MOLBIOL 3II3 Final exam
1/440
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
441 Terms
HP1
Binds to H3K9me3
Highly repressive
Mediates:
Self-aggregation
HP1 mols bind each other
Once its on chromatin, clusters nucleosomes and promote local compaction
Chromatin condensation by recruit chromatin remodeling complexes (BRG1/BRM)
Histone deacetylation - recruit HDACs
Remove acetyl grps
Acetylation = open chromatin
Deacetylation = repression
DNA methylation - recruiting DNMTs
More stable repression
Nuclear lamina interaction
Bind lamins
HP1 structure
Two key domains
N-term chromodomain (CD)
reader
recognizes repressive histone marks H3K9me3
C-term chromoshadow domain (CSD)
protein-protein interaction
recruits additional repressive machinery
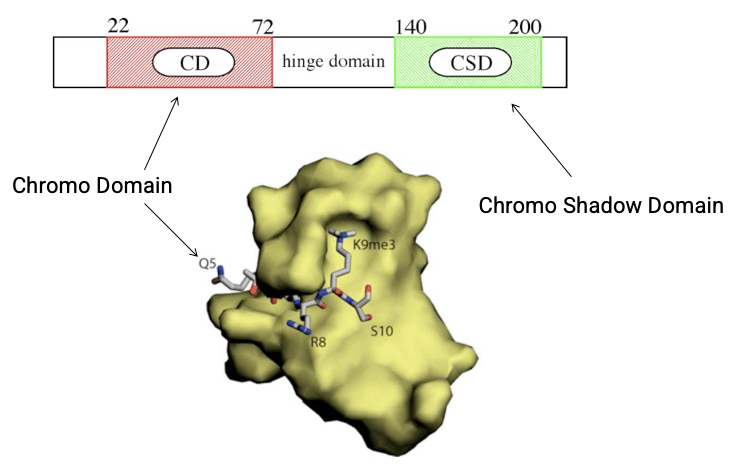
How HP1 compacts chromatin?
H3K9me provides a docking site for HP1
Once HP1 is anchored, the CSD promotes HP1-HP1 interactions (dimerization/oligomerization)
HP1 self-aggregation can bridge neigbouring nucleosomes and drive local chromatin condensation
Trigger → Bridge mechanism of HP1-mediated silencing
Trigger: H3K9me occurs, creating a docking site
Docking: HP1 CD anchors to methylated histone
Aggregation: CDS interacts with the CD, causing HP1 molecules to stick together
Compaction: the oligomerization bridges nucleosomes together, physically condensing the chromosomes
How does HP1 connect histone modification to DNA methylation?
HP1 acts as a recruitment platform
After binding to H3K9me2/3, HP1 physically recruits DNMTs to the chromatin
Repression becomes DNA-based signal, more stable and long-term
DNMT3 vs DNMT1
DNMT3 (3A/B): “WRITER” de novo methylation, establishes new methylation patterns on previously unmethylated DNA
DNMT1: “COPIER” maintenance methylation, copies existing methylation patterns to the nascent strand after DNA replication to ensure inheritance of the OFF state
both establish and preserve epigenetic memory
DNA methylation: long-term silencing in heterochromatin STEPS
H3K9 methylation - histone mark is written
HP1 binding - reader anchors to the mark
DNMT recruitment - HP1 brings in DNMTs
CpG methylation - DNA itself is methylated
Stable silencing - reinforcement through additional repressive proteins
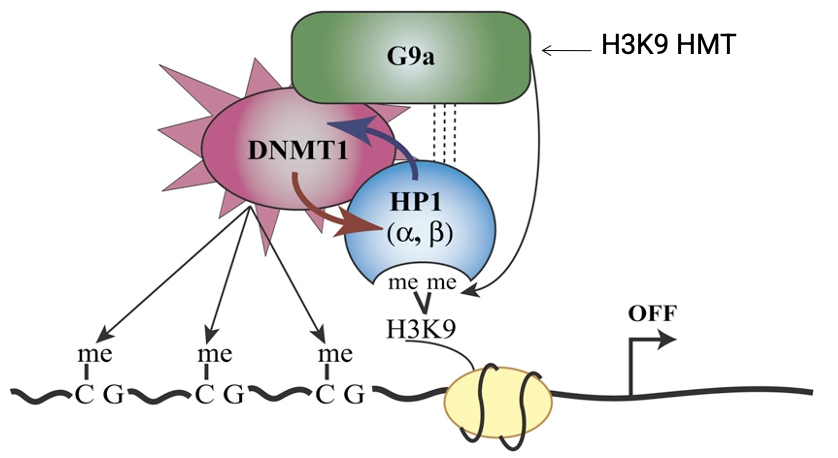
Why is the recruitment of DNMTs particularly important for repeat-rich regions like centromeres and telomeres?
These regions require long-term, stable silencing that can be inherited across many cell divisions
DNA methylation provides a more permanent “lock” than histone marks alone, preventing the mobilization of repetitive elements or genomic instability
Which specific histone mark does HP1 have the highest affinity for when acting as a platform for DNMTs?
H3K9me3 (TRI-METHYLATED FORM)
What is the specific biological role of the positive feedback loop bw H3K9me3 and DNA methylation?
To stabilize and reinforce gene silencing
This self-reinforcing loop ensures that repressive marks are maintained and spread across genomic regions like pericentric heterochromatin, keeping them condensed and stably silent
Sequence the 6-step positive feedback loop that maintains heterochromatin, starting from H3K9me3
H3K9me3 - histone mark
HP1 - reader/platform recruitment
DNMT3 - de novo DNA methyltransferase
DNA methylation - CpG sites
MeCP2 - methyl-DNA binding protein
SUV39H1/2 - histone methyltransferase, writes more H3K9me3
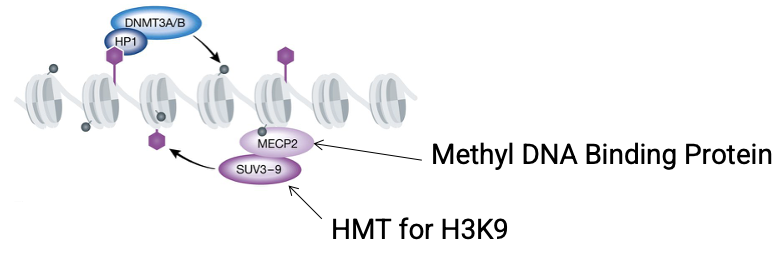
What is the role of MeCP2 in heterochromatin stabilization cycle?
