BCH 5413 - Exam 2, Module 1
1/87
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
88 Terms
What does transgenic mean?
Refers to an organism that has been genetically altered to contain additional DNA (from the same organism or others) in order to alter expression of specific phenotypes to compare with wild type expression
When creating a transgenic mouse, how many copies are inserted and where will they insert into the genome?
Between 1-200 copies of the transgene can be inserted into the genome at random.
Which of the following statements is true about a transgenic mouse?
The transgenic mouse may have multiple copies of the transgene
3 multiple choice options
Where is the transgene injected into the mouse?
Into a fertilized oocyte (DNA directly injected within the male pronucleus) or embryo (using a retrovirus). The surviving embryos are then implanted back into the donor female
Once offspring are obtained from the donor mouse, how are they determined to have the transgene(s)?
Through the use of PCR (using transgene specific primers) followed by southern blotting and comparing the transgenic mice genes to the wild type (endogenous) genes. This process can confirm transgene presence as well as the copy number of the gene.
When determining transgene presence using PCR, how can you distinguish between the wild type and transgenes?
Specific primers can be used - one primer will be shared between the transgene and the endogenous gene and one will be unique to the transgene/cloning vector (another option would be to add one unique to the endogenous gene, if possible, for hetero/homozygous comparisons). Using these primers should create banding that can be used to identify the presence or absence of the transgene.
What is used as a control for the PCR reactions for determining wild type or transgene presence?
An unrelated universal mouse gene. This gene will be a good negative control for transgene presence because it will not react with primers specific to the transgene/vector and not produce a band to showcase lack of contamination. Additionally, primers for this gene can be used as a control for to illustrate lack of contamination when comparing to the wild type because it will have banding with the universal gene but not with the transgene primers.
When determining transgene presence using southern blot, how can you distinguish between the wild type and transgenes?
Specific restriction enzyme sites can be used - the use of different sites between the transgene and endogenous gene will produce different band sizes (depending where the sites are flanking) for comparison. The intensity of the bands in the transgenes can also be used to determine the amount of copies present when compared to the wild type.
What controls can be used during the southern blot to determine transgene presence/quantity?
Wild type (or no clone) DNA. This DNA will not produce a band for the transgene restriction enzymes because the sites will not be present (negative control). The band intensity for the wild type can also be used as a base line to compare the intensity of the transgenes to (greater intensity than the wild type indicates more gene copies)
What is a reporter gene and how can be used in a transgenic system?
A reporter gene is an expression vector that is used in concert with a target regulatory sequence that is used to study gene expression as opposed to gene function. This is significant because transgenic mice in these systems can be used to study changes in gene expression relative to changes in their promoters or repressors.
How does creating a knockout mouse differ from creating a transgenic mouse?
A knockout mouse cannot have any copies of the target sequence while a transgenic mouse can have multiple copies of an altered sequence. Therefore, embryonic stem cells must be used to create knockout genes, via homologous recombination, that can be passed on in the germ cell line.
What is the Tk gene in relation to the knockout selection process?
Thymidine kinase gene which results in sensitivity to ganciclovir. Normally, mammalian cells are resistant to ganciclovir. If non-homologous recombination occurs within the ES cells, they will be susceptible to ganciclovir.
Why does non-homologous recombination result in ganciclovir susceptibility?
The Tk gene is present on the vector plasmid containing the knockout of the target gene. However, it is located outside of the target gene so it will not be present during homologous recombination. If non-homologous recombination occurs, a larger chunk of the vector will recombine containing the Tk gene.
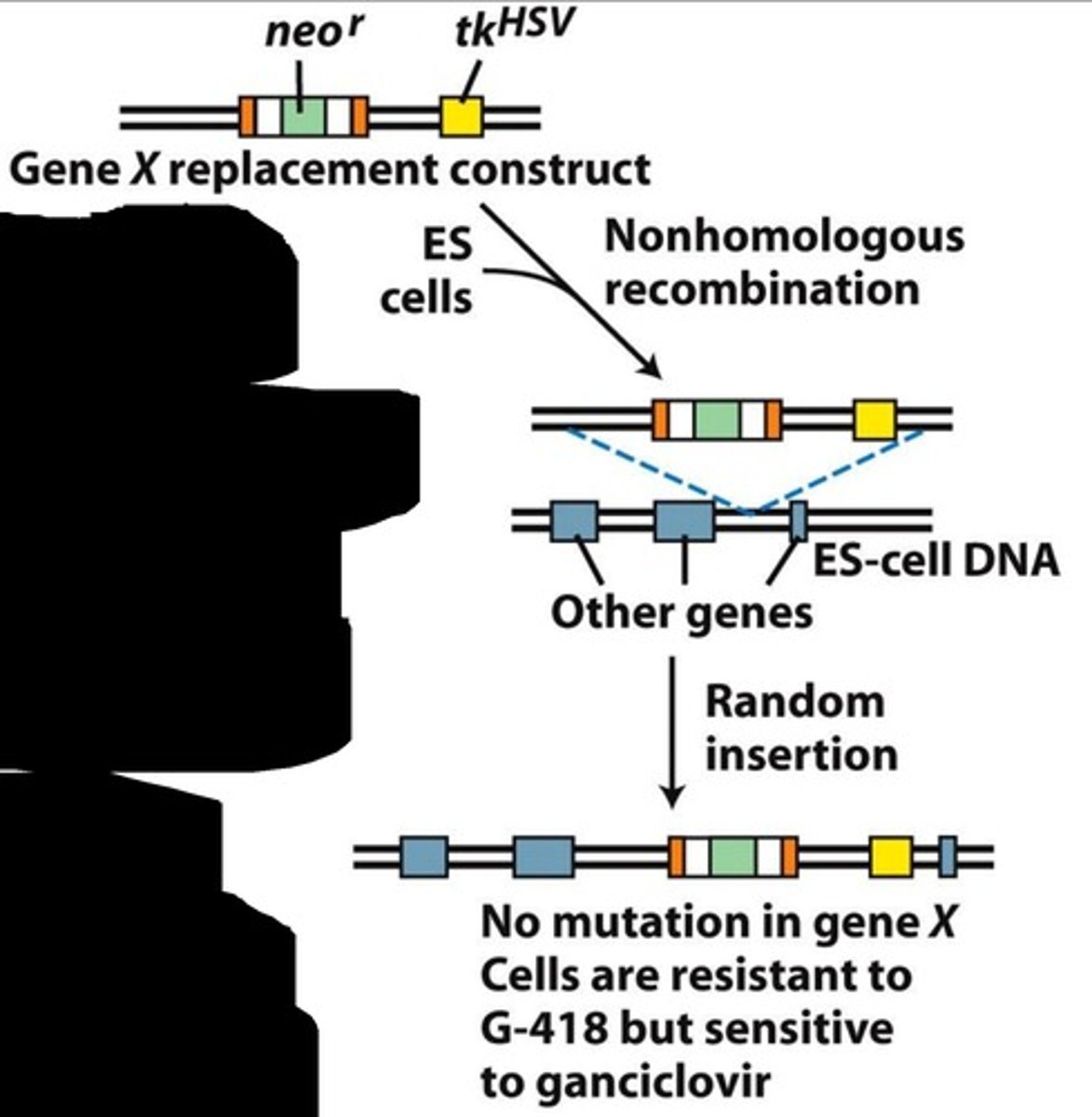
What is the Neo gene in relation to the knockout selection process?
Neomycin resistance gene which makes cells resistant to G-418/neomycin. Normally, mammalian cells are susceptible to neomycin. If any recombination occurs (homologous or not) within the ES cells, they will be resistant to neomycin.
Why does any recombination result in G-418 resistance?
The Neo gene is inserted in the middle of the target gene within the vector construct. This leads to a knockout of the target gene and provides neomycin resistance. If either type of recombination occurs, this gene will be present and provide drug resistance.
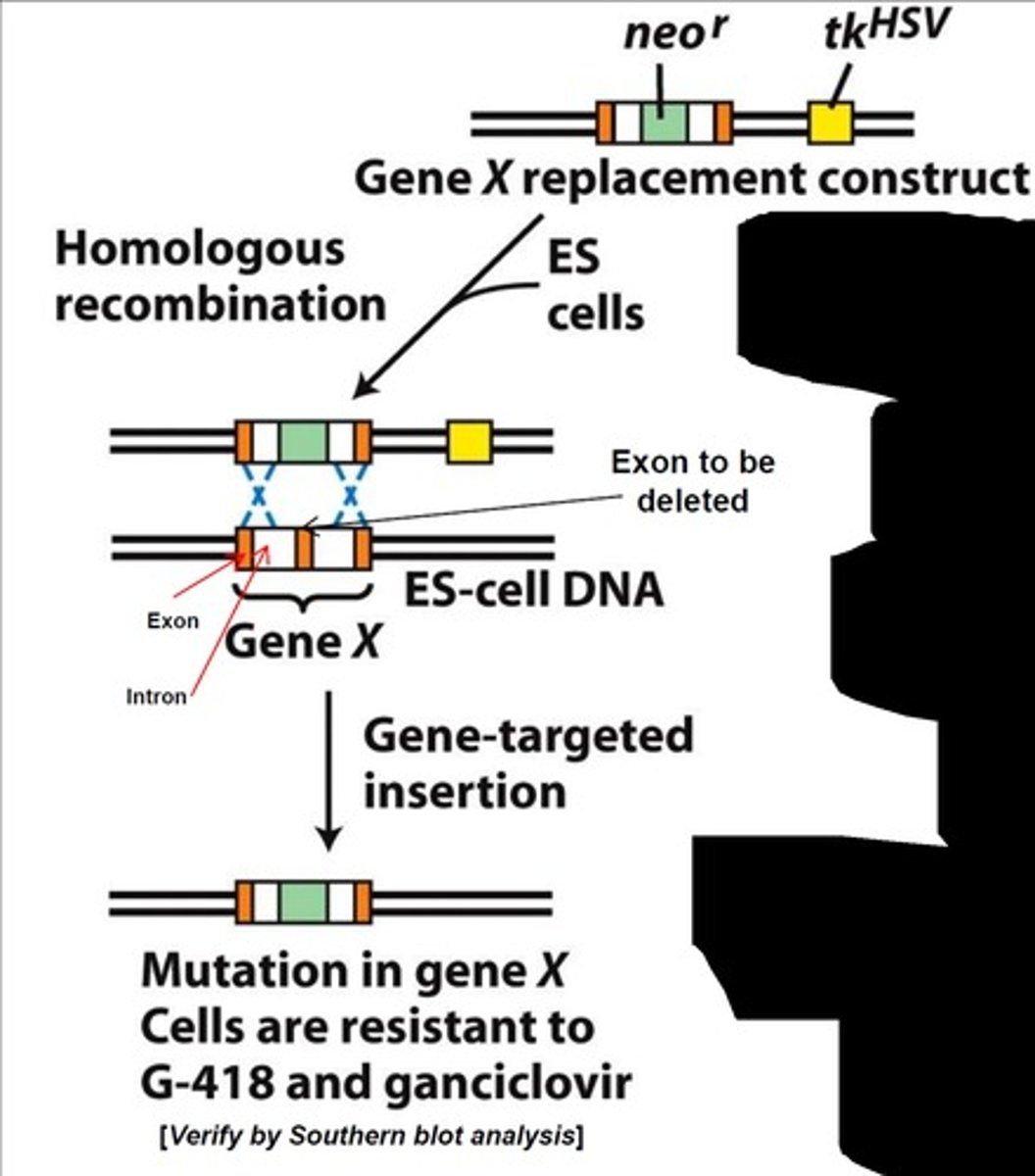
How are knockout embryonic stem (ES) cells selected?
Through the use of ganciclovir and G-418/neomycin. During the recombination process 3 cell types will be formed: homologous recombined, non-homologous recombined, and non-combined ES cells. The first will be resistant to both drugs. The second will be susceptible to ganciclovir. The third will be susceptible to G-418/neomycin.
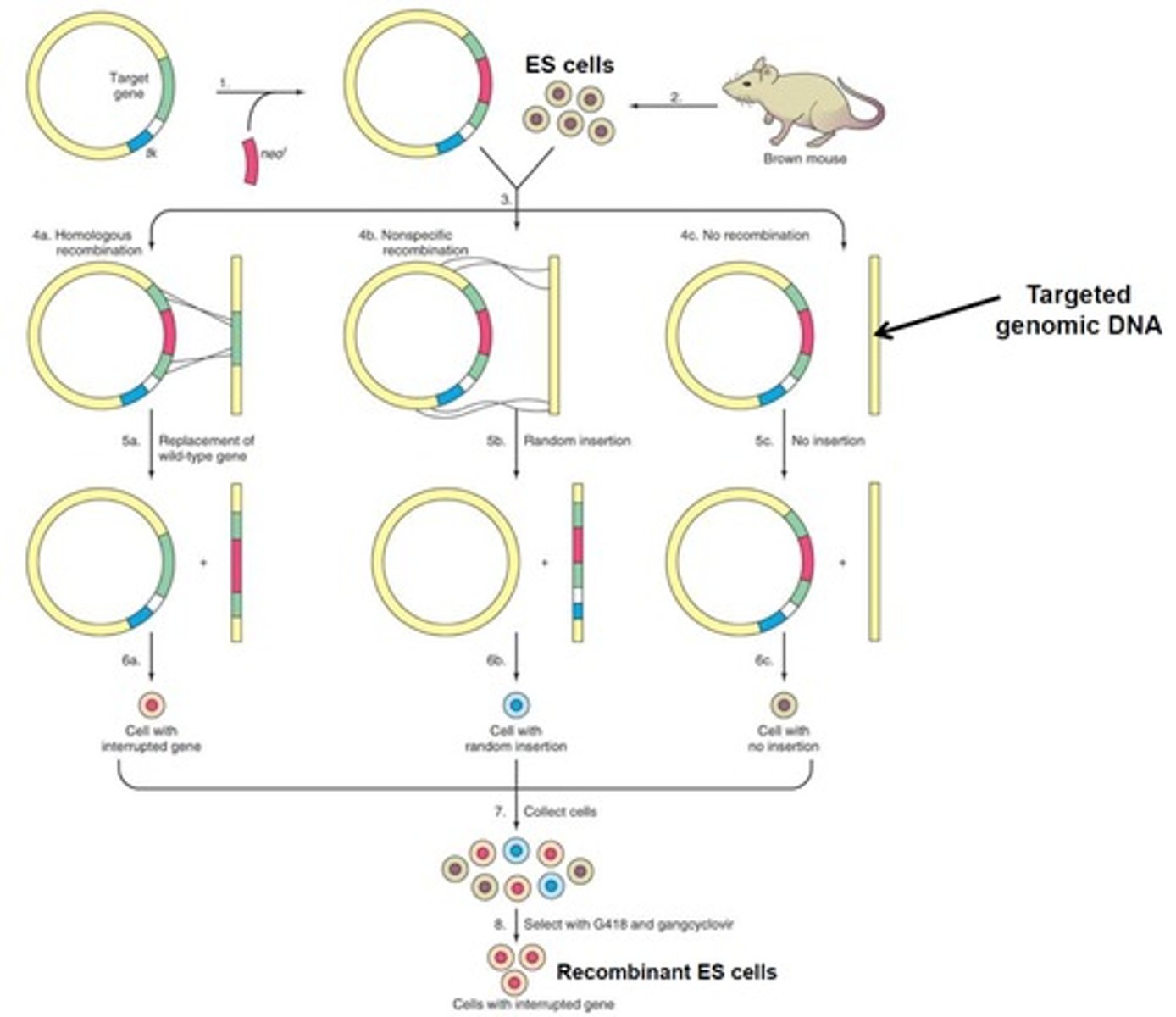
After knockout ES cells are selected, how are they used to create knockout offspring?
They are injected into the blastocoel cavity of early embryos from black wild type mice. These embryos will contain both the brown knockout cells and the black wild type cells and can be implanted into a foster mother to produce progeny.
How are progeny contain knockout genes determined?
Through the use of recessive vs dominant fur color genes. The brown gene is dominant while black is recessive. Therefore, progeny that came from the embryo containing both cell types will be chimeric and have both colors in their fur.
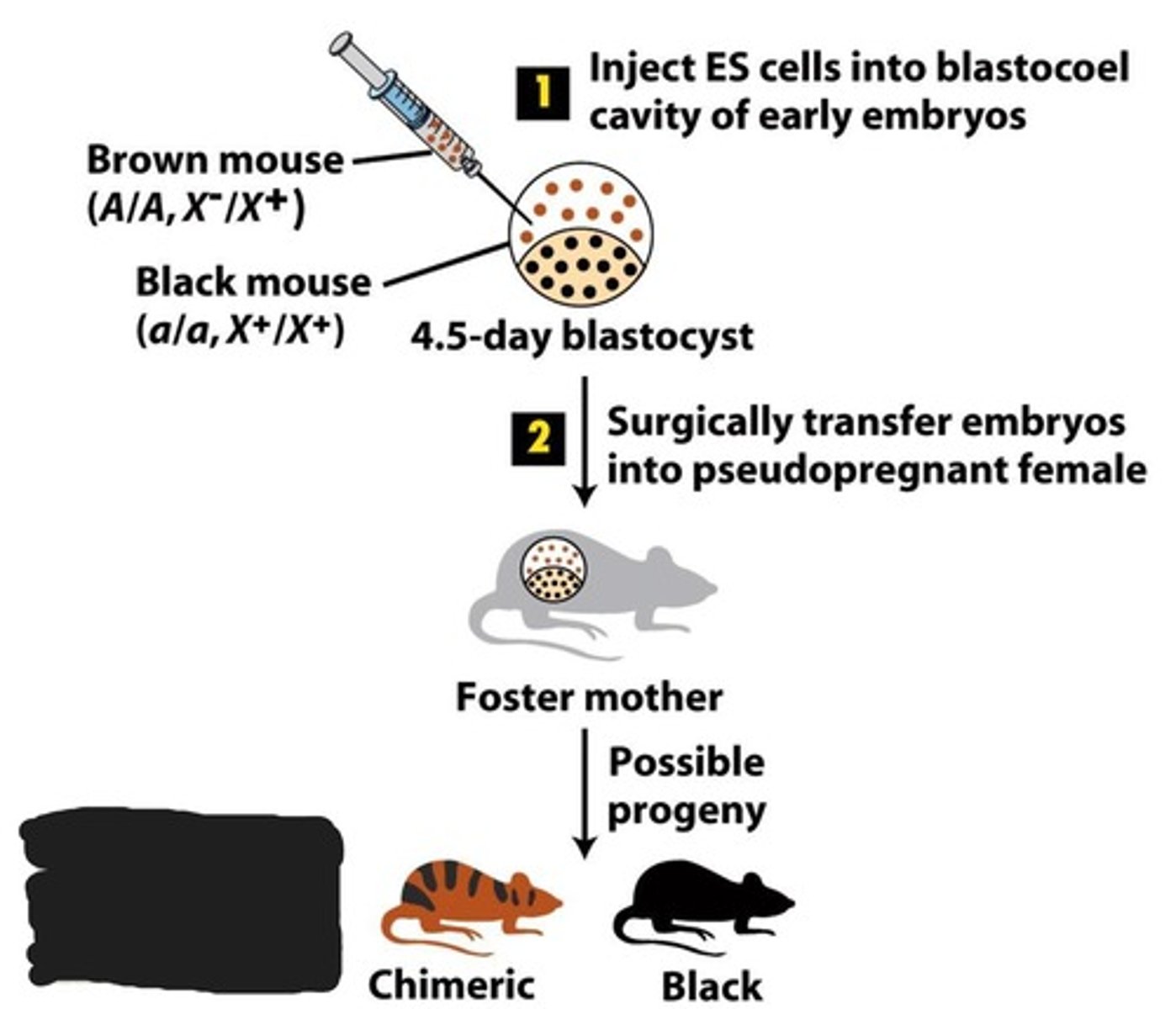
Do chimeric mice have homozygous knockout genes?
