structure of the substance.
Properties & Importance of Hydrogen Bonding:Stronger than dipole-dipole and London dispersion forces, but weaker than covalent or ionic bonds.
Increases boiling and melting points (e.g., water has a higher boiling point than expected due to hydrogen bonding).
Affects solubility (substances that form hydrogen bonds dissolve well in water).
Essential for biological molecules like DNA and proteins.
Electronegativity
Electronegativity (EN) is the ability of an atom to attract shared electrons in a covalent bond. It determines how strongly an atom pulls electrons toward itself when bonded with another atom.
Importance of Electronegativity:
Determines Bond Polarity:
Higher EN difference → More polar bond (e.g., H₂O).
Lower EN difference → Non-polar bond (e.g., O₂).
Affects Molecular Properties:
Polar molecules dissolve in water (H₂O).
Non-polar molecules dissolve in non-polar solvents (oil).
Explains Reactivity:
Highly electronegative elements (e.g., Fluorine) are very reactive because they strongly attract electrons.
Electronegativity Difference (ΔEN)
Bond Type
Example
ΔEN = 0.0 - 0.4
Non-polar Covalent
O₂, N₂ (equal sharing of electrons)
ΔEN = 0.5 - 1.7
Polar Covalent
H₂O, NH₃ (unequal sharing of electrons)
ΔEN ≥ 1.8
Ionic Bond
NaCl, KBr (electron transfer)
Structure of Water (H₂O) and Explanation
Final Summary: Why Water Has Its Structure
Bent shape due to lone pair repulsion.
Polar molecule due to oxygen’s high electronegativity.
Hydrogen bonding explains its high boiling point, solvent abilities, and density anomaly.
Essential for life due to its unique structural properties.
How Water’s Structure Determines Its Properties
1. High Heat of Vaporization
Definition: The amount of energy required to convert 1g of liquid water into vapor without a temperature increase.
Cause: Hydrogen bonds hold water molecules together, requiring a large amount of energy to break these bonds before water can transition to a gaseous state.
Effect:
Water absorbs heat efficiently, preventing overheating in living organisms (sweating cools the body as evaporation removes heat).
Stabilizes climate by absorbing excess heat from the atmosphere.
2. High Specific Heat
Definition: The amount of energy needed to raise 1g of water by 1°C.
Cause: Hydrogen bonding restricts molecular movement, so more energy is required to increase kinetic energy (temperature).
Effect:
Water resists sudden temperature changes, making it essential for climate stability.
Maintains homeostasis in organisms by regulating body temperature.
3. Less Density in the Solid State (Ice Floats)
Definition: Unlike most substances, solid water (ice) is less dense than liquid water.
Cause: When water freezes, hydrogen bonds form a stable hexagonal lattice, keeping molecules farther apart than in liquid form.
Effect:
Ice floats, insulating water bodies and protecting aquatic life in winter.
Prevents entire lakes from freezing solid, maintaining ecosystems.
4. Polarity
Definition: Water has a partial positive charge (δ⁺) on hydrogen and a partial negative charge (δ⁻) on oxygen due to oxygen’s high electronegativity.
Cause: The bent shape prevents charge distribution from being symmetrical, making water a polar molecule.
Effect:
Dissolves polar and ionic substances (like NaCl, glucose), making it the “universal solvent”.
Helps with nutrient transport in living systems (e.g., blood plasma).
5. Cohesion
Definition: Water molecules stick to each other due to hydrogen bonding.
Cause: Each water molecule can form multiple hydrogen bonds with its neighbors, creating strong cohesion.
Effect:
Creates surface tension, allowing small insects to walk on water.
Enables capillary action, helping plants pull water upward through roots and stems.
pH
pH is a measure of the acidity or alkalinity (basicity) of a solution, based on the concentration of hydrogen ions (H⁺) in the solution. It is defined mathematically as:
pH=−log[H+]
where [H⁺] represents the concentration of hydrogen ions in moles per liter (M).
Buffer: Definition and Function
What is a Buffer?
A buffer is a solution that resists changes in pH when small amounts of an acid (H⁺) or base (OH⁻) are added. It works by using a weak acid and its conjugate base (or vice versa) to neutralize pH changes and maintain stability.
