Chapter 11: Chromosome Structure and DNA sequence organization
1/26
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
27 Terms
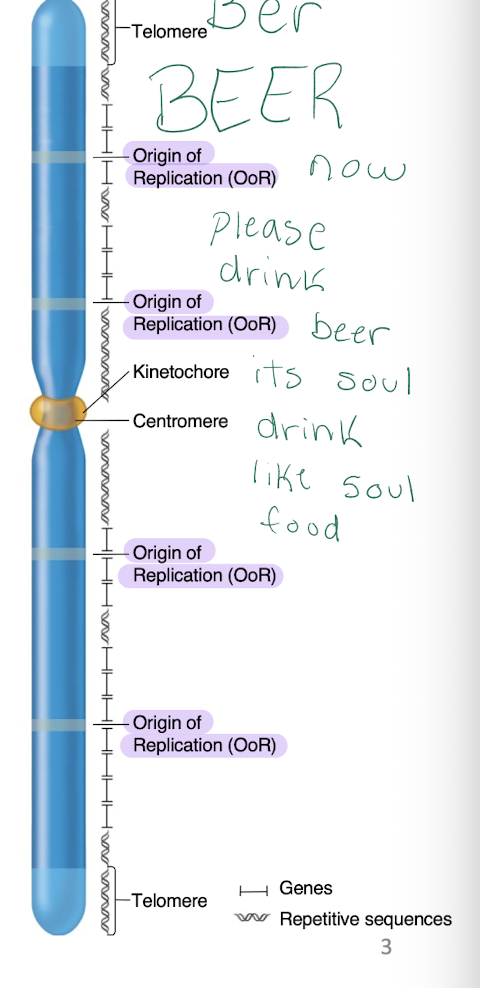
Eukaryotic chromosomes
-Only visible during mitosis
-10-100 million base pairs in length
-Several 100-1000 genes interspersed throughout
-Origin of Replication are about 100,000 bp apart
-Centromere is the recognition site for kinetochore microtubules
-Telomeres have repetitive sequences at both ends
-Other repetitive sequences near centromere and interspersed throughout
Human nucleus
-5-10 um, 46 chromosomes
-contain enough DNA to extend to more than 2 meters
Linear DNA
interacts with proteins, which affect the degree of chromatin compaction
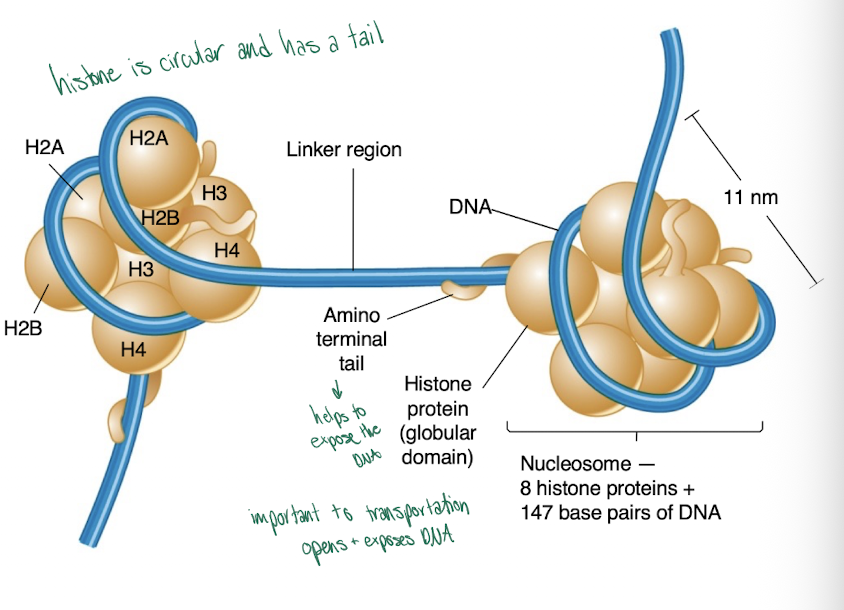
Histones
basic, positively charged proteins that play a critical structural role in chromatin
-have a globular domain and a charged, flexible histone tail
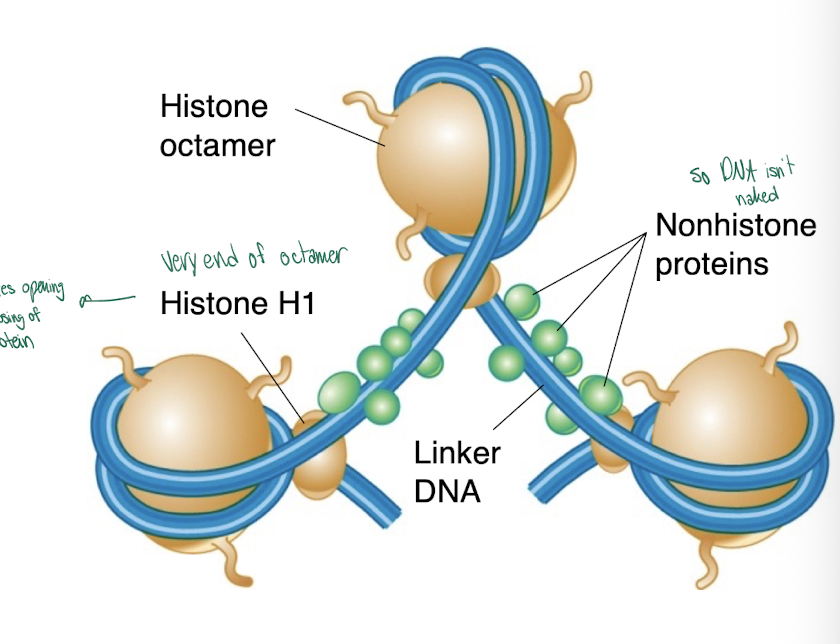
Non-histones
less positively charged proteins, dont make DNA so compact but keeps DNA together
-bind the linker region and also aid in chromatin compaction, they also may effect gene expression
Linear double stranded DNA
-wraps around an octamer (8 histones) of 2 copies each of 4 histones to form nucleosome
-H2A, H2B, H3 and H4 are the core histones
Core histones
H2A, H2B, H3 and H4
Nucleosomes
-repeating structural unit of chromatin and condense its structure by 7-fold
-147 base pairs of DNA winds around the histone octamer and resembles beads on a string
H1
the linker histone that binds to spacer DNA between nucleosomes, but not as tightly as the core histones
30-nm solenoid