Acts as a reader of DNA methylation
It specifically recognizes and binds to methylated CpG sites, subsequently recruits SUV39H1/2 to write more H3K9me3 marks on nearby nucleosomes
Which enzyme in the pericentric silencing loop is responsible for “closing the circle” by writing more histone methylation?
SUV39H1/2
It is recruited by MeCP2 and catalyzes the trimethylation of H3K9 => recruits more HP1 to restart the cycle
Why is this silencing loop particularly more common in pericentric heterochromatin and repeat-rich domains?
These regions must remain condensed and transcriptionally inactive most of the time
The loop creates a robust, stable OFF state, resistant to accidental activation
Why is the nuclear lamina described as a “repressive envr” for chromatin?
It physically sequesters heterochromatin at the nuclear periphery, away from translational machinery of the nucleoplasm
Exposes it to high conc of repressive factors like HDAC3
What are the two molecular tethers that lamin B receptor (LBR) uses to anchor heterochromatin to the inner nuc mem
Direct histone binding - TUDOR DOMAIN binds to the H4K20me2 mark
Protein bridging - It binds HP1, which is already anchored to H3K9me3 marks on the chromatin
What is the functional role of HDAC3 at the nuclear periphery?
Removes acetyl groups from histones
Maintains a deacetylated, compact state, reinforcing the low-transcription envr of the lamina-associated heterochromatin
Recruitment chain for spatial silencing: Inner nuc mem → LBR → ?? → H3K9me3
HP1.
LBR tethers HP1, which in turn is bound to the H3K9me3 mark on the nucleosome
True or False: LBR is a soluble protein that floats in the nucleoplasm to find heterochromatin
FALSE. LBR is an INM protein that binds to lamin B, providing a fixed physical anchor point at the edge of the nucleus
How does the "Tethering Model" relate to the "Feedback Loop" (H3K9me3/DNA Methylation)?
Complementary
The feedback loop establishes and stabilizes the chemical marks of repression
Tethering provide the physical organization to keep those repressed regions compartmentalized in a silent zone “the nuclear periphery”
What specific chemical state characterizes the CpG islands of actively transcribed mammalian promoters?
They are UNMETHYLATED
Not just lack of repression, acts as a functional signal to recruit specific chromatin-modifying complexes
Which protein is recruited to unmethylated CpGs to facilitate the deposition of the H3K4me3 active mark?
CFP1
A component of the SET1/COMPASS (trithorax-related) complex, which writes H3K4me3 at the promoter
What is the function of KDM2A when it is recruited to an active CpG island?
It acts as a demethylase that removes H3K36 methylation
Erasure helps maintain the specific permissive chromatin envr required for the promoter to function
Describe the recruitment chain for an Active Promoter starting from the DNA status
Unmethylated CpG island - the signal
CFP1/KDM2A recruitment - the readers
H3K4me3 deposition/H3K36 removal - histone remodelling
Transcriptionally permissive state - OUTCOME
How does Dense CpG Methylation contrast with Unmethylated CpG Islands in terms of gene expression?
DENSE methylation: recruits MeCP2 and DNMTs => stable silencing
UNMETHYLATED islands: recruits CFP1 and KDM2A => active transcription
What is the specific biochemical rxn catalyzed by DNMTs?
They transfer a methyl group onto 5-C of the cytosine base, producing 5mC
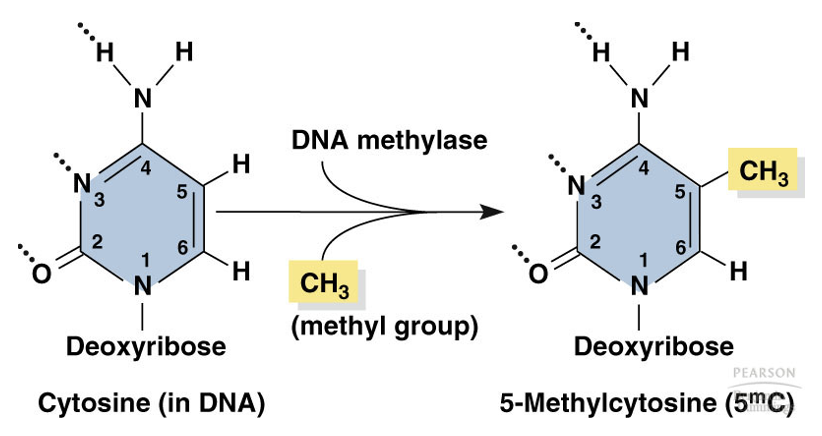
Why is the symmetry of CpG dinucleotides significant for epigenetic inheritance?
Allows methylation patterns to be maintained as paired marks.
After replication, the “old” strand remains methylated while the “new” strand is not (hemimethylated), providing a template for restoration
DNMT1 vs DNMT3A/3B in maintaining the cellular OFF state
DNMT1 - for maintenance methylation, restores the symmetry by methylating the new strand after DNA replication “COPY CAT”
DNMT3A/3B - for de novo methylation, establish entirely new patterns for 5mC on previously unmethylated DNA “TRENDSETTER”
How does the stability of DNA methylation fundamentally differ from most histone marks?
DNA methylation = more stable
Unlike histones, which have dedicated demethylase enzymes like KDM2A
NO OFFICIAL DIRECT DNA DEMETHYLASES
DNA methylation is usually removed either passively (diluted over replication) or through complex, multi-step pws
What is the primary functional outcome when DNMTs install 5mC marks at promoter CpG sites?
Transcriptional silencing
The presence of 5mC at CpGs is strongly associated with a stable, repressed chromatin state
Maintenance methylation in DNA replication
DNMT1 recognizes hemimethylated DNA (methylated on parent strand only) and adds a methyl group to the daughter strand = ensure epigenetic state is inherited by the progeny cells
Three primary ways DNA methylation (5mC) promotes transcriptional silencing?
