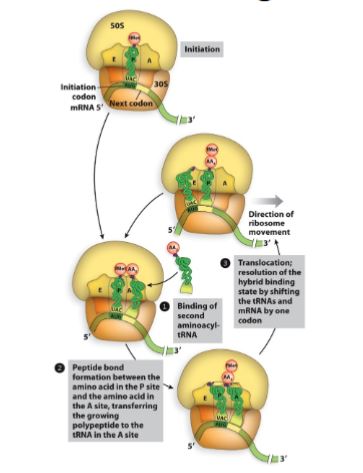
Molecular bio translation
1/62
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
63 Terms
Why is translation the most important step of the central dogma?
It produces proteins, which carry out all cellular functions.
Functions of proteins:
Catalytic
Transport
Structural
Regulatory
Why would a mutation in DNA polymerase be catastrophic?
Because proteins are required to execute DNA replication.
What are the 4 primary components of protein synthesis?
Ribosome (machinery)
mRNA (message)
tRNA (adapter)
Aminoacyl-tRNA synthetases (decision maker)
Which component did she explicitly call the most important?
Aminoacyl-tRNA synthetases.
What do aminoacyl-tRNA synthetases do?
They link the correct amino acid to the correct tRNA.
Why are aminoacyl-tRNA synthetases critical for accuracy?
They determine which amino acid corresponds to each codon.
Does the ribosome check if the correct amino acid is attached?
No
Where is translation fidelity primaryily determined?
Aminoacyl-tRNA synthetases.
Why is it significant that proteins make up 44 percent of the dry weight of a human body?
There is a huge investment in making proteins.
Why is protein synthesis considered complex?
Requires hundreds of proteins and RNAs.
What does “linear march” refer to?
Sequential addition of amino acids during translation.
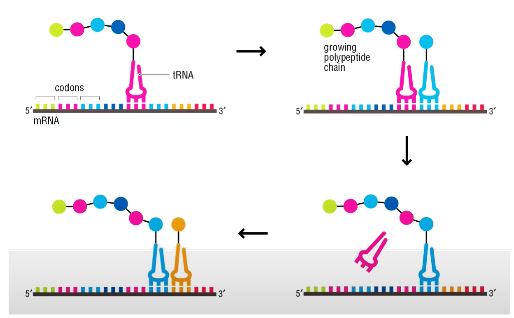
Is the genetic code overlapping?
No.
Directionality of protein?
It goes from the N-terminus → C-terminus, not 5’ → 3’ like DNA. Met is on N-terminus (at the top of the chain, amino acids are being added on the bottom (C-terminus))
What happens to the internal amino groups?
They form peptide bonds.
What are the components of translation?
The genetic code
mRNA
tRNA (aminoacyl-tRNA)
rRNA ( and like alot of translational factors)
Phases of translation?
Initiation (identifying the start site)
Elongation (speed vs accuracy) (the balance check step)
Termination
Recycling
Why can DNA replication proofread but translation cannot?
DNA uses base pairing template; translation does not.
Does translation have proofreading?
Very limited.
What is the start codon?
AUG.
What are the stop codons?
UAA
UAG
UGA
Which codon position allows wobble?
The third, the first two are very important.
think of the chair analogy → you can still sit on a chair with one broken leg, the third codon is not as important.
tRNA structure.
Is a cloverleaf structure:
Anticodon reads the code (mRNA)
3’ CCA tail carries the amino acid
Amino acids are attached here by aminoacyl-tRNA synthetase
T-loop and D-loop ensures stability and structure (heavily modified)
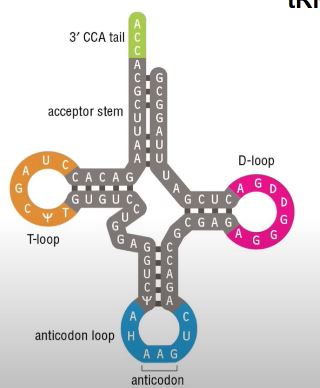
What modification did she mention (that modifies the T and D-loop)?
Pseudouridine, allows for stability.
What sequence orients mRNA in prokaryotes?
Shine-Dalgarno sequence. This is exclusive to only prokaryotes, eukaryotes do not have this.
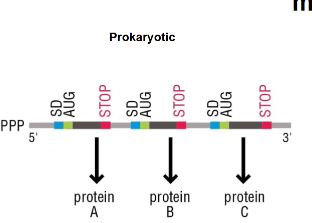
What is a ribosome?
