Nucleic acids and protein synthesis
1/10
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced |
|---|
No study sessions yet.
11 Terms
Requirements of a Genetic Material
Must have:
Ability to store information → instructions to control cell behaviour.
Ability to copy itself accurately → ensures no loss of information during cell division.
Early assumption (before 1940s): proteins carried genetic info (thought too complex for DNA).
1940s–50s: Experiments proved DNA is the genetic molecule.
Structure of DNA and RNA
DNA → Deoxyribonucleic acid.
RNA → Ribonucleic acid.
Both are nucleic acids (originally found in nucleus).
Both are polymers (polynucleotides) → built from nucleotides (monomers).
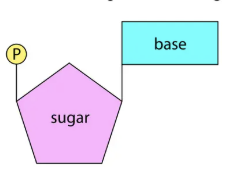
Nucleotides components
Nitrogen-containing base
DNA: A (adenine), G (guanine), T (thymine), C (cytosine).
RNA: A, G, C, U (uracil replaces T).
Purines (2 rings): A, G.
Pyrimidines (1 ring): T, C, U.
Pentose sugar
Ribose → RNA.
Deoxyribose → DNA (one oxygen atom less).
Phosphate group → gives acidic nature.
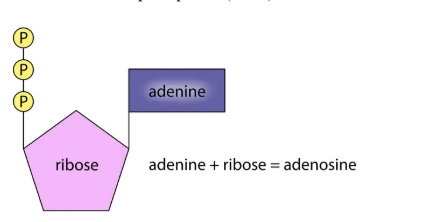
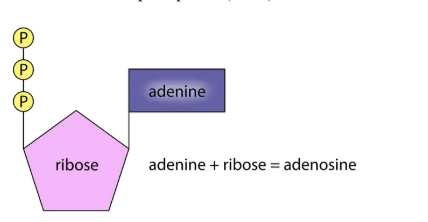
ATP
A nucleotide, not part of DNA/RNA.
Structure: adenine + ribose + phosphate groups.
Forms:
AMP (adenosine monophosphate).
ADP (adenosine diphosphate).
ATP (adenosine triphosphate).
⚠ Don’t confuse:
Adenine (base) vs. Adenosine (adenine + sugar).
Thymine (base) vs. Thiamine (vitamin).
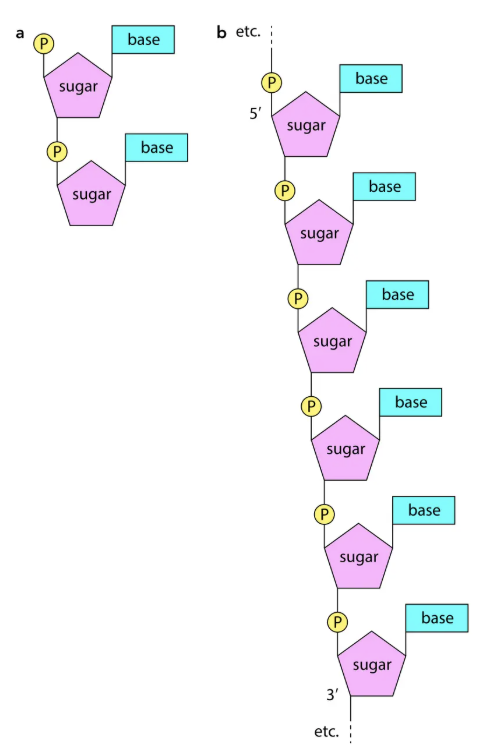
Polynucleotides
Nucleotides join via condensation → phosphodiester bonds (phosphate + sugar).
Structure: sugar-phosphate backbone with bases projecting sideways.
Key terms:
Dinucleotide → two nucleotides joined by phosphodiester bond.
Phosphodiester bond → joins nucleotides, two ester bonds (one on each side).
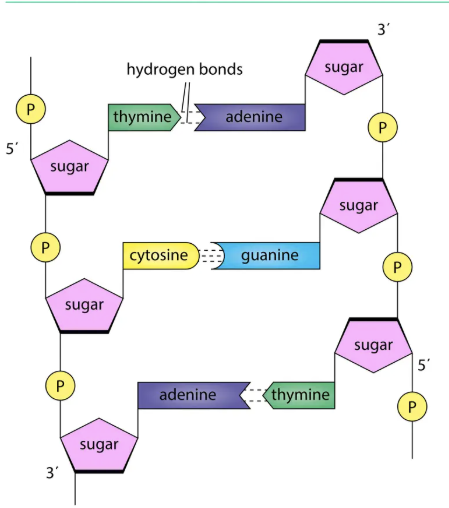
Structure of DNA
Discovered by Watson & Crick (1953).
Features:
Two anti-parallel polynucleotide chains.
Twisted into double helix.
Held by hydrogen bonds between bases.
Complementary base pairing:
A pairs with T (2 hydrogen bonds).
G pairs with C (3 hydrogen bonds).
Purine always pairs with pyrimidine → width constant (3 rings).
One complete turn = 10 base pairs.
Base ratio rule (Chargaff):
%A ≈ %T, %G ≈ %C.
Information storage: sequence of bases = coded message.
Replication: possible by unzipping → each strand acts as template.
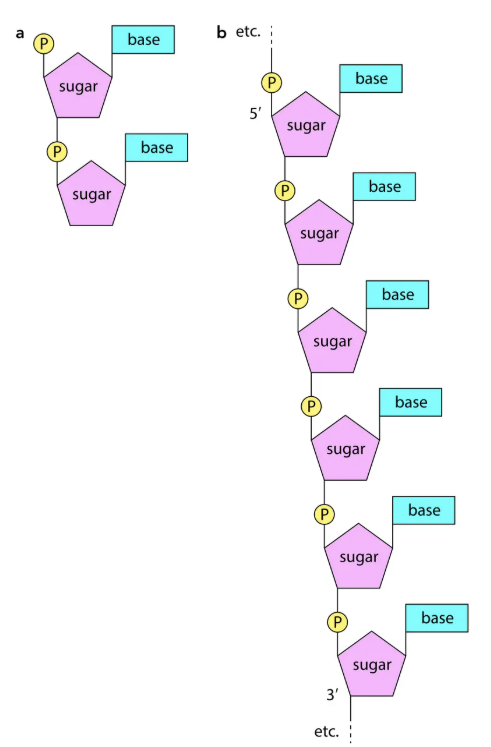
Structure of RNA
Single polynucleotide strand.
Types:
mRNA → messenger, carries code to ribosome.
tRNA → