NE102 Midterm 2 Study Guide: RNA and Macrobiology
1/103
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
104 Terms
What is RNA?
single stranded protein copied from DNA that can be used in the cytosol
What is messenger RNA (mRNA)?
transcribed from DNA, and helps with translation to proteins
What do the different types of RNA (tRNA, mRNA, rRNA) do broadly?
they regulate the presence (expression) of proteins in a cell
What are the three main differences between RNA and DNA?
RNA has an OH group (ribose), while DNA only has an H group (hence, deoxyribose)
RNA uses uracil (U), while DNA uses thymine (T)
RNA is single-stranded, while DNA is double-stranded
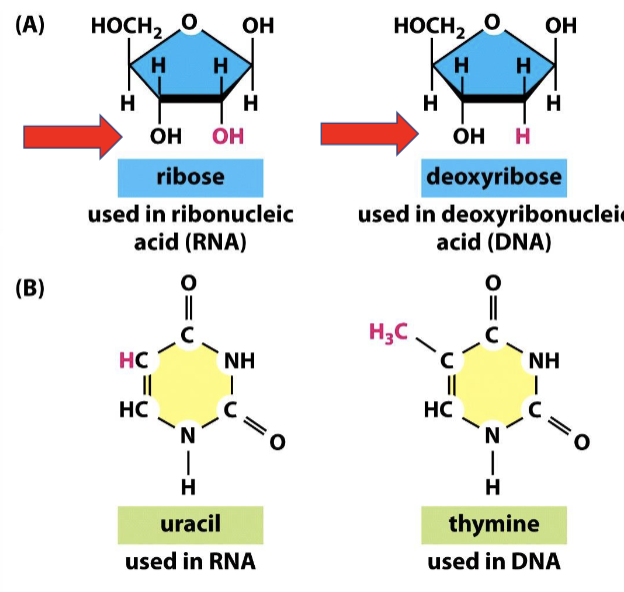
How are RNA and DNA similar?
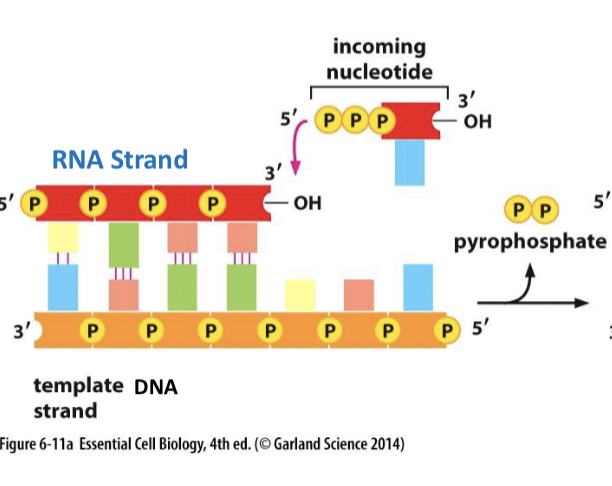
they use the same phosphodiester bonds, are transcribed from 5’ to 3’, and are nucleic acids made up of nucleotides
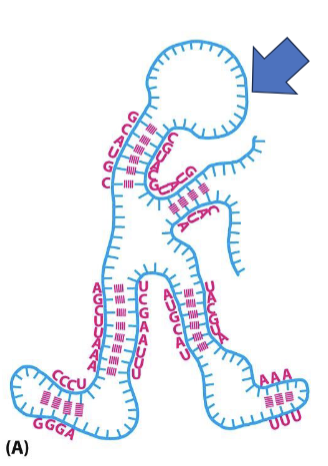
What are the advantages of RNA as a single-stranded molecule?
can easily fold in different confirmations
different RNA molecules can make interactions via H bonds with complementarity, as seen during protein translation (codons/anticodons)
What is RNA polymerase?
transcribes genetic information into RNA. it does NOT need primers, helicase, topoisomerase, and copies only selected DNA (genes)
What makes transcription specific, and not just chance, as RNA polymerase collides with RNA all the time?
promoters (TATA box) and transcription factors allow for specificity in trancription
What do promoters do?
they are needed for RNA polymerase to “recognize” the gene (at the 5’ side) to be transcribed (promoter is NOT copied)
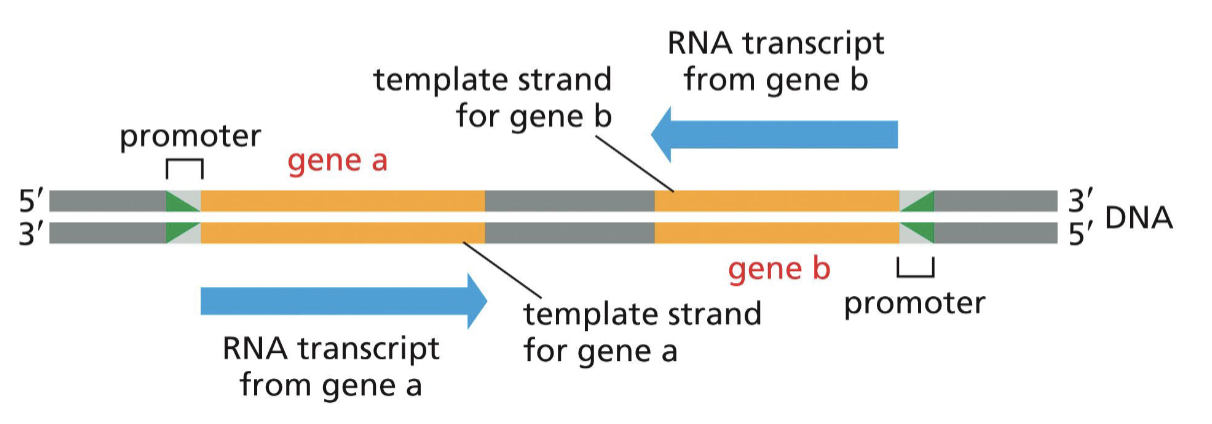
What do terminators do?
they are needed for RNA polymerase to “recognize” when to stop trancription
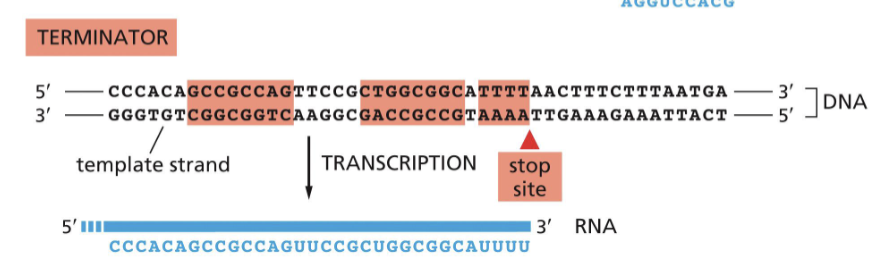
What is a gene?
a sequence of nucleotides on one DNA strand that is transcribed into an RNA molecule
What is transcription?
rewriting DNA to match the RNA language (T —> U)

What is special about the TATA box as a promoter?
the T and A regions only have two hydrogen bonds, which are easier to pull than the C and G which have three
What are transcription factors broadly?