A macromolecular machine responsible for protein synthesis.
Where does transcription and translation occur in eukaryotes?
Transcription occurs in the nucleus
Translation occurs in the cytoplasm
What are the three functions of a ribosome?
Bind rRNA and identify the start codon (where translation begins)
Facilitate complementary base pairing of mRNA codons and tRNA anticodons
Catalyze peptide bond formation
Prokaryotic ribosome size?
70s
Large subunit: 50S
Small subunit:30S
%rRNA vs protein in bacterial ribosomes?
60 percent rRNA, 40 percent protein.
Eukaryotic ribosome size
80S
large subunit: 60S
small subunit: 40S
Key difference between bacteria and eukaryotic ribosome?
Eukaryotic ribosomes are more complex.
How does complexity increase?
It increases as we evolve = the more RNA and protein complexes → the higher the complexity.
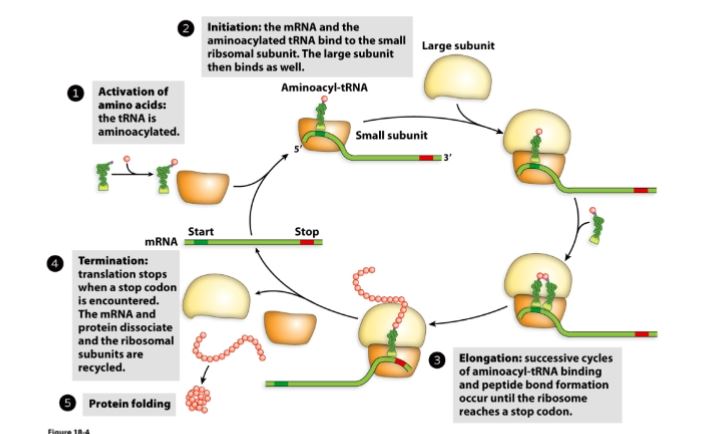
Association and Dissociation of Ribosomal Subunits
Activation of amino acid: amino acid attacks ATP → releases inorganic pyrophosphate (PPi) → forms aminoacyl-AMP. (this process is called adenylylation)
tRNA charging (transfer to tRNA): aminoacyl-AMP is transferred to 3’ CCA tail of tRNA forming aminoacyl-tRNA (charged tRNA).
Initiation: mRNA and aminoacylated-tRNA bind to the small subunit (FIRST), large subunit binds after (SECOND).
Elongation: successive cycles of aminoacyl-tRNA binding and peptide bond formation occur (until they reach the stop codon)
Termination: Stop codon is encountered and translation stops. mRNA and protein dissociate, the ribosomal subunits are recycled.
Protein folding (hydrophobic collapse).
Ribozyme experiment.
They degraded and denatured the protein, yet they still saw synthesis. This indicated that RNA was the one carrying out the catalytic function.
What catalyzes peptide bond formation?
rRNA.
What is a ribozyme?
RNA with catalytic activity.
The catalyst is RNA itself, not a protein.
What does puromycin do?
Inhibits translation
mimics the 3’ end of aminoacylated tRNA
binds the ribosome and causes chain termination
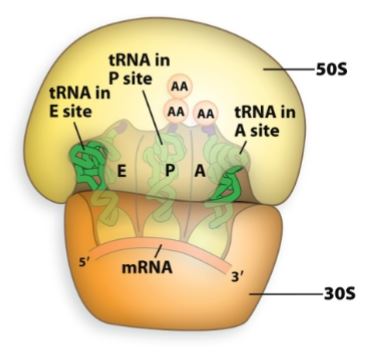
Aminoacyl site (A-Site)
Binds a new tRNA molecule containing an amino acid (aminoacyl-tRNA) to be added to the growing polypeptide chain.
enters and brings in correct amino acid.
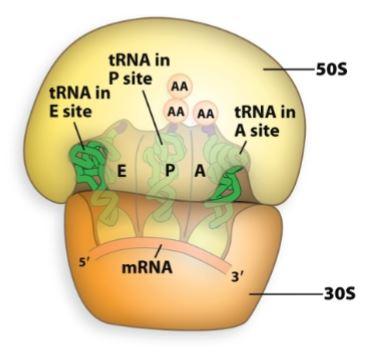
Peptidyl site (P-site).