No. Chimeric mice have a mix of germ cells due to their initial embryos containing both knockout and wild type ES. Therefore, they have genes from the heterozygous knockout and homozygous wildtype.
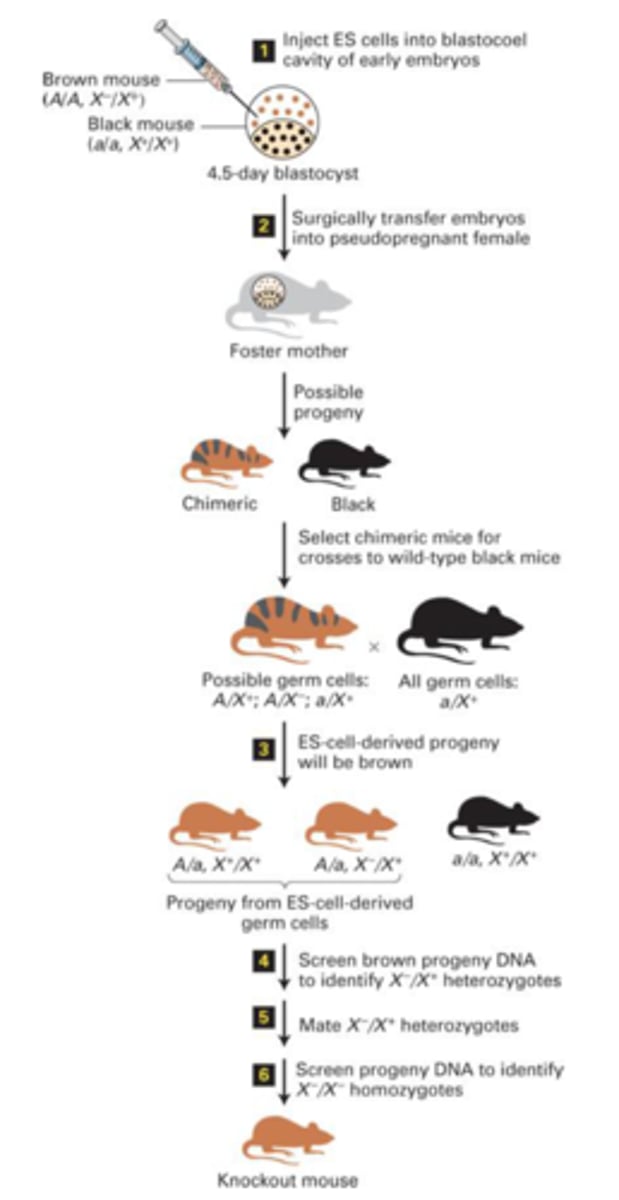
How are homozygous knockout mouse created?
Breeding a chimeric mouse with a wild type black mouse, then breeding the heterozygous progeny with another.
The initial breeding will result in three progeny options: 1 brown with wildtype homozygous, 2 brown with heterozygous wildtype/knockout, and 3 black homozygous wildtype.
The second breeding will be two of the 2nd progeny. The progeny of this second breeding can be screened for homozygous knockouts.
Keep in mind, germ cells only have one copy of each gene from the parent
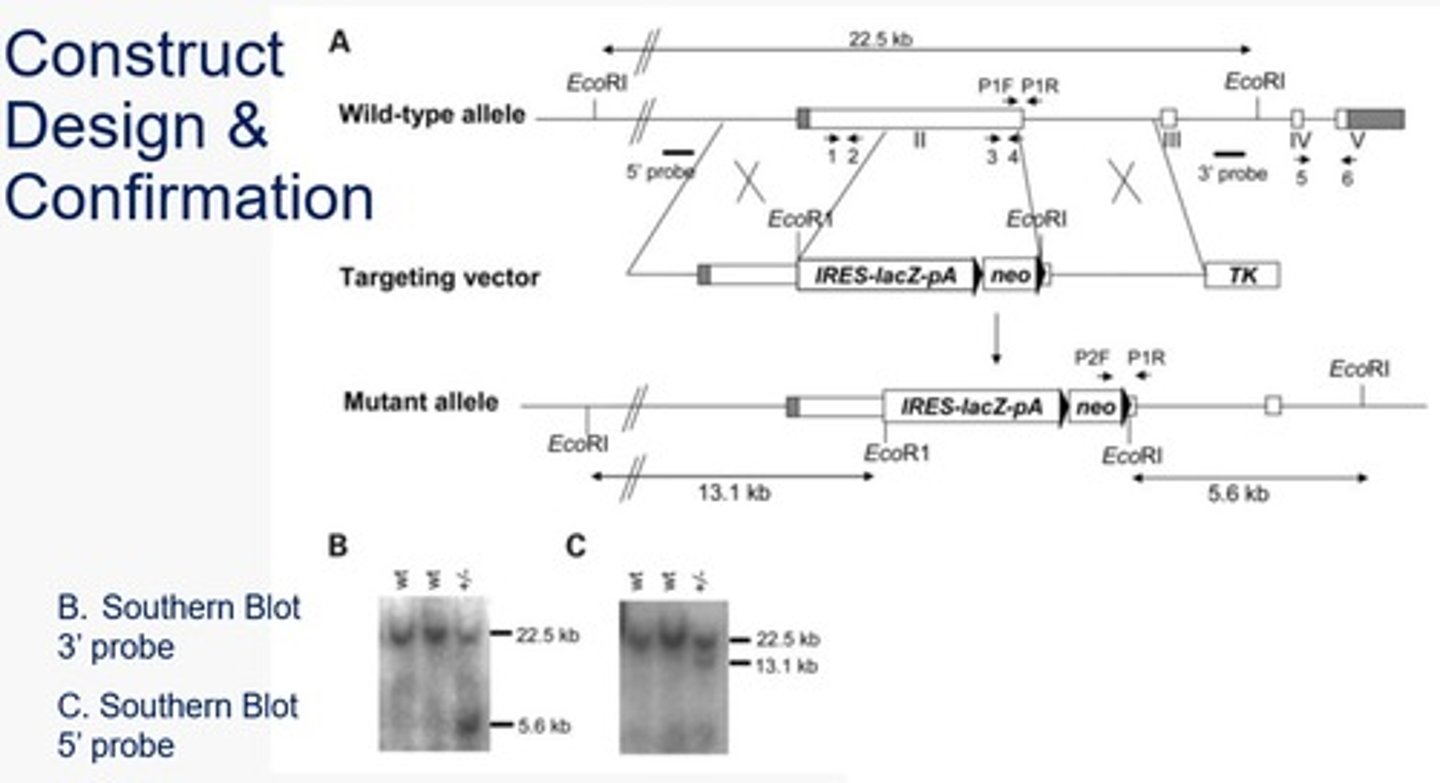
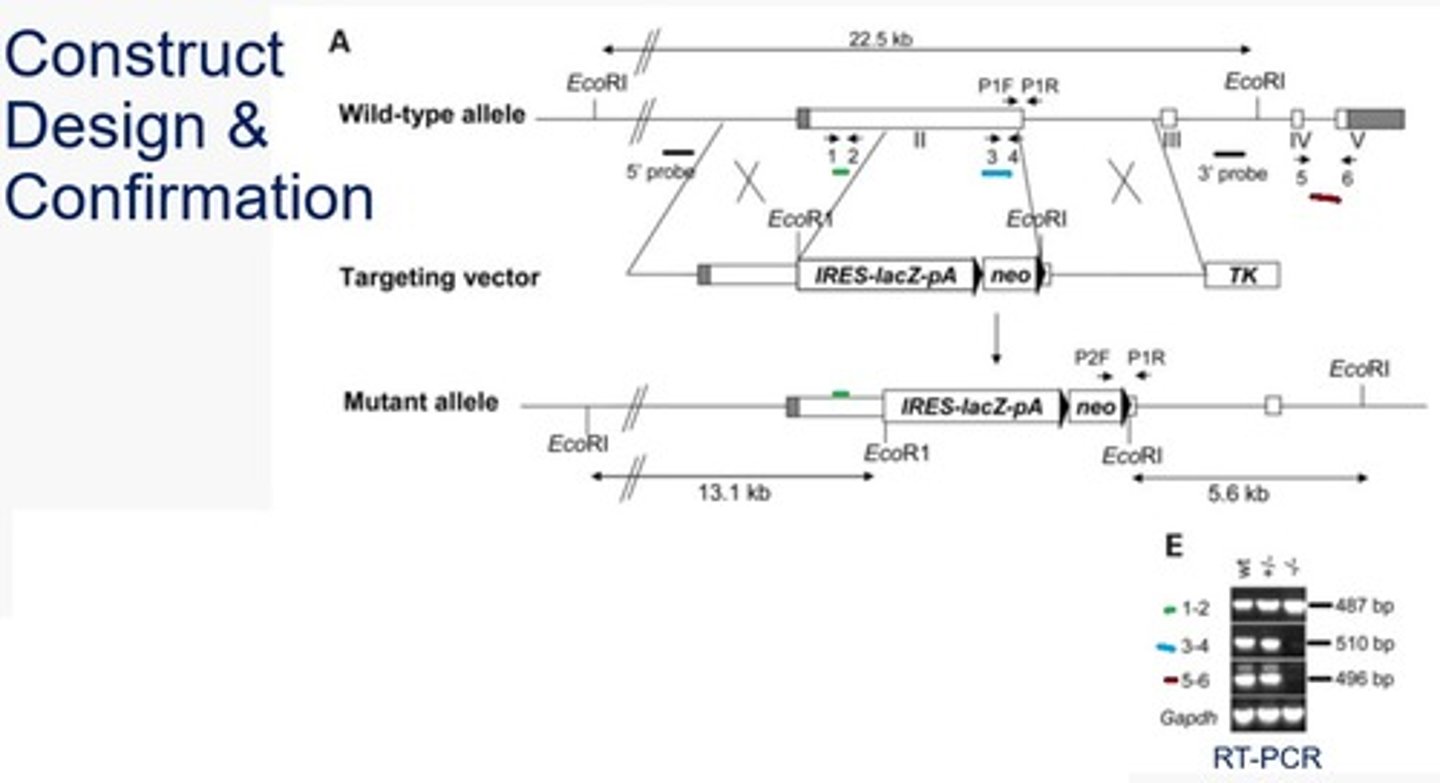
Summarize and explain the RAI1 knockout example southern blot figure (after the using EcoR1 restriction enzyme digestion).
Wildtype = 2 fragments produced - one contains both probe sites and the other has none >> one band produced when using either probe
Mutants = 4 fragments produced - one contains both probe sites, one contains the 5' probe site, one contains the 3' probe site, one has none >> two bands produced using each probe (larger band is from the wild type allele, the smaller band is from the mutant allele)
In the RAI1 knockout experiment, how were homozygous gene knockouts determined?
PCR using two sets of primers. Each set used the same reverse primer while two different forward primers were used based on the wildtype vs knockout. The wildtype forward produced a small band while the mutant produced a larger band. In a heterozygote, two bands would be produced. In a homozygote (wildtype or mutant), a single band would be produced.
*The wildtype F primer was within the region the knockout occurred and the mutant F primer was within the NeoR gene.
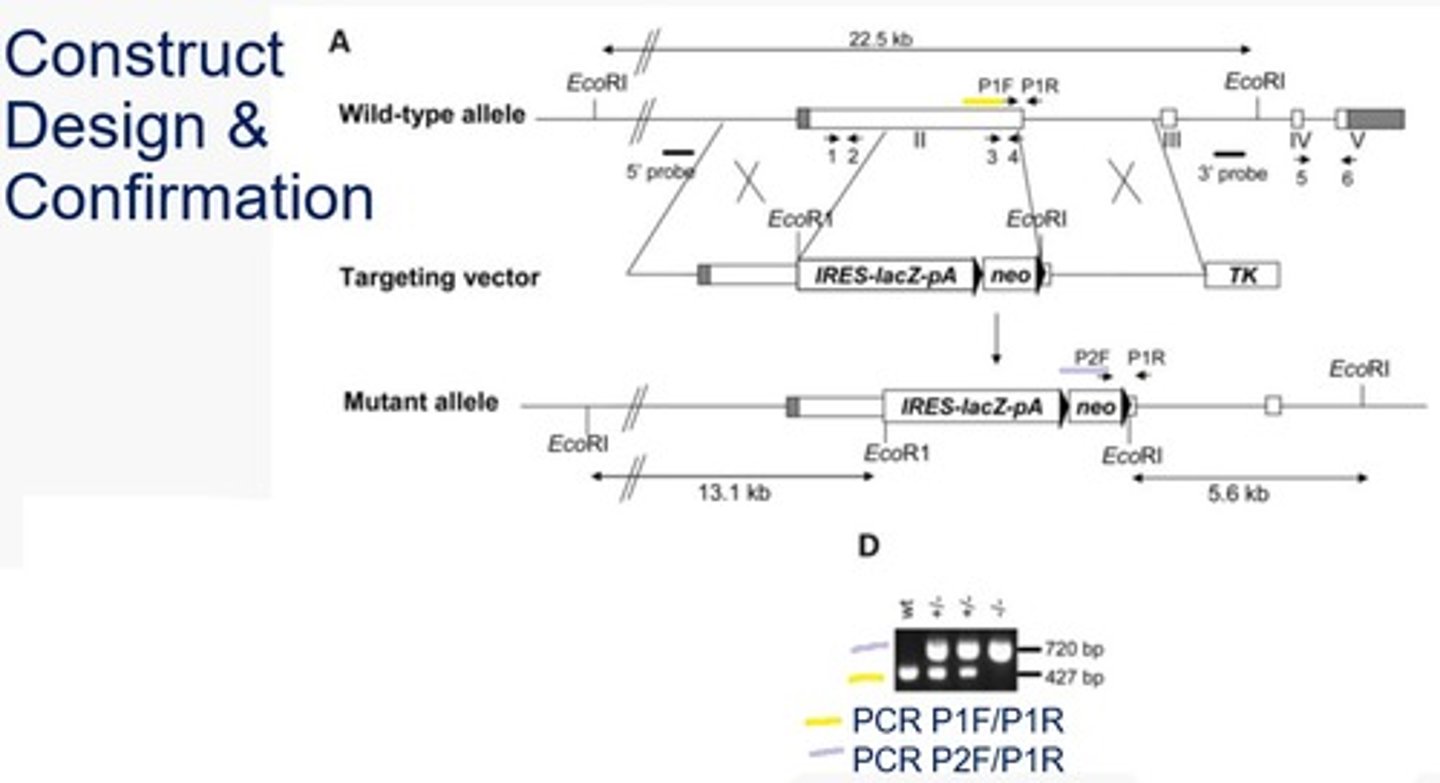
In the RAI knock out experiment, how was transcription knockout confirmed?
RT-PCR using 3 sets of primers (1&2, 3&4, and 5&6).
1-2 (5' end of the target gene) - the smallest banding, band will occur on all because it is on the target gene and outside the mutated region
3-4 (within the target gene, replaced by neomycin resistance in the mutant gene) - band will only occur in the wildtype gene because the primer site is not present in the mutant
5-6 (3' end of the target gene) - this band will only occur in the wildtype gene because the mutant gene contains transcription termination sites at the end of the LacZ and Neomycin resistance genes, so the 5-6 primer region will not be transcribed and therefore no band will be produced.
*Gapdh - this control is used because it is a transcriptionally active gene that will be unaffected by the mutation of the gene of interest
Which of the following statements is true about embryonic knockout cells?
The disrupted gene will replace the endogenous gene
3 multiple choice options
What is the advantage of using the Cre/Lox knockout system?
Instead of having the gene knockout occurring at birth, it can occur conditionally (e.g., specific time, location, or condition). This is especially useful for genes (like the RAI1 example) in which the homozygous knockout results in fetal death. It can also help avoid the potential for compensation of the progeny in response to the knockout or it can help to study the local effects of a knockout without the possibility of causing global systemic change.
When using the Cre/LoxP system, where are the LoxP sites located?
Flanking either side of the target gene/sequence (aka a floxed gene). This is significant because when the Cre recombinase is activated, recombination between the LoxP sites on the same allele occurs thus leading to target sequence conditional removal but the gene can still be transcribed up until that point.
