7.1 Using Gene Sequences
1/35
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
36 Terms
What is gel electrophoresis? (2)
- It is a laboratory technique used for separating macromolecules like DNA.
- The separation is based on molecular size and charge, achieved by applying an electric field to move the molecules through a gel matrix.
What is the role of restriction endonucleases in DNA analysis? (2)
- They are enzymes that cut DNA at specific recognition nucleotide sequences.
- This cutting action produces DNA fragments of varying sizes, which are then separated using gel electrophoresis.
What are the main components of a gel electrophoresis system? (3)
- An agarose gel that acts as a molecular sieve through which the DNA fragments migrate.
- A buffer solution that conducts the electric current and maintains a constant pH.
- A fluorescent dye that binds to the DNA and allows it to be visualised under UV light.
How are DNA fragments separated by size during gel electrophoresis? (3)
- When an electric field is applied, the negatively charged DNA migrates towards the positive electrode.
- The agarose gel acts as a matrix, resisting the movement of the DNA fragments.
- Smaller fragments move through the gel more easily and quickly than larger fragments, resulting in separation based on size.
How are DNA fragments visualised after gel electrophoresis? (2)
- The agarose gel is exposed to ultraviolet (UV) radiation.
- A fluorescent dye bound to the DNA absorbs the UV light and fluoresces, revealing the position of the separated DNA fragments.
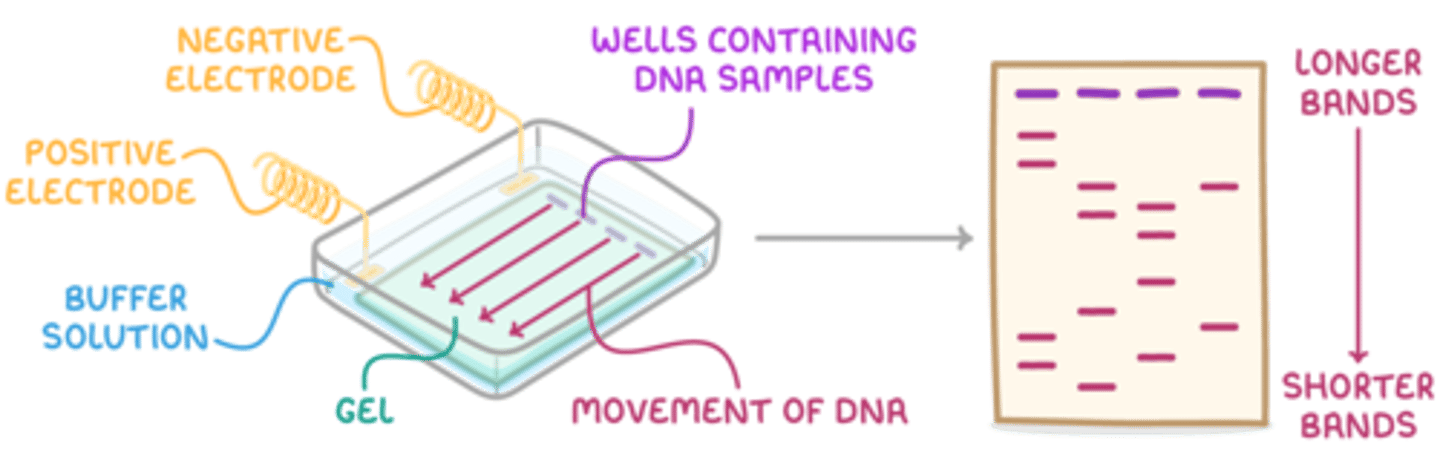
Draw a labelled diagram showing a gel electrophoresis setup? (5)
- A correctly drawn diagram should include the electrophoresis tank containing the buffer solution.
- The agarose gel should be shown submerged in the buffer.
- Wells for sample loading should be indicated at the end of the gel near the negative electrode (cathode).
- The positive electrode (anode) should be at the opposite end of the gel.
- A power supply must be shown connected to both the cathode and anode.
