exam 2
1/44
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
45 Terms
Use of recombinant DNA?
-determine protein funcation
-study genome organization
-genetically modify animals
Restriction enzyme
DNA endonuclease that recognizes specific DNA sequences and preform hydrolysis of the backbone
Which are prefered? Sticky or blunt ends
Sticky , Base pairing stabilizes interaction, makes it easier to ligate
how is recombinant DNA made
cloning vector-> restriction enzyme cleaves bond→ the gene of interst is inserted and ligated → recombinant DNA
whole process is also called molecular cloning
components of a cloning vector
Origin of replication: plasmid replicates inside bacteria, with this million of DNA is copied
Restriction enzyme: hold different cutting enzymes for DNA , where DNA of interest is inserted
Antibiotic resistance(selection method): DNA is grown on antibiotic medium, only one w/ plasmid survive and grow
each bacterium takes up __ recombinant DNA molecule to construct a genomic library
one
cDNA process
lyse tissue or cultured cells and purify MRNA
poly T primer is added
DNA copy gets made with reverse transcriptase ( DNA/RNA double helix)
RNA partially degraded withe RNase
DNA pol synthesizes a new strand of DNA to join
Whats type of library is used for eukaryotic protien in bacteria
cDNA library, cDNA is already processed, the presence of other components would make it difficult for bacteria to express protein
Nucleic acid hybridization
used to detect specfic DNA/RNA sequence
Denature DNA
Heat → double-stranded DNA separates
Add probe
Short labeled DNA/RNA sequence (complementary)
Annealing (hybridization)
Probe binds ONLY to matching sequence
Detection
Fluorescent/radioactive signal shows where binding occurr
High temp → only perfect matches bind
Low temp → mismatches can bind
In situ hybridization tell you,,,
where gene/mRNA is
RNA- seq tells you,,,
how much gene expression
reporter gene tell you,,,
when/where gene is active
GFP fusion tell you,,,
where protein goes (intercellular location)
shotgun approach
used to determine nucleotide sequence of entire genome
repetive sequences created problems
process: multiplie copies of genome → random breakage → a common end and start is found -> those strands are assembled
SNPs
single difference in nucleotide
1 in every 1000bp
two unrelated individuals maybe have 3×10^6 ( 3 million) nucleotide differences in their genomes
Transgenic model organisums can be used to study gene function, what are the types?
Gene replacement:Replacing a normal gene with a modified version (mutated, tagged, or altered) , used when studying effects of specific sequences ex 5bp amino acid
gene knock out: Completely inactivating (removing) a gene, tests what happens when a entire protein is not active
gene addition: Adding an extra copy or expressing a gene in a new context
You want to know which genes are actively expressed in liver cells but not brain cells. What do you use?
cDNA library or RNA-seq → only expressed genes (from mRNA) are analyzed.
what is cDNA vs Genome ibrary
Genome library : DNA fragments that represent the entire genome
cDNA:represents specific genes at specific time
You want to study regulatory regions controlling gene expression. Genomic or cDNA library? Why?
Genomic library → contains promoters and regulatory DNA (cDNA does not)
You want to visualize where a gene is expressed in an embryo. What method?
In situ hybridization
You want to determine if two DNA sequences are complementary. What principle is used?
Nucleic acid hybridization.
Why is antibiotic resistance essential in cloning vectors?
To select only bacteria that took up the plasmid
After 10 PCR cycles, how many copies theoretically exist?
2¹⁰ = 1024 copies
Why do ddNTPs terminate DNA synthesis?
No 3’ OH → no elongation
Why is NGS faster than Sanger sequencing?
Parallel sequencing of millions of fragments.But due ti shorter reads their high error
You discover a new protein but don’t know its function. What is an approach?