Direct interference - physically blocks TFs and promoter-binding proteins
Polymerase inhibition - prevents RNAP from using the DNA as a substrate for transcription
Protein recruitment - acts as a “landing pad” for methyl-DNA binding proteins (MeCP2, kaiso), which recruit corepressors
How does DNA methylation affect the assembly of the pre-initiation complex (PIC) at promoters?
The presence of the methyl group on the 5-C of cytosine can directly interfere with the binding of TFs, preventing the PIC from assembling and starting transcription
Even if RNAP is successfully recruited to a methylated promoter, what prevents gene exp?
RNAP is inhibited by the methylated DNA
It cannot effectively use a methylated template as a substrate to transcribe RNA
What type of corepressor machinery is typically recruited by proteins like MeCP2 and Kaiso to methylated DNA?
HDACs and other chromatin-condensing complexes
Ensure region remains in a closed, “heterochromatic” state
Approximately what percentage of silenced DNA regions in mammals exhibit CpG methylation?
~90%
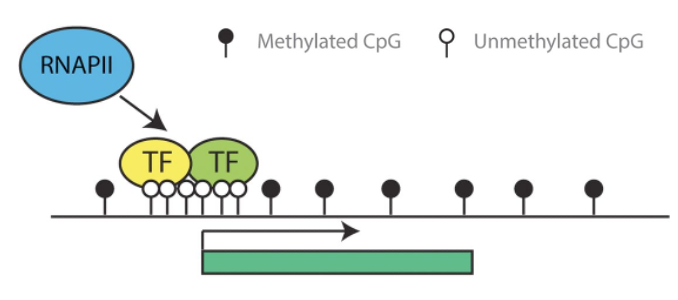
What are the characteristic epigenetic and molecular features of a GENE ON state at a CpG-island promoter?
LOW DNA methylation (open circles = unmethylated CpG)
TFs bind to specific DNA motifs
Activators recruit coactivators and RNAP II machinery (GTM) = Pre-initiation complex
Result: transcription proceeds
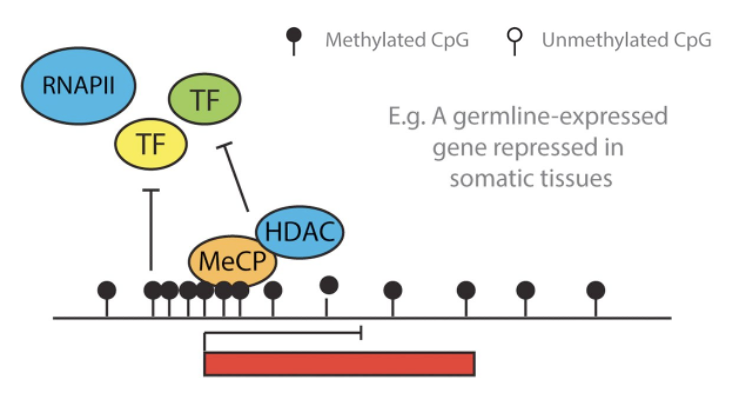
How does DNA methylation at CpG siles lead to a GENE OFF state and chromatin compaction?
Direct inhibition - methylated CpG (filled circles) prevent the binding of TFs = blocked RNAPII recruitment
Methylated DNA acts a binding site for MeCP
MeCP recruits HDACs and methyltransferases (like SUV3-9)
HDACs remove acetyl groups from histones => chromatin compaction and a reinforced silenced state
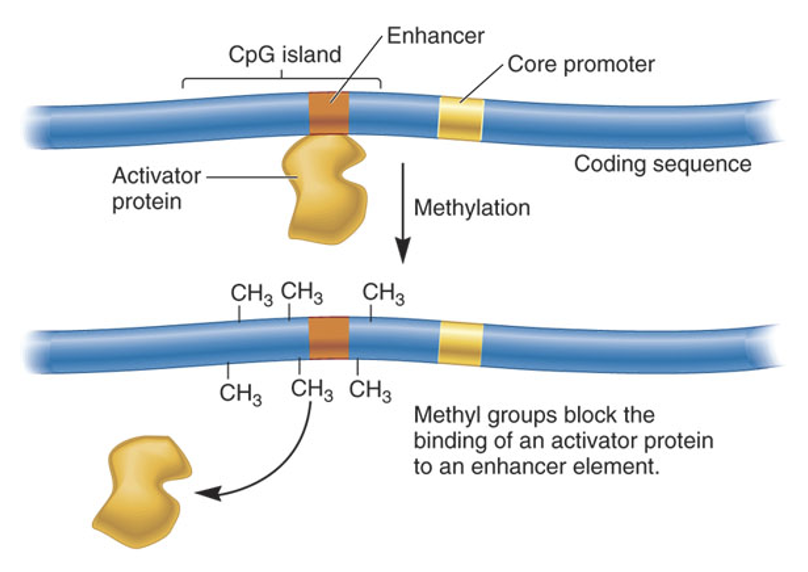
What are the two primary reasons why activator binding and recruitment of the initiation machinery fail at methylated promoters?
Poor binding affinity - some TFs bind boopry or not at all to their motifs when they contain CpG methylation
Physical obstruction - methyl groups can sterically hinder the assembly of PIC
Two primary mechanisms by which DNA methylation inhibits gene expression at enhancers and promoters
Passive/direct - direct blocking of activator proteins from their DNA motifs
Active/indirect - recruitment of methyl-DNA readers (repressors) that modify chromatin structure
How does 5mC physically prevent TF binding?
It changes the chemical shape of the binding surface within the DNA motif
Causes the TF to either bind much more weakly or lose its ability to bind entirely
Which family of proteins acts as the bridge bw methylated DNA and the recruitment of corepressors?
Methyl-CpG binding proteins or methyl-DNA readers, MeCP family
Describe the sequence of events that transforms open chromatin into closed chromatin via DNA methylation
CpG sites are methylated
MeCP proteins dock onto methyl-CpG sites
MeCP brings in HDACs
HDACs remove acetyl groups from his tails
Loss of acetylation = nucleosome-DNA interactions to tighten
How does the recruitment of SUV39H by methyl-DNA binding proteins create a self-reinforcing repression cycle?