What is Southern blotting? (2)
- It is a laboratory technique used to detect a specific DNA sequence within a complex DNA sample.
- It involves transferring DNA fragments separated by gel electrophoresis onto a filter membrane and then identifying the sequence of interest.
Why is an alkaline buffer solution used in Southern blotting? (2)
- Its purpose is to denature the double-stranded DNA fragments, separating them into single strands.
- This ensures that a DNA probe can bind to the exposed bases on the single-stranded DNA.
How is DNA transferred from the gel to a filter in Southern blotting? (3)
- An absorbent nitrocellulose filter paper is placed directly on top of the gel.
- Capillary action draws the buffer through the gel and onto the filter.
- This process transfers the single-stranded DNA from the gel onto the surface of the filter in the same pattern.
What is the purpose of a labelled probe in Southern blotting? (3)
- A probe is a short, single-stranded DNA sequence that is complementary to the target DNA sequence.
- It is labelled with a radioactive or fluorescent tag for detection.
- The probe hybridises with its complementary sequence on the filter, allowing the location of the gene to be identified.
How is the final result of Southern blotting visualised? (2)
- After hybridisation, the filter is placed on X-ray film if a radioactive probe is used.
- The radiation from the labelled probe exposes the film, creating a dark band that corresponds to the location of the gene of interest.
What is the polymerase chain reaction (PCR)? (2)
- PCR is an automated laboratory procedure used to amplify a specific DNA sequence.
- It is a method for making a large number of copies of a DNA sequence from a small initial sample.
Why is the polymerase chain reaction (PCR) used? (2)
- It is used to amplify a specific segment of DNA from a given sample.
- The process involves making millions of copies of the target DNA.
What are the three main stages of a PCR cycle? (3)
- Denaturation, which occurs at approximately 95°C.
- Annealing, which occurs between 55-65°C.
- Elongation, which occurs at approximately 72°C.
What happens during the stages of a PCR cycle? (3)
- During denaturation, the DNA is heated to separate it into two single strands.
- During annealing, short DNA primers bind to the single-stranded DNA templates.
- During elongation, Taq DNA polymerase synthesises new DNA strands by adding nucleotides.
How does the polymerase chain reaction (PCR) work? (3)
- A reaction mixture is heated to 95°C to break the hydrogen bonds and separate the DNA into two single strands.
- The mixture is then cooled to between 50-65°C, which allows primers to bind to the DNA templates.
- The temperature is increased to approximately 72°C, the optimal temperature for Taq polymerase to synthesise new DNA strands.
What is the function of primers and Taq polymerase in PCR? (2)
- Primers are short DNA sequences that provide a starting point for DNA synthesis.
- Taq polymerase is a thermostable enzyme that catalyses the formation of new DNA strands by adding nucleotides.
What are the uses of DNA amplified by PCR? (2)
- Amplified DNA samples can be used in DNA profiling.
- Amplified DNA samples can be used in DNA sequencing.
What is required for a polymerase chain reaction? (3)
- A DNA template that contains the target sequence to be amplified.
- A sufficient supply of primers that are specific to the start and end of the target sequence.
- A supply of free DNA nucleotides and a thermostable enzyme, such as Taq polymerase.
What is a genome? (1)
A genome is the complete set of genetic material within an organism.
What is the difference between introns and exons? (3)
- Introns are non-coding regions of a gene and are not translated into a protein.
- Exons are the regions of a gene that are expressed and code for the amino acid sequence in a polypeptide.
- During RNA modification, introns are removed, and the exons are spliced together.
What are variable number tandem repeats (VNTRs)? (2)
- VNTRs are short sequences of DNA that are repeated one after another in tandem.
- They are found in the non-coding parts of DNA and are highly variable between individuals.
How does DNA profiling work? (3)
- The principle is based on comparing the highly variable, non-coding regions of an individual's DNA.
- By analysing the unique pattern of repeats in VNTRs, a distinctive profile can be created for an individual.
