BS2092 - Intracellular Compartments and Protein Sorting
1/45
Earn XP
Description and Tags
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
46 Terms
Define co-translational insertion
the process where ER-bound ribosomes synthesise proteins and simultaneously translocate them into the ER
What is required for protein sorting to the correct membrane?
a signal, intrinsic to the protein
a receptor that recognises the signal and directs it to the correct membrane
translocation machinery
energy to transfer the protein to its new place
Signal for nucleus importation
a sequence of 5 consecutive positively charged amino acids anywhere in the protein
Signal for mitochondria importation
positively charged amino acids spread out by 2-3 other amino acids
Signal for ER importation
a stretch of between 5-12 hydrophobic amino acids present at the N-terminus of the protein (within the N-terminal 16-35 residues)
What are the 3 positively charged amino acids
Lysine (Lys/K)
Arginine (Arg/R)
Histidine (His/H)
What is TOM (translocase outer membrane) complex
the receptor that recognises the matrix protein
What does TOM do
binds to the signal sequence
delivers protein through a protein conducting channel in the complex
What is the TIM23 (Translocase of the Inner Membrane 23) complex
a mitochondrial protein machine that transports proteins across the inner membrane into the matrix
What happens to the signal sequence after the protein has been successfully delivered into the matrix
it is cleaved
Role of cytosolic hsp70 chaperone proteins
keep the protein unfolded to be delivered through the protein channel into the matrix
Role of mitochondrial hsp70 chaperone proteins
binds to the protein as it enters the matrix
causes a conformational change in the import ATPase (with aid of ATP hydrolysis) which helps pull the matrix protein into the matrix
What does NPC stand for
nuclear pore complexes
How many NPCs in a typical cell
3000-4000
What is an NPC
an aqueous pore lined by proteins with extended conformation, which facilitates the bidirectional transport of molecules in and out of the nucleus
Purpose of the proteins lining the pore
they act as a sieve/meshwork to block diffusion of large molecules
How do larger molecules move across the nuclear envelope
active transport
How does nuclear-cytoplasmic transport work
transported molecules contain sorting signals
importins and exportins (nuclear transport receptors) recognise the signals and carry the molecules through the pore
proteins are transported against their concentration gradient using energy from GTP hydrolysis
Examples of proteins which undergo nuclear import
structural proteins of the nucleus
histones, nuclear lamins
DNA/RNA polymerases
ribosomal proteins
gene regulators
transcription factors, chromatin modifying enzymes
What regulates nuclear import and export
Ran
Features of Ran
involved in signal transduction
acts as on/off molecular switches
activity is controlled by their GTP/GDP bound status
How does their GTP/GDP bound status control the activity of Ran
GDP-bound = inactive
GTP-bound = active
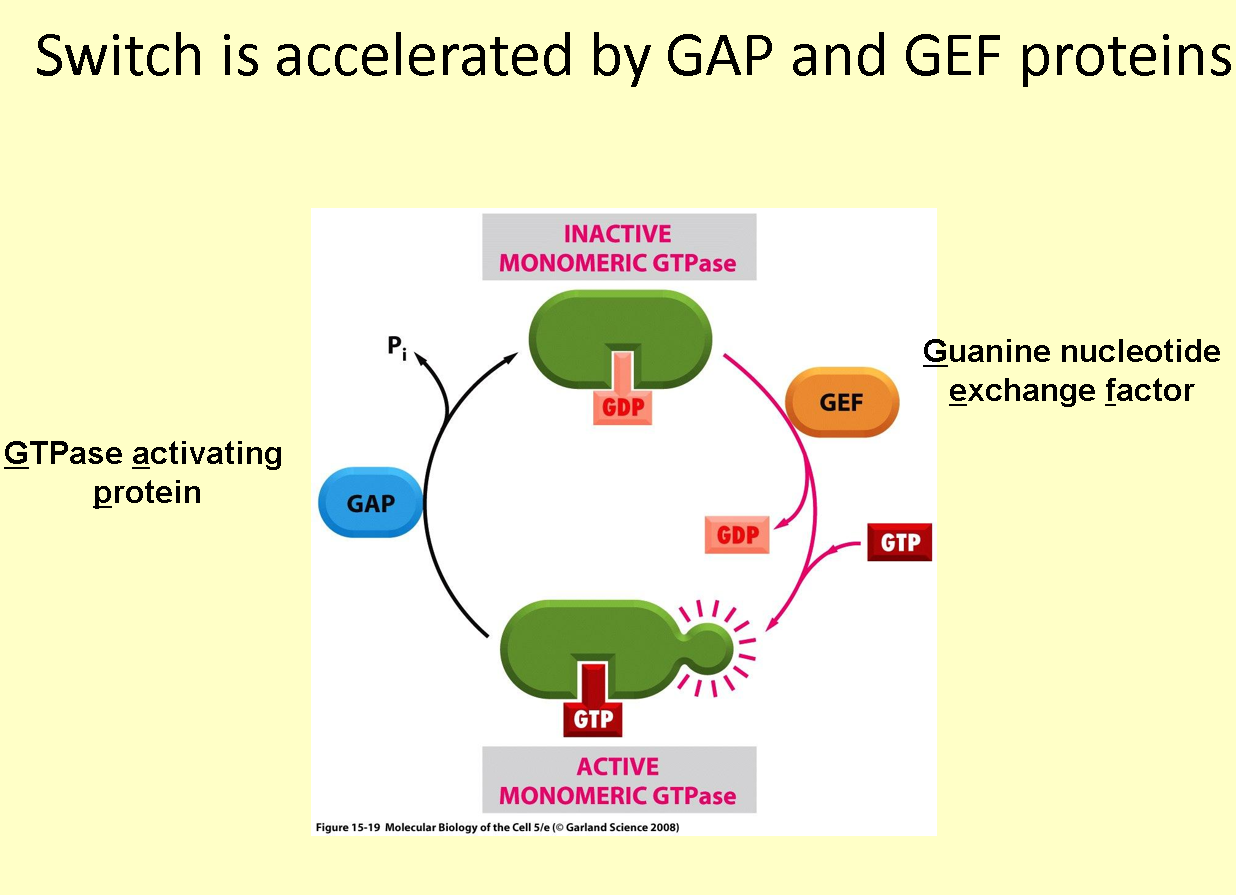
What catalyses the activation of Ran
Guanine nucleotide exchange factor (GEF)
How does GEF catalyse the activation of Ran
binds to Ran, stimulating the release of GDP
GTP binds in place of GDP, due to a higher concentration of GTP in the nucleus
Ran-GTPase intrinsic activity
long enough time allows it to hydrolyse GTP to GDP itself
The effect of GAP (GTPase activating protein) on Ran-GTPase intrinsic activity
accelerates the hydrolysis of GTP to GDP to switch off Ran
Which types of Ran are more abundant in the nucleus and in the cytoplasm
Ran-GTP is more abundant in the nucleus
Ran-GDP is more abundant in the cytoplasm
How does Ran allow importins/exportins to sense whether they are in the nucleus or cytoplasm
Ran-GAP and Ran-GEF are tethered in the cytoplasm and nucleus, respectively