they help RNA bind to the promoter, RNA polymerase bind to the DNA, and generally prepare for transcription
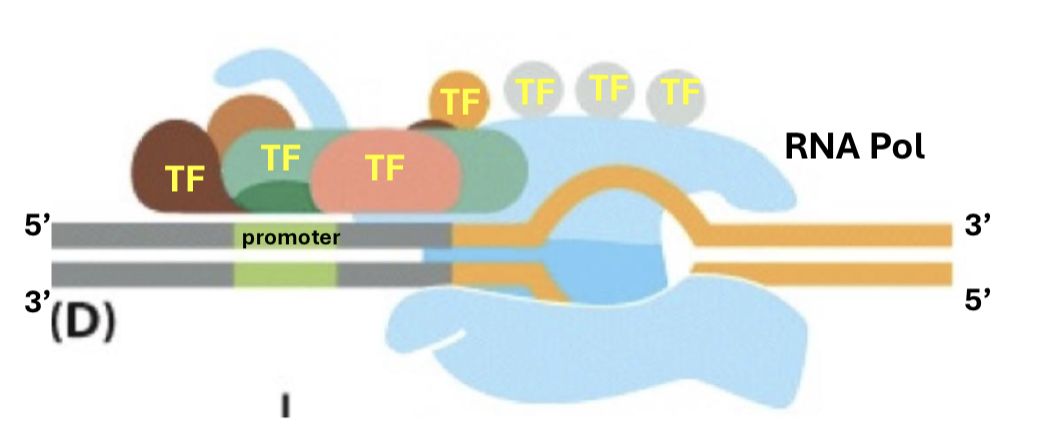
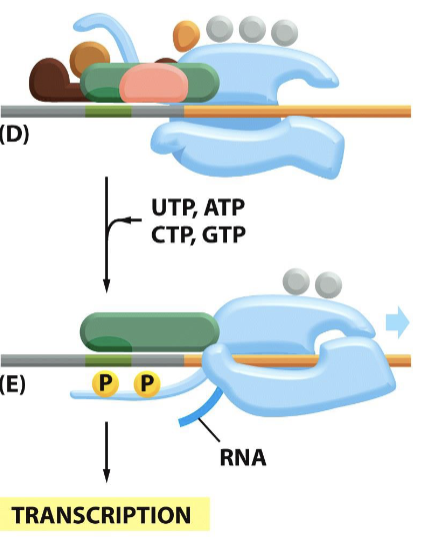
What are the specific steps of transcription?
TFIIB binds to TFIID and the double helix is “bent” to access DNA
TFIIE facilitates the assembly of TFIIH, which unwinds DNA and phosphorylates it for activation
the newly bent DNA orientation signals TFIIF to bind to RNA polymerase, optimizing recognition and binding to the TATA box
transcription factors are detached from RNA polymerase, and transcription begins
What is mRNA processing?
allows the newly transcribed “micro” segments of RNA to leave the nucleus with only the coding segments (exons)
What are the three modifications made to mRNA?
capping, polyadenylation, and splicing
What is the process of mRNA phosporylation?
negative charges are brought to the tail, which changes the confirmation of “loch ness monster” and attracts RNA-modifying molecules to cap, splice, and add poly-A tail
What is the process of mRNA capping?
while RNA starts building its mRNA, the cap is added right away to facilitate transport to the cytosol and ensure correct translation
What is the general process of mRNA splicing?
exon coding regions are extracted from eukaryotic gene and introns selectively removed using specific sequences of nucleotides
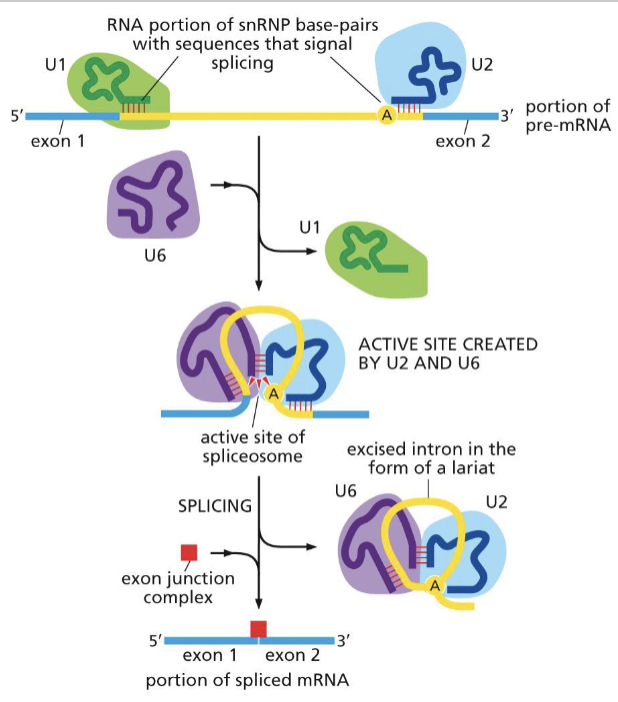
What is the specific process of mRNA splicing?
semi-lariots are made w/ introns adjoining in loops (5’ binding to adenosine residual end)
spliceosomes (snRNA + snRNP) tag the beginning and end of introns for removal
catalyzes intron U2 + U6 to excise intron in lariat
exon junction complex marks the adjoining of two exons
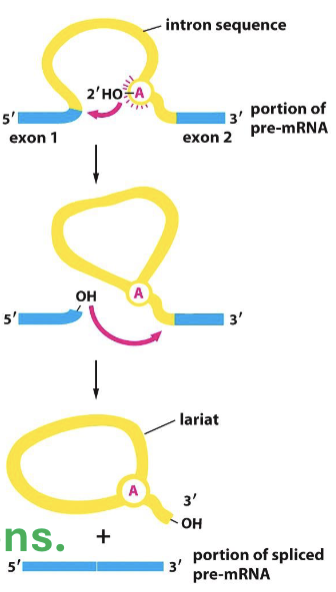
What do spliceosomes do?
the snRNA and snRNPs tag the beginning and end of introns for removal and U6 and U2 excise lariat
What is the process of mRNA polyadenylation?
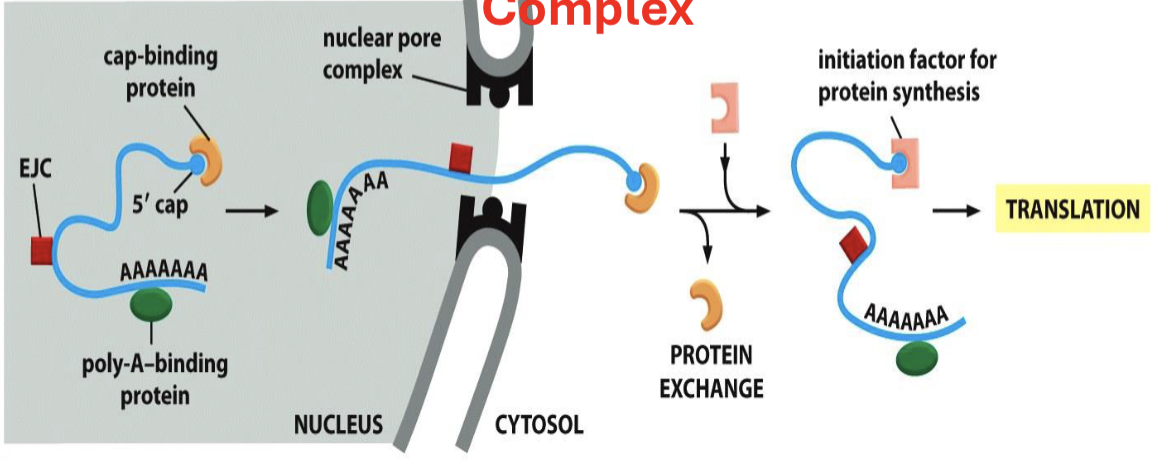
after the entire mRNA molecule is synthesized, a poly-A tail is added to the 3’ end, protecting the mRNA from degradation and aiding in nuclear export
What happens directly after the mRNA is transported out of the nucleus?
the cap-protein is replaced with the initiation factor to confirm correctness
What is the difference in molecular conversion in transcription and translation?
transcription: nucleic acid to nucleic acid
translation: nucleic acid to amino acid
How does translation occur broadly speaking?
mRNA binds to ribosome, which moves along the mRNA as it adds amino acids to the chain in the large ribosomal subunit
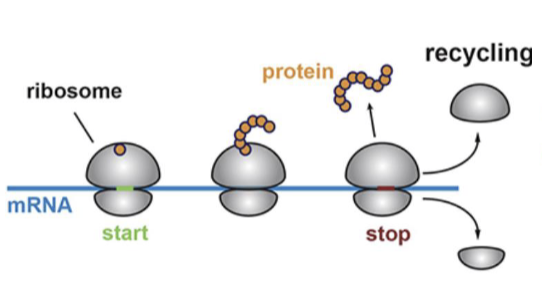
What are ribosomes comprised of?
rRNA
How is nucleotide genetic information turned into a protein?
sequences of 3 nucleotides in the mRNA that code for one amino acid “codon”
How many codons are there?
