Genetics Exam 2
1/47
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
48 Terms
Structural differences between RNA vs. DNA
RNA has ribose (with -OH) and Uracil (U); DNA has deoxyribose and Thymine (T)
General structure and secondary features of RNA
Usually single-stranded; can form functional secondary structures like hairpins or stem-loops
Functions of mRNA, tRNA, and rRNA
mRNA: Encodes proteins
tRNA: Adaptor between mRNA and amino acids
rRNA: Core of the ribosome
How do bacteria initiate transcription?
A holoenzyme is formed, composed of the Core enzyme + Sigma factor; the Sigma factor controls promoter binding to specific consensus sequences
Bacterial Promoter Consensus Sequences
The -10 Pribnow box and the -35 region
How Do Bacteria Terminate Transcription?
Rho-dependent: Uses Rho protein to unwind the DNA-RNA hybrid.
Rho-independent: Uses inverted repeats (hairpin) and a string of U nucleotides to destabilize pairing.
Eukaryotic Chromatin Modification for Transcription
Histones must be modified (e.g., by acetyltransferases) to "open" the DNA structure
Role of Eukaryotic RNA Polymerase II
Specifically synthesizes pre-mRNAs
How do eukaryotes initiate transcription?
TFIID (a general transcription factor) binds to the TATA box to bend and unwind the DNA
How do eukaryotes terminate transcription?
The torpedo method in which mRNA is cleaved and Rat1 exonuclease "chews up" the trailing RNA until it reaches the polymerase to stop it.
Mechanism of $\alpha$-amanitin (Death cap mushroom)
Specifically inhibits RNA Polymerase II, halting mRNA synthesis and causing cell death
Mechanism of Rifamycin (Tuberculosis antibiotic)
Specifically targets and inhibits bacterial RNA Polymerase
Differences between Exons and Introns
Exons are coding sequences that remain in the mature mRNA
Introns are noncoding sequences that are removed (common in eukaryotes, rare in bacteria)
Consensus sequences for ribosome recruitment: Bacteria vs. Eukaryotes
Bacteria: Shine-Dalgarno sequence (in 5’ UTR).
Eukaryotes: Kozak sequence (in 5’ UTR).
5’ Cap and 3’ Poly-A Tail
5’ Cap (guanine): Protection from degradation, increased stability, and initiation of translation.
Poly-A Tail (adenine): Facilitates nuclear export, increases stability, and helps ribosome attachment.
What is the Spliceosome?
A large complex of 5 snRNAs and proteins (snRNPs) that removes introns from pre-mRNA
Alternative Splicing vs. Multiple 3’ Cleavage Sites
Alternative Splicing: Splicing different combinations of exons to create different proteins from one gene.
Multiple 3’ Cleavage: Cleaving at different points to produce mRNAs of different lengths.
What is RNA Interference (RNAi)?
Eukaryotes use RNAi to defend against foreign genes (like viruses) and regulate their own gene expression
What is the mechanism of RNA Interference?
Dicer chops double-stranded RNA into small pieces; these pieces join Argonaute (AGO) proteins to form the RISC complex, which targets and silences specific mRNA
Gene “Knockdown” Using RNA Interference
Temporary decrease in gene expression.
miRNA vs. siRNA
miRNA (mine and multiple targets): Encoded by their own genes, come from a single RNA molecule forming a stem-loop, and regulate different genes
siRNA (stranger and same targets): Originates from a foreign organism (i.e., viruses), comes from long RNA duplexes, silences the same genes from which they originated
What are lncRNAs?
Long noncoding RNAs (>200 nucleotides) that do not make proteins but regulate gene expression (e.g., Xist for X-chromosome inactivation).
Basic Amino Acid Structure
A central carbon bonded to an Amino group (NH3+), a Carboxyl group (COO-), and a variable R (radical) group
Key Characteristics of the Genetic Code (3 Rules)
Triplet: 3 nucleotides = 1 codon (64 combinations).
Degenerate: Multiple codons can code for the same amino acid (synonymous codons).
Nonoverlapping: Each nucleotide is part of only one codon.
Stop Codons
UAG, UAA, and UGA. They do not encode amino acids and have no corresponding tRNAs.
What is tRNA Charging?
Attaching the correct amino acid to tRNA, catalyzed by Aminoacyl-tRNA Synthetases (requires ATP).
Initiation of Translation: Bacteria vs. Eukaryotes
Bacteria: Small subunit binds Shine-Dalgarno; initiator tRNA carries fMet.
Eukaryotes: Ribosome recruited by 5' cap/Poly-A tail; scans for AUG within the Kozak sequence.
The three ribosome sites (A, P, E) in order
A (Aminoacyl): Arrival of new tRNA.
P (Peptidyl): Where the peptide bond forms/protein is built.
E (Exit): Where the empty tRNA leaves.
Energy and Factors for Elongation of Translation
Requires GTP for energy and Elongation Factors (EF-Tu brings tRNA to A site; EF-G moves the ribosome/translocation).
Translation Termination Mechanism
A stop codon enters the A site; Release Factors (RF-1, 2, or 3) bind and trigger the cleavage/release of the polypeptide chain.
Nonsense-mediated mRNA decay (NMD)
Destroys mRNA with a Premature Stop Codon (PTC)
Bacterial vs. Eukaryotic stalled ribosome recovery
Bacteria: Use tmRNA to clear stalled ribosomes.
Eukaryotes: Use Nonstop decay (missing stop codon) or No-go decay (stalls from damage/structure).
Correct order of Prokaryotic Translation events
1. 30S initiation complex
2. fMet-tRNA (binding)
3. 70S initiation complex (assembly)
4. EF-Tu (delivery of tRNAs)
5. EF-G (translocation)
6. RF1 (termination)
General structure and characteristics of a Bacterial Operon
Includes structural genes, a promoter, and an operator; produces polycistronic mRNA (one mRNA for multiple proteins); contains no introns.
What is an operon?
Bacterial organization of related genes into a single transcriptional unit
Pros and Cons of organizing genes into Operons
Pros: High efficiency (fast on/off), coordinated control of pathways, energy conservation.
Cons: No precise control over individual genes, one mutation can block the whole pathway, vulnerable to gene movement.
Inducible vs. Repressible Operons
Inducible: Usually OFF; turned ON by an inducer that binds to the repressor on the operator, removing it and allowing transcription.
Repressible: Usually ON; turned OFF by a co-repressor that binds to the inactive repressor to block the operator
Functions of lacZ and lacY
lacZ: Encodes B-gal (cleaves lactose into glucose and galactose)
lacY: Encodes Permease (transports lactose into the cell)
Role of lacI and lacO
lacI: Regulatory gene encoding the repressor
lacO (Operator): Binding site for the repressor
What is the Inducer for the lac operon?
Allolactose (a lactose derivative); it binds and inactivates the repressor to allow transcription.
Positive Control: cAMP and CAP
When glucose is low, cAMP levels rise. cAMP binds CAP, which binds DNA to help RNA Polymerase bind and increase transcription.
What does a lacOc mutation signify?
A constitutive operator mutation; the repressor cannot bind, so the operon is structurally always "ON".
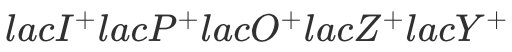
No Lactose: No B-gal or Permease
With Lactose: B-gal + Permease
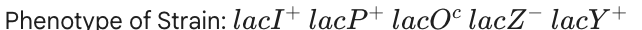
No Lactose: Permease only
With Lactose: Permease only
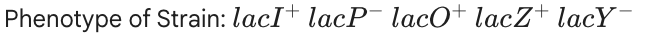
No Lactose: None.
With Lactose: None. The promoter is broken ($lacP^-$), so RNA Polymerase can never bind.
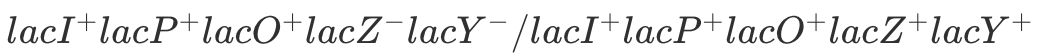
No Lactose: No B-gal or Permease.
With Lactose: B-gal + Permease. The second set of genes is perfectly normal and will function properly when lactose is present.