creating a gradient so that Ran-GDP is predominantly in the cytoplasm and Ran-GTP is predominantly in the nucleus
What is Ran-GAP tethered to in the cytoplasm?
cytoplasmic fibrils
What is Ran-GEF tethered to in the nucleus
chromatin
How does Ran act as a molecular switch for importins
Ran-GTP binds to importins, changing their confirmation to allow cargo release
Ran-GDP dissociated from importins, changing their conformation to cause cargo binding
Why can importins change their conformation
they consist of multiple stacked α-helices that act like a flexible spring-like structure
How does Ran-GTP binding allow for cargo release
binding of Ran-GTP moves a loop, forcing the release of cargo
Nuclear import summary in cytoplasm
Ran-GDP predominates
Ran-GDP does not bind importin
Free importin can bind cargo
Nuclear import summary in nucleus
Ran-GTP predominates
Ran-GTP binds to importin
Conformational change displaces cargo
What is the SRP (signal recognition particle)
a multidomain riboprotein that mediates a 3-way association with its receptor in the ER, the ribosome and the signal peptide
Pathway of SRP-mediated delivery of ribosomes to ER membrane
as the protein is synthesised it is immediately recognised by the SRP
this stops further translation occuring so that the protein is not produced within the cytosol
SRP then binds to its receptor in the ER (this receptor is in close association with the protein translocator in the ER membrane)
Once the SRP binds to its receptor, together with its ribosome, ER protein synthesis occurs
Pathway of SRP-mediated delivery of ribosomes to ER membrane (part 2)
After SRP is bound to its receptor, the ribosome is placed on top of the protein translocator, causing it to open
GTP hydrolysis occurs from both subunits of the SRP receptor to provide the energy required
protein synthesis carries on, proteins are translocated through the translocator until the fully synthesized protein is delivered into the ER lumen
the signal peptide is cleaved when it protrudes through the membrane by the signal peptidase
Comparison of Targeting to different organelles
Organelle:
ER
Mitochondrial matrix
Nucleus
Signal:
Hydrophobic signal sequence (N-term)
amphipathic helix (N-term)
short stretch Arg/Lys (anywhere)
Signal removed?
Yes
Yes
No
Receptor:
SRP/SRP Receptor
TOM Complex receptor
importin
Translation machinery:
Sec61 protein translocator
TIM/TOM complex
nuclear pore complex
Energy source:
GTP hydrolysis
ATP hydrolysis and membrane potential
GTP hydrolysis
What types of ER membrane proteins insertion involves SRP
Types I-IV
Type I ER membrane protein
N-terminus in ER lumen
C-terminus in cytosol
N-terminus signal sequence is cleaved during insertion
Type II ER membrane protein
N-terminus in cytosol
C-terminus in ER lumen
anchored by a signal-anchor sequence close to the N-terminus
Type III ER membrane protein
N-terminus in ER lumen
C-terminus in cytosol
anchored by a signal-anchor sequence close to the N-terminus
Type IV ER membrane protein
multi-pass proteins which span the membrane multiple times
divided into subtypes IV-A (N-terminus in cytosol/C-terminus in ER lumen) and IV-B (N-terminus in ER lumen/C-terminus in cytosol)
contains multiple signal-anchor/stop-transfer sequences that allow them to span the bilayer several times
Mechanism of synthesis of Type I membrane protein at the ER
similar pathway to the soluble example until a 2nd hydrophobic sequence is reached
when the 2nd hydrophobic sequence is reached, translocation is stopped in the lumen and the protein is anchored into the membrane
the rest of the protein synthesis continues in the cytosol until a STOP codon is reached to form the C-terminal
Insertion of Type II membrane protein into ER membrane
SRP recognises an internal hydrophobic stretch of amino acid
protein is delivered into membrane by SRP
protein synthesis continues and the protein is inserted such that its N-terminus is in the cytosol and C-terminus in the ER lumen