64 codons, but only 61 used to encode for amino acids
What is tRNA?
speaks both “languages” to catalyze translation. using its anticodon, tRNA binds to the correct complimentary codon to form the amino acid chain from mRNA
Why is RNA very good at folding?
only 4 options for complementary nucleotides to form H bonds (AUCG)
easy to anneal to other proteins as single strand
Which end of the tRNA molecule holds the amino acid?
amino acid - 3’ end
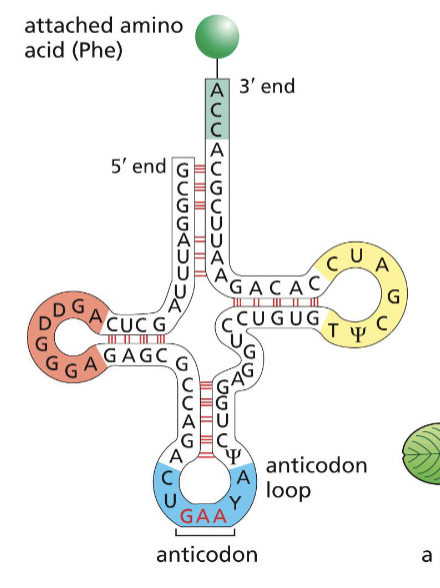
What is the direction of protein synthesis?
from the N terminal of the amino group to the C terminal of the carboxyl chain
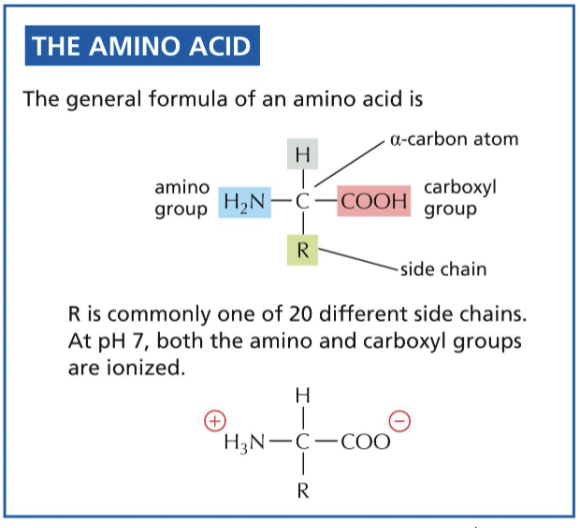
What is the anticodon?
complementary strand to the codon, found on tRNA, to interact with the mRNA for synthesis to occur
What is aminoacyl-tRNA synthetase?
can “host” in its structure one amino acid and one tRNA (based on anticodon), and bonds them together with ATP
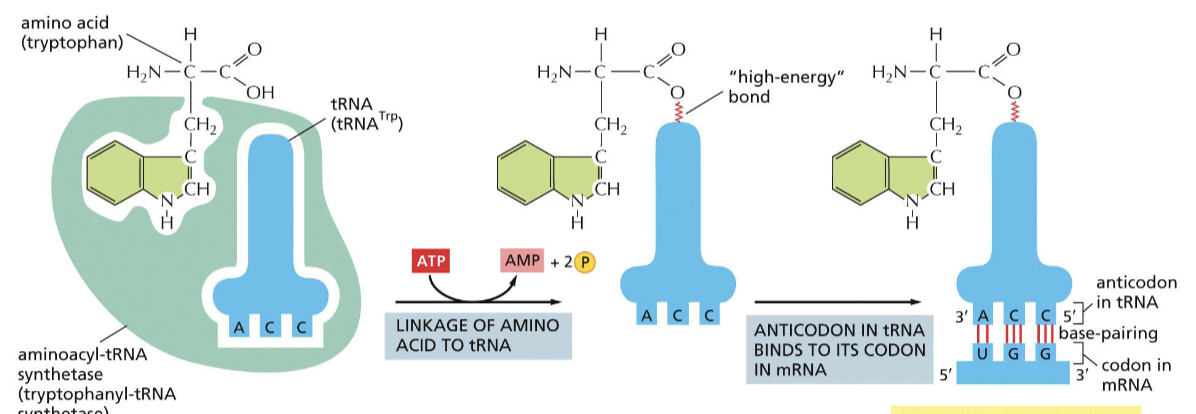
How can one tRNA read multiple codons?
wobbling, which allows for mismatching on the 3rd position in some codons
What is the structure of the ribosome?
rRNA is the core, forming the A, P, and E sites, and it is surrounded by proteins
What is the first codon that is translated?
AUG
How does translation occur along the ribosome?
translation begins when a Met tRNA, using translation initiation factors, lands at the 5’ site of mRNA and starts searching for the AUG codon
when it finds AUG, the reading frame begins, translation initiation factors dissociate, and the large ribosomal subunit binds to the P site
tRNA starts at the P site with the ribosome, and mRNA, starting from the A site, is dragged along the reading frame until exit at the E site
How does the polypeptide chain grow during translation?
newly bound complementary charged tRNA enters at the A site codon, and is bound to the P site tRNA with peptidyl transferase
the ribosome translocates, the P site tRNA is moved to E site and ejected, detaching from its amino acid in the chain
after the ribosome shifts, the A site is now free and a new complementary tRNA is found and added
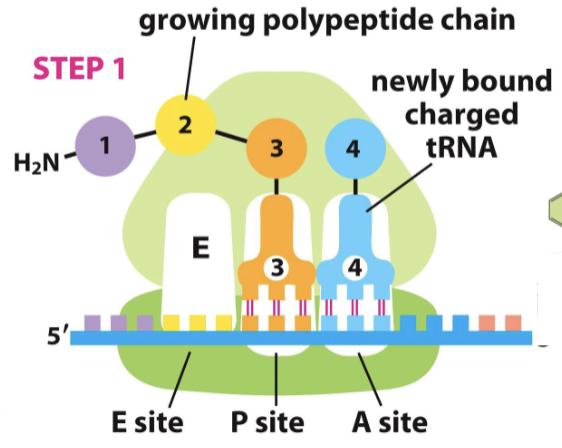
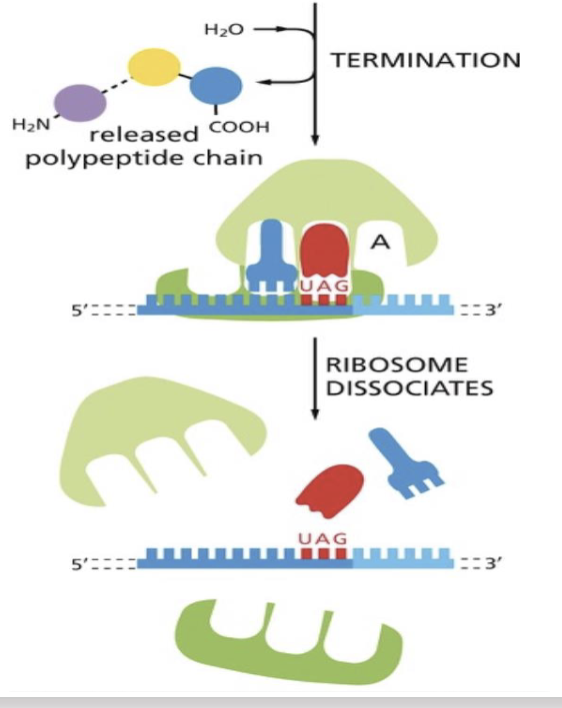
What happens when mRNA to amino acid translation ends?
when tRNA reaches stop codons, it cannot recognize them, and translation stops
release factor changes the activity of peptidyl transferase
one H2O molecule is added, releasing the amino acid chain from the tRNA
What are the post-translational modifications?
phosphorylation and acetylation
What is the general process of phosphorylation?
addition of a phosphate group (negative charge) when protein is fully folded, helping to attract other proteins and stopping transcription factors. can only happen on the Ser, Thr, or Tyr amino acids.
What is the specific process of phosphorylation?
protein kinase “picks up” on a target substrate that can be phosphorylated with ATP.