SUV39H deposits H3K9me, which then recruits HP1
This links the DNA methylation signal to the histone-based heterochromatin machinery, making the OFF state stable and self-sustaining
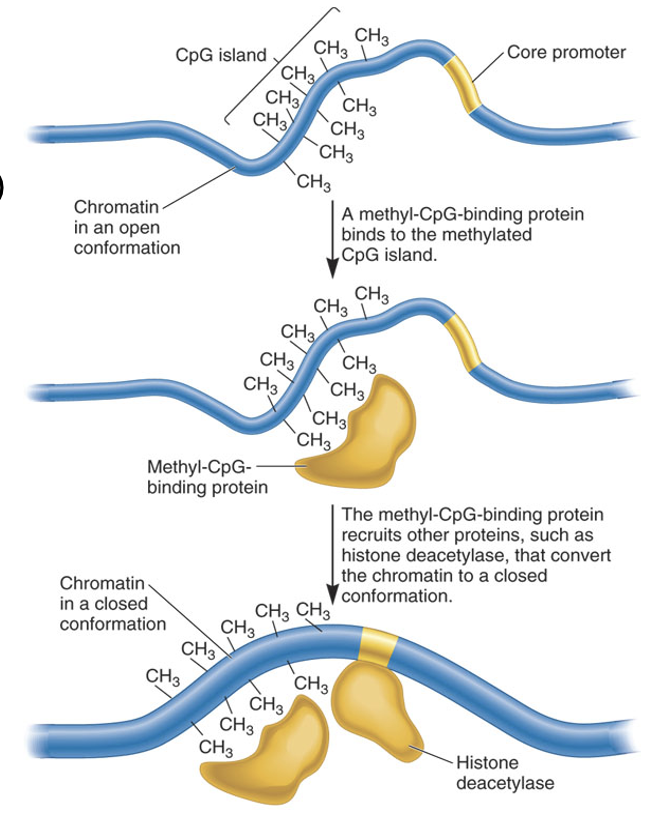
Why does the removal of acetyl groups from histone tails by HDACs result in a "closed" chromatin state?
Acetylation normally neutralizes the positive charge of histone tails
Weakens their interaction with negatively charged DNA = keeps it open
Removing these groups restores attraction, promoting tighter packing and reduced accessibility
In the transition from open to closed chromatin, what is the structural outcome of MeCP binding?
The recruitment of corepressor complexes that physically condense the chromatin => making the DNA inaccessible to RNAP II
What is the specific protein domain used by MeCP2 to recognize and bind to methylated DNA?
Methyl-CpG binding domain
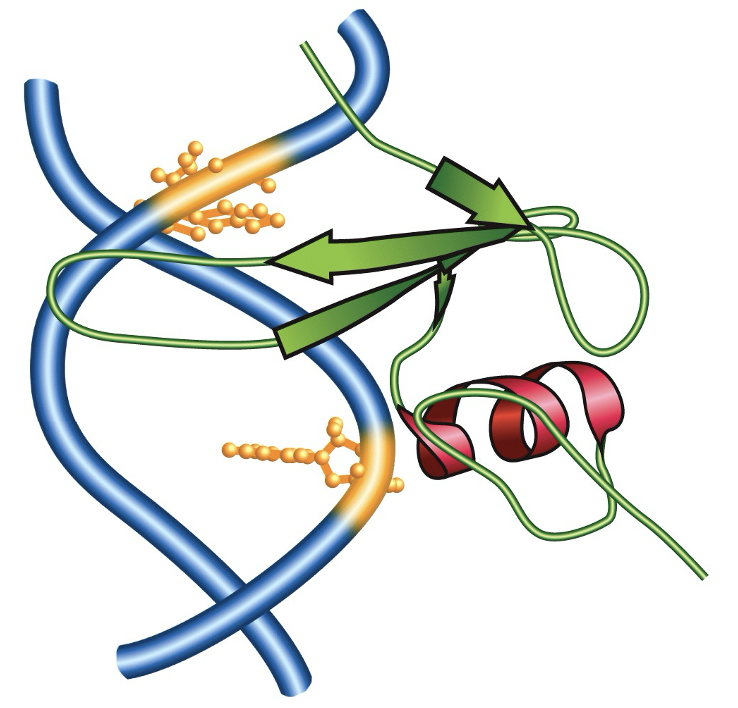
How does a Methyl-CpG-binding domain (MBD), such as the one found in MeCP2, physically recognize methylated DNA?
Target: MBD specifically reads 5mC within a CpG context
Structural fit: the protein uses specific loops that sit against the DNA helix
Shape complementarity: binding interface is a complementary shape that fits the methy group (yellow bulges) on the dna
allows the protein to bind preferentially only when methylation is present
What is the general cross-talk rule bw histone methylation and DNA methylation long-term silencing?
Repressor complexes that deposit histone marks (H3K9me or H3K27me3) act as recruitment signals for DNMT3, which then installs DNA methylation to lock the silenced state
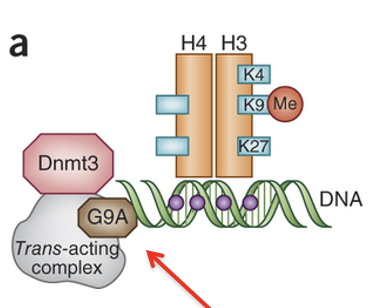
Describe the specific cross-talk pw for G9a-associated repression
G9a (histone methyltransferase) deposits H3K9me1/2
These marks recruit DNMT3
DNMT3 established de novo DNA methylation at nearby CpG sites
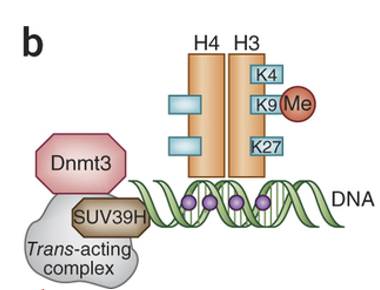
How does the SUV39H pathway differ from G9a in terms of the histone marks used to recruit DNMT3?