-nucleosomes closely associate with each other form this and is compacted 50 fold
-histone protein H1 plays a key role in this packaging
-This structure is characteristic of the uncoiled chromatin fiber in interphase
-Going into metaphase, the 30-nm structures are folded into a series of lop domains that are 300 nm structures
Euchromatin
-not tightly coiled
-less condensed regions of chromosomes that are transcriptionally active
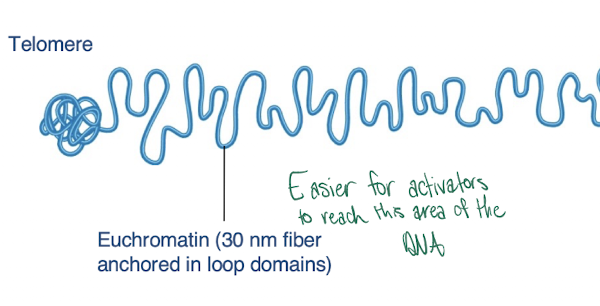
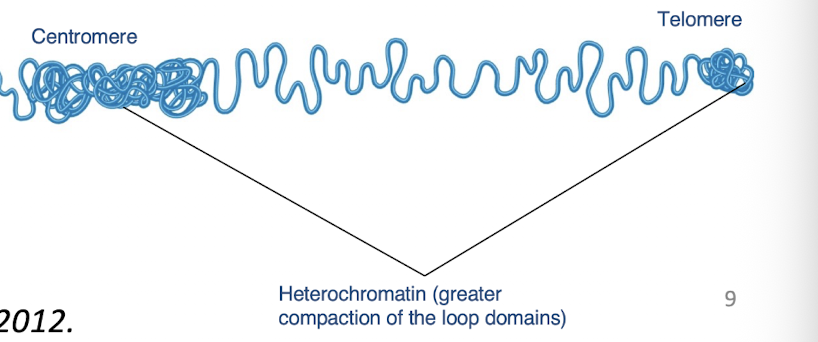
Heterochromatin
-condensed
-tightly compacted regions of chromosomes that are, in general, transcriptionally inactive
-Unique to eukaryotes
-replicates later during S phase than euchromatin
-Important in maintaining the structural integrity of chromosomes
-Ex: inactive X chromosome is condensed into heterochromatic Barr body
-Translocation of genes into heterochromatic regions can alter gene expression- if too tightly wound, unable to be expressed
Chromatin remodeling
-when chromatin can be induced to change its structure
-histone tails are targets for a variety of chemical modification that regulate chromatin folding and gene expression
-Acetylation, methylation and phosphorylation of amino acids in histone tails
-makes chromatin unwind for DNA replication or for it to become tightly wound, makes it active or inactive
-Open: transcription happens
-Close: transcription doesn’t happen
Repetitive DNA
-59%
-regulatory DNA
-dont know its function
Unique noncoding DNA
-15%
-makes muscle cells, dont produce stomach enzymes
Introns and other parts of genes
-24%
-form preRNA
Regions of genes that encode proteins
2%
Middle repetitive sequences