What allows for the removal of phosphates from a protein’s tail?
phosphotase proteins
What is the process of acetylation?
adding an acetyl group (negative) to the tails of nucleosomes unwraps the DNA (negative) from being sandwiched to the histones (positive)
What are lysosomes?
lysosomes perform bulk degradation of long-lived proteins
What are proteasomes?
proteasomes can bind to proteins (with a ubiquitin), denature them, and break their bonds. structure contains a regulatory and core particle.
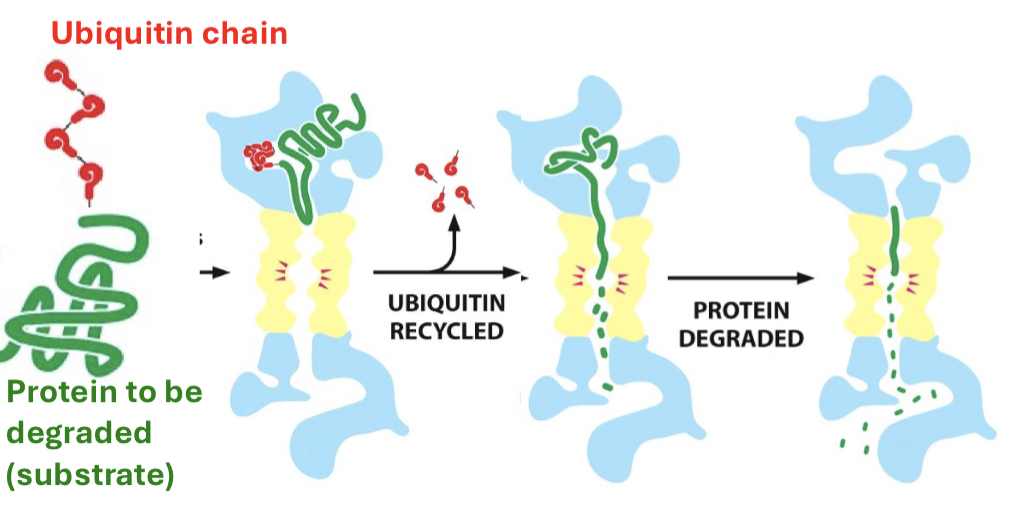
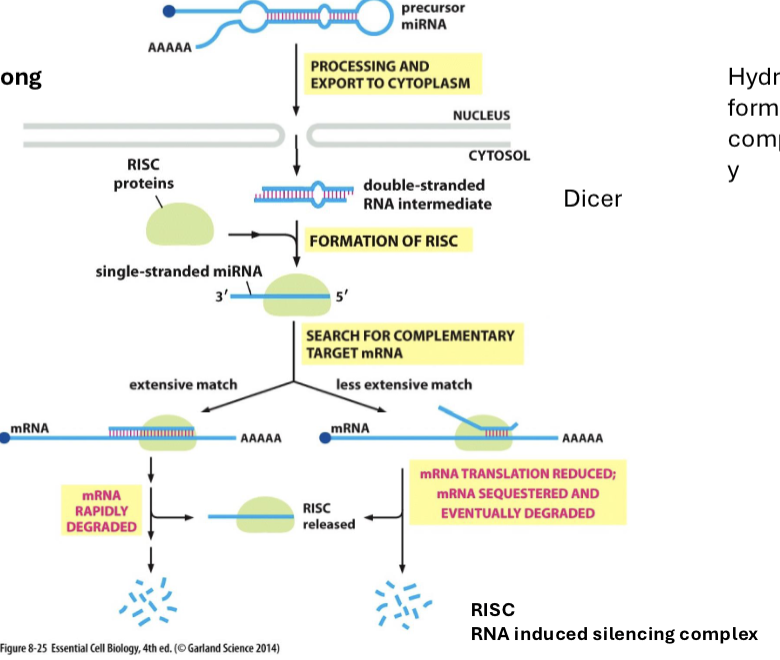
What are microRNAs?
small (22 nucleotides long) sequences of RNA that bind to RISC and help RISC find species of complementarity in the cytosol
How does transcription control cellular function and development?
although DNA is the same in all cells, the ACCESS to that DNA is not the same. only genes that encode for that gene’s function can be accessed for transcription.
What are transcription regulators?
proteins that make the region of the DNA within a gene of interest available (or not) for transcription. REGULATE the TRANSCRIPTION of SPECIFIC proteins/genes
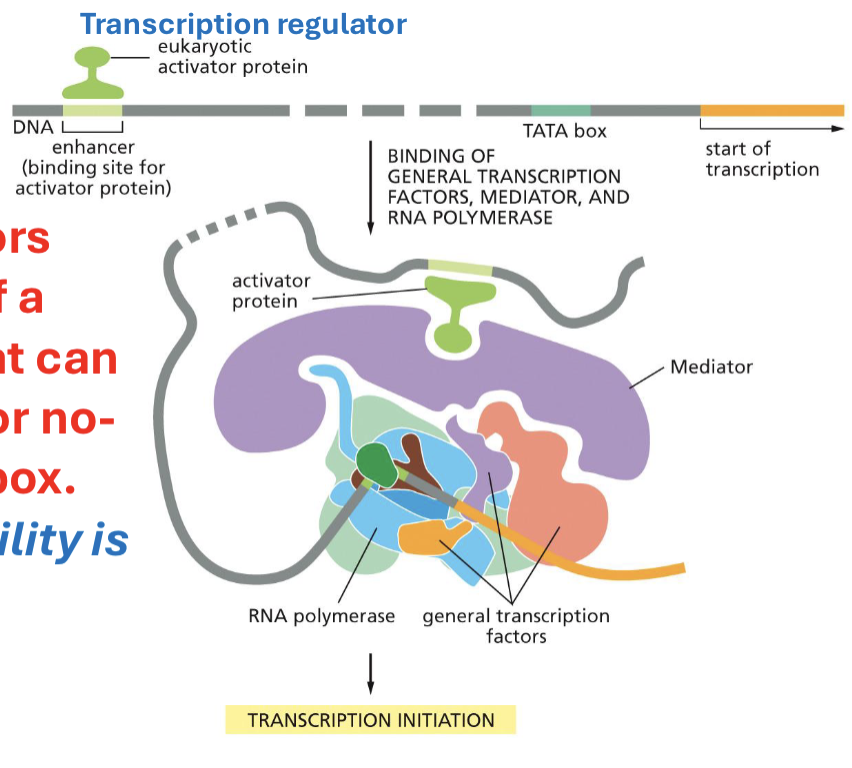
What was the transplanted nuclei and cell differentiation experiment?
Sir Gurdon generated a tadpole: the nucleus of a cell from a frog’s intestine was transplanted into the egg of a normal frog (frog generated in ABSENCE OF FERTILIZATION)
How are transcription regulators different from transcription factors?
transcription regulators regulate the transcription of specific genes/proteins and are important during development/cell differentiation
transcription factors help to initiate or inhibit transcription, acting as switches
What is transcriptional control?
regulatory DNA sequences (element) are located to turn genes on or off
when transcription regulators bind to the dedicated element, the formation of a mediator occurs
mediator complex of proteins bind to chromatin modifying enzymes as a bridge
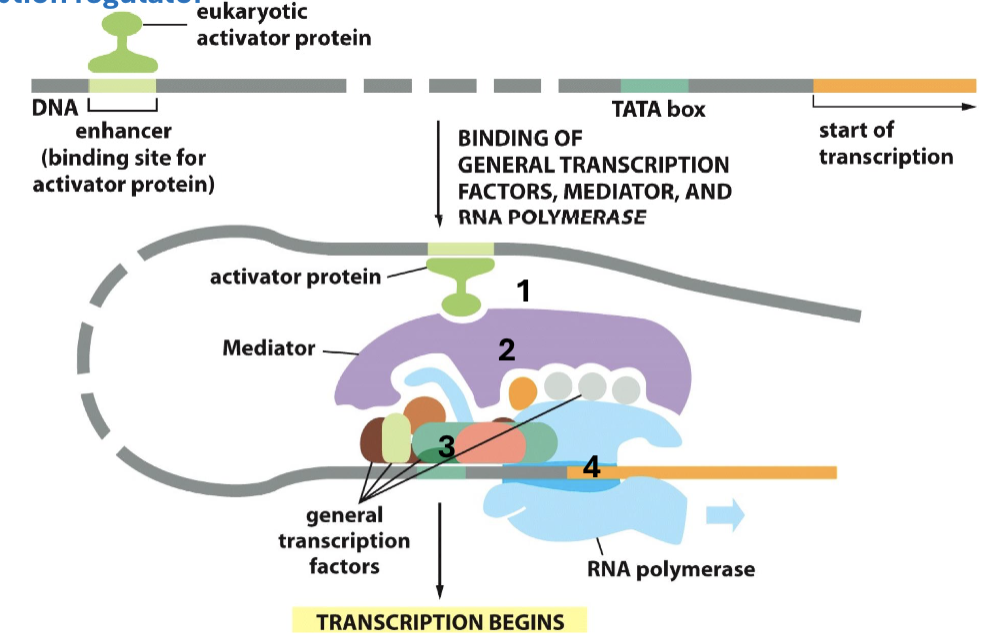
How can chromatin be tightened with mediators?