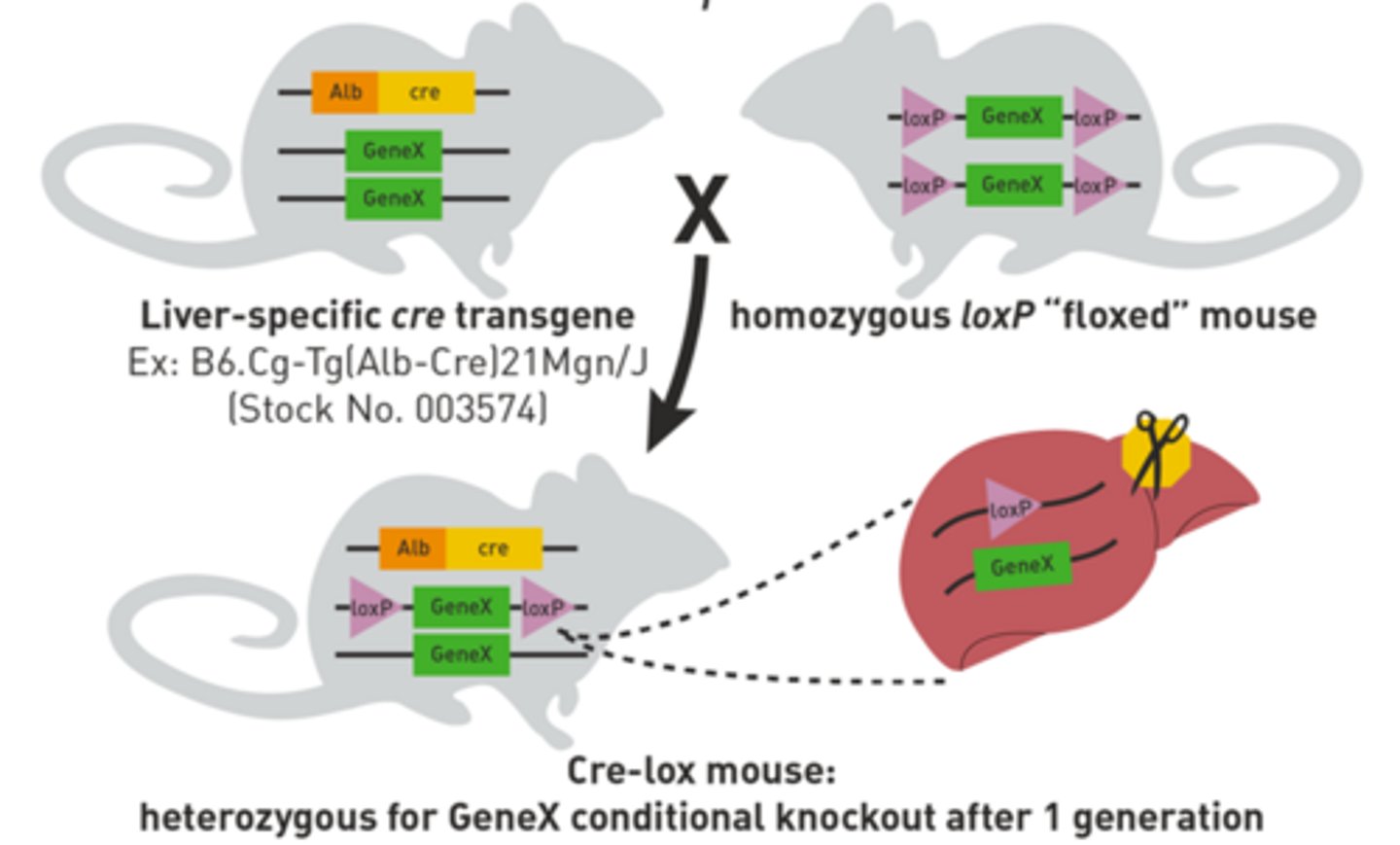
Why does the LoxP mouse need to be crossed with the Cre mouse?
The LoxP sites will not recombine without the Cre recombinase. Therefore the LoxP mouse needs to cross with a Cre mouse so that the progeny will obtain both the Cre recombinase gene and the floxed target gene. However, a single cross will only result in a heterozygous floxed gene. Therefore, the first progeny will need to be crossed with a LoxP mouse again in order to produce Cre as well as homozygous conditional knockouts.
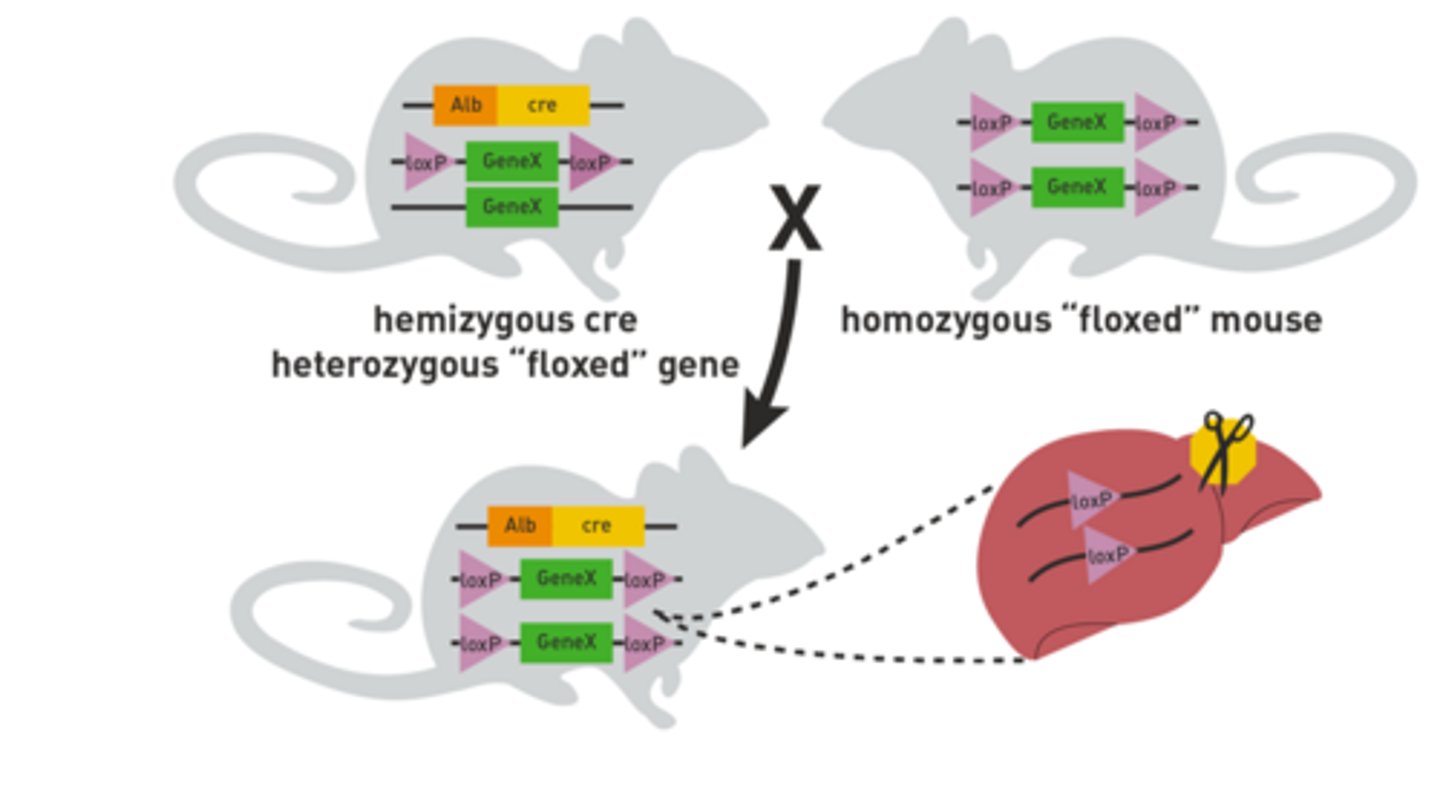
What feature of the Cre/Lox system allows for a conditional knockout of a target gene?
The Cre recombinase gene is placed next to the gene necessary for the conditional knockout. The example used placed the Cre gene next to a liver-specific gene. Therefore, the Cre gene would only be transcribed within the liver so the recombinase will only be produced and be active within the liver. Without the recombinase activity, the target gene remains active so the knockout only occurs in the liver where the recombinase is produced/active.
What is CRISPR?
Clustered Regularly Interspaced Short Palindromic Repeats - a revolutionary discovery that lead to the ability to edit genes in vivo. This can help with genetic diseases, Aids, cancer, etc.
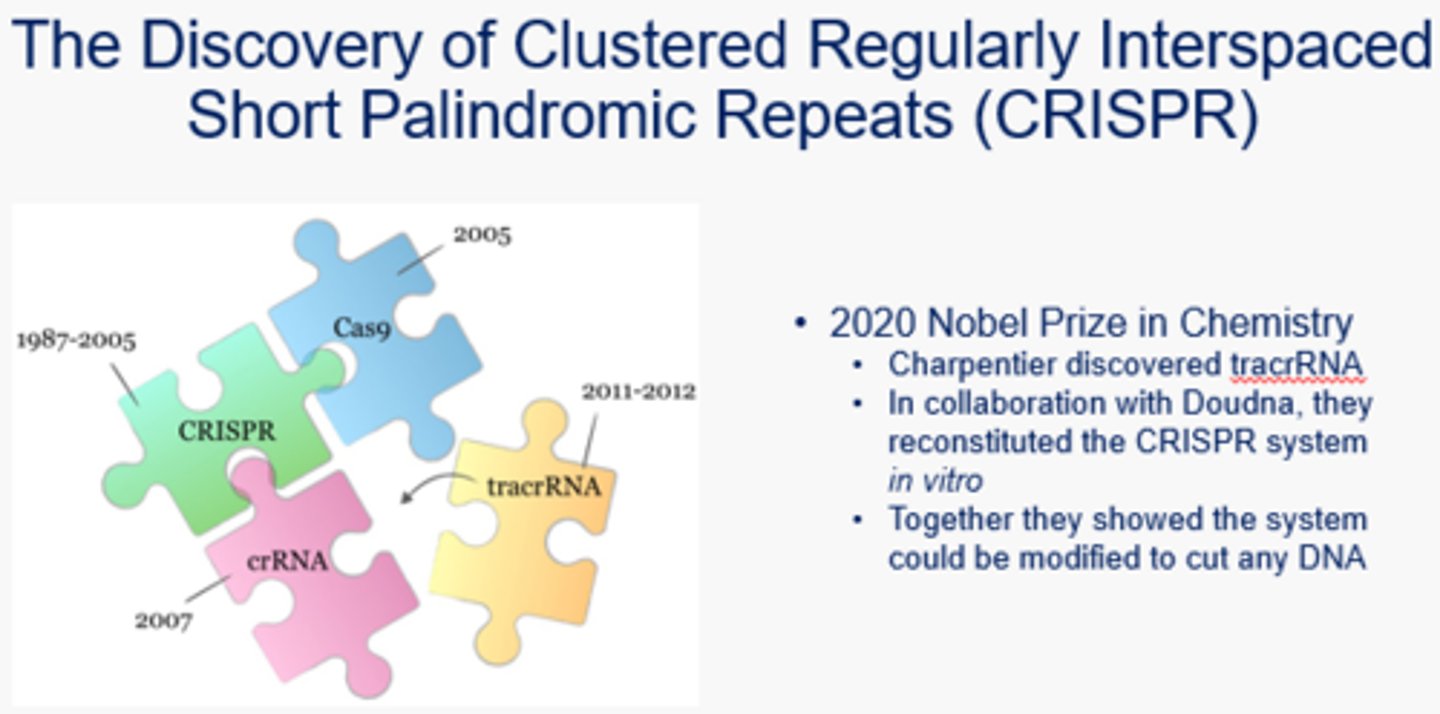
What is the immunization phase of CRISPR-Cas9 in bacteria?
Aka the acquisition phase - the Cas Complex, formed from the transcription of the Type II CRISPR locus, contains a Crispr-associated Endonuclease that chops up viral/foreign DNA and inserts a fragment into the end of the CRISPR locus as a "spacer." There is a spacer for every past viral DNA encounter, and they are separated by repeat sequences.
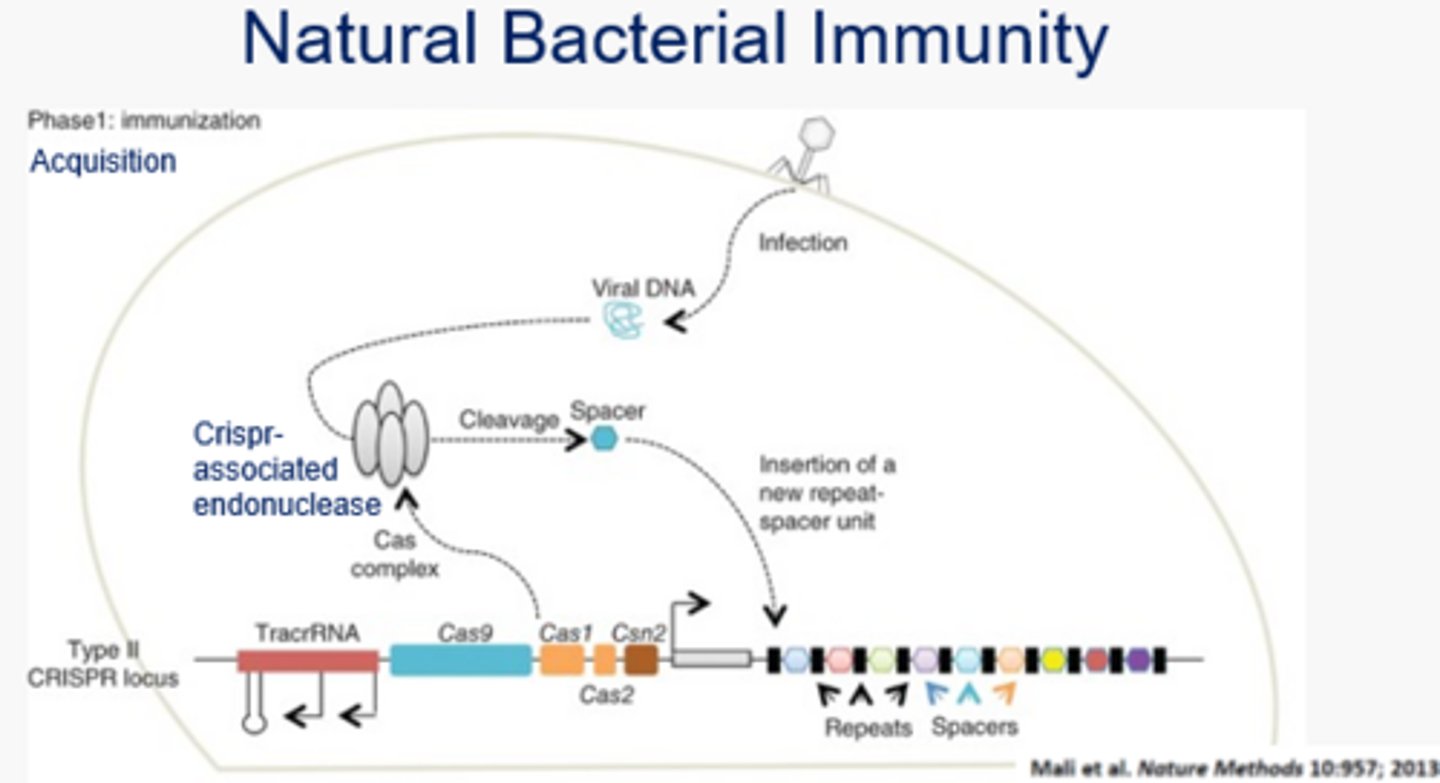
What is the immunity phase of CRISPR-Cas9 in bacteria?
Aka the interference phase - this occurs during reinfection in which viral DNA is recognized using the previous "spacers" and leads to viral DNA destruction
What RNAs are present in the Cas9 complex and what is their role?