Holds the tRNA to which the polypeptide is attached.
holds the growing polypeptide chain.
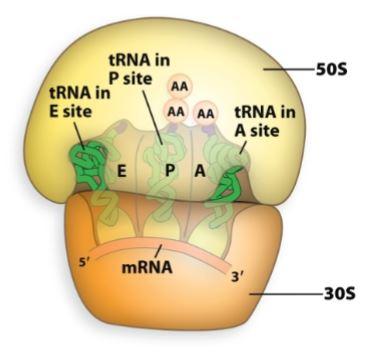
Exit site (E-site).
Provides an avenue for the exit of the tRNA after its amino acid had been added to the chain.
tRNA exits.
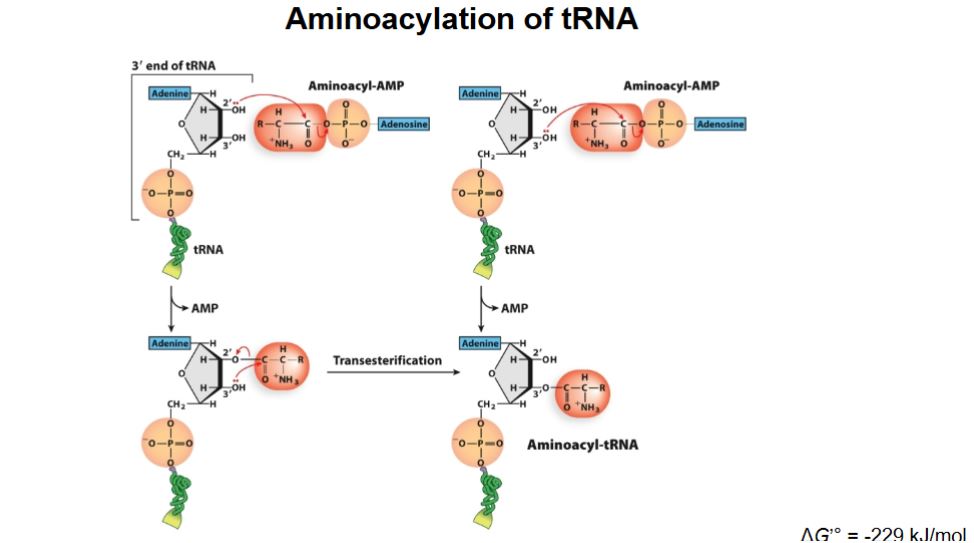
Describe Aminoacylation of tRNA
Adenylylation: Amino acid + ATP → Aminoacyl-AMP + PPi
Transfer of Aminoacyl-AMP to tRNA: Aminoacyl-AMP transfers amino acid to 3’ end of tRNA (3’ CCA tail), then briefly separates. Aminoacyl group reconnects with the amino acid and loses AMP → becoming aminoacyl-tRNA. (Class I amino acids have an extra step → transesterification)
This happens before translation and uses energy (ATP).
There is a specific aminoacyl-tRNA synthetase for each amino acid and its matching tRNA.
True.
What is the “second genetic code”?
The molecular recognition between a specific tRNA by a specific synthetase.
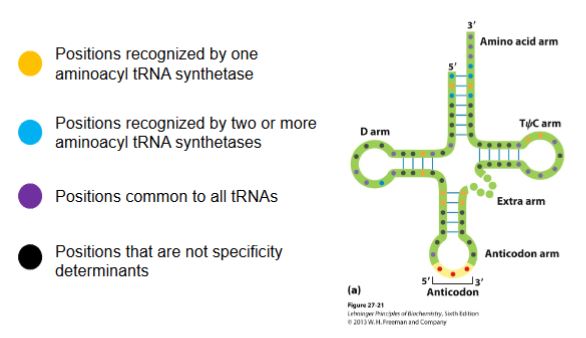
How do synthetases recognize tRNA?
Specific nucleotide identity elements.
What are the phases of translation?
Initiation, elongation, and termination.
What are the steps for Translation initiation?
Recruitment of small ribosomal subunit to the mRNA.
Identification of the start codon.
Association of the charged initiator tRNA with the mRNA.
Recuitment of the large ribosomal subunit to form an active ribosome.
Which site does the initiator tRNA bind first?
P site.
What exactly does the Shine-Dalgarno sequence do?
It base pairs with 16S rRNA which aligns the ribosomes correctly/positions the AUG correctly. It recruits ribosomes to the mRNA.