SUV39H = classic writer for H3K9me3 in constitutive heterochromatin
Like G9A, this H3K9me3 mark serves as a signal to recruit DNMT3 for DNA methylation
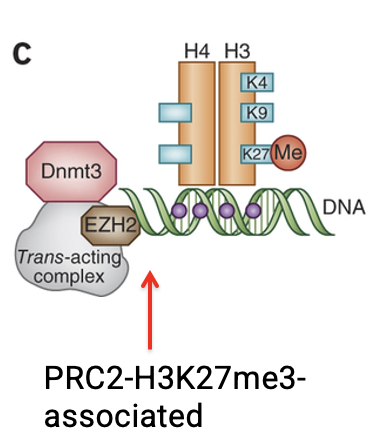
In polycomb-associated repression (PRC2), which catalytic subunit and histone mark are linked to DNA methylation?
The catalytic subunit EZH2 deposits H3K27me3
In certain contexts, this complex is coupled to DNMT recruitment to stabilize repression
According to the schematic, how can DNMTs be recruited in the Polycomb (PRC2/EZH2) context?
DNMTs can arrive pre-assembled within the PRC2/EZH2 complex, allowing DNA methylation to occur simultaneously with H3K27me3
What is the primary mark of G9a in DNMT3 recruitment chain?
H3K9me1/2
What is the primary mark of SUV39H in DNMT3 recruitment chain?
H3K9me3
What is the primary mark of EZH2 (PRC2) in DNMT3 recruitment chain?
H3K27me3
Why are HOX gene clusters frequently used as the classic example for Polycomb-mediated repression?
HOX genes require precise, tightly regulated developmental timing
They must be expressed only in specific cell types at specific times
Once a developmental decision is made to turn them off, they must stay stably and heritably silenced via polycomb complexes
Polycomb Bodies
Visible nuclear clusters or neighbourhoods where polycomb-repressed chromatin regions physically aggregate
They are enriched for polycomb proteins (PRC1/2) and repressed chromatin = localized envr dedicated to silencing
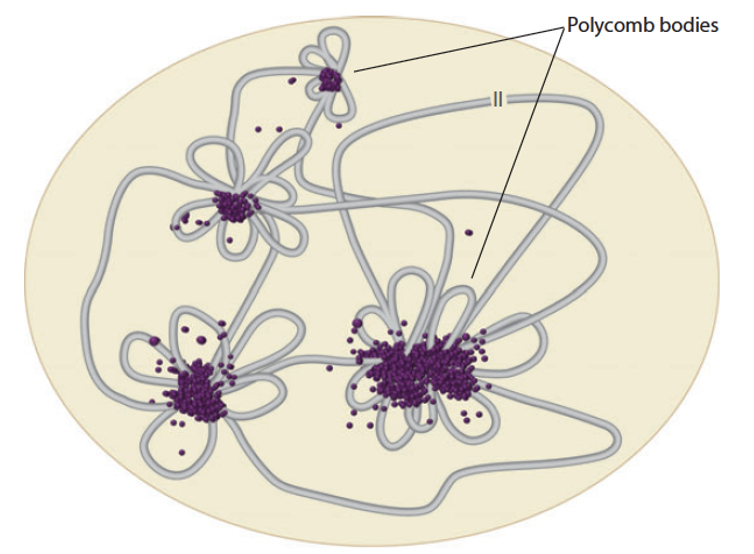
What is a major spatial function of the Polycomb complexes PRC1 and PRC2?
Promote the physical clustering of polycomb-bound genomic regions
This moves repressed loci from isolated positions into organized, collective territories (polycomb bodies)
PRC2 vs PRC1 in establishing repressive territories
PRC2 - places the H3K27me3 histone mark
PRC1 - recognizes the mark and helps compact the chromatin and facilitate the physical clustering of repressed regions
What is the functional advantage of clustering repressed genes (like HOX clusters) into a Polycomb body?
Creates a self-reinforcing envr
By concentrating repressive factors and silenced chromatin in one territory, the cell ensures the OFF state is stable and protected from accidental activation until a specific developmental signal occurs
If a specific HOX cluster is observed within a Polycomb body in a certain tissue, what does that indicate about its transcriptional state?
It indicates the cluster is in a state of stable, heritable repression
It is "marked" and "parked" in a silent nuclear neighbourhood
How does an intact lamin network support the stable silencing of developmental genes via polycomb groups (PcG) proteins?
Clustering - the lamin network allows polycomb proteins to aggregate into localized PcG bodies within the nucleus
Spatial organization - target gene loci are physically sequestered into these hubs
Functional outcome - these dense chromatin interactions maintain a stable OFF state, ensuring developmental genes remain repressed until specifically needed
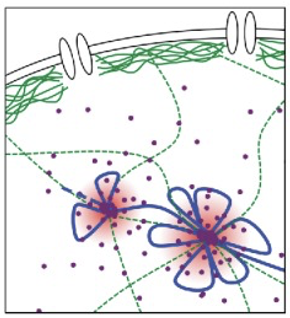
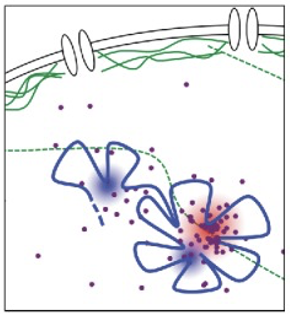
What happens to polycomb-mediated gene repression when the lamin A/C network is depleted or disorganized?
Structural decay - PcG bodies become disorganized and less clustered within the nucleus
Weakened interactions - higher-order chromatin interactions that maintain the repressive hubs are significantly weakened
Functional failure - polycomb-mediated repression becomes unstable
Consequence - target genes (often developmental) are misregulated, frequently turning ON when they should remain silenced (bigla silang nagparty ganon)
During DNA replication, why is the Gene OFF methylation state at risk of being lost?
Problem - DNA rep is semi-conservative, the parental strand retains its methyl marks but the newly synthesized daughter strand is unmethylated
Risk - without a maintenance mechanism, the OFF signal would be diluted by 50% every cell division, eventually disappearing and causing a loss of stable gene silencing
Hemimethylated DNA
DNA that is methylated on only one of its two complementary strands
Occurs immediately after DNA rep, where parental strand keeps its 5mC marks but the newly synthesized daughter strand is unmethylated
Which specific enzymes are the primary de novo DNMTs in mammals?