transcription regulators can bind to repressor binding sites, which also have mediator, but TURN OFF genes w/ HDAC (takes off a negative charge and tightens DNA)
How can chromatin be loosened with mediators?
transcription regulators can bind to activator binding sites which have mediator and TURN ON genes w/ HAT (adds a negative charge and loosens DNA)
What happens in Huntington’s disease?
the gene that controls the motor cortex striatum is degenerated w/ dysregulation, leading to the inhibition of BDNF (essential for neuronal integrity)
What is the relationship between REST and BDNF in Huntingtons?
In Huntington’s cells, there is more REST (a repressing transcription regulator) in the nuclei, which means that there is less transcription of the BDNF gene. LESS BDNF —> less neuronal integrity
What is cell differentiation?
the process by which stem cells and controlled gene expression allow a cell to specialize with specific transcription regulators (e.g., muscle cells, fat cells, neurons)
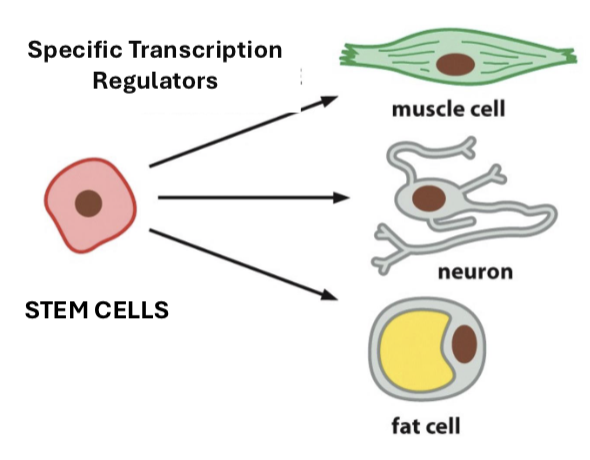
What are stem cells?
very early cells with no specific function yet
What are the three important stages of embryonal development?
zygote —> blastocyst —> gastrula (with ecto, meso, endoderm)
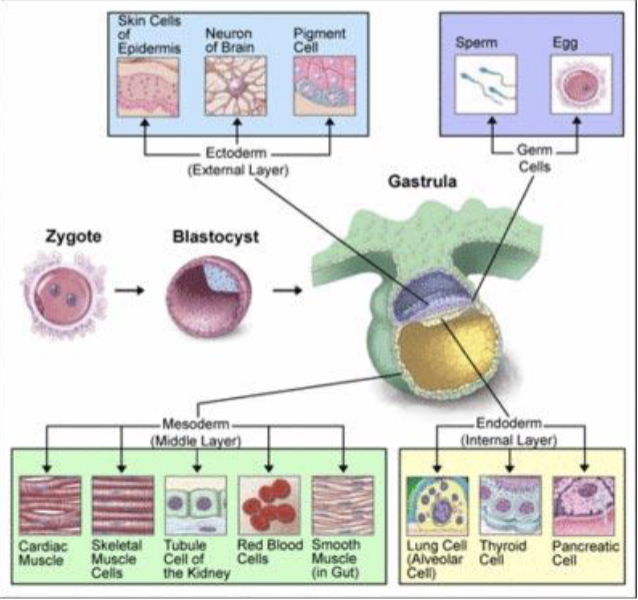
What are the four types of stem cells and their abilities to differentiate?
totipotent - can become embryo and placenta
pluripotent - can become embryo (not placenta)
multipotent - can become endo/meso/ectoderm
non-embryonic stem cells (in adults) - organ repair
What is combinatorial control?
during the first stages of development, few transcription regulatory proteins are used to control protein and gene expression for developing cells (cascading umbrella with increasing complexity)
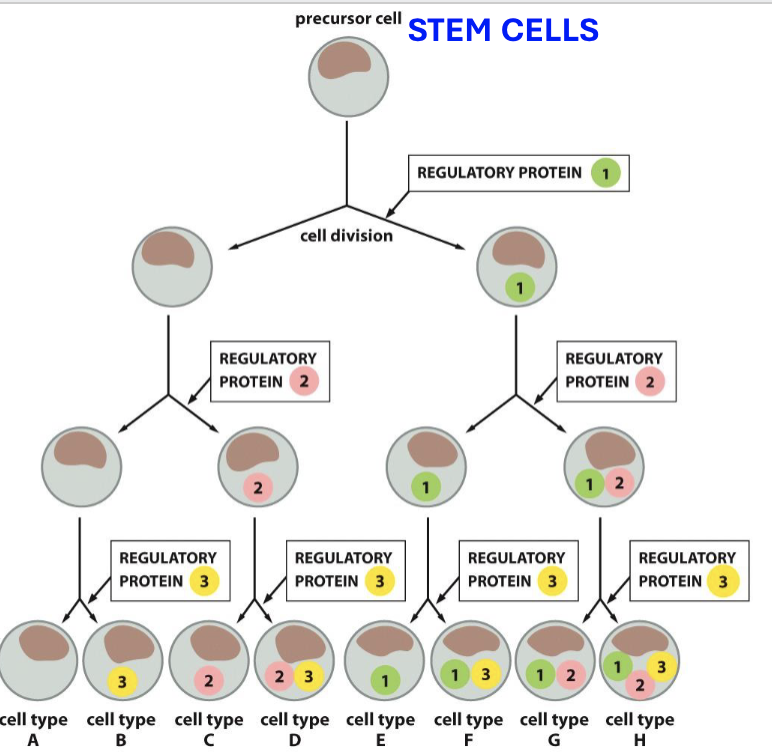
What is the zygote?
fertilized egg which differentiates to a blastocyst using transcription factors
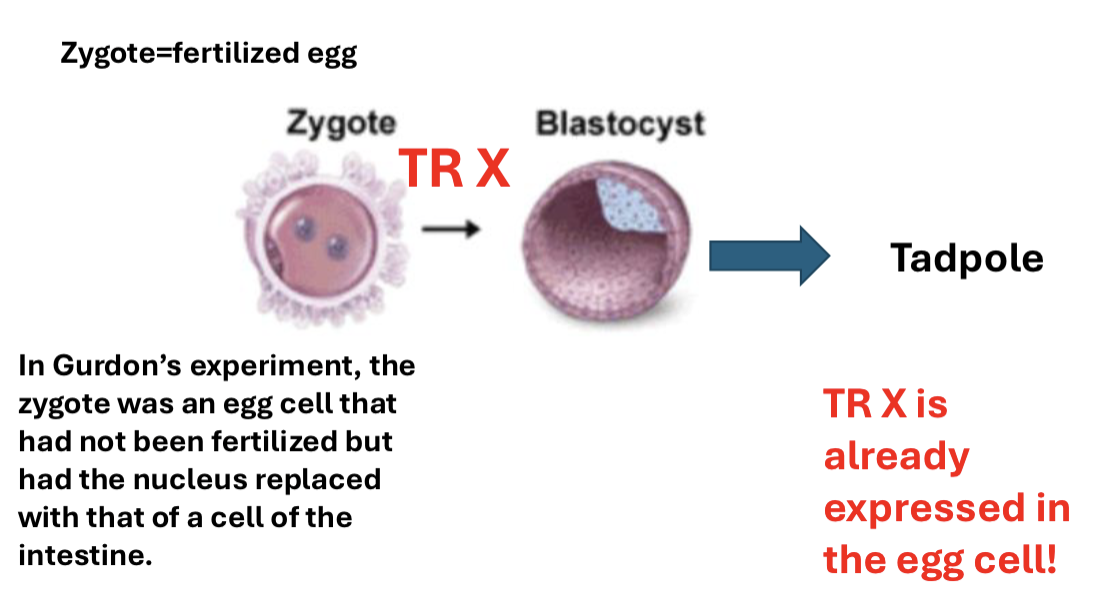
What is cell memory?