Replaces the need for 5’ cap (found on eukaryote mRNAs).
What is the first amino acid in all proteins?
Methionine.
In bacteria, what are the specific tRNAs for Methionine?
tRNAmet → internal AUG (methionine)
tRNAfmet→ 5’ start AUG (methionine)
In bacteria, the initiation methionine is formylated.
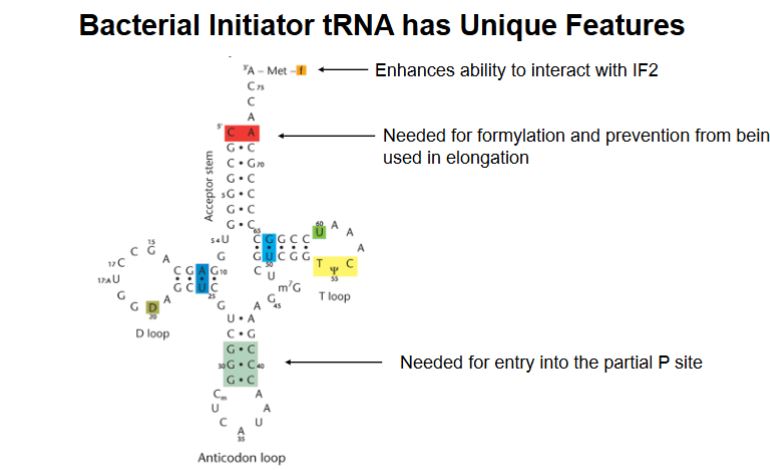
What are some unique features that the bacterial initiator has?
the formyl modification at the top enhances the ability to interact with IF2
CA in the middle is needed for formylation and prevention from being used in elongation.
Bottom base pairs needed for entry directly into the partial P site.
The initator tRNA is specialized for starting translation and is structurally prevented from participating in elongation.
What does bacteria require for initiation?
30S ribosomal unit (small subunit)
mRNA
fMet-tRNA
Initiation factors - IF-1, IF-2, and IF-3
GTP (This step requires energy)
50S ribosomal unit (large subunit)
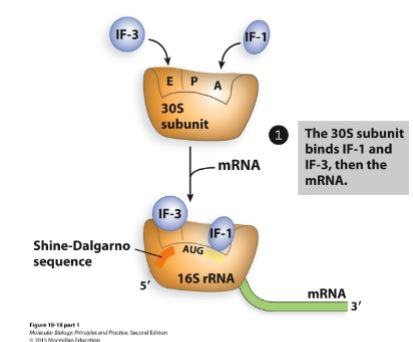
First step of initiation (bacteria).
30S subunit (small subunit) binds IF-1 and IF-3, then binds the mRNA.
IF-3 and IF-1 force the first tRNA to bind at the P site.
16S rRNA (apart of 30S subunit) binds to the RBS/SD sequence to position AUG correctly at the P site.
Second step of initation (bacteria).
GTP bound IF-2 and fMET-tRNA join the small subunit.
The other IFs block the A and E site making it bind to the P site.
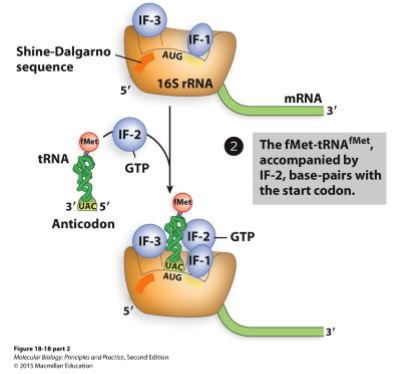
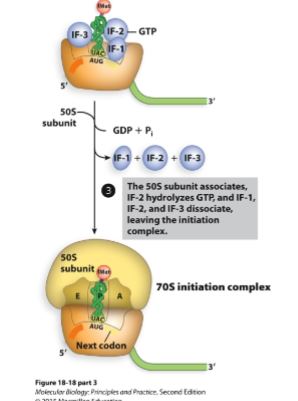
Third step of initiation (bacteria).
IF-3 is released
50S (large subunit) subunit joins
GTP is hydrolyzed to GDP and Pi
IF-1 and IF-2 are released
70S initiation complex is now ready for elongation
What is the difference between prokaryotic and eukaryotic initiation?