Importance of Buffers:
Maintain biological homeostasis (e.g., blood pH must stay between 7.35 - 7.45).
Prevent extreme pH changes in cells, blood, and body fluids.
Used in chemical reactions and industrial processes where pH control is necessary.
Bicarbonate/Carbonic Acid Buffer System (HCO₃⁻ / H₂CO₃)
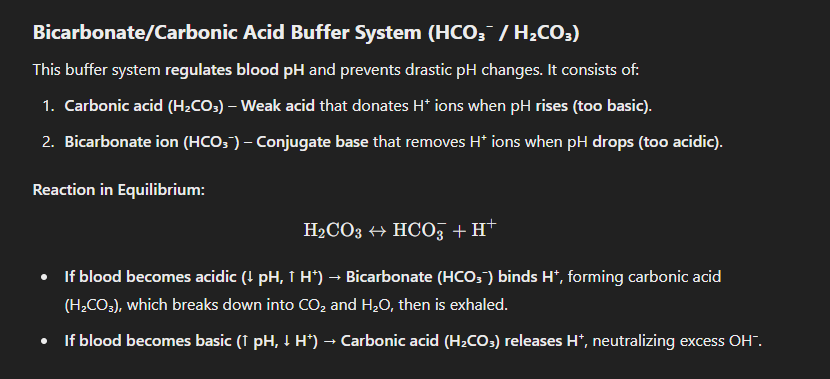

examples (buffers)
1. After Eating Something Too Acidic (e.g., Citrus, Soda, Vinegar):
Acids release excess H⁺ ions, lowering pH.
The buffer system reacts by:
Bicarbonate (HCO₃⁻) binds to excess H⁺, forming H₂CO₃.
H₂CO₃ dissociates into CO₂ and H₂O, and CO₂ is exhaled.
Blood pH remains stable, preventing acidosis.
2. After Eating Something Too Basic (e.g., Antacids, Baking Soda, Egg Whites):
Bases remove H⁺, raising pH.
The buffer system reacts by:
H₂CO₃ dissociates, releasing H⁺ to neutralize the excess base.
This prevents alkalosis (excessively high pH).
Blood pH stays within its normal range.