once cells differentiate, they can aid in the process of new cell differentiation with already known transcription factors forming the positive feedback loop
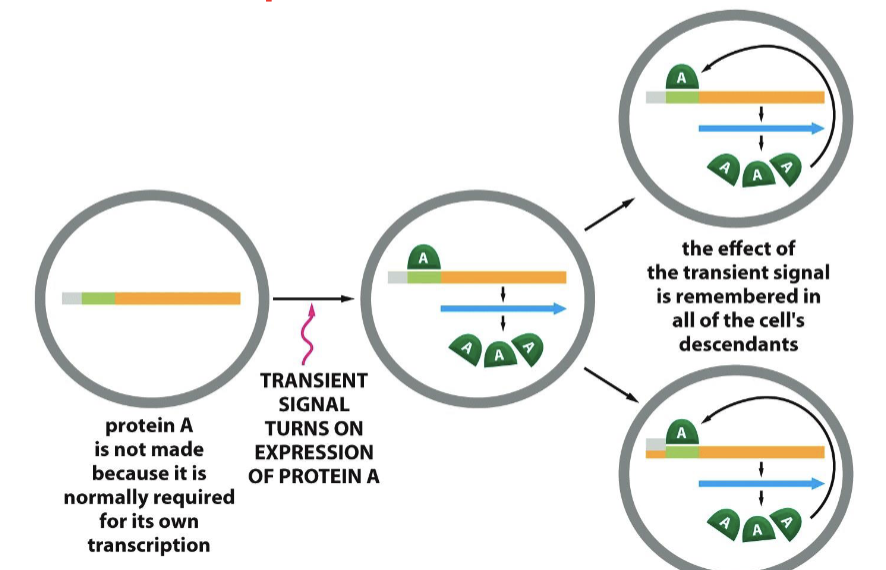
Can we turn differentiated cells to stem cells?
yes, with induced pluripotent stem cells (iPSCs), using cell memory. in 2012, the nobel prize was awarded to the discovery of three transcription regulators that reversed fibroblasts into iPSCs.
What is the current law on embryonic handling?
14 days, around when gastrulation and cell differentiation begins, is the maximum that embryos can be studied
What are organoids?
once converted back to stem cells, iPSCs can be reprogrammed back into desired organs using specific transcription regulators to study diseases or personalize medicine
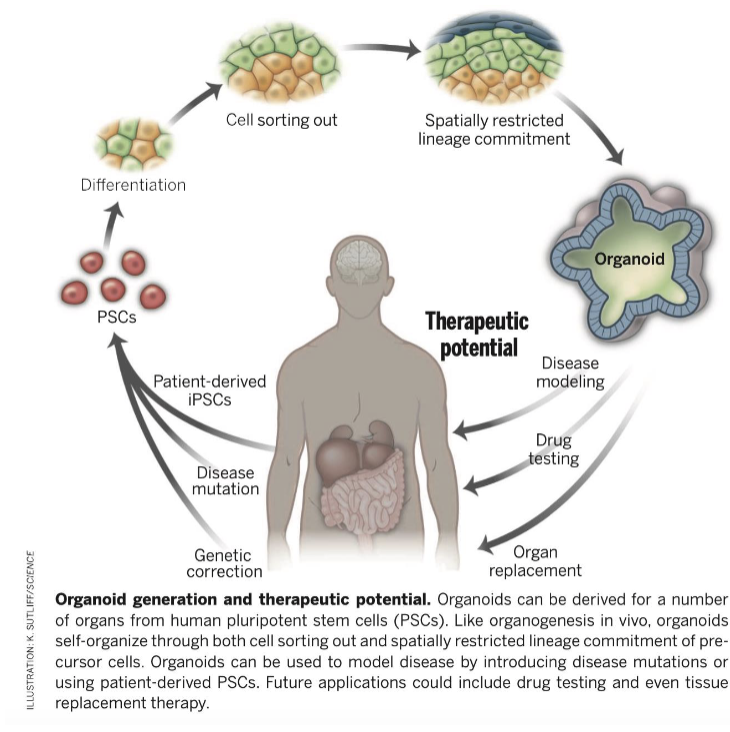
What are brain organoids?
the iPSC cells differentiate and organize in layers similar to the brain using neural induction
What are some applications for the use of organoids?
to find effect of cell oxidation/aging
can study diseased organoids (Alzheimers)
drug screening + testing
autologous and allogenic cell reprogramming + implantation
What are autologous and allogenic cell therapy cells?
autologous - iPSCs using differentiation of patient’s own cells (lowered chance of immune rejection)
allogenic - reprogrammed iPSCs from a donor with healthy cells (inserted cells without Parkinson’s disease)
How do cells get passed down by generation?
the information in germ line cells will be passed on to the next generations, defining their somatic and germ cells
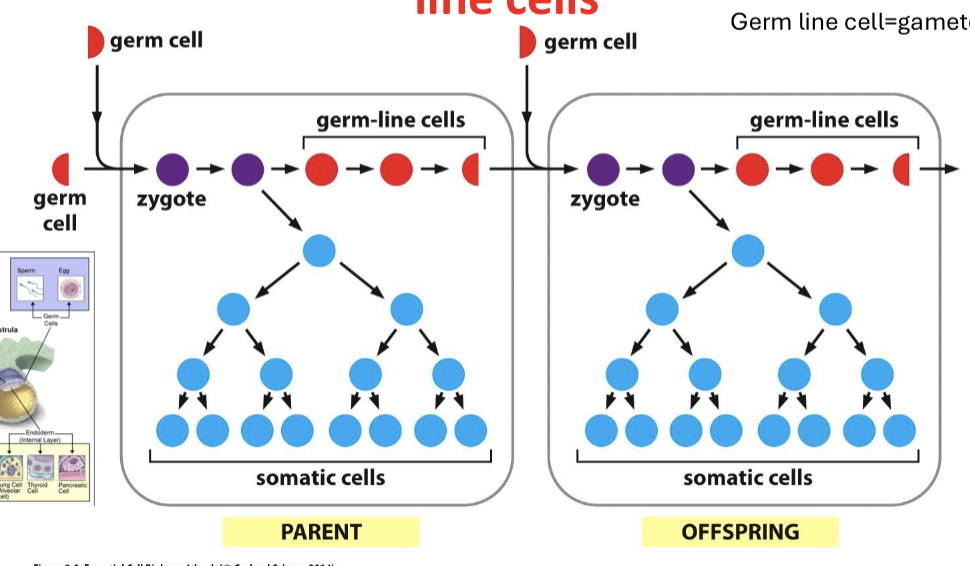
How do mutations affect the next generation of daughter cells?
if there is a mutation in a germ line cell, it can continue as duplication continues and change the genome. however, is there is a mutation in a somatic cell, it will not be passed on.