gene knockout ( remove gene and visualize changes) , localization (where does this act)
functions of plasma membrane
receiving info, import and export of small molecules capapctiy for movement and expansion
proteins constitute _% of plasma membrane mass
50
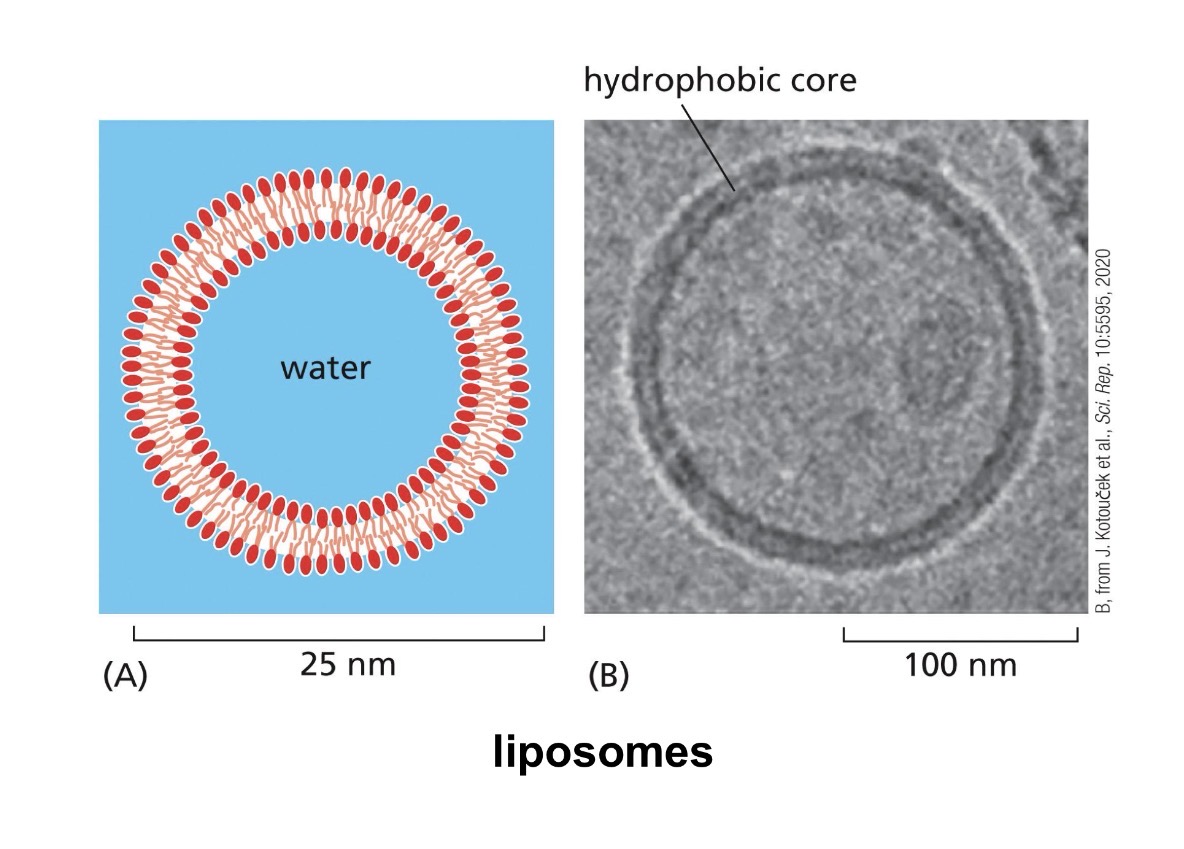
lipid bilayer
main component of membranes
consists of phospholipids
hydrophilic head, hydrophobic tale
double bond in tail created kink
ampihatic (hydrophillic & hydrophobic )
the lipid bilayer is the most…
energetically favorable way for phospholipids to exist in water
heads maintain contact w/ water while tails and toward eachother and stay dry
how is membrane fluidity demonstrated?
fusion of cells → incubation→ with time the lipids mix and have lateral membrane fluidity
How is the rate of diffusion measured in cells?
FRAP: fluorescence recovery after photo bleaching
light shines of section of cell causing photo bleaching
use recovery of color to make conclusions on how fast cells are moving
why is membrane fluidity important?