Isomers: Definition and Types
Isomers are compounds with the same molecular formula but different structures or spatial arrangements, leading to different physical and chemical properties.
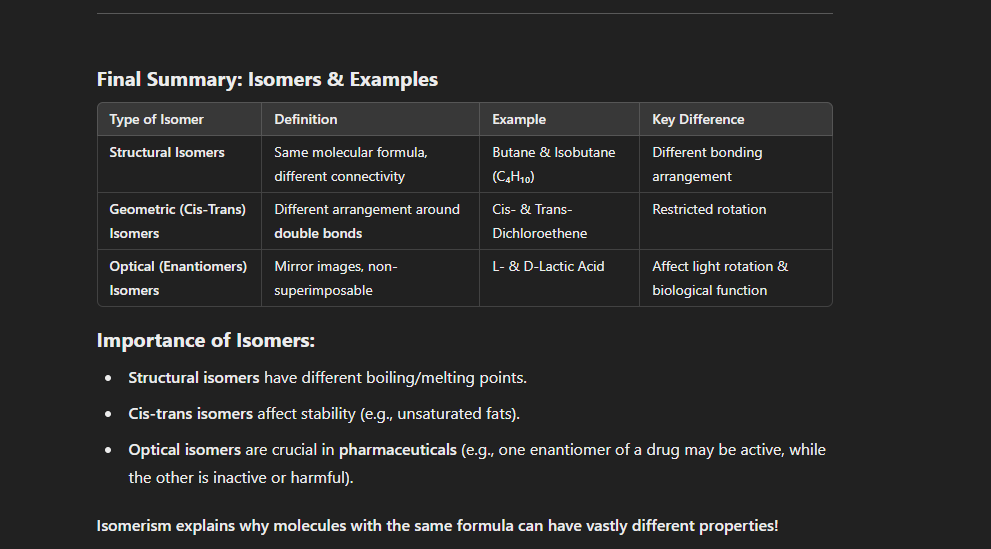
Distinguish between polar and non-polar
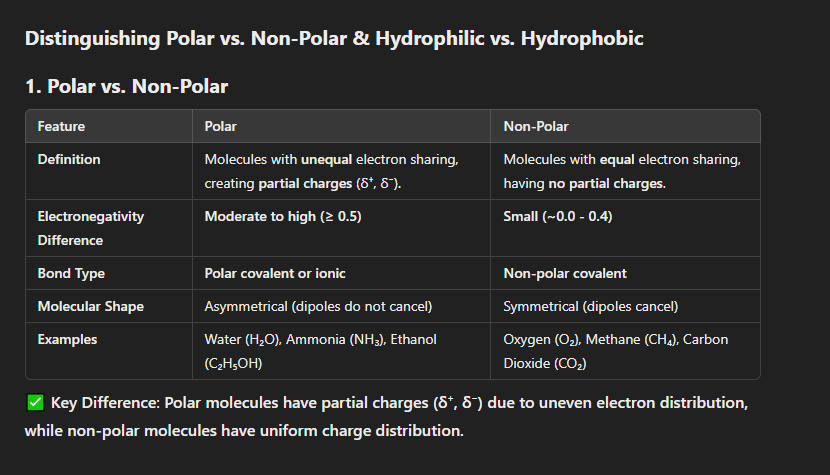
; hydrophilic and hydrophobic

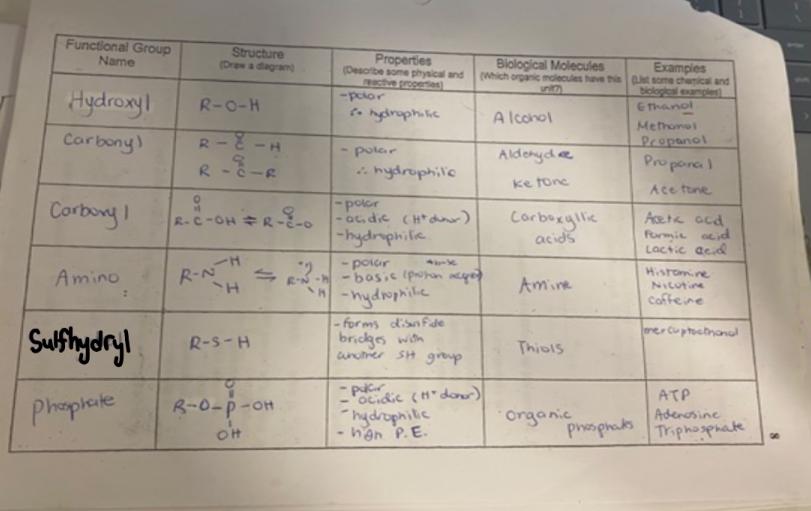
identify all functional groups
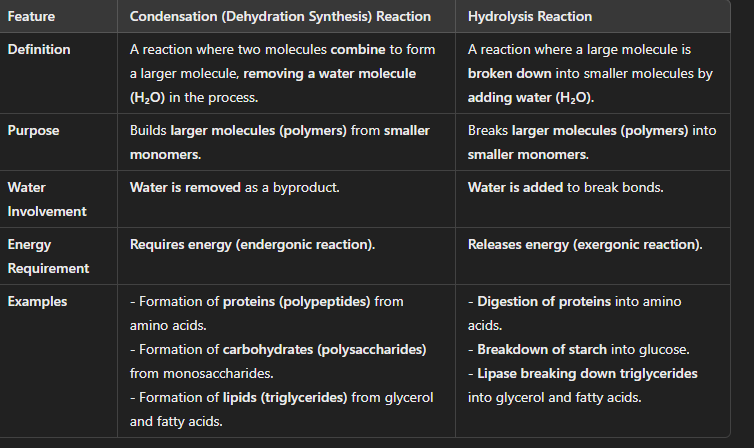
Distinguishing Condensation (Dehydration Synthesis) vs. Hydrolysis Reactions
Condensation builds molecules by removing water, while hydrolysis breaks molecules by adding water.
Condensation requires energy, while hydrolysis releases energy.
Definitions & Relationships of Macromolecules, Polymers, and Monomers
1. Macromolecule
A large, complex molecule made up of thousands or millions of atoms.
Essential for biological functions (e.g., DNA, proteins, carbohydrates, lipids).