DNMT3A
DNMT3B
What is the "preferred substrate" for the maintenance methyltransferase DNMT1?
Hemimethylated DNA
Describe the mechanical role of DNMT1 immediately following DNA replication
DNMT1 identifies hemimethylated CpG sites and methylates the new daughter strand at matching positions
THis restores the symmetric methylation pattern and preserves epigenetic memory
At what stage of the cell cycle is DNMT1 activity most critical?
S-phase
Immediately following the passage of the replication fork
This is when hemimethylated DNA substrates are generated
Two major patterns of aberrant DNA methylation found in cancer
Global hypomethylation - pericentromeric heterochromatin
Focal hypermethylation - tumour suppressor genes TSGs
Contrast the global vs. focal changes in DNA methylation typically seen in cancer cells.
Global hypomethylation - pericentromeric heterochromatin
normal: heavily methylated, compact, and transcriptionally quiet
in cancer: loss of methylation = open chromatin
consequence: promotes promiscuous recombination bw repetitive sequences => chromosomal instability
Focal hypermethylation - tumour suppressor genes TSGs
normal: CpG island promoters are unmethylated and transcriptionally permissive
in cancer: gain of methylation at these specific promoters
consequence: stable transcriptional silencing, functional loss-of-function, effectively turning off genes that normally prevent tumour growth
Why is hypomethylation in pericentromeric heterochromatin dangerous for genomic stability?
Loss of methylation makes repetitive sequences more available to the cells DNA machinery
These repeats can acts as promicuous substrates for recombination, where the repair machinery incorrectly pairs similar sequences on different chromosomes
What are the potential structural outcomes of "promiscuous" recombination at repetitive elements?
Mitotic recombination
Deletions
Duplications
Translocations
Overall chromosomal instability
In a "Normal" cell, what is the typical distribution of DNA methylation across a gene's promoter versus its gene body?
Promoter - typically unmethylated, maintains a transcriptionally permissive state = gene exp
Gene body and distal regions - higher levels of methylation, stable & healthy genomic organization
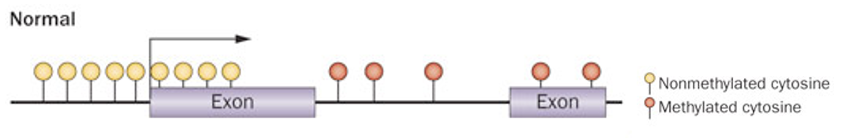
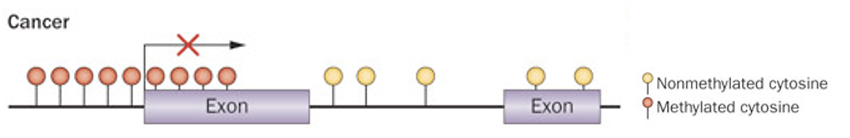
What is the functional consequence of promoter hypermethylation in cancer cells, and how does it mimic genetic mutation?
normally unmethylated promoter becomes hypermethylated
Methylation at the TSS acts as a red X, completely blocking expression
mechanism:
physically blocks TF binding
recruits methyl-binding proteins
attracts corepressors to close chromatin
equivalent to loss-of-function mutation, gene is present but permanently silenced
CpH motif (in DNA methylation)
A cytosine followed by any base except guanine (H = A, C, or T)
Methylation at these sites is referred to as mCH
In which two specific cellular contexts is mCH most highly enriched?
Pluripotent stem cells
Nervous sytem
How does the establishment of mCH in the brain relate to development?
It is developmentally regulated and becomes established postnatally as neurons mature
It is a molecular hallmark of neuron maturation, synapse formation, and plasticity
What is the general "rule of thumb" for the functional effect of mCH on gene expression?
It behaves as a repressive mark.
Higher levels of mCH, particularly across gene bodies, are typically associated with lower gene expression
How does the cell-type specificity of mCH compare to standard CpG methylation?
mCH is highly cell-type and development-specific
CpG methylation is the dominant and relatively stable form across most somatic cell types.
Besides "repressive," what other biological processes is mCH frequently associated with in the brain?
neuron maturation
synapse formation
synaptic plasticity
How do the developmental trajectories of mCG and mCH differ in the mammalian brain (human and mouse)?