Single spacer RNA sequence - complementary binding with foreign DNA
Single repeat sequence - binding to the TracrRNA
TracrRNA - binding to the repeat sequence to stabilize the RNA-Cas9 complex
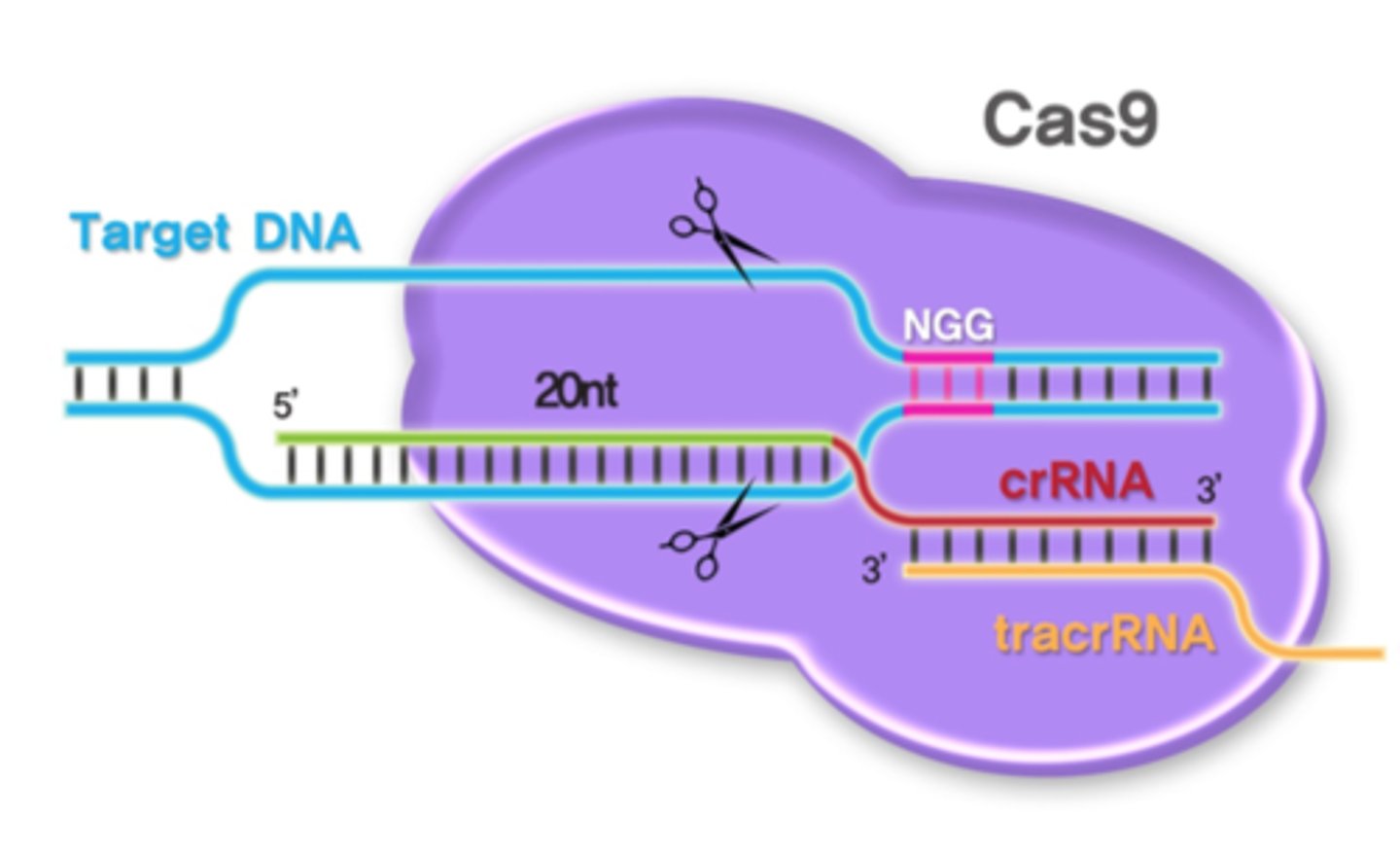
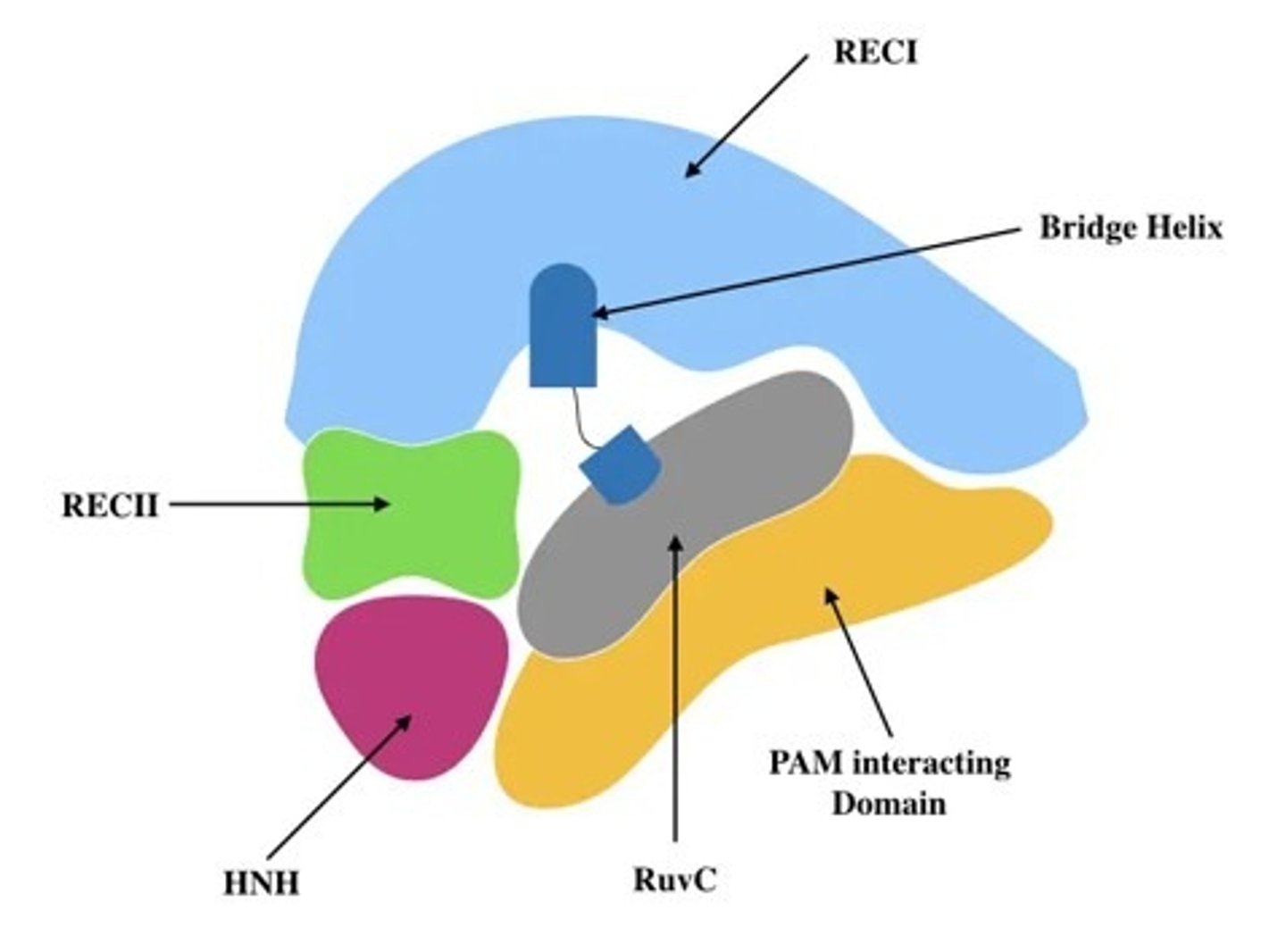
What are the significant Cas9 protein elements and their functions?
Rec I and II - foreign recognition, complementary base pairing between the spacer element and the viral dsDNA
HNH and RuyC - nuclease, each cuts one of the strands of the viral DNA to result in the dsDNA break
Pam domain - interacts with the PAM sequence to confirm foreign DNA presence, without this sequence no strand breaks would occur
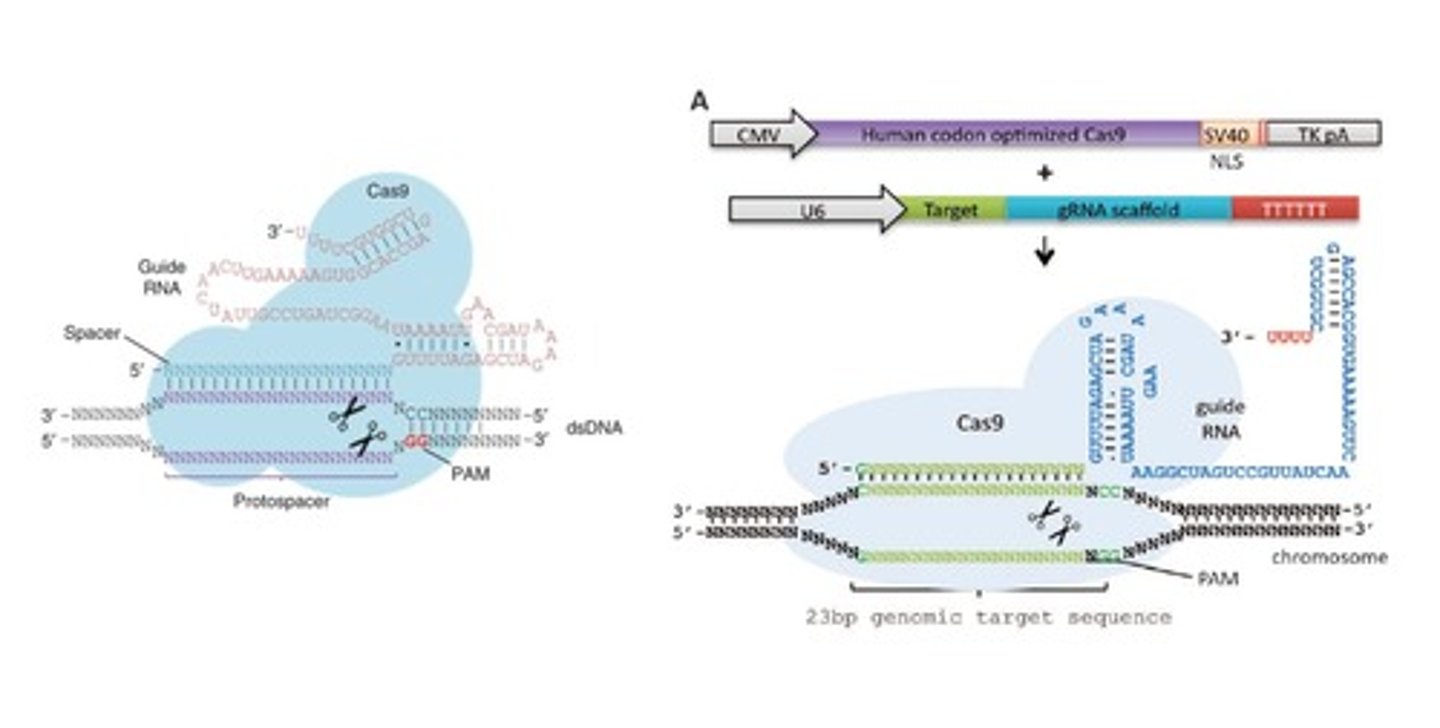
Why is the PAM sequence significant in relation to CRISPR-Cas9 technology?
In order to properly edit genes, breaks within the target DNA must be formed. Without the PAM sequence, no breakage will occur and thus no gene alteration can happen. Therefore, the target genes must be located near a PAM site in order for recombination of the target-mutant to occur.
In regards to using Cas9 as a gene editing system, what does guide RNA (gRNA) do?
Single guide RNA (sgRNA) is the complex of complementary RNA for the target gene with the guide RNA. The gRNA acts in place of the TracrRNA in that it stabilizes the RNA-Cas9 complex without the need for a second strand of RNA (the repeat element is not longer necessary)
What are the 2 possible repair mechanisms of a double stranded break?
Non-homologous end joining (NHEJ) - error prone repairs that likely result in deletions or insertions
Homology directed repair (HDR) - recombination of the target gene with donor DNA
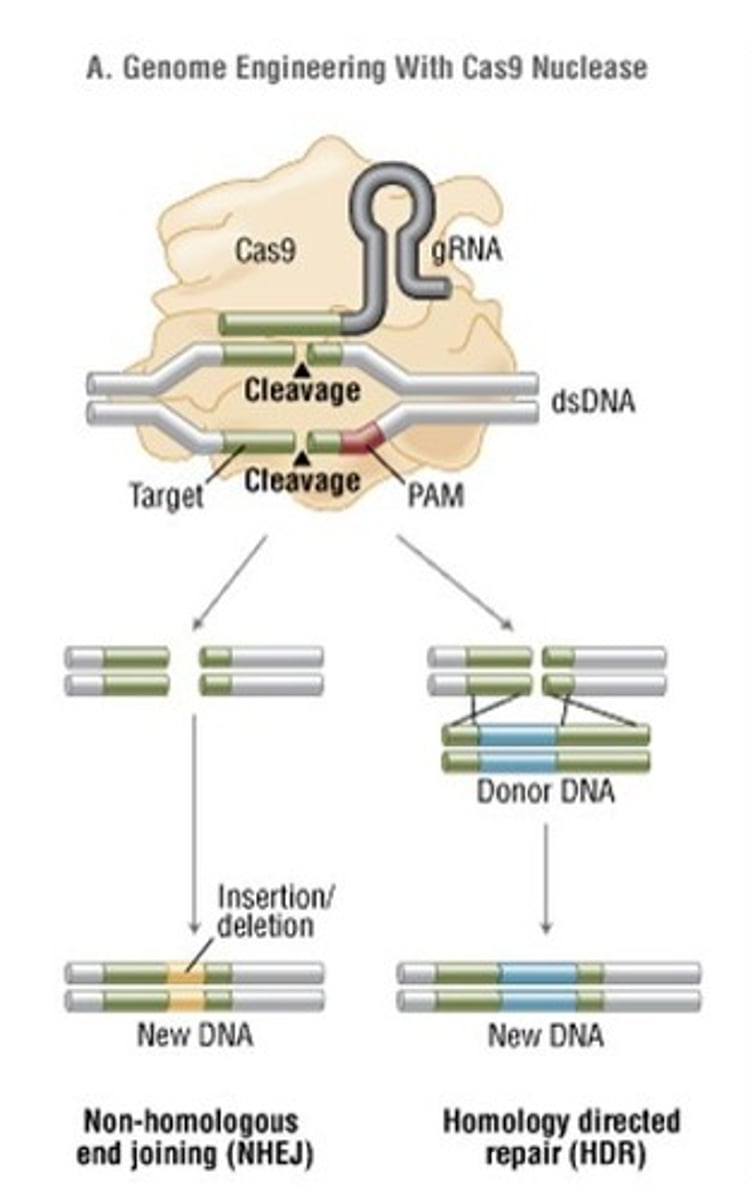
Which double stranded repair mechanism is best for a 3 nucleotide change in the target gene?
Homology directed repair (HDR) is best for sequence alterations and it is very likely to occur in both alleles because the double stranded break occurs in both regions of the target gene
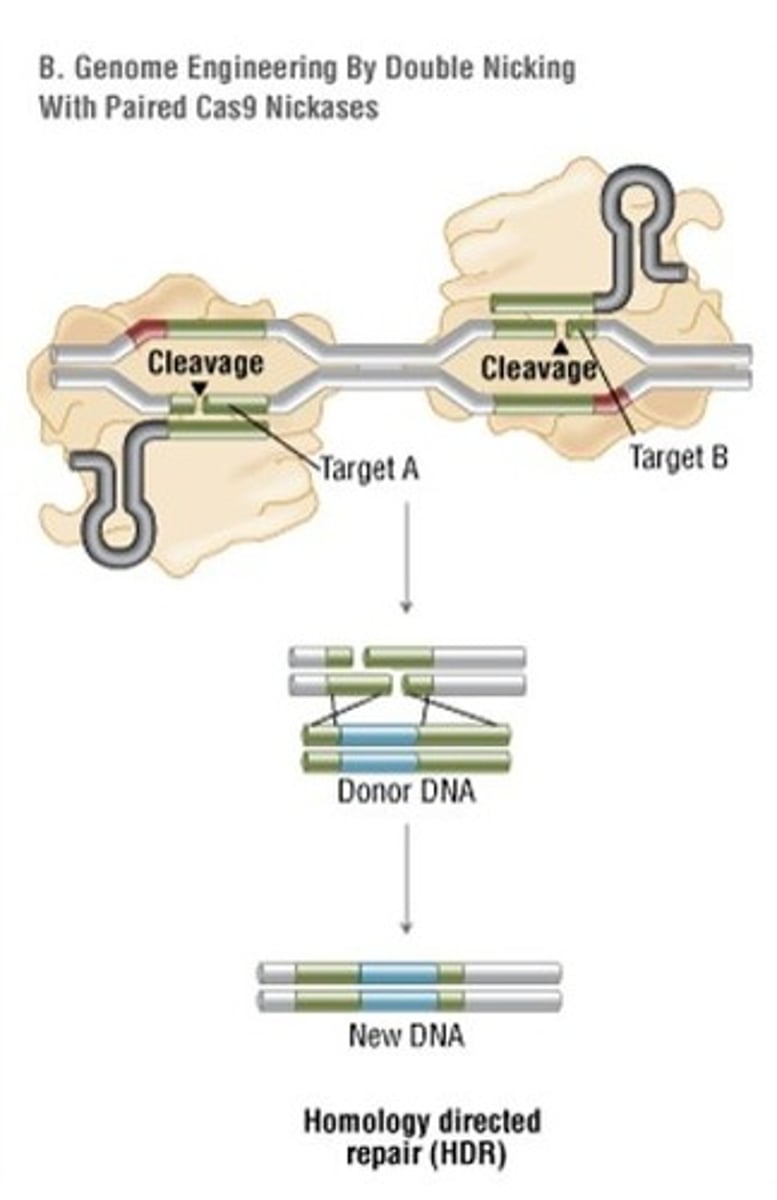
How can the Cas9 system be made even more gene specific?
Using a mutated Cas9 with paired nickases. The mutated Cas9 will only cause a single stranded break rather than a double stranded break. there are two mutated Cas9's present and complexed with different sgRNA's for the same gene. The result being the dsDNA break will only occur if both binding and cutting events occur (as opposed to a single binding event). This effectively eliminates NHEJ because dsDNA can only occur with the target gene.
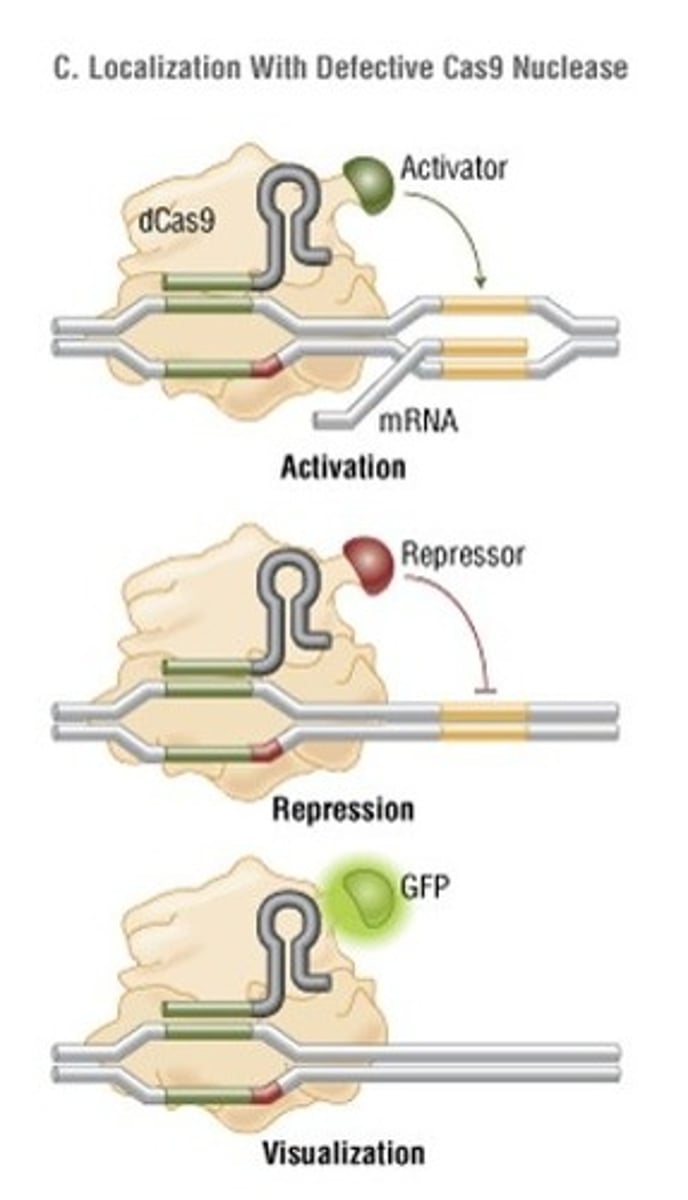
Aside from gene recombination and editing, what is another way in which the Cas9 system can be used to influence or study genes?