In initiation (for translation) for eukaryotes:
The initiator tRNA, Met-tRNAi is not formylated
More eukaryotic initiation factor proteins are required (~12 eIFs)
There is no Shine-Dalgarno sequence
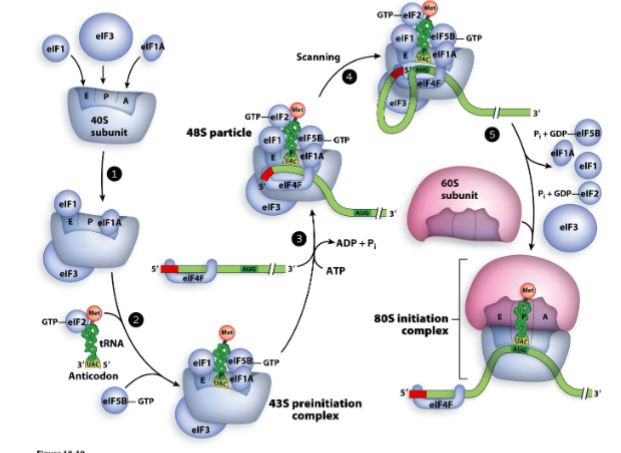
What are the steps to Eukaryotic translation initiation?
Binding of translation factors (eIFs) to the 40S ribosome subunit (small subunit). These eIFs make sure that the aminoacyl-tRNA binds to the P site first.
GTP bound EIF associates with initiator tRNA as well other eIFs join the small subunit to form the 43S preinitiation complex. A eIF4F mediates binding of mRNA to the 43S preinitiation complex.
43S pre-initiation complex scans mRNA (in the 5’ → 3’ direction) for the first AUG.
Base pairing of initiator tRNA with the start codon
Release of eIFs and association of 60S subunit (large subunit) with 40S subunit (small subunit) to form the active 80S ribosome
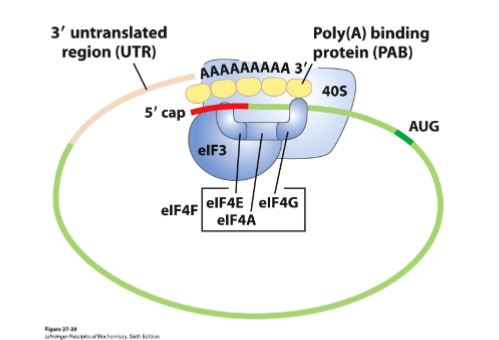
What is the eIF4F complex (in eukaryotic initation)?
This complex mediates the binding of mRNA to the 43S preinitiation complex. It contains:
eIF4E - (cap binding protein) that binds to the 5’ cap
eIF4A - is an ATPase and RNA helicase
eIF4G - binds to poly(A) binding protein (PABP)
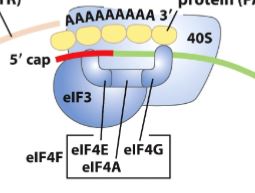
mRNA circulization
eIF4E binds the cap and eIF4G is bound to PABP which binds to the poly A tail → circulizing mRNA
good for orientation
efficient translation
(not in bacteria)
What is the Kozak sequence.
It is like the Shine-dalgarno sequence, it is a consensus sequence that holds the start codon (AUG).
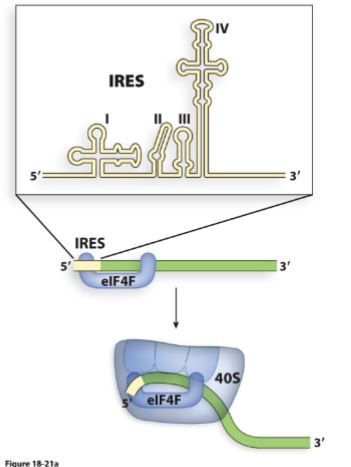
What is an Internal Ribosome Entry Site (IRES)?
Normal eukaryotic translation requires a 5’ cap + uses scanning.
This is an alternative mechanism: it is an RNA segment located on the 5’ side to the start codon.
Directly recruits:
40S ribosomal subunit
Eukaryotic Initiation Factors (eIFs)
What are the three steps involved in translation elongation?
Aminoacyl tRNA is “loaded” into the vacant A site.
Peptidyl transferase reaction results in transfer of peptide from tRNA in the P site to amino acid of charged tRNA in the A site (transfer of chain to new amino acid).
Resulting “new” peptidyl-tRNA with its associated codon is translocated from A site to P site.