-include tandem repeats and interspersed retrotransposons
Tandem repeats include:
-Multiple copy genes
-Variable number of tandem repeats (VNTRs)
-Short tandem repeats (STRs)
Multiple copy genes
-E.g., rDNA- clustered on the short arm of acrocentric chromosomes (13, 14, 21, 22)
Variable number of tandem repeats (VNTRs)
-15-100 bp repeated sequences within and between genes that extend for 1-20 kb in length
-Clusters of VNTRs throughout the genome are called minisatellites
Short tandem repeats (STRs)
-Di-, Tri-, tetrea-, penta repeats (2-6 bp long)
-Clusters of STRs throughout the genome are called microsatellites
-Most common is (CA)n, where n is 5 to 50 repeats

Retrotransposon
-Copy and paste
-replicates and inserts DNA somewhere else
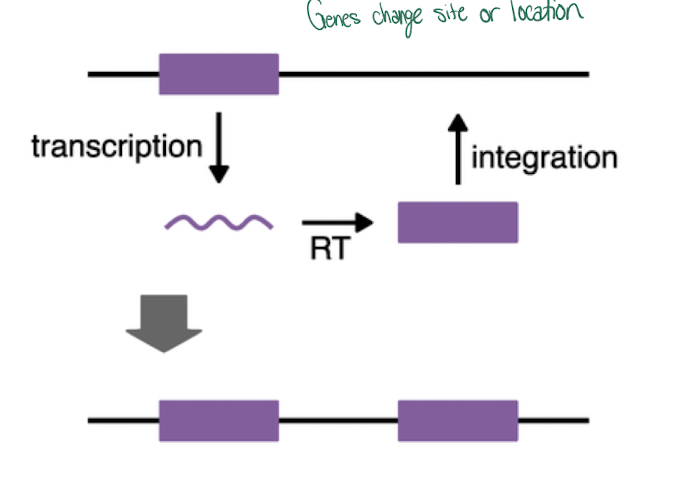
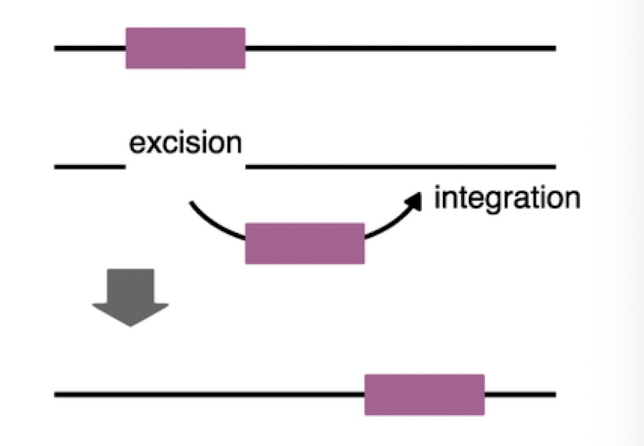
DNA transposon
-cut and paste
-gene changes site or location
Interspersed retrotransposons
-Transposable sequences are interspersed and can relocate within the genome
-Short interspersed elements (SINEs) make up 13% of genome
<500 bp long present in 50,000+ times in human genome, best characterized is the ALU family, which are 200-300 bp long and make up 5+% of the human genome
-Long interspersed elements (LINEs) make up 21% of genome; ~6 kb long and present ~850,000 times in human genome
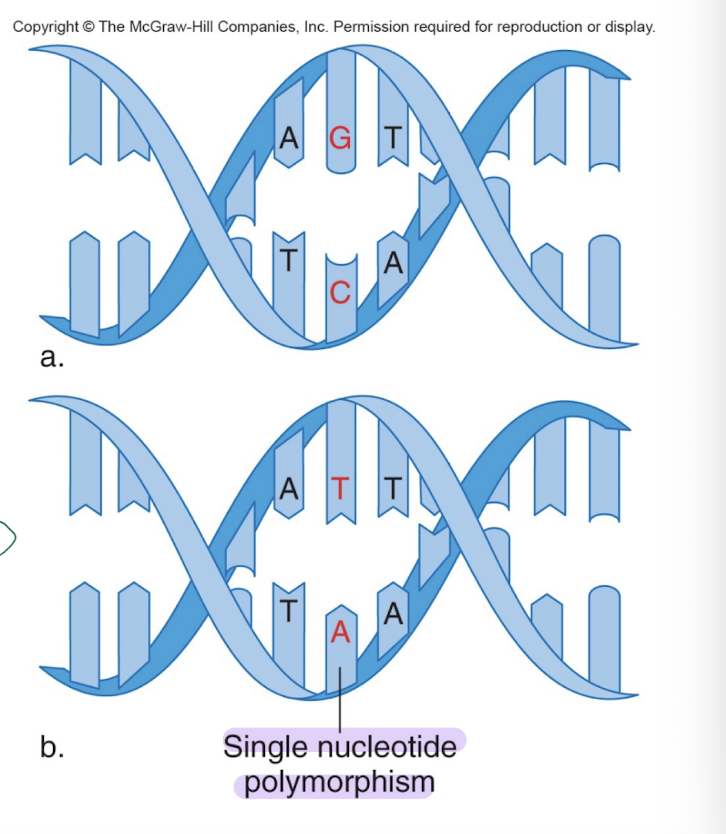
Single nucleotide polymorphisms (SNPs)
-the most common type of genetic variation
-Occur once in every 500-1000 bp; 10 million SNPs in the human genome
-Most are between genes and have been used to identify genes in the human genome
-are also used for personalized genomic medicine