mCG (global/CpG) - remains relatively high and steady across the entire lifespan
mCH (non-CG) - starts nearly undetectable, rises sharply during early postnatal development and childhood, plateaus in adolescence/adulthood
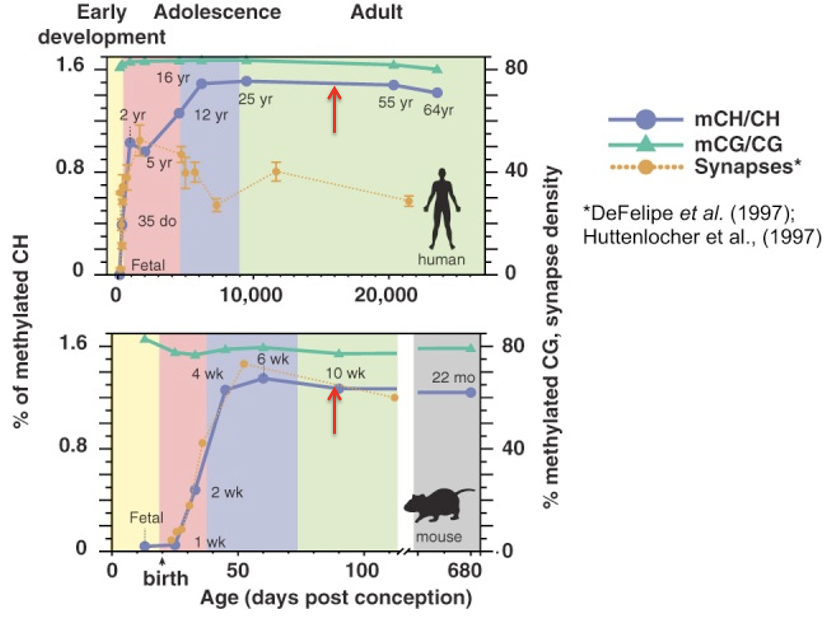
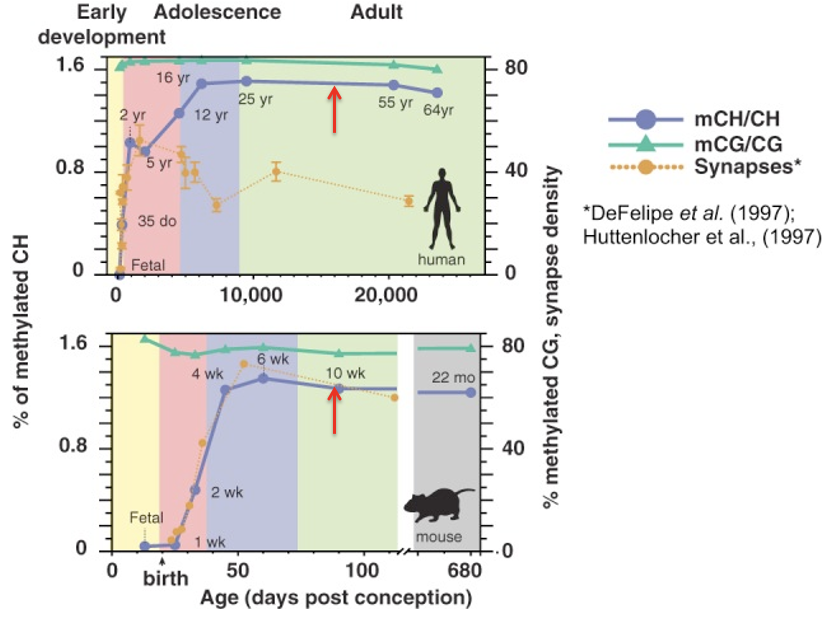
What critical neurodevelopmental process is represented by the dotted yellow line in these lifespan figures?
Synapse density
It increases rapidly during early development and then declines as the brain refines its circuitry through synaptic pruning
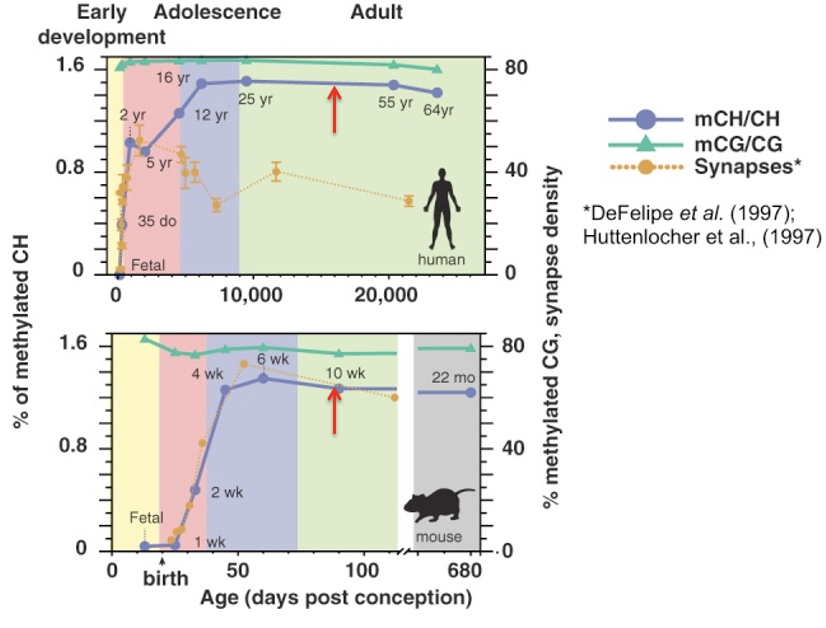
Why is the timing of the sharp rise in mCH biologically significant in the brain?
It synchronizes w the periods when neural circuits are being built and refined
The accumulation of mCH as a molecular marker for the maturation and stabilization of these circuits
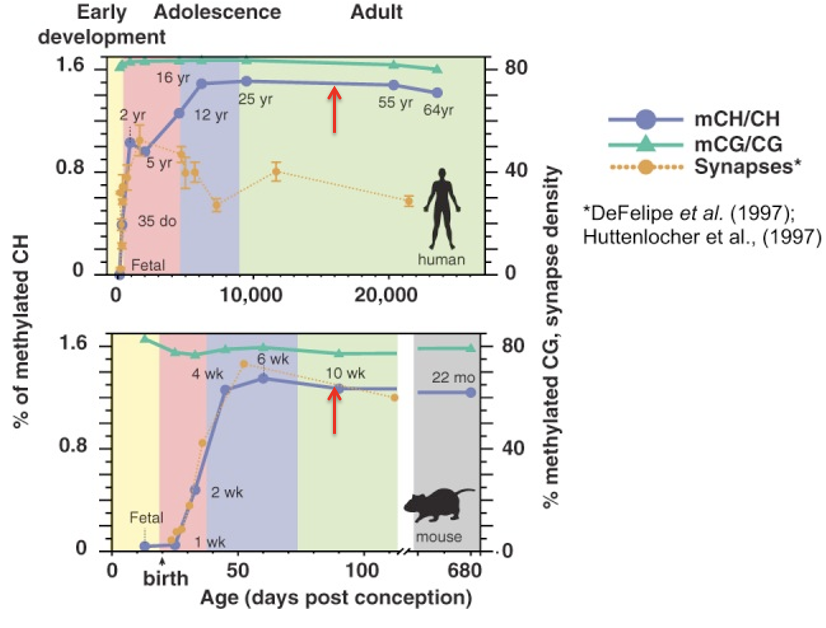
In the context of human brain maturation, how is the decline in synapse density interpreted?
As normal developmental modelling
Synapses are selectively stabilized or eliminated to fine tune neural networks for adult function
Describe the relationship between mCH accumulation and synaptic pruning
mCH rises sharply while the brain is most active in circuit refinement. As synapses are pruned and networks are stabilized, mCH levels eventually reach a steady plateau.
What is the specific biochemical precursor for 5-hydroxymethylcytosine (5hmC)?