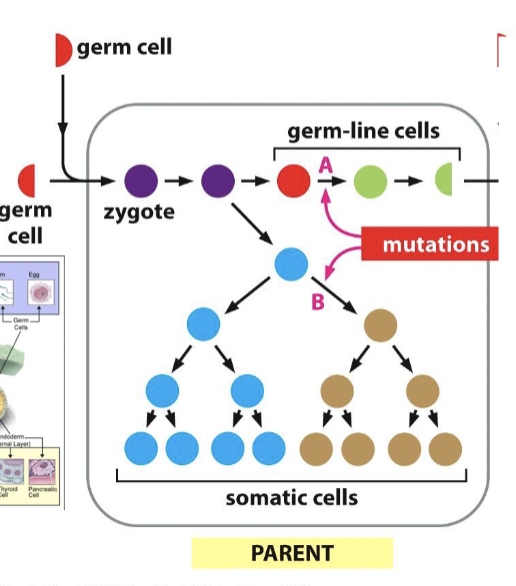
What are two ways new genes can be formed from mutations?
single nucleotide changes in the DNA (point mutations)
exon shuffling or other rearrangements of DNA sequences
How do point mutations occur in a gene?
the presence of SINGLE nucleotide changes in exons, introns, or regulatory regions of DNA
What are the implications of an edited gene on DNA?
changes in genes will change the codons of RNA, which CAN change the amino acids coded
this can be toxic (incorrect proteins made) or benign (lactose tolerance)
What is CRISPR?
clustered regularly interspaced short palindromic repeats, which are regions found in bacterial DNA and allow us to edit the genome
What are the CRISPR sequences useful for in bacteria?
CRISP sequences are designed to integrate genetic material from viruses into bacterial DNA, acting as its “immune system”
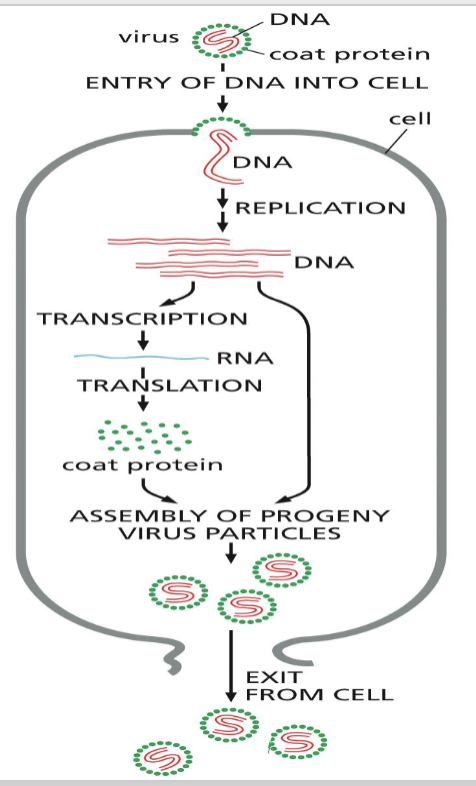
How do viruses invade prokaryotic cells?
once bacteria have been infected with viruses:
viral DNA is transcribed and viral proteins are made
viral DNA is also copied by DNA polymerase and viruses are multiplied, propagating the infection
Where is the immunity of viral DNA in bacterial cells found?
the repeats (found in many different types of bacteria) and spacers (viral DNA)
What are CRISPR Associated Proteins (CAS)?
transcribed and translated proteins found just upstream of CRISPR regions
What happened with the DANISCO yogurt company?
they found that their bacterial cultures were dying, and discovered that the bacteria that incorporated viral sequences in their CRISPR regions were the ones that survived
Where is the terminator found on CRISPR?
the very end of CAS proteins
What are the roles of different CAS proteins?
CAS1/2 - break phosphodiester bonds
CAS3/5/7 - helicase and single-strand nuclease activity
CAS9 - RNA-guided DNA endonuclease (cleave WITHIN a DNA chain)
What are the steps to and function of mature CRISPR RNA?
crispr locus —> pre-crRNA transcript —> mature crRNA (binds to CAS9 and breaks down viral DNA)
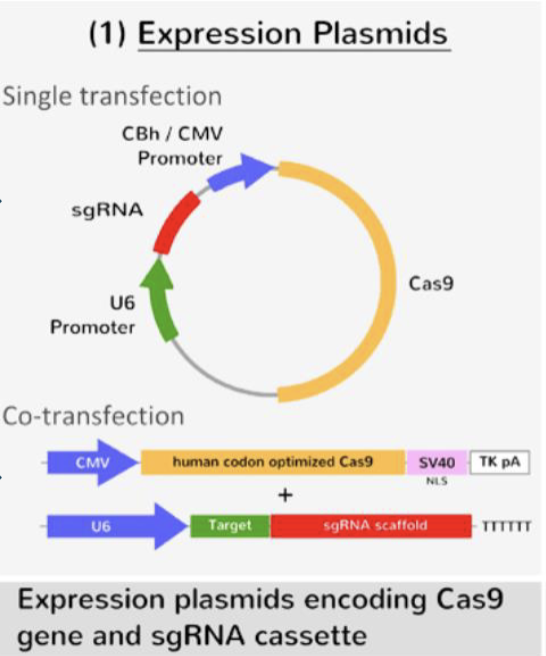
How can CRISPR be used in the human genome?
as the human genome does not encode for CAS proteins, scientists must transfect DNA with expression plasmids containing CAS and crRNA
What is the potential to “break” DNA in selected regions of a eukaryotic cell’s genome?
erase genes carrying mutations (chop the AUG start codon)
replace incorrect portions of selected genes with correct ones
What are the risks of CRISPR?
the guide RNA is precise, but the cut may not be precise, leading to:
imprecise cuts imply imprecise editing, which can change the reading of nucleotides
can lead to changes in regulation of expression of genes that respond to DNA being cut (possible cancer risk??)
What are PAM sequences?
allow CAS proteins to excise at the region of interest, but may not be precise at the nucleotide level
What is HIV?
human immunodeficiency virus (HIV) destroys immune system T cells lymphocytes
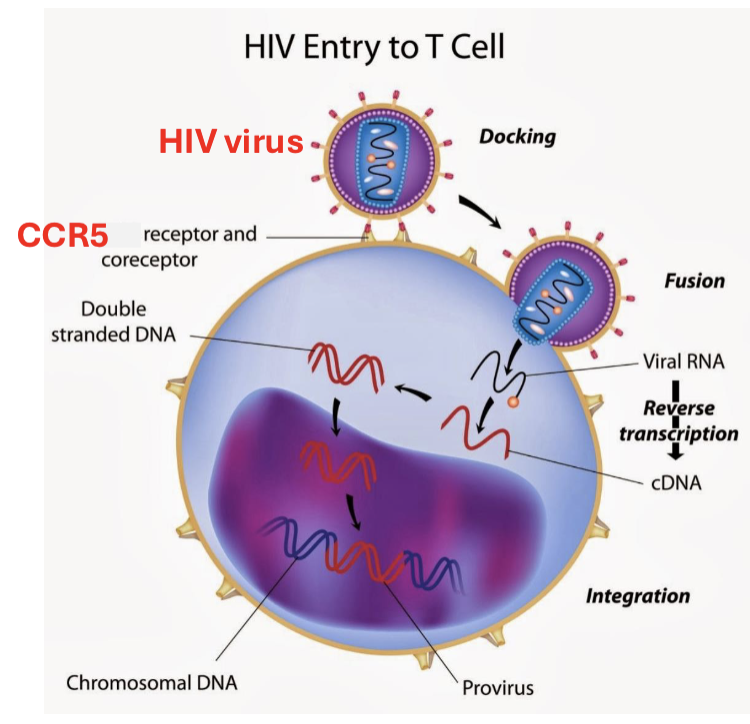
What is the process of HIV virus cell infection?
the HIV virus touches down (at CCR5 receptor), fuses, and integrates into the genome of T cells
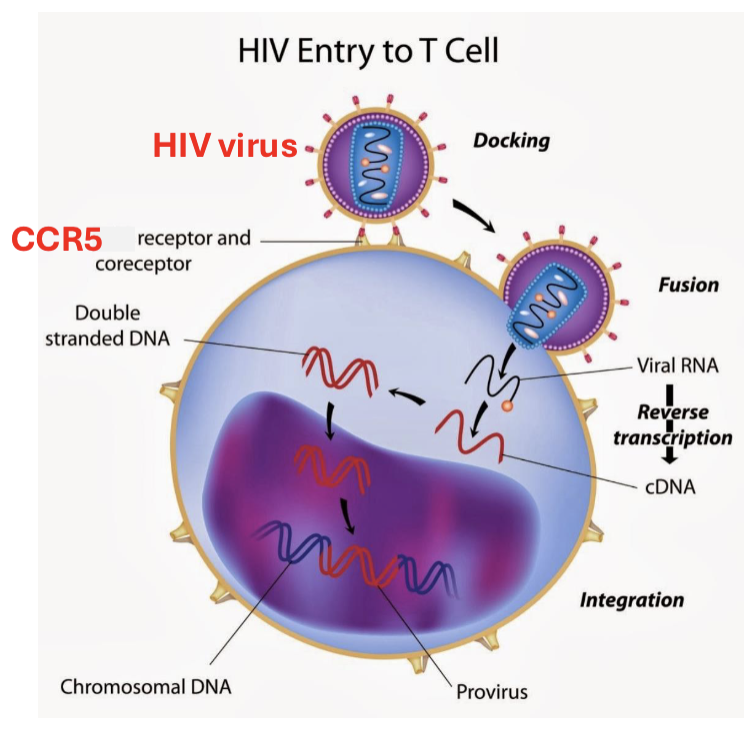
How can T cells be modified with CRISPR?