Cas9 can be mutated to no longer cut (defective nuclease) and complexed with an activator or inhibitor element to affect gene transcription. Additionally it can be complexed with a fluorescent molecule instead in order to visualize where genes are located in the nucleus.
Why is sickle cell anemia a good candidate for CRISPR treatment?
This disease results from a single base-pair change in the gene that causes the amino acid to change from a hydrophilic one to a hydrophobic one. This mutation is only actively expressed in the RBC. This means the only cell type needing the gene alteration are RBC. Therefore, red blood stem cells can be removed, altered, confirmed for correct integration, grown, and replaced within the body to fix the issue. Not every cell in the body needs to have this gene altered for it to work.
What are the two possible CRISPR treatments for sickle cell anemia?
1 - bypassing the beta globin gene via inactivation of the fetal (gamma) globin repressor, healthy gamma globin will over power the defective beta globin and prevent RBC sickling
2 - swapping the beta globin gene with a healthy beta globin gene so no defective beta globin is produced
What was the defective beta globin gene swapped with when treating sickle cell anemia?
The healthy beta globin gene, without introns, was inserted in the center of the defective beta globin gene. This effectively knocked out the defective gene and provided all the necessary exons for the healthy gene rather than just replacing the single altered base pair.
How were the researchers able to determine if the wild type beta-globin gene was targeted by the donor sequence?
Using PCR and 2 primer sets - one set of primers is for the wild-type (defective) beta globin gene and will result in a larger band size while the other set of primers will be for the integrated sequence that will have a smaller band. This will result in a banding pattern that will indicate homologous WT, heterozygous (mono-allelic integration), or homologous integration of the corrected beta globin gene.
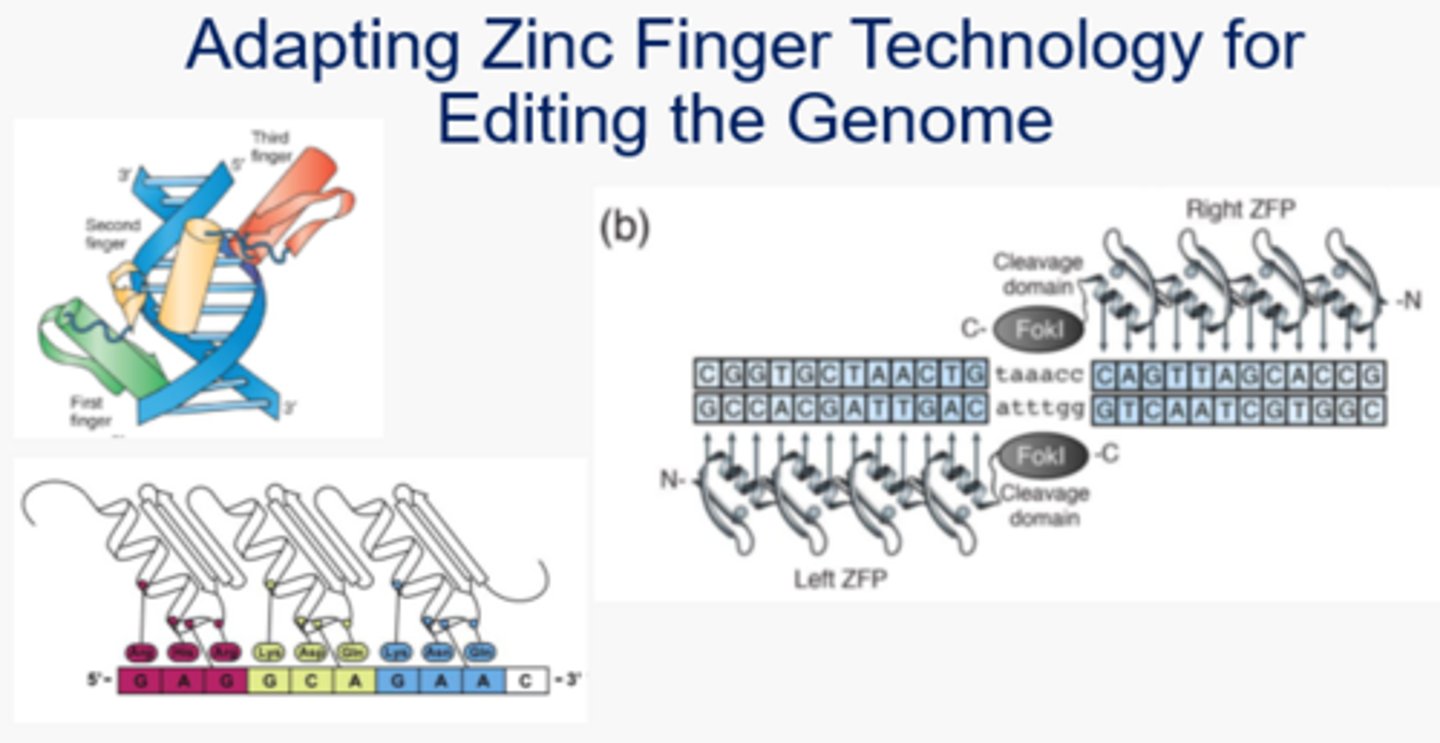
In the Zinc Finger approach to gene editing, what would be the equivalent of the gRNA from CRISPR? Why?
Zinc finger - binds to target DNA sequence based on the amino acid sequence that interacts with the DNA directly (specific amino acids will react with specific nucleotides)
FokI enzyme bound to Zinc finger - performs the double stranded break at the 3' end of the target DNA.
In this approach, the sgRNA is replaced with the Zinc finger and the FokI replaces the Cas9. The FokI enzyme can only cause a single stranded break, so two different complexes (right and left ZFPs) will need to be used to cause a full double stranded break.
When using the Zinc Finger technology to treat HIV, what is the importance of targeting the CCR5 transporter protein gene?
The CCR5 transporter protein is the target for entry of HIV. Researchers found that mutations in CCR5 prevents HIV infection of cells.
Briefly describe how Zinc finger technology is helpful in controlling the replication of HIV-1.
ZFPs were created that target the CCR5 protein gene. Then CD4 T stem cells are removed and isolated from the patient infected with HIV and are used as targets for genetic manipulation. When the ZFPs bind to the gene within those stem cells, they create a double stranded break which either results in NHEJ or replacement with donor sequence DNA that contains a mutation in the CCR5 gene. Both these results lead to knockout of the gene, so cellular viral resistance occurs since the virus can no longer infect their target cells.
**Another reason this is a huge advance in HIV treatment because the same patient's cells are being altered and returned, so there is no chance for a donor reaction like there would be if the cells were being replaced with a different donor's cells.
What are the differences between prokaryotic and eukaryotic transcription?
1 - transcription and translation occur at the same time in prokaryotes because they don't have a membrane bound nucleus
2 - prokaryotes do not have post-transcription processing mechanisms
3 - prokaryotes do not wrap their DNA around hsitones
What is the difference between the template and the coding strands?
Template - the DNA that is used to build the RNA
Coding - the same sequence as the RNA
What are the two components of the prokaryotic RNA pol holoenzyme?
Core polymerase - transcription elongation
Sigma factor - promoter recognition and transcription initiation (transcription bubble formation)
What are the steps to promoter recognition and binding?
The holoenzyme searches the DNA for the promoter sequence then loosely binds. The sigma factor binds to the promoter tightly and begins melting the DNA to form the transcription bubble
What happens when sigma-70 exactly matches versus partially matches?
Exact/perfect match = high affinity, strong promoter with high rate of transcription initiation
Not exact = low affinity match, weak promoter with lower rate of transcription initiation
What are the three perfect sigma-70 binding sequences?
TTGACA at -35 (aka -35 box), TATAAT at -10 (-10 or Pribnow box), and has a spacer region of 17nt
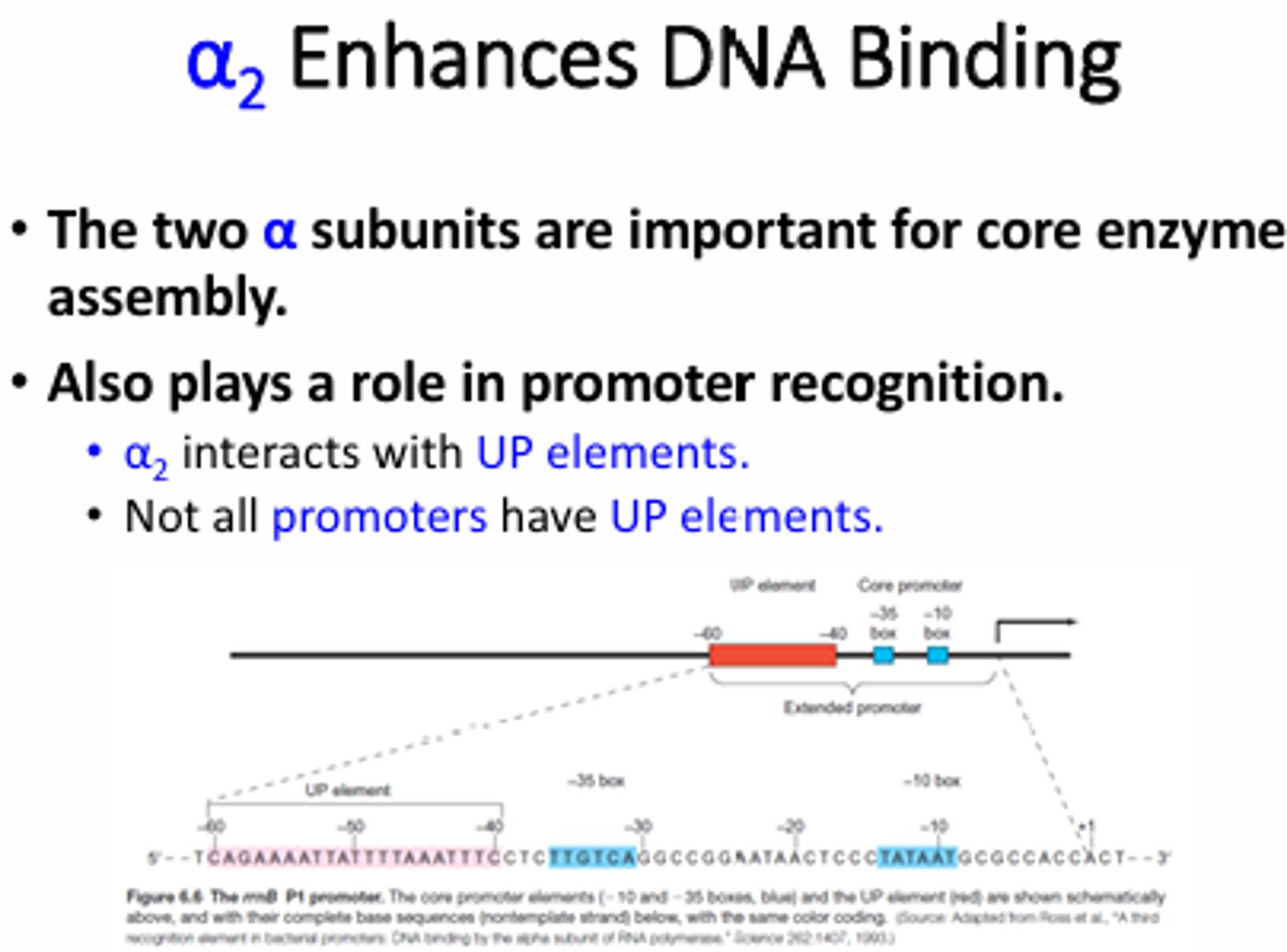
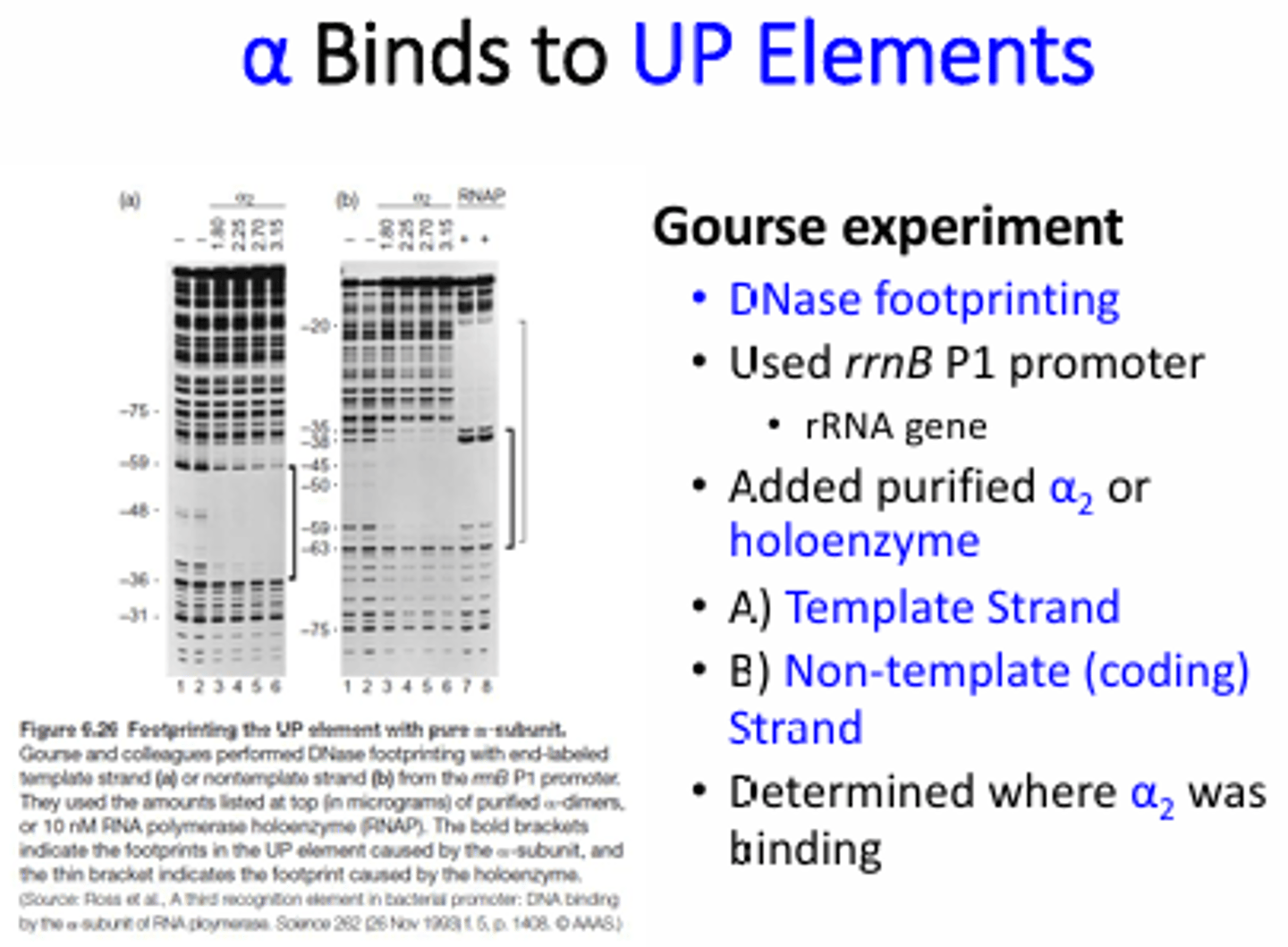
What regions of the promoter were shown to be important for RNA pol binding based on the DNA footprint results?
-40 and -60 on the template and coding strands ("UP" elements). There are 2 alpha subunits in the RNA pol core enzyme so each binds to one strand of the DNA.

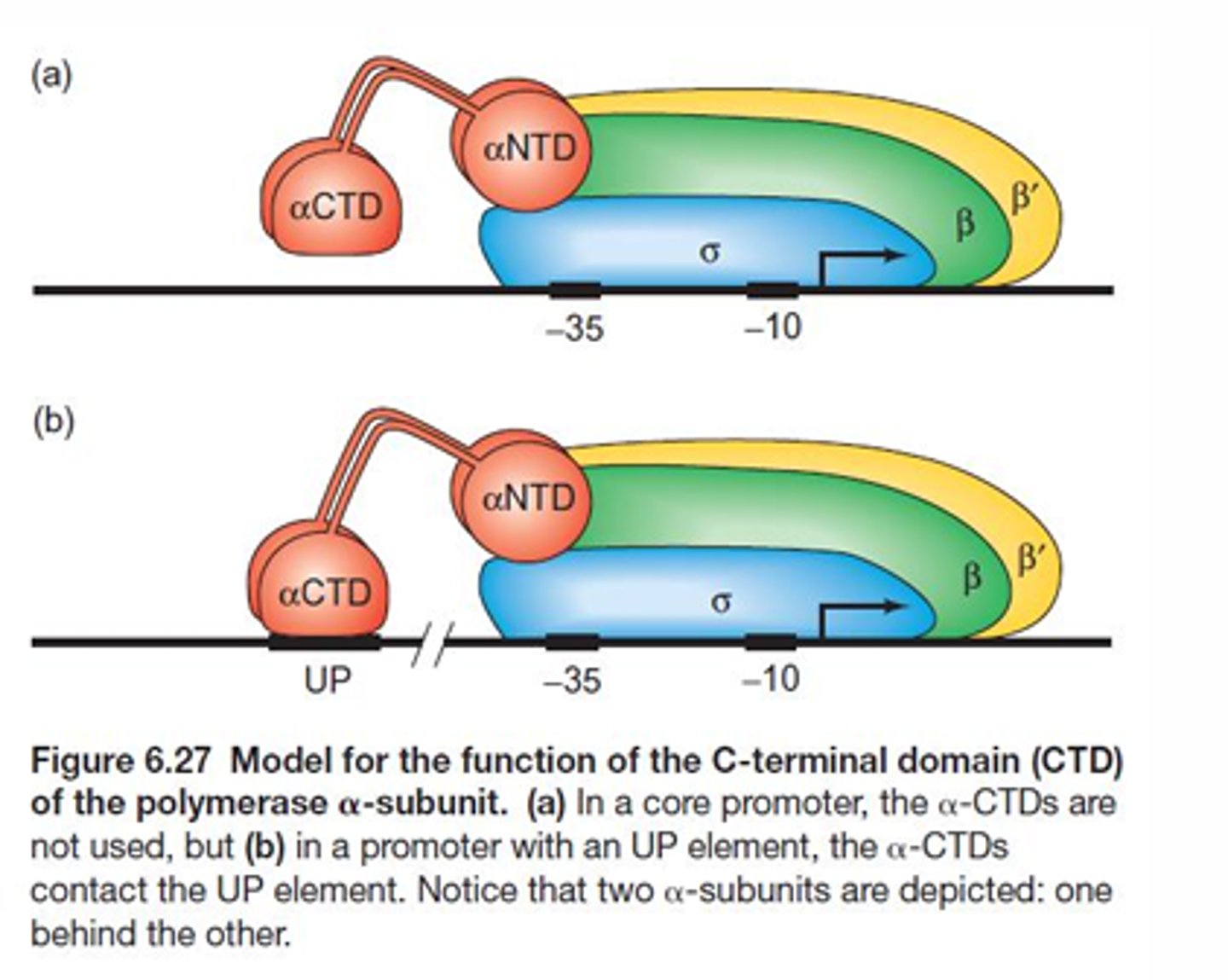
What region sof the RNA pol were shown to be important for DNA binding?