Example:
Proteins (e.g., hemoglobin)
Nucleic acids (DNA, RNA)
Polysaccharides (starch, glycogen)
2. Polymer
A large molecule made up of repeating smaller units (monomers) linked by covalent bonds.
Not all macromolecules are polymers (e.g., lipids are macromolecules but not true polymers).
Example:
Proteins → Polymer of amino acids.
Carbohydrates → Polymer of monosaccharides.
Nucleic acids (DNA, RNA) → Polymer of nucleotides.
3. Monomer
The smallest repeating unit that joins together to form a polymer.
Monomers bond through condensation (dehydration synthesis) and break apart through hydrolysis.
Example:
Amino acids → Monomers of proteins.
Glucose → Monomer of starch (carbohydrates).
Nucleotides → Monomers of DNA and RNA.
How They Relate to Each Other
Monomers are the building blocks of polymers.
Polymers are a type of macromolecule when they grow large and complex.
Macromolecules can be made of one or multiple types of polymers.
Biological Macromolecules: Constituent Atoms, Functional Groups, Monomers, and Functions
Carbohydrates are primarily for energy storage and structural support.
Proteins serve enzymatic, structural, and transport roles.
Lipids are hydrophobic and play roles in energy storage, cell membranes, and signaling.
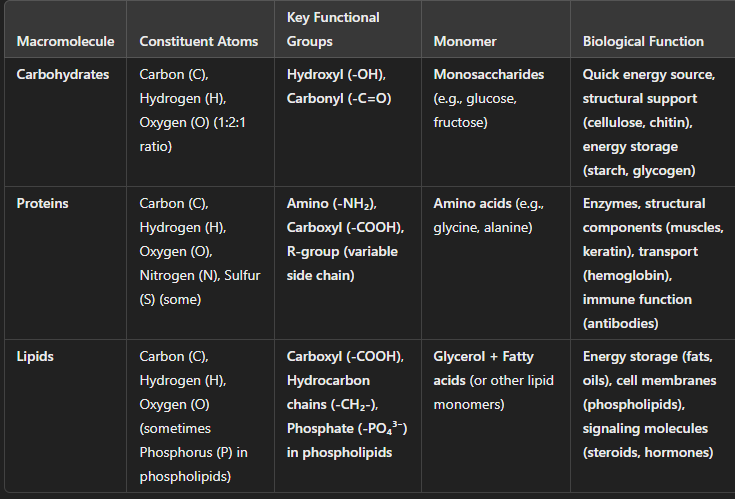
alpha & beta glucose
right side placement (difference)
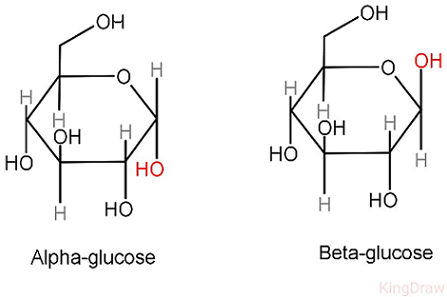
amino acid
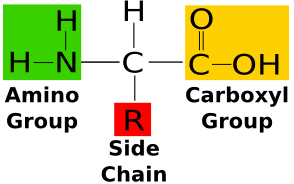
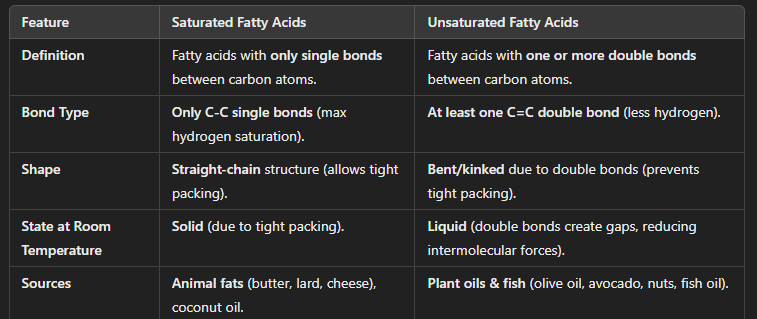
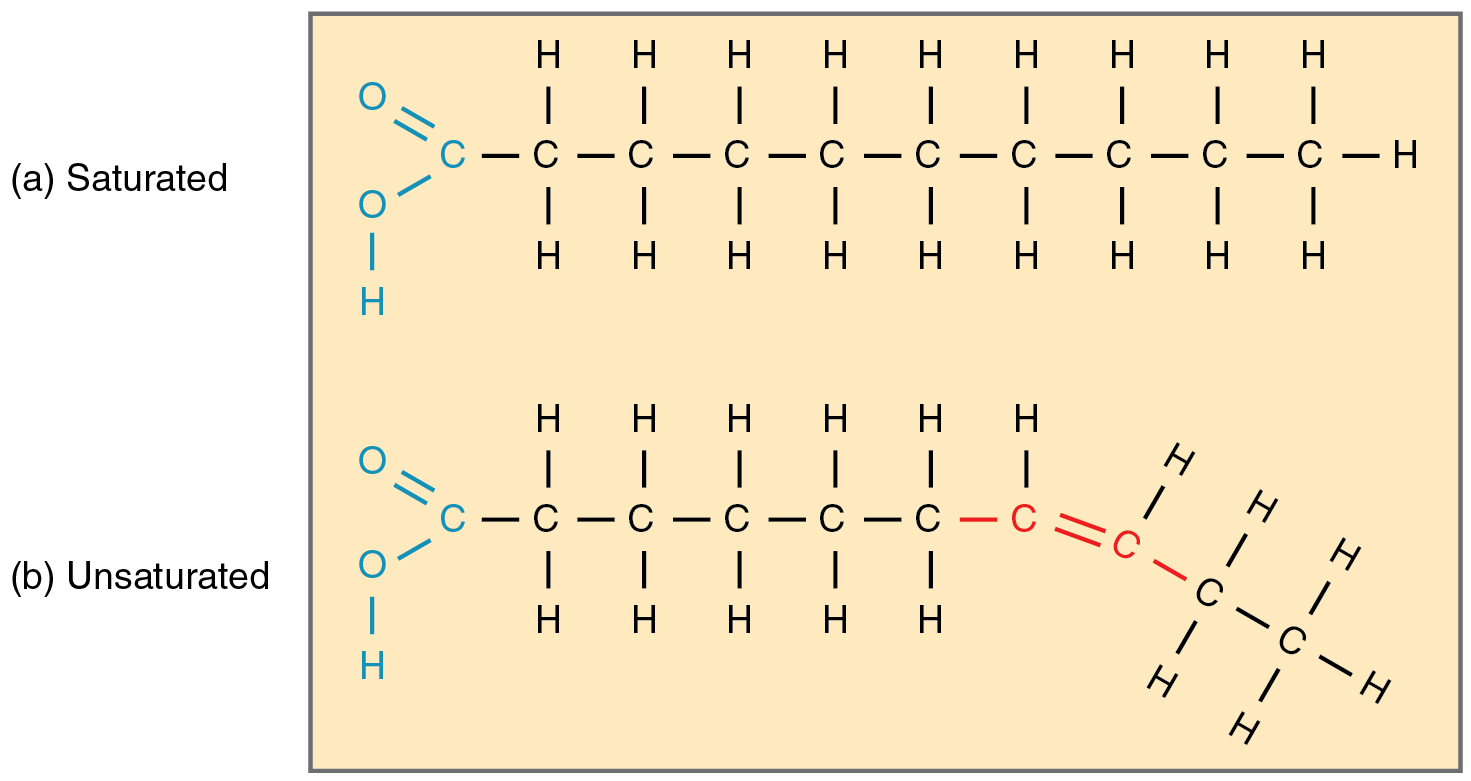
saturated vs unsaturated fatty acid
give the word equation for forming maltose, lactose, sucrose
Maltose: Glucose + Glucose → Maltose + H₂O Lactose: Glucose + Galactose → Lactose + H₂O Sucrose: Glucose + Fructose → Sucrose + H₂O
1. Classification by Number of Carbon Atoms
The number of carbon atoms in a sugar determines its name:
Type | Number of Carbons | Example |
|---|---|---|
Triose | 3 | Glyceraldehyde, Dihydroxyacetone |
Tetrose | 4 | Erythrose, Erythrulose |
Pentose | 5 | Ribose, Ribulose, Xylose |
Hexose | 6 | Glucose, Fructose, Galactose |
Heptose | 7 | Sedoheptulose |
✅ Key Rule: More carbon atoms = More complex sugar.