CRISPR transfection of T cells that are resistant to the R5 receptor, but not 100% efficient and can cause harmful immune response with imprecise cuts
How can CRISPR eradicate HIV?
embryonic stem cells can be modified to not have the CCR5 gene, which hypothetically would eradicate the binding of the HIV virus to T cells
What are the concerns for the CRISPR eradication of HIV?
CRISPR isn’t precise, and could cause unwanted deletions
cutting the DNA could trigger cell death mechanisms
not super ethical to manipulate embryos
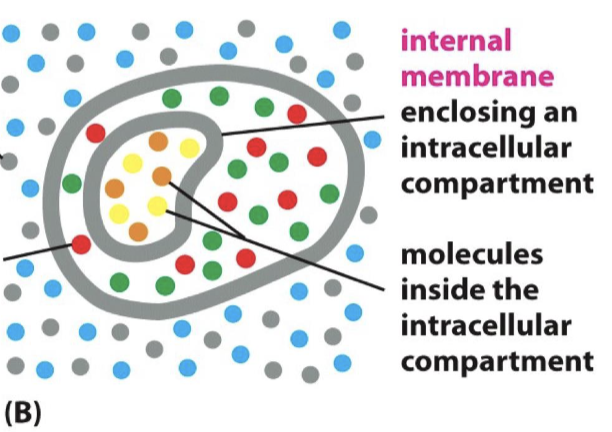
What do membranes do?
contain and delimit cells with water impermeability
help to isolate intracellular organelles
allow cells to receive information and import/export molecules with embedded proteins
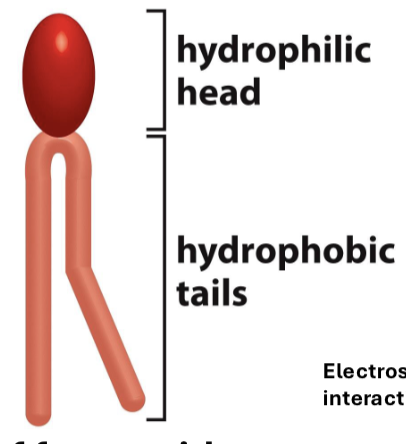
What are phospholipids?
amphiphatic (has both hydrophobic/phillic) molecule containing two nonpolar lipids (tails) with a phosphate group head (polar) arranged feet to feet
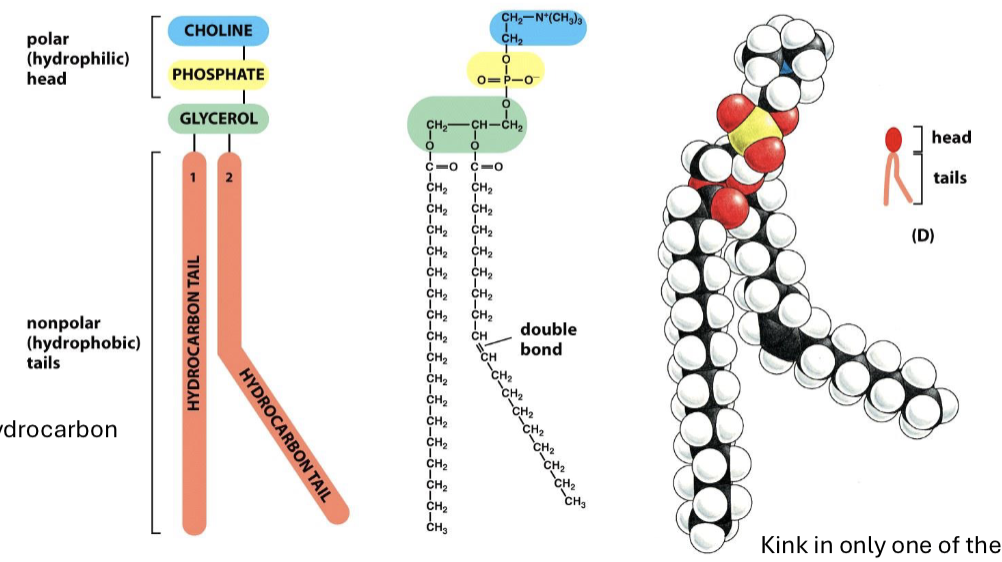
What is phosphatidylcoline?
the most abundant phospholipid, made up of a head with a glycerol, phosphate, and choline group and hydrocarbon nonpolar chains (one straight and one with double bond kink)
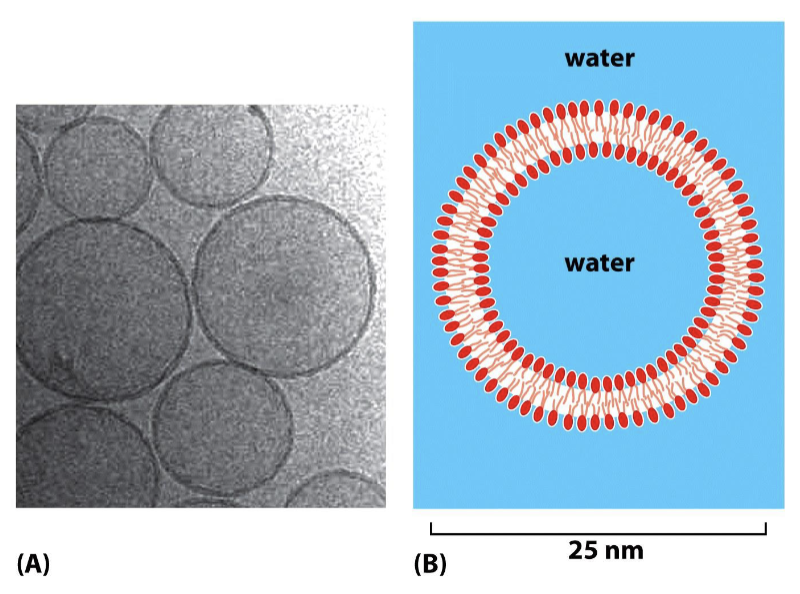
What are liposomes?
synthetic circular phospholipids without proteins
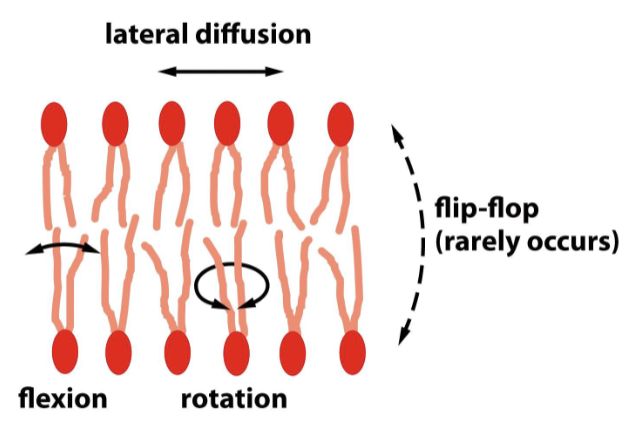
What is phospholipid fluidity?
lipids can…
rearrange within their own monolayers (lateral diffusion for transport)
show rotation across their axis
show flexion of hydrocarbon chains
What are the effects of longer/more fatty acids on fuidity?
long fatty acids → more interactions → less fluidity
more fatty acids → less interactions → more fluidity