- This profile can be used to compare individuals for forensic purposes or to determine paternity.
What is the process of DNA profiling? (3)
- Restriction enzymes are used to cut DNA into fragments at sites surrounding the VNTRs.
- The resulting DNA fragments are then separated according to their size using gel electrophoresis.
- Gene probes, which are labelled complementary DNA sequences, are used to visualise the specific VNTR patterns.
What is dye-terminator sequencing? (2)
- It is a method used to determine the precise order of nucleotides within a DNA molecule.
- It works by creating copies of a DNA sequence that terminate at specific bases due to the incorporation of terminator nucleotides.
What components are required for dye-terminator sequencing? (3)
- A DNA template and a primer to initiate DNA synthesis.
- A set of four normal deoxynucleotides (dNTPs).
- A small proportion of four fluorescently labelled dideoxynucleotides (ddNTPs).
Why are dideoxynucleotides (ddNTPs) used in sequencing? (3)
- Dideoxynucleotides lack the 3'-hydroxyl group necessary for forming a phosphodiester bond.
- When a ddNTP is incorporated into a growing DNA strand, it immediately terminates the synthesis of that strand.
- Each of the four ddNTPs is labelled with a different coloured fluorescent dye for identification.
Why is DNA sequencing performed? (2)
- It is used to help predict the amino acid sequence of proteins.
- It is also used to identify possible links to genetically determined conditions.
What are the main steps in dye-terminator sequencing? (3)
- A reaction is set up with DNA polymerase, a primer, standard nucleotides, and fluorescently labelled terminator nucleotides.
- The reaction produces DNA fragments of varying lengths, each ending with a specific labelled terminator nucleotide.
- High-resolution gel electrophoresis is used to separate these fragments by size, and the sequence is read by detecting the fluorescent labels.
How is a DNA sequence determined after a sequencing reaction? (3)
- The resulting mixture, containing DNA fragments of varying lengths, is separated by size using capillary electrophoresis.
- An electric voltage draws the negatively charged DNA fragments through a narrow capillary tube towards the positive anode.
- A laser at the end of the tube excites the fluorescent dye on each fragment as it passes, and a computer records the sequence of colours to reveal the DNA sequence.
How can DNA sequencing provide genetic information about an individual? (3)
- The sequence of bases in a DNA sample can be determined through a process called sequencing.
- This base sequence can then be used to identify the specific amino acids encoded by successive DNA triplets within a gene.
- This information can reveal if a person is susceptible to, or will develop, a genetically determined condition like cystic fibrosis.
What information can DNA sequencing provide about a protein? (2)
- It provides the necessary information to determine the primary structure of a protein.
- It also helps in determining the stereochemical properties that the protein is comprised of.
Why are DNA base sequences specific to an individual? (2)
- Particular regions within a genome are known to be variable between individuals.
- Consequently, the base sequences in these regions are highly likely to be unique and specific to a certain individual.
How is DNA sequencing used to determine the relationship between different species? (3)
- DNA sequencing allows for the comparison of DNA base sequences between two or more species.
- The greater the similarity between the base sequences, the more closely related the species are considered to be.
- An exception is found in highly conserved genes, which are essential for life and change very little over evolutionary time.
Why is the polymerase chain reaction (PCR) used before gel electrophoresis to produce DNA profiles? (3)
- The polymerase chain reaction is used because the initial DNA sample needs to be amplified or replicated.
- This is often necessary as only very small samples of DNA may be available, and a larger quantity is needed for analysis.
- The process ensures that all the copies of the DNA produced are identical to the original sample.