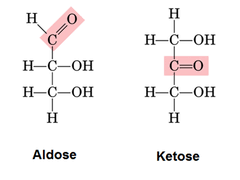
2. Classification as Aldose or Ketose
Sugars are classified as aldoses or ketoses based on the functional group they contain:
Type | Functional Group | General Structure | Example |
|---|---|---|---|
Aldose | Aldehyde (-CHO) at C1 | H-C=O at the end | Glucose, Galactose, Ribose |
Ketose | Ketone (C=O) at C2 | C=O in the middle | Fructose, Ribulose, Dihydroxyacetone |
✅ Key Rule:
Aldoses have C=O at the end of the carbon chain.
Ketoses have C=O within the chain (usually at C2).
Final Classification of Common Sugars
Sugar | Number of Carbons | Aldose or Ketose? |
|---|---|---|
Glucose | Hexose (6) | Aldose |
Fructose | Hexose (6) | Ketose |
Ribose | Pentose (5) | Aldose |
Ribulose | Pentose (5) | Ketose |
Glyceraldehyde | Triose (3) | Aldose |
Dihydroxyacetone | Triose (3) | Ketose |
Summary: How to Classify a Sugar
Count the carbon atoms → Name it triose (3C), pentose (5C), hexose (6C), etc.
Look for the carbonyl group (C=O):
If it’s at C1 → Aldose.
If it’s at C2 → Ketose.
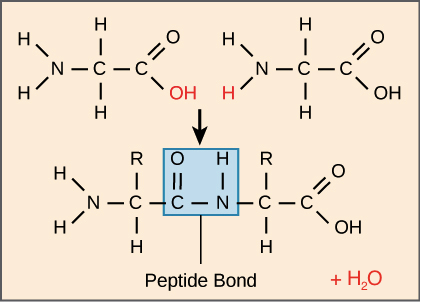
formation of peptide bond
1. Substrate Specificity
Definition: The ability of an enzyme to recognize and bind to a specific substrate (reactant).
Example: Sucrase only breaks down sucrose, not lactose.
Reason: The active site is shaped to fit only certain substrates.
2. Active Site
Definition: The region on an enzyme where the substrate binds and the reaction occurs.
Key Features:
Has a specific shape complementary to the substrate.
Can undergo conformational changes (see induced fit model).
Example: The active site of amylase binds to starch and breaks it into maltose.
3. Optimal Temperature
Definition: The temperature at which an enzyme works most efficiently.
Effect:
Too low → Enzyme activity slows (molecules move less).
Too high → Enzyme denatures (loses shape and function).
Example:
Human enzymes: ~37°C (body temp).
Bacteria in hot springs: ~80°C.
4. Optimal pH
Definition: The pH at which an enzyme is most active.
Effect:
pH too high or low → Changes enzyme structure, disrupting the active site.
Examples:
Pepsin (stomach enzyme): pH ~2 (acidic).
Trypsin (intestinal enzyme): pH ~8 (basic).
5. Saturation
Definition: When all enzyme active sites are occupied by substrates, enzyme activity cannot increase further even if more substrate is added.
Graph Effect: Reaction rate levels off when saturation is reached.
Solution: Adding more enzymes increases reaction rate.
6. Induced Fit Model
Definition: The enzyme’s active site adjusts its shape slightly to fit the substrate more snugly.
Key Idea: Enzymes are flexible, not rigid.
Example: Hexokinase (binds glucose and slightly changes shape for a tighter fit).
7. Lock-and-Key Model
Definition: The enzyme’s active site is a perfect fit for the substrate, like a key fitting into a lock.
Key Idea: No shape change occurs.
Example: Maltase only binds to maltose.
Difference from Induced Fit Model:
Lock-and-key suggests a static shape,
Induced fit suggests slight flexibility in the enzyme
8. Activation Energy
Definition: The minimum energy required for a chemical reaction to occur.
How Enzymes Help: Enzymes lower activation energy, allowing reactions to happen faster.
Example: Catalase lowers activation energy for the breakdown of H₂O₂ into water and oxygen.
9. Allosteric Sites
Definition: A secondary binding site on an enzyme, different from the active site.
Function: Molecules bind here to activate or inhibit enzyme activity.
Types of Regulation:
Allosteric activation → Increases enzyme activity.