5mC
5hmC is generated by modifying an existing 5mC mark, rather than being added to an unmodified cytosine
Which family of enzymes is responsible for the oxidation of 5mC into 5hmC?
The TET family of dioxygenases
In which specific neuronal population is 5hmC strikingly abundant, reaching roughly 40% of the level of 5mC?
Cerebellar Purkinje Cells
Contrast the typical genomic localization and functional association of 5hmC vs 5mC in the brain
5mC - transcriptional silencing at promoters/enhancers
5hmC - enriched over the gene bodies of active genes and positively associated with transcriptional levels
How do TET enzymes facilitate the conversion of 5mC, and what are the resulting oxidized derivatives?
5mC → 5hmC
5hmC → 5fC
5fC → 5caC
significance: this pw is a key mechanism for active dna demethylation
5fC and 5caC can be recognized and excised by TDG and replaced w unmethylated cytosine via base excision repair pw
Intergenic CG methylation vs mCH/5hmC (in stability and timing in the NS)
Intergenic CG - widespread and stable, involved in long-term genome regulation
mCH and 5hmC - cell type and time dependent, accumulate specifically during brain development and circuit refinement
Roles of mCG in intergenic regions
maintaining repression of repetitive elements
supporting stable cell identity
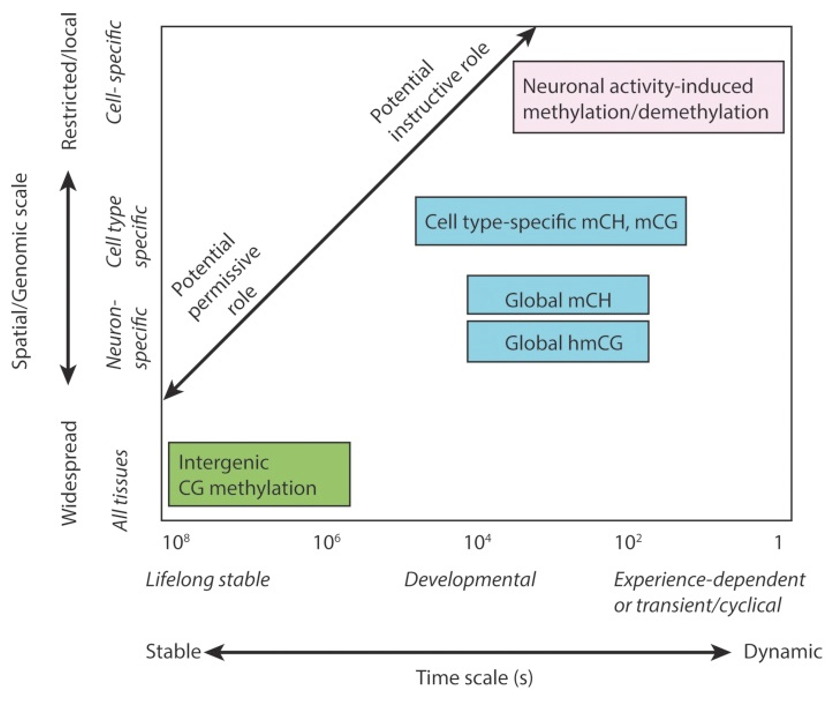
According to the "Big Picture" schematic, what is the relationship between methylation patterns and neurons?
Methylation patterns are strongly linked to neuronal identity and activity, though their exact functional roles are highly cell-specific
What is the primary functional domain in MeCP2 that allows it to target methylated cytosines?
methyl-CpG binding domain MBD
Structurally, how do the MeCP2-e1 and MeCP2-e2 splice isoforms differ from one another?
They differ at their N-terminus, but they both retain the essential binding and regulatory domains
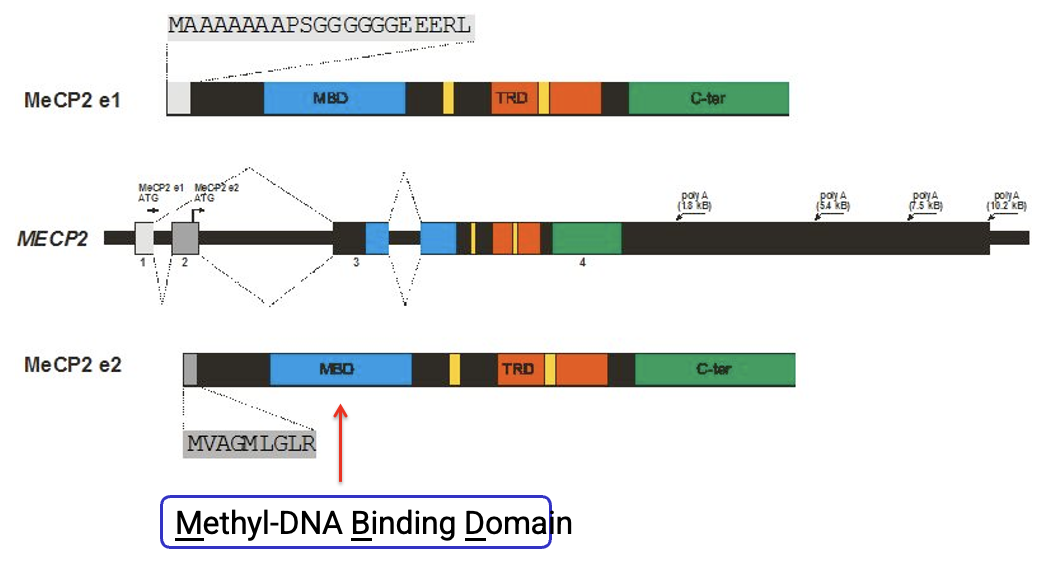
Why are two different protein "versions" of MeCP2 mapped to the same single gene?
Due to alternative splicing, which allows one gene to produce multiple isoforms (e1 and e2) with distinct N-terminal sequences
On which chr and specific locus is the MeCP2 gene located?
X-chromosome, tip of the long arm, specifically Xq28
What percentage of Rett syndrome cases are caused by de novo loss-of-function mutations in MeCP2?
95% of cases. Most arise as new mutations rather than being inherited from parents