The c-terminal domain of the alpha subunit will bind to eh UP element of the promoter region while sigma binds to the -35 and -10 regions.
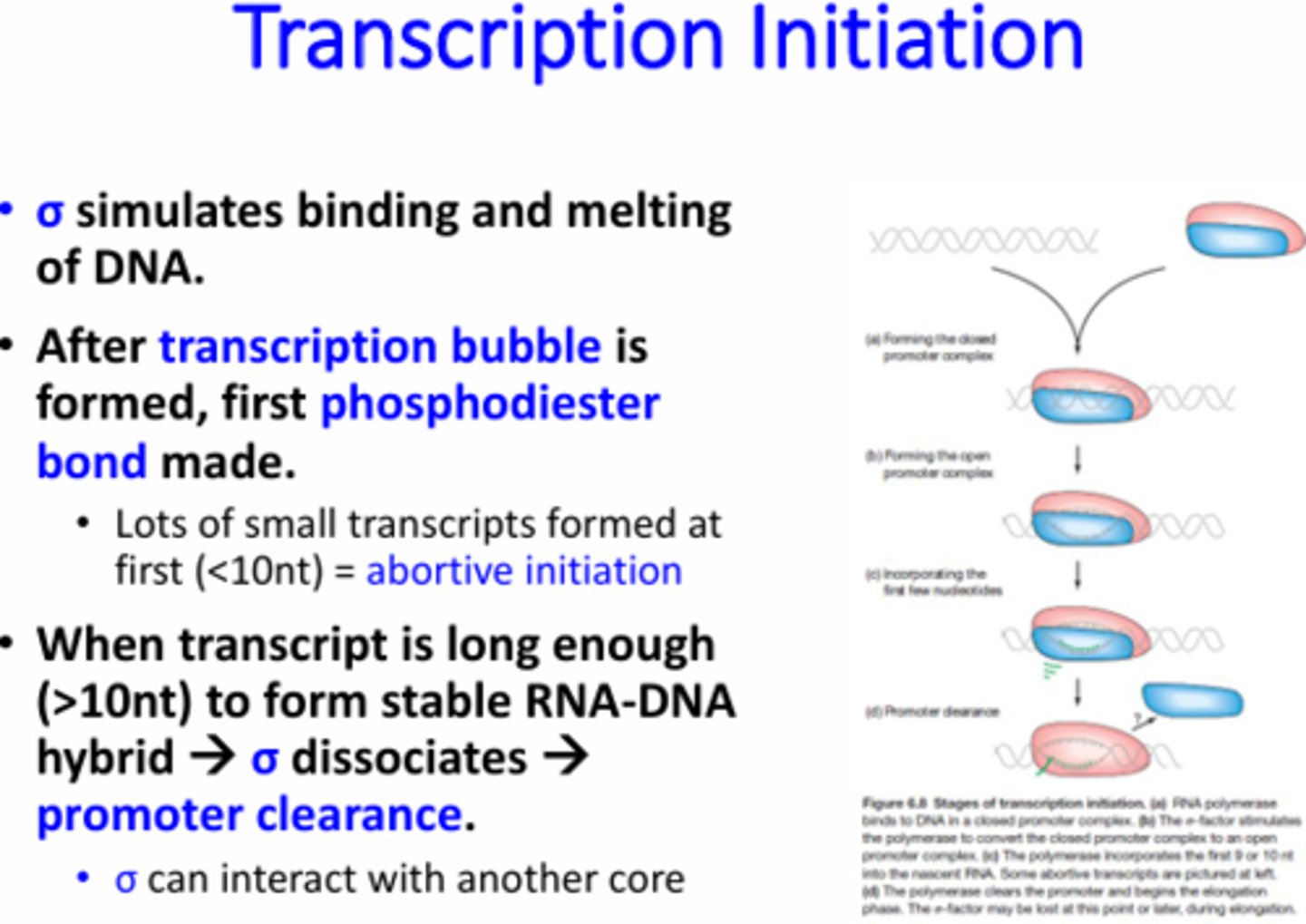
What is promoter clearance and why is it necessary?
The removal of the Sigma factor from the RNA pol complex. Sigma has a really strong affinity for the promoter sequence so RNA pol will not be able to continue down the DNA to transcribe without sigma dissociation.
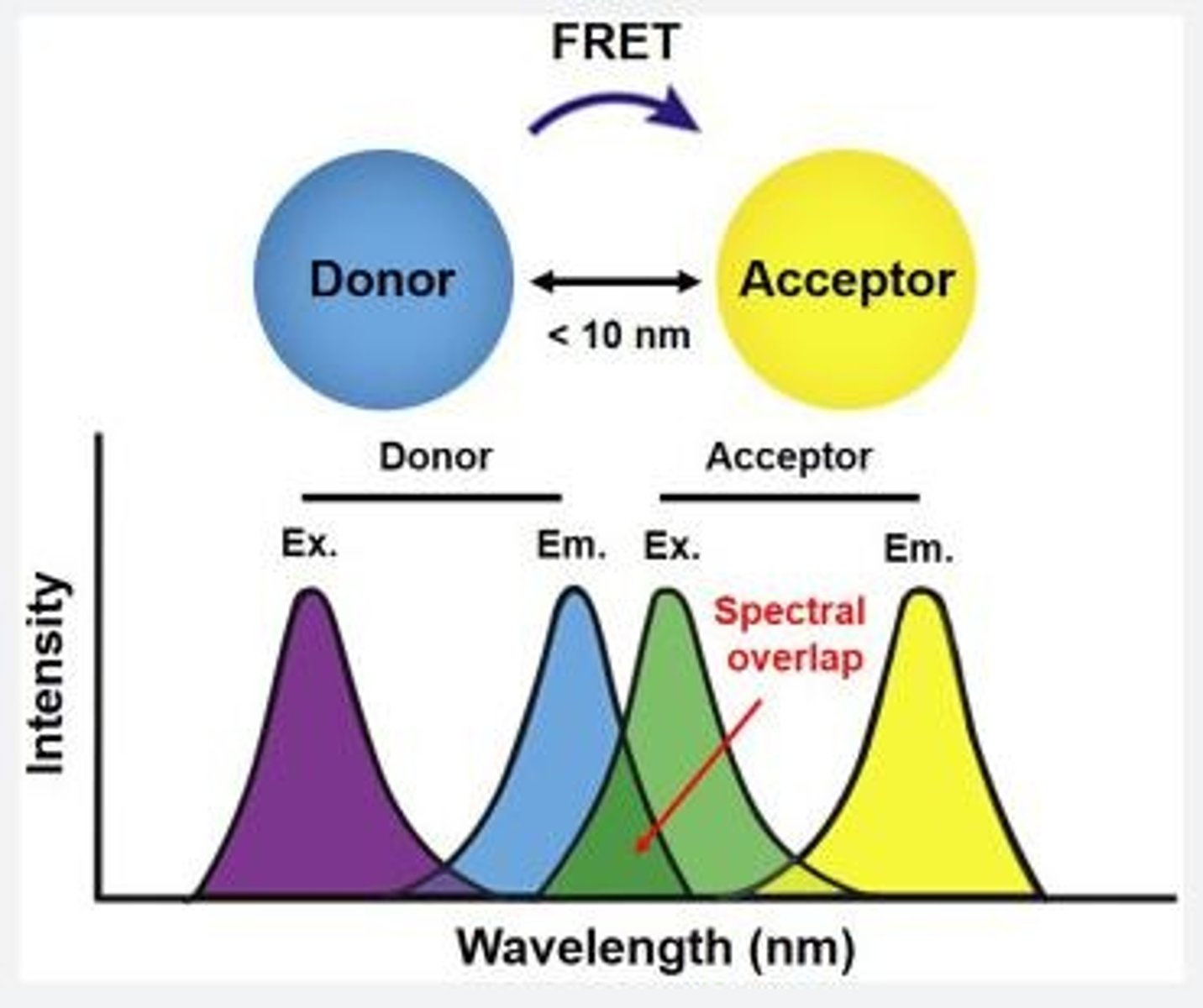
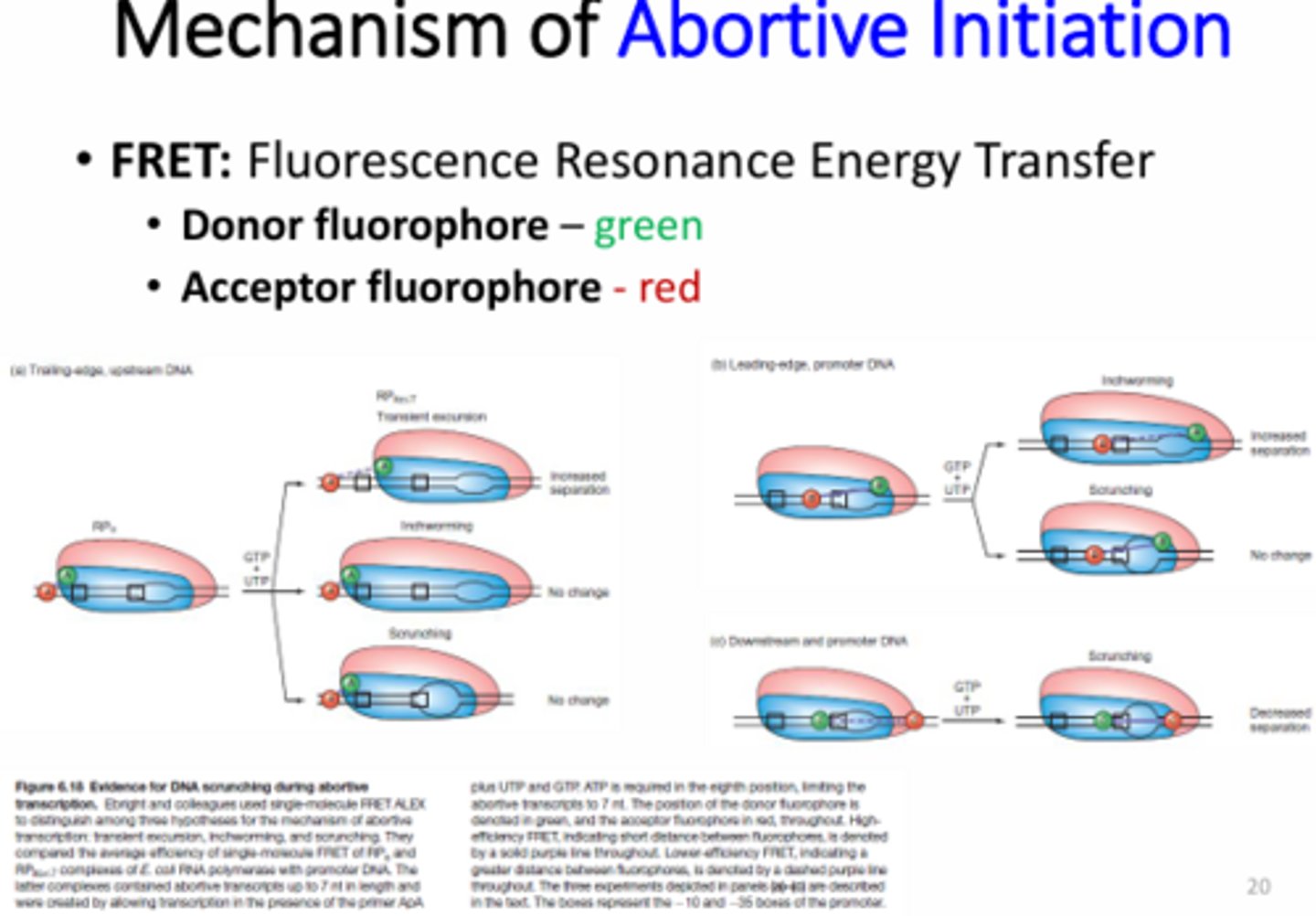
What is FRET?
Fluorescence Resonance Energy Transfer - uses the addition of two different fluorophores to identify a change in distance. The donor fluorophore will donate energy to the acceptor if they are in close contact, which results in an increase in fluorescence. In contrast, if the fluorophores are far away from each other there will be a decrease in fluorescence.
What type of promoter clearance was supported by FRET and why?
Scrunching - because no color change was observed when the fluorophores were on the back of RNA pol and upstream of the RNA pol binding site, no color chance was observed when they were moved to the front of sigma and between -35 and -10, and finally color change was observed when the fluorophore on sigma was moved downstream within the gene.
What are the reasons for RNA pol pausing?
1 - allows the translation to keep up with transcription
2 - this is the first step in stopping transcription
3 - allows for backtracking and fixing errors if needed
What are the reasons for RNA pol backtracking?
1 - low nucleotide concentration (unable to add new nucleotides)
2 - wrong nucleotide incorporation (GreA and GreB stimulate RNase activity to cleave off error - transcription is either resumed or terminated altogether)
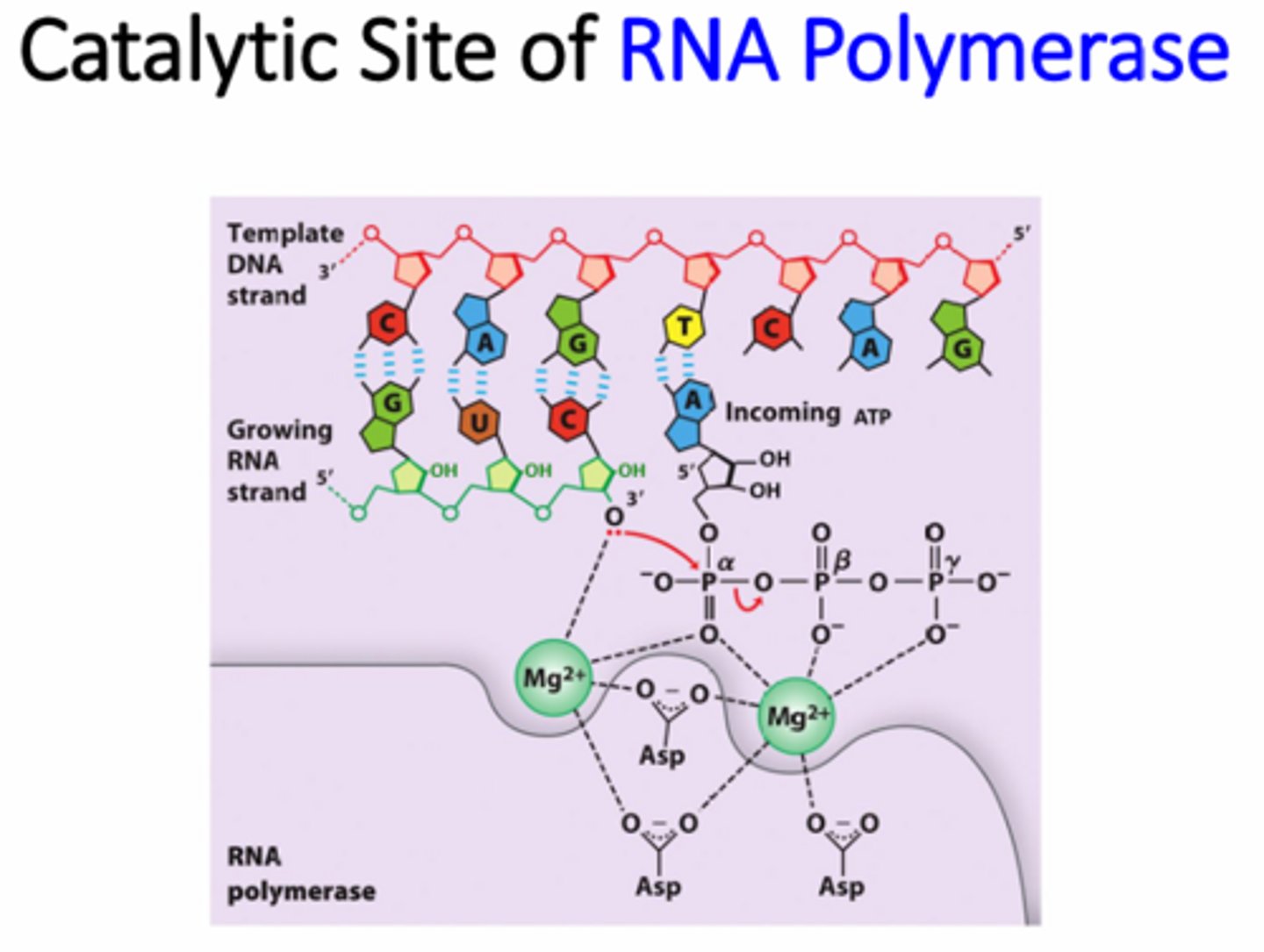
Why are magnesium ions necessary in the RNA pol complex?