Allosteric inhibition → Decreases enzyme activity.
Example: Phosphofructokinase (PFK) is inhibited allosterically by ATP when energy levels are high.
How Enzymes Work: Step-by-Step Process
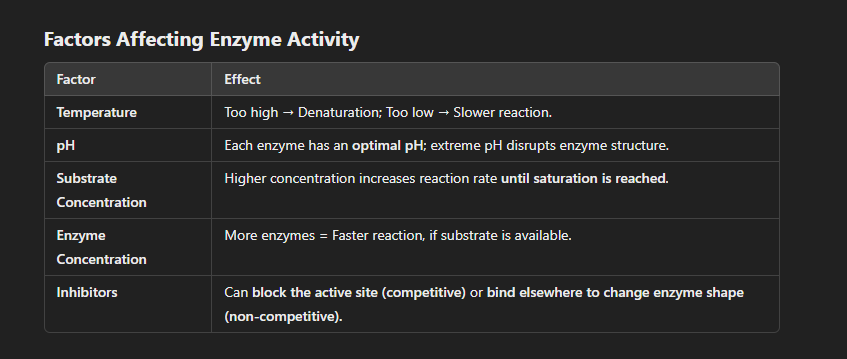
Enzymes are biological catalysts that speed up chemical reactions by lowering activation energy without being consumed. The process begins when the substrate binds to the enzyme’s active site, forming an enzyme-substrate complex. The active site, which is highly specific, either fits the substrate exactly (lock-and-key model) or adjusts slightly (induced fit model). Once bound, the enzyme stabilizes the substrate and weakens its bonds, making it easier for the reaction to occur. The substrate undergoes a chemical transformation, forming products that are then released, leaving the enzyme unchanged and ready to be reused. Enzyme activity is affected by temperature, pH, substrate concentration, enzyme concentration, and inhibitors. If conditions are optimal, the enzyme functions efficiently, ensuring biochemical reactions happen quickly, which is essential for life processes such as digestion, energy production, and DNA replication.
how inhibitors work
Enzyme inhibitors are molecules that slow down or stop enzyme activity by interfering with the enzyme's function. There are two main types: competitive and non-competitive (allosteric) inhibitors.
Competitive inhibitors resemble the substrate and bind to the active site, preventing the substrate from attaching. Since both the inhibitor and substrate compete for the active site, increasing substrate concentration can reduce the effect of competitive inhibition. An example is malonate, which competes with succinate in the Krebs cycle.
Non-competitive inhibitors bind to an allosteric site (a different location on the enzyme), causing a shape change that makes the active site less effective or completely inactive. Since the substrate and inhibitor bind to different sites, increasing substrate concentration does not reverse inhibition. An example is cyanide, which inhibits cytochrome c oxidase, preventing ATP production.
feedback inhibs
How Feedback Inhibition Works
Feedback inhibition (end-product inhibition) is a regulatory mechanism in which the final product of a metabolic pathway inhibits an earlier enzyme in the pathway, preventing overproduction and conserving resources.
Mechanism of Feedback Inhibition:
A substrate enters the metabolic pathway and undergoes a series of enzyme-catalyzed reactions, forming intermediates.
The pathway continues until the final product is synthesized.
If the product accumulates in excess, it binds to an allosteric site on an enzyme early in the pathway (usually the first enzyme).
This changes the enzyme’s shape, reducing or stopping its activity.
As product levels decrease, the inhibitor detaches, allowing the pathway to resume.
Example: Feedback Inhibition in Cellular Respiration
Enzyme Inhibited: Phosphofructokinase (PFK) in glycolysis.
Inhibitor: ATP (high levels signal sufficient energy).
Effect: If ATP is abundant, it binds allosterically to PFK, slowing glycolysis and conserving glucose.
Importance of Feedback Inhibition:
Prevents wasteful overproduction of molecules.
Maintains homeostasis by regulating metabolic pathways.
Allows cells to respond to environmental changes efficiently.
This self-regulating mechanism ensures that metabolic pathways only produce what is needed, preventing energy loss and imbalance in biological systems.
Commercial and Industrial Uses of Enzymes
Enzymes are widely used in industries because they speed up reactions, work under mild conditions, and are biodegradable, making them more efficient and environmentally friendly than chemical catalysts.