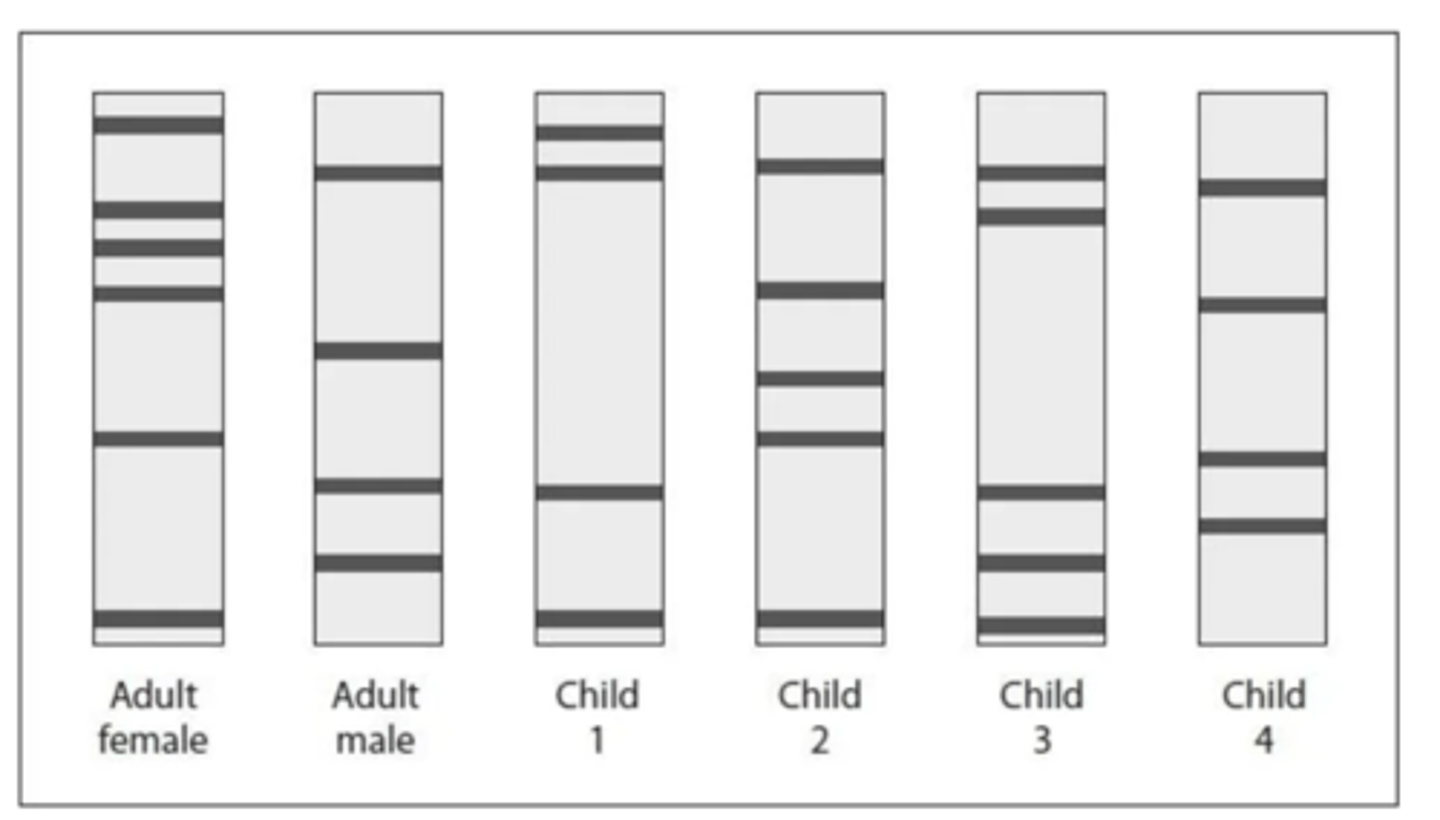
How can the parentage of the four children be determined from the DNA profiles provided, and which children belong to the adult male and female? (4)
- To determine parentage, every band in a child's DNA profile must match a corresponding band in either the adult female's profile or the adult male's profile.
- Child 1's bands all have a corresponding match with a band from either the adult female or the adult male.
- Child 3's bands all have a corresponding match with a band from either the adult female or the adult male.
- Child 2 and Child 4 both have bands that do not match either adult, therefore they cannot be the biological children of this pair.