Catalyzing the phosphodiester bonds between the nucleotides and stabilizing the phosphates
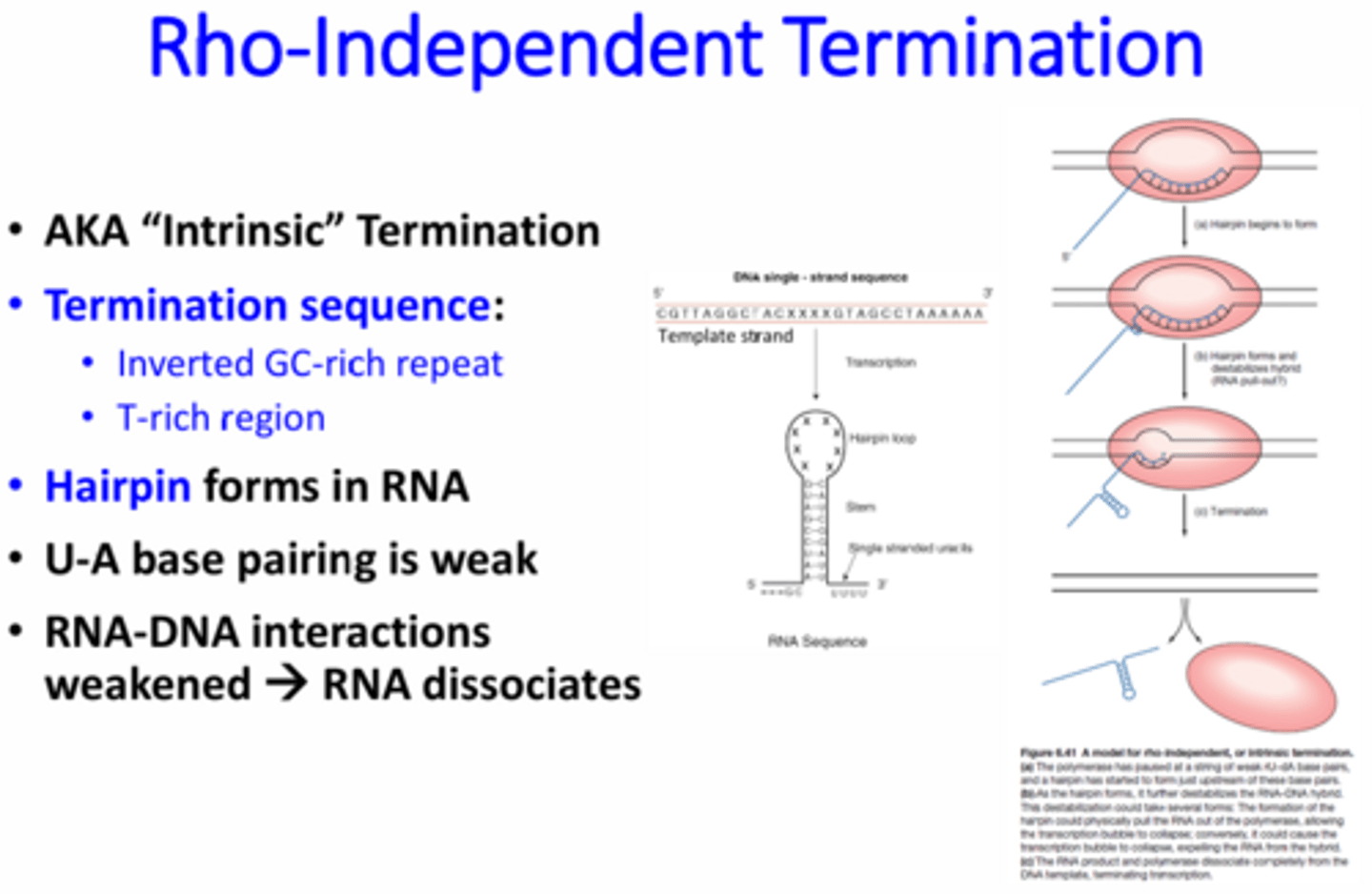
What is Rho-independent transcription termination?
Aka intrinsic termination, is the use of hairpin structures within the RNA for transcription termination - A termination sequence is transcribed that contains an inverted GC-rich repeat and T-rich regions. This forms a strong hairpin loop followed by a Poly-U tail. The bonds between the UA are much weaker than the GC bonds. The GC hairpin loop essentially pulls the Poly-U's off of the A's to remove the RNA from the DNA.
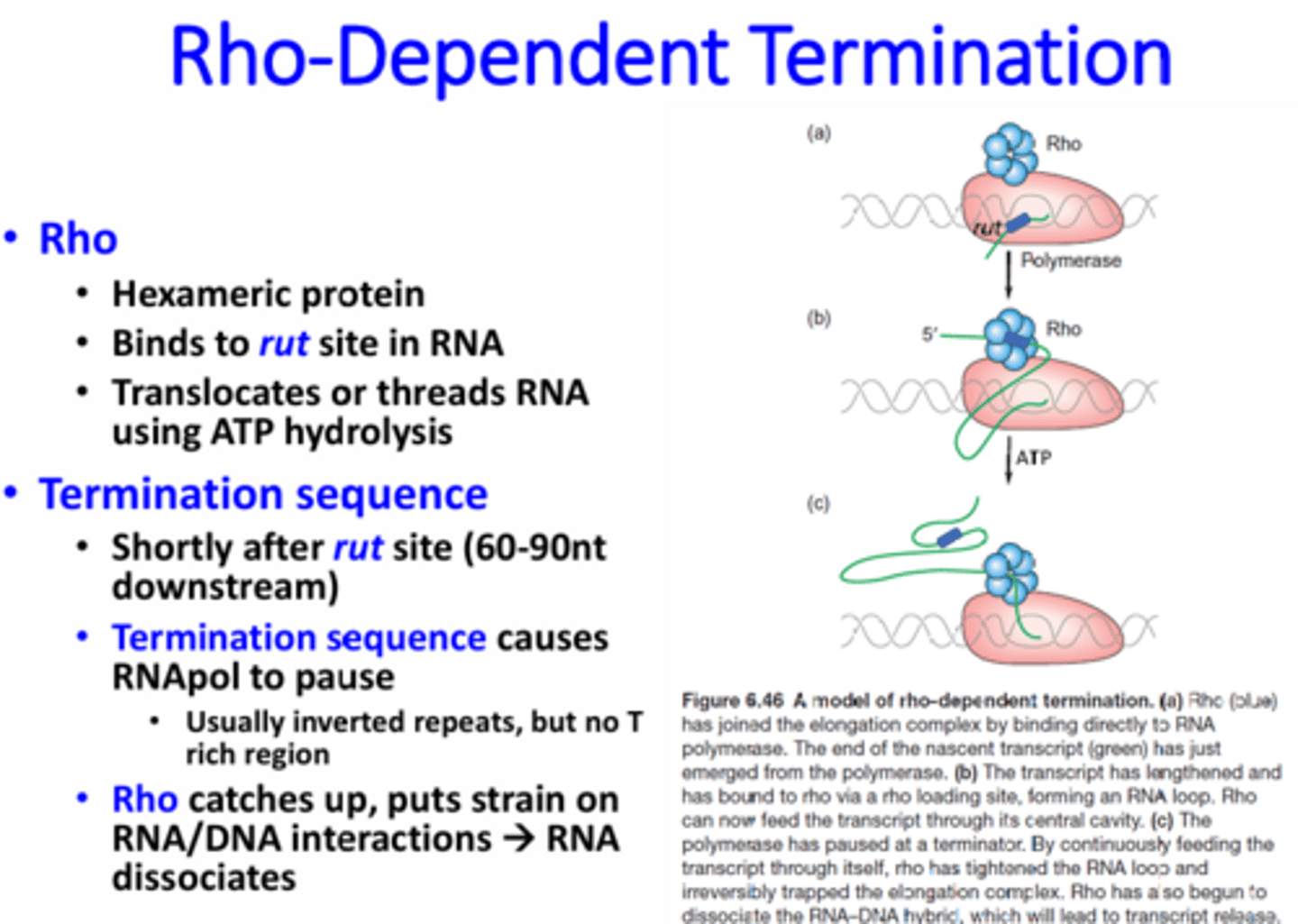
What is Rho-dependent transcription termination?
The use of Rho protein that binds to rut sites on RNA. After binding to the rut site, the protein travels down the RNA until it reaches a termination sequence where the RNApol is stalled. Since the RNApol is stopped, the Rho protein will catch up and put strain on the RNA-DNA bond eventually causing the RNA to be pulled off.
What is an operon?
A prokaryotic genetic element that contains several genes that are functionally related which are controlled by the same regulatory elements
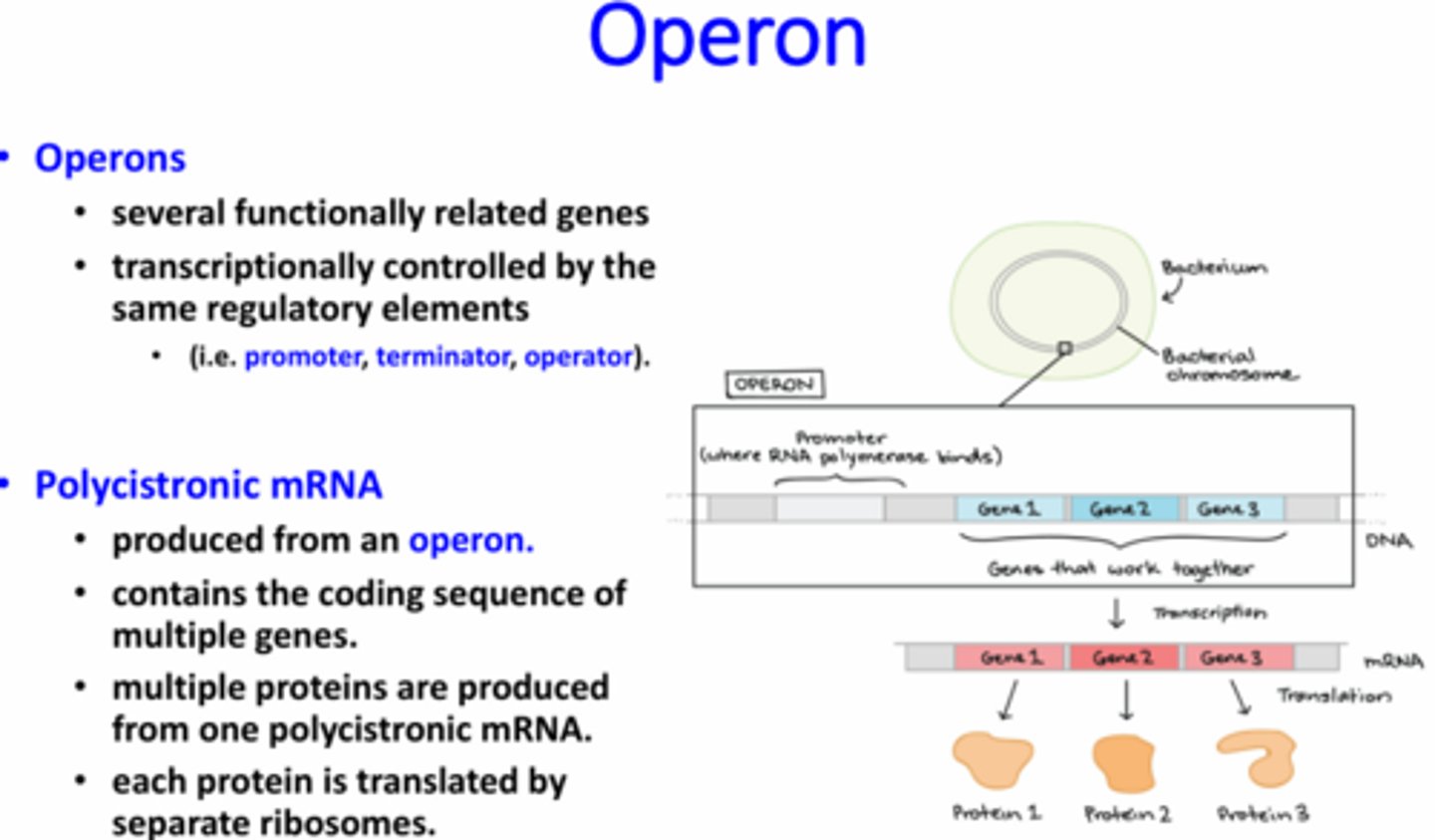
What is an operator?
A sequence of DNA (cis-acting) where a repressor protein binds (trans-acting) to reduce transcription rates
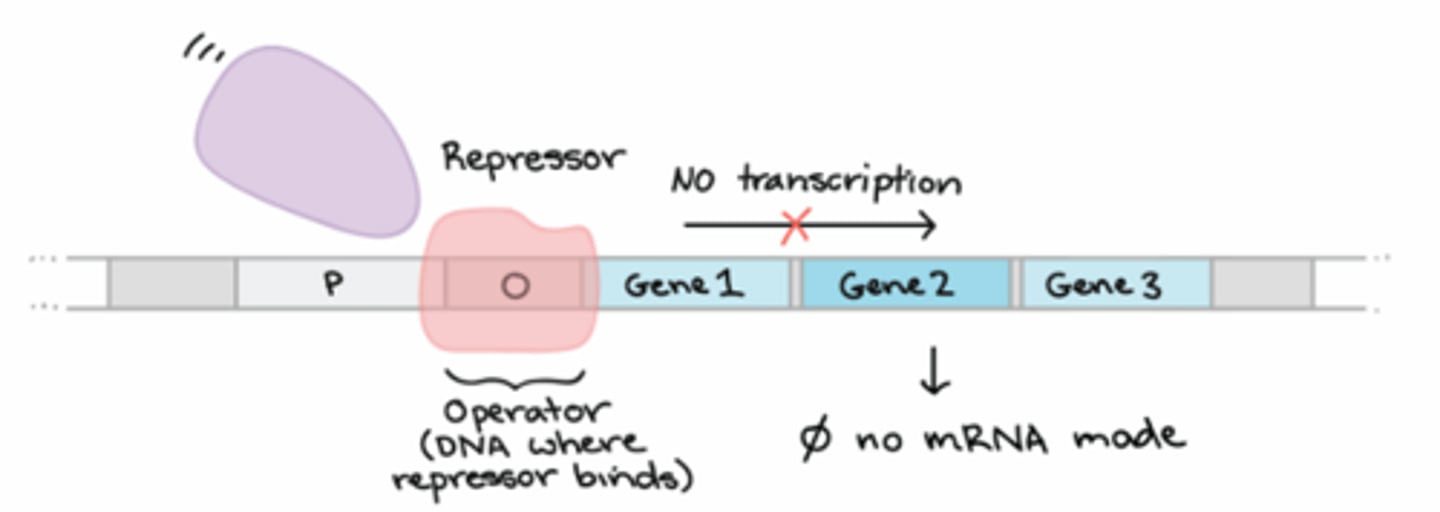
What is a repressor?
A trans-acting element that reduces transcription rates by binding to the operator in order to block RNA pol binding or moving past the promoter region (usually stimulated by an environmental cue like another small molecule presence or absence)
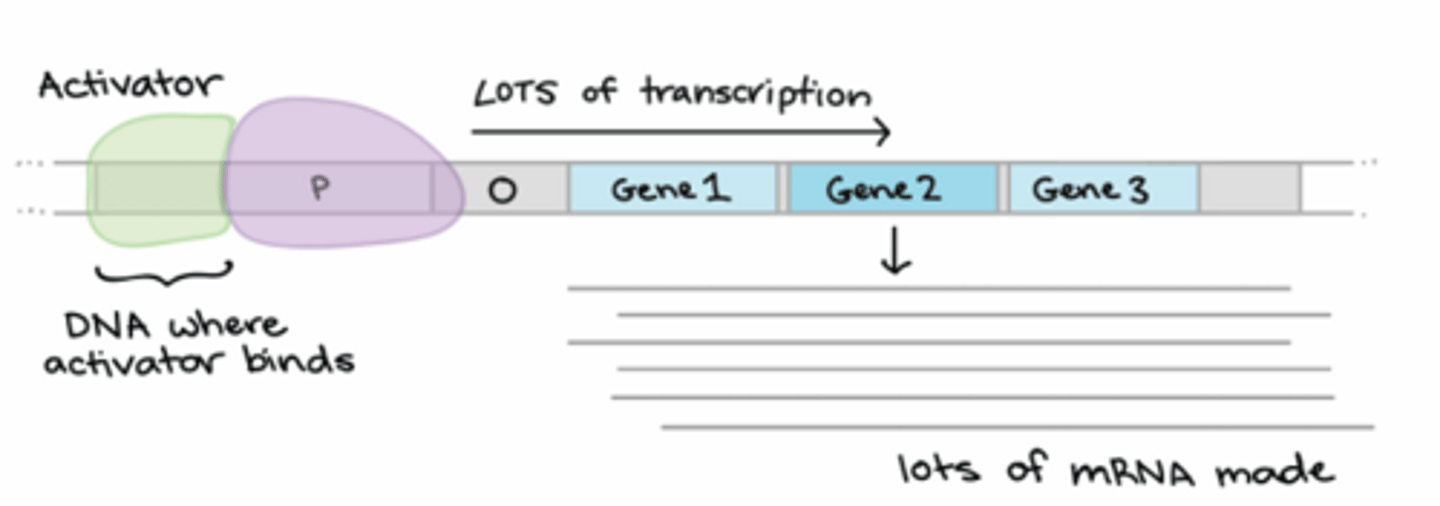
What is an activator?
A trans-acting element that increases transcription rates by helping to recruit RNA pol to the promoter. The bind to an activator-binding site (cis-acting element) upstream of the promoter
What is an inducer?
A trans-acting element that stimulates an activator or inactivates a repressor
What is a trans-acting factor?
A diffusible protein that can function at multiple sites in the genome (usually DNA binding proteins like transcription factors)
What is a cis-acting element?
Typically a DNA sequence that is closely tied to the gene
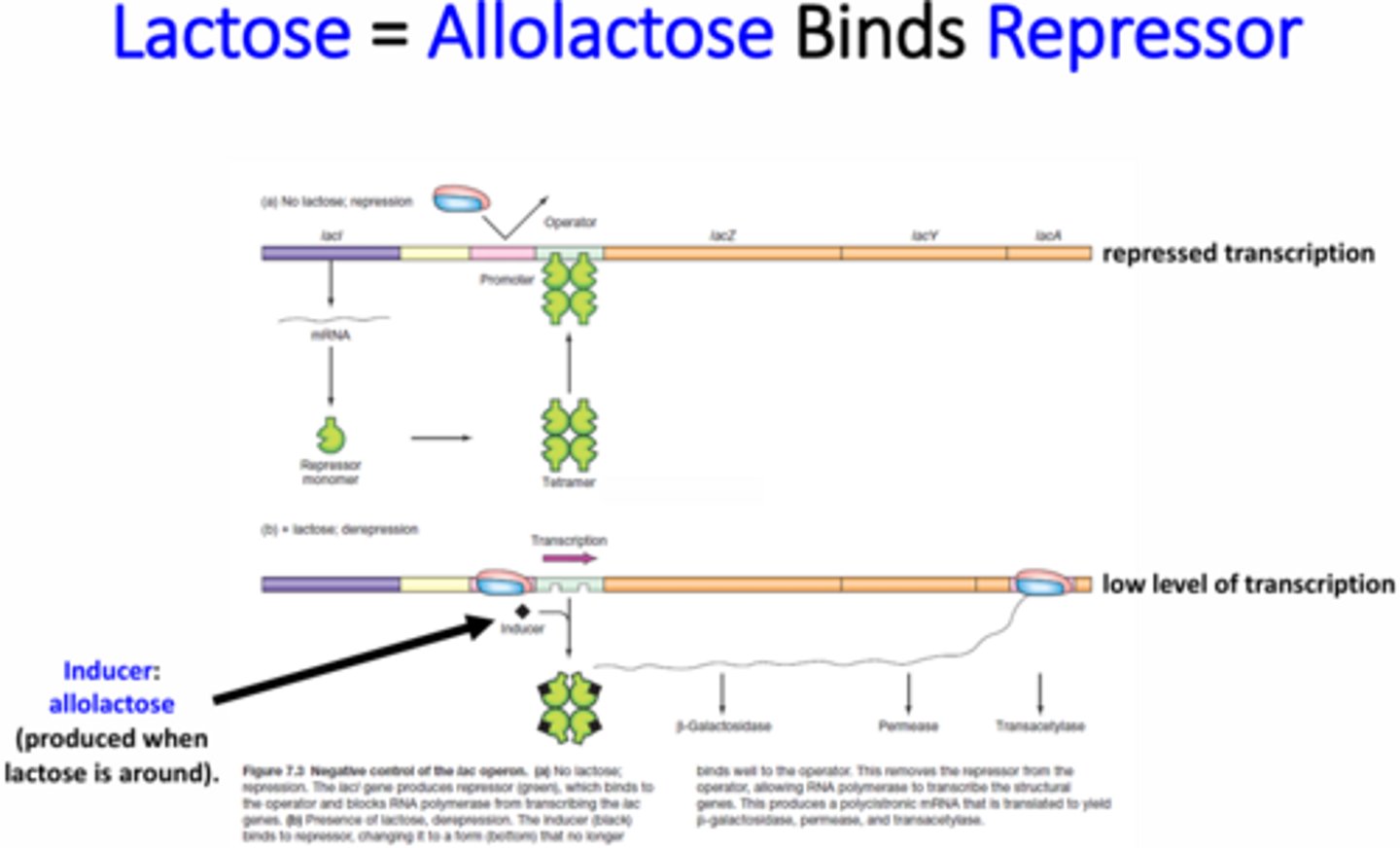
What is the Lac operon?
Encodes the proteins necessary for lactose breakdown (catabolic operon)
What is the structure of the Lac operon repressor?
Two dimers (aka tetramer) - one binds to the Operator 1 (O1) site and the other binds to the O2 or O3 site to create a loop that hides RNA pol binding site
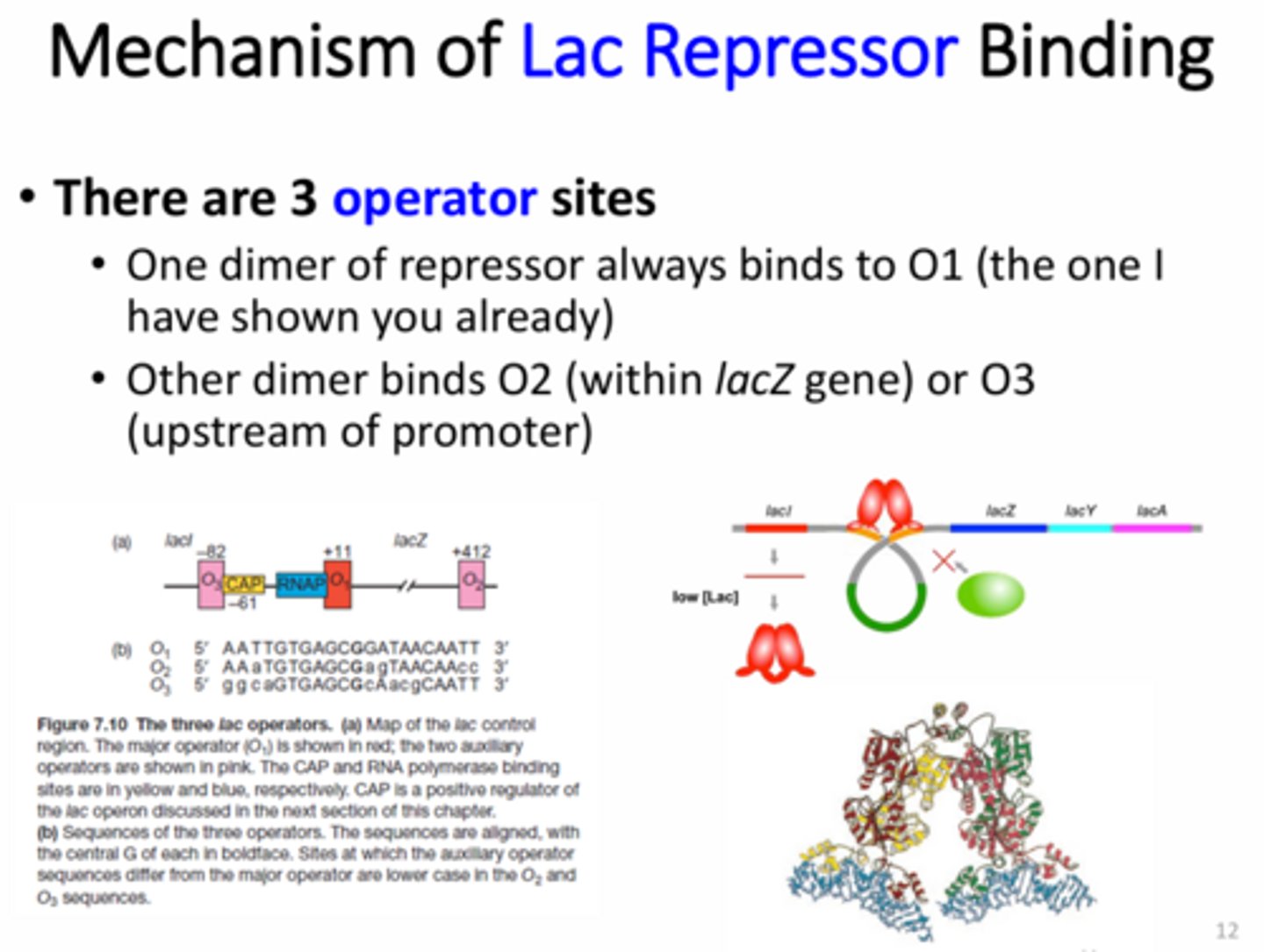
When is the Lac operon repressor active and how is it inactivated?
Active when there is no lactose present
Turned off by the inducer allolactose (a product of lactose breakdown)
What is the Lac operon activator and when is it active?
Catabolite activator protein (CAP) - aka cAMP receptor protein (CRP)
Active when there is low glucose levels
What is cAMP?
Cyclic adenosine monophosphate - produced by adenylate cyclase produces cAMP from ATP when glucose is low to signal the need for a carbon source
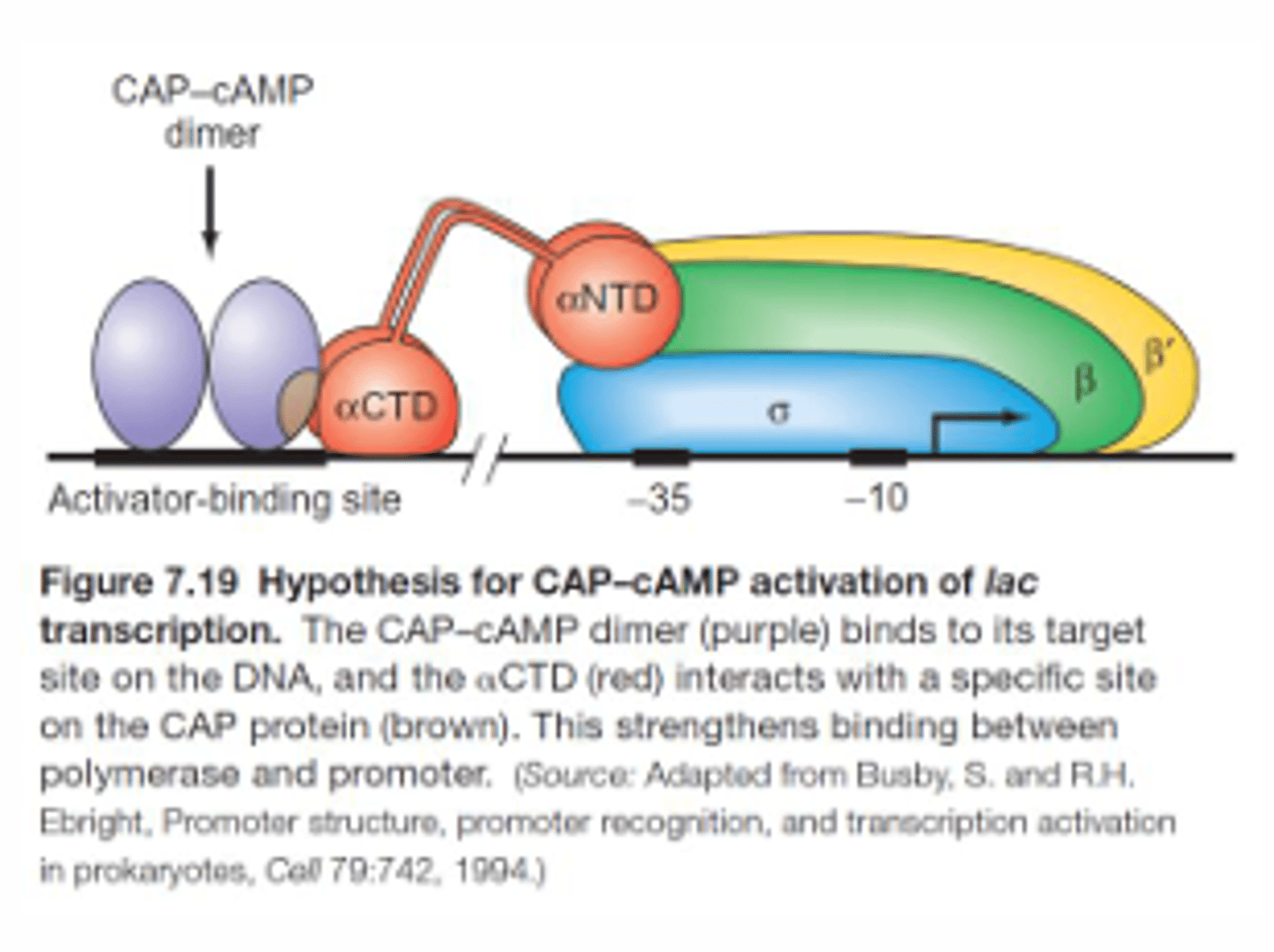
What is the mechanism of CAP binding?
cAMP induces a conformational change in CAP to increase CAP's affinity for the activator binding site. CAP then makes protein-protein interactions with the alpha-CTD subunit of RNA pol to increase transcription initiation by creating a more open complex formation
What are the Lac operon inducers
Allolactose - induces repressor inactivation
cAMP - stimulates activator
Why are RNA pols sensitive to Rifampicin and how does this relate to CAP activation?
Stops transcription by blocking the nucleotide channel at the very beginning of transcription
CAP helps to start transcription before Rifampicin can bind (blocked by the multiple phosphodiester bonds that have formed)
In the presence of both glucose and lactose, is the Lac operon transcribed?
Low level transcription -
Neither activator nor repressor
*the repressor is active but there is still lactose present which will lead to small amounts of allolactose production that can bind to the repressor and allow for transcription of the Lac operon. However, there is glucose present so there is no cAMP present to induce the activator action
In the presence of glucose but not lactose, is the Lac operon transcribed?
No (extremely low level) -
Repressor without activator
*the repressor is active and there is no lactose present to make allolactose which means the repressor is not removed so the only transcription that is occurring is accidental. Additionally, there is glucose present so there is no cAMP available to stimulate the activator for the Lac operon
In the presence of lactose but not glucose, is the Lac operon transcribed?
high level transcription -
Activator without repressor
*there is no glucose present so there is a high amount of cAMP present to stimulate the activator and there is lactose present to be made into allolactose to remove the repressor from the operator
In the absence of glucose and lactose, is the Lac operon transcribed?
No (extremely low level) -
Repressor active
there is no lactose present to form allolactose to remove the repressor so the only transcription that would take place is accidental (even though cAMP is present for CAP activation, it is still blocked by the repressor - the repressor is stronger than the activator)
Why is it good that the Lac operon promoter deviates from the consensus sigma-70 transcription?
This allows for varying levels of transcription based on the repressor and activators (presence/absence of glucose/lactose) rather than an all or nothing situation.
Why does the lac operon exhibit a very low (basal) level of transcription even in the absence of lactose?
The Lac repressor binds to the operator reversibly, allowing occasional transcription to occur if it dissociates (rare).
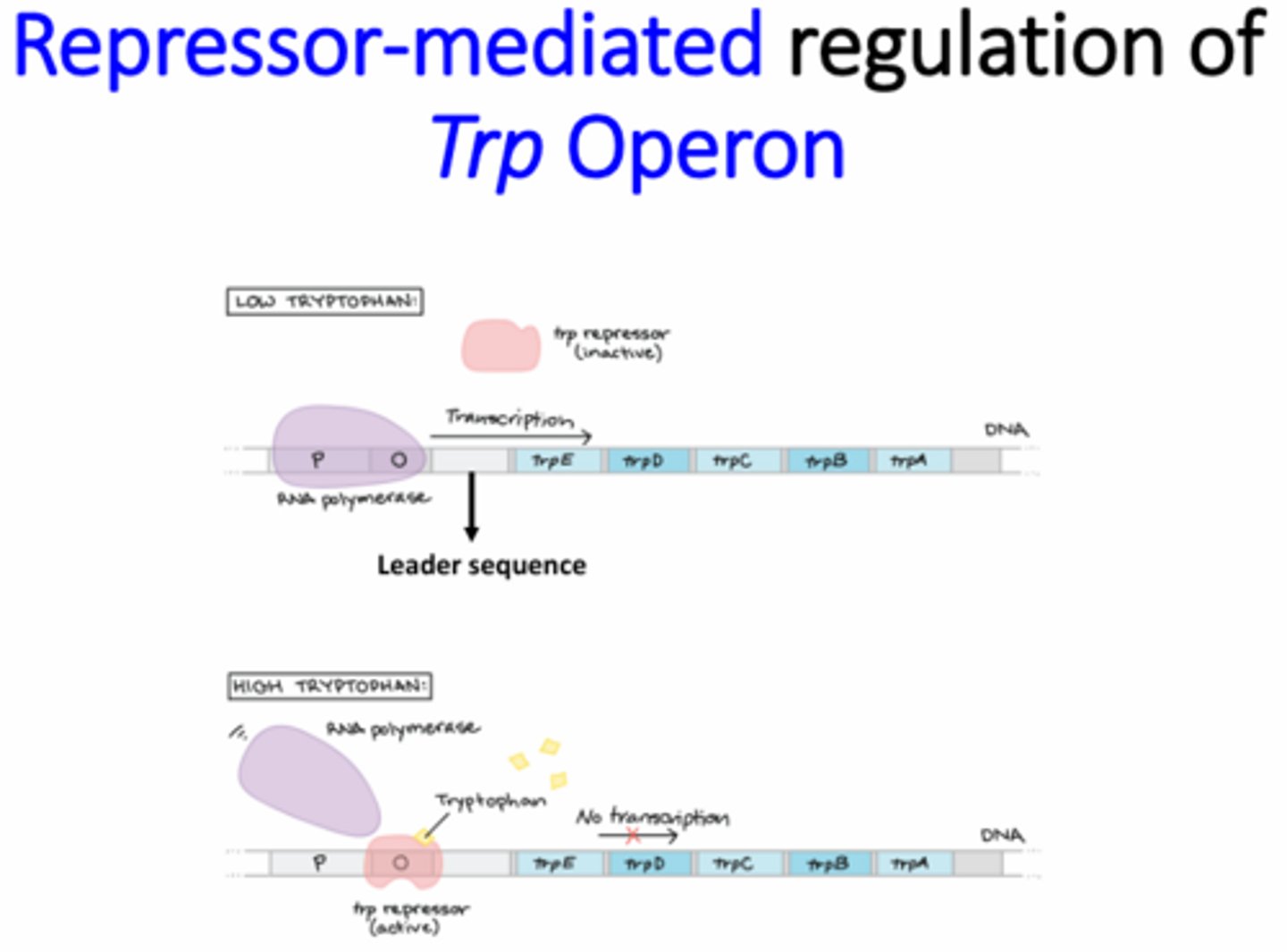
What is the Trp operon?
Encodes the proteins needed for tryptophan synthesis (anabolic operon)
What is the Trp operon inducer?
Tryptophan - activates the repressor
When is the Trp repressor active and how is it inactivated?
Active - High intracellular tryptophan levels
Inactivated by lack of tryptophan binding
What is attenuation in the context of the trp operon?
A mechanism of transcriptional control based on the fact that transcription and translation occur simultaneously. It helps to ensure that the operon is only active when tryptophan levels are low.
How does Trp operon attenuation work?
There are two tryptophan codons side-by-side in the first coding region of the mRNA. When tryptophan levels are low the ribosome stalls, which allows for alternate stem loop formation between regions 2 and 3. This conformation allows RNA pol to continue translating the Trp genes.
If tryptophan levels are high and are quickly added to the polypeptide during translation, the loop between regions 3 and 4 remain and act as a transcription termination signal.