unit 5
1/56
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
57 Terms
mendelian inheritance
traits coded for by one gene w/ two alleles, where one allele is fully dominant over the other
law of segregation
genes on different chromosomes must segregate equally into gametes
each gamete receives one gene copy
homozygous
organism inherited two identical alleles at the same gene
heterozygous
organism inherited two different alleles at the same gene
genotype
organisms underlying genetic make-up consisting of both physically visible & non-expressed alleles
phenotype
the observable traits expressed by an organism
physical traits
monohybrid cross
two organisms differing in one trait
law of independent assortment
alleles for different traits are inherited independently of one and other during the formation of gametes
karyotype
visual rep of an organisms chromosomes
down syndrome
triploidy 21
triple X syndrome
XXX (three x chromosomes)
klinefelters syndrome
XXY
males are born with an extra X
turners syndrome
X
jacobs syndrome
XYY
males born with an extra Y
autosomal recessive inheritance pattern
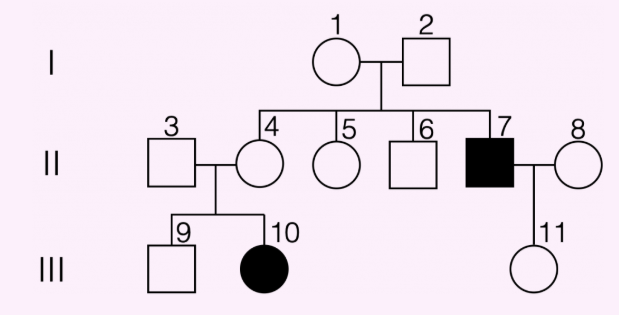
autosomal dominant inheritance pattern
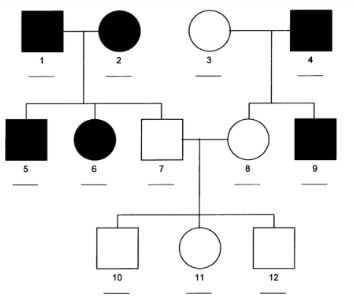
codominance
both alleles for the same characteristic are simultaneously expressed in the heterozygote
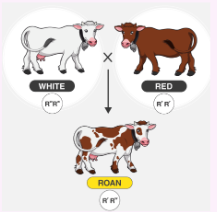
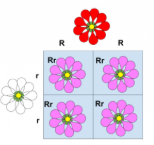
incomplete dominance
expression of two contrasting alleles such that the heterozygous individual displays an intermediate phenotype
blending of alleles is expressed
multiple alleles
single trait controlled by more than two alleles
ex- blood type (A, B, AB, O)
polygenetics
single trait is controlled by more than one gene
seen in traits with a “sliding scale” → hair color, eye color, weight, height
often influenced by environmental factors
epistasis
epistasis
one gene masks/ interferes with the expression of another
ex- black and brown mice
pleiotropy
expression of multiple traits by a single gene
occurs because genes code for proteins, and proteins themselves perform functions
non-nuclear inheritance
female gametes in animals and plants contain the majority of cellular component inherited by the zygote
this is because male gametes are are small and only contain nucleus & cytoplasm
mitochondrial & chloroplast DNA is
exclusively maternal
linked genes
genes inherited together because they are nearby and on the same chromosome
result: genes are rarely separated by crossing over & random assortment
do not follow expected ratios (alternative hypothesis)
recombination frequency of <50%
map distance
how close together 2 genes are, determined by crossing over frequency
ex- pair of genes have a recombination frequency of 10%, so they are 10 map units apart
sex-linked genes
traits determined by genes located on sex chromosomes
most of these traits are coded for on the X chromosome since its bigger
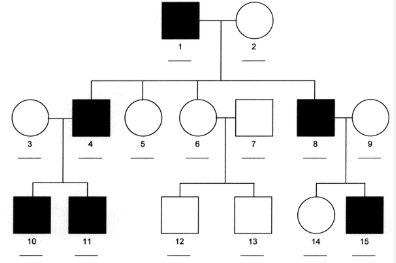
Y-linked chromosomes
all males of a family will express the trait, no females
X-linked genes (female)
operate the same way as Mendelian traits
carriers
carriers
heterozygous individual who carries a recessive gene but doesn't express it
X-linked genes (males)
will only express the dominant or recessive trait
cant be heterozygous
hemizygotes
hemizygote
having only one allele for a trait
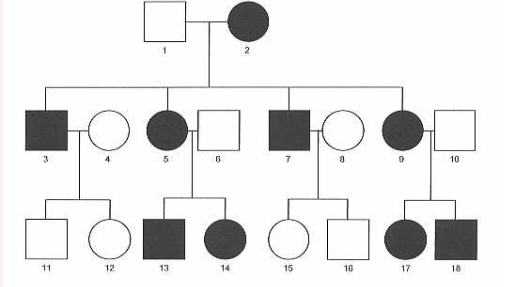
X-linked recessive
more males are effected than females, males cannot inherit the trait from their fathers
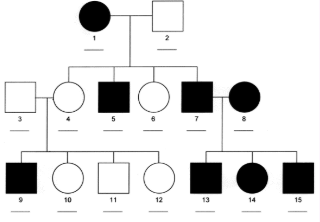
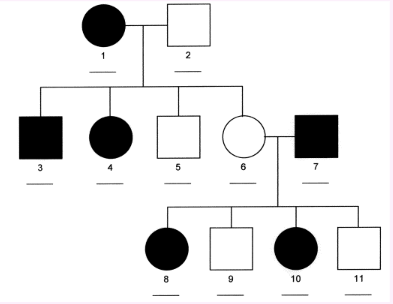
X-linked dominant
affected fathers pass on trait to all daughters
phenotypic plasticity
more than one phenotype can be expressed from one genotype, depending on gene expression & environmental conditions
prophase I
chromatin condenses into chromosomes
homologous pair up
nuclear envelope dissolves
spindle fibers form from centrosomes & attach to the kinetochores at the chromosomes
each homologous chromosome attached to fibers from opposite poles
homologous chromosomes pair up and undergo recombination
synapse allows this to happen
4 different chromosomes result
2 recombinant, 2 non-recombinant
locus (loci)
location of a gene on a chromosome
recombinant
recombination of genetic material
crossing over (recombination) happens at ___
chiasma/chiasmata
metaphase I
homologous pairs line up along the center of the cell
either chromosomes of the pair may face either side of the cell
this gives more genetic variation
anaphase I
spindle fibers pull homologous pairs apart
no longer identical sister chromatids stay tg
spindle fibers must pull each pair to opposite side & break the synapse
goes wrong = nondisjunction
nondisjunction pattern in anaphase I
n+1, n+1, n-1, n-1
nondisjunction
wrong number of chromosomes in each daughter cell
telophase I
meiotic spindle breaks down
nuclear envelope reappears to fully separate the homologous pairs
each nucleus contains a haploid # of chromosomes
chromosomes uncoil into chromatin
cytokinesis
cleavage furrow/ cell plate forms to full separate animal cells
meiosis I creates ___
2 haploid cells from 1 diploid cell
prophase II
chromatin condenses into chromosomes + nuclear envelope breaks down
new spindle fibers form
metaphase II
non-identical sister chromatids line up at the center of the cell
each chromatid is attached to a spindle fiber from opposite poles of the cell
anaphase II
non-identical sister chromatids are pulled to opposite poles of the cell be shortening spindle fibers
goes wrong = nondisjunction
telophase II
chromosomes uncoil into chromatid
nuclear envelope forms around each haploid set of single chromatid DNA
all genetically unique
meitotic machinery breaks down
cytokinesis II
4 genetically unique haploid cells are formed
null hypothesis
there is no relation/ difference between two groups of data
independent variable has no effect on dependent variable
alternative hypothesis
observed results are due to non-random cause
null hypothesis in genetic problems
there is no relation between genes (they are unlinked and randomly assorted)
rejecting the null hypothesis
when the critical value is lower than the chi-squared value
“if the p is low, reject that ho”
means that genes are linked
failing to reject the null hypothesis
when the critical value is higher than the chi-squared value
means that there is no significant difference in the data, results are due to chance
insufficient evidence to prove linkage
degrees of freedom
number of distinct possible values minus 1