allows interactions between protiens
membrane fusion and budding
membrane repair
etc
membrane fluidity regulated
length of hydrocarbon tails (longer tails make a more rigid membrane)
degree of saturation ( saturated fatty tails make a more rigid membrane)
cholesterol (in animals cells): more rigid membrane
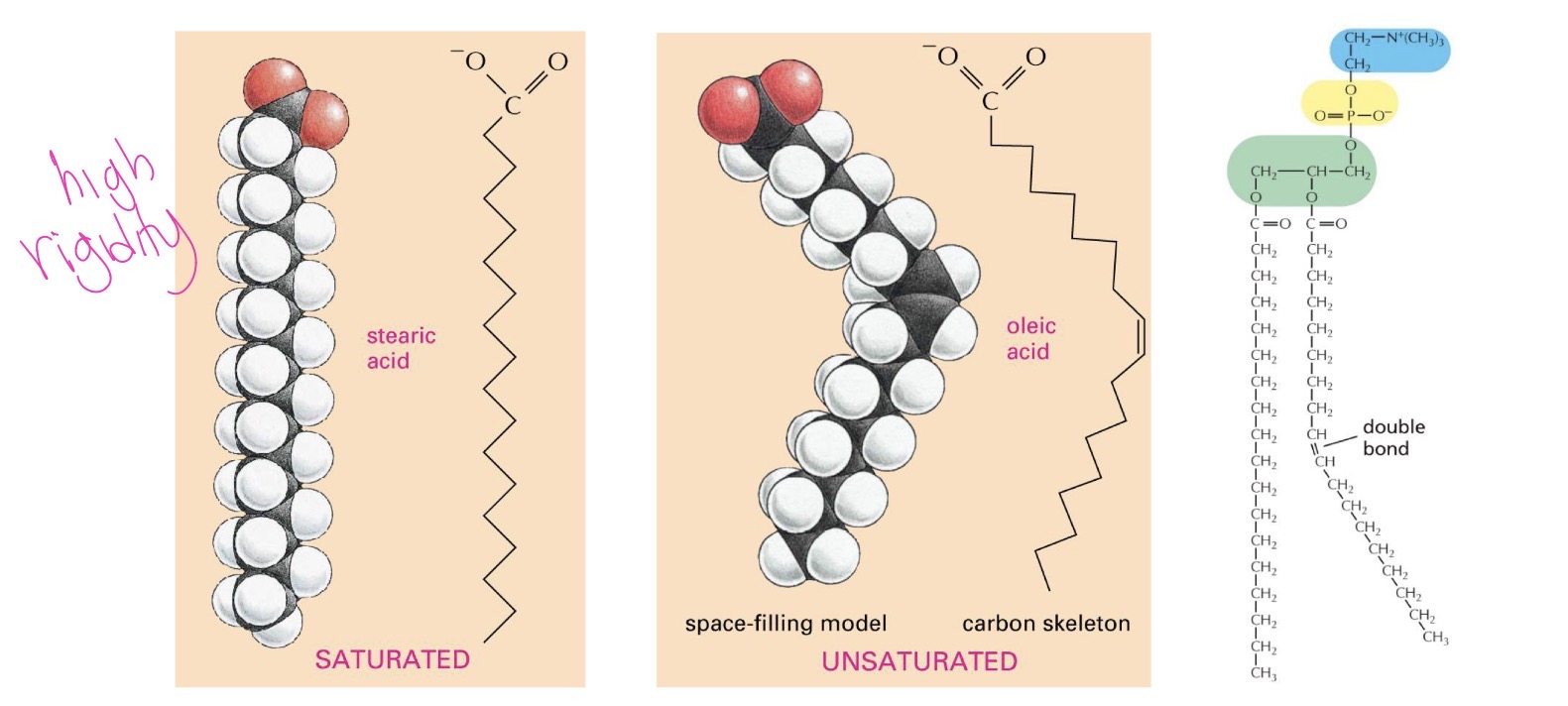
fatty acids can be…
saturated or unsaturated
saturadted has no double bonds between hydrocarbons, have max hydrogens, increase rigidity
unsaturated at least one double bond, reduce rigidity
enzymes of the ER membrane
preform scrambles
distributes phospholipids randomly
randomly redistributes phospholipids between both leaflets
enzymes of the golgi apparatus
preforms flippase
distributes phospholipids aymmetric
requires ATP
selectively moves specific phospholipids from the outer leaflet to the inner leaflet
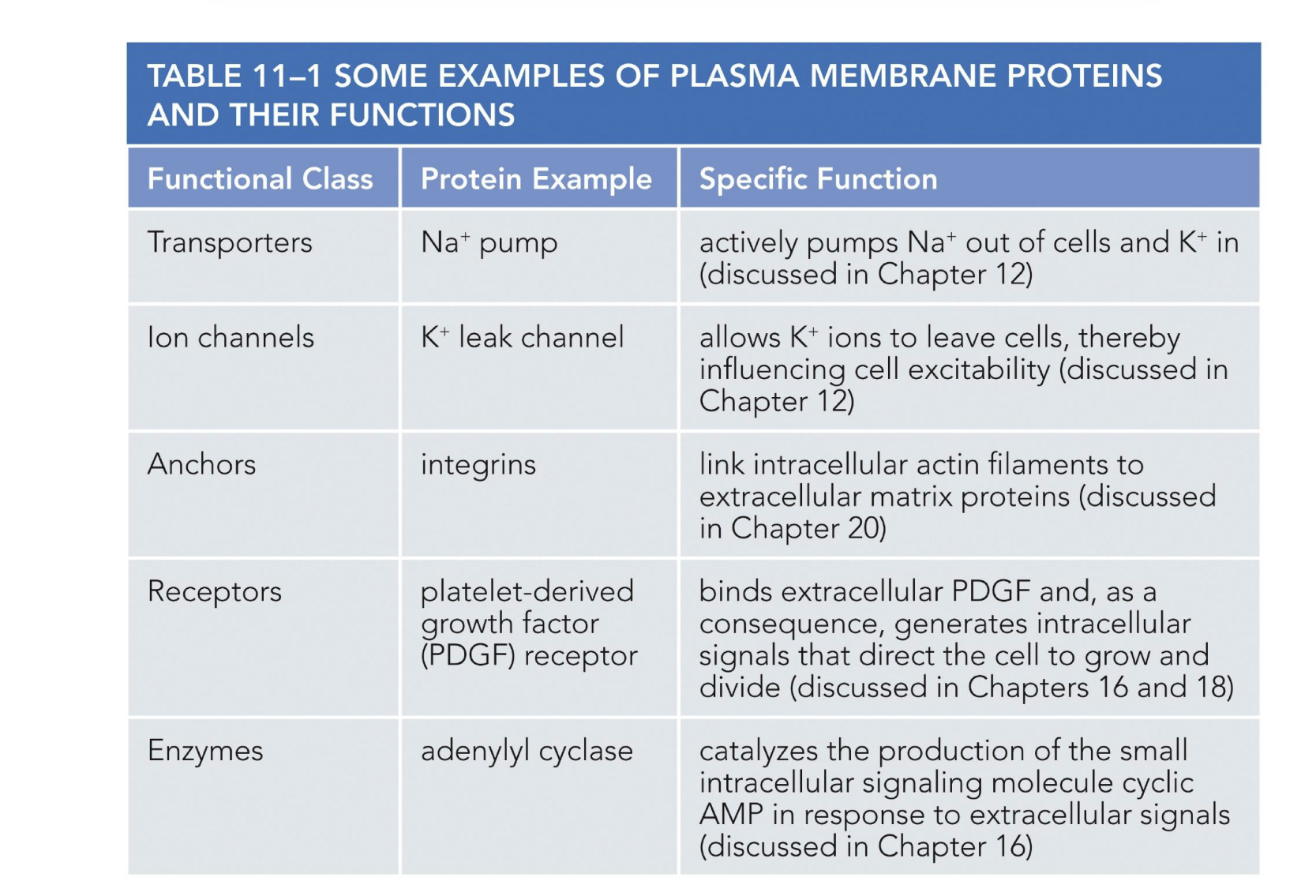
types of membrane protiens
transporters and channels
anchors
receptors
enzymes

transmembrane regions are formed by…
short a-helices
single & multi pass
proteins pass about 11-13 times
multi-pass transmembrane proteins can form
aqueous pores to allow passage of water soluble molecule across lipid bilayer
hydrophilic side chains form aqueous pore
hydrophobic side chains interact with phospholipid tails
eukaryotic cells are coated with sugars, why?
provide cell movement and recognition, creates proper signals for cells to go through
Why do phospholipids spontaneously form bilayers instead of random aggregates?
Because it minimizes free energy:
Hydrophobic tails avoid water (hydrophobic effect)
Hydrophilic heads interact with water
Bilayer eliminates exposed hydrophobic edges → more stable than micelles for phospholipids
Sealing into vesicles (liposomes) removes edge instability completely
Why do flat lipid bilayers tend to form closed vesicles?
hydrophobic tails are exposed to water→ energetically unfavored
Membrane bends to eliminate edges → Forms sealed compartments
If fluorescence does NOT recover after photobleaching in a FRAP experiment, what does that suggest?
Proteins/lipids are not mobile
Membrane may be:
Anchored to cytoskeleton
Highly rigid (e.g., lots of cholesterol or saturated fats)
Proteins confined to domains
👉 FRAP = measures lateral diffusion
Why are β-barrel proteins common in bacterial outer membranes?
Form rigid, stable pores
Allow passive diffusion of small molecules
Ideal for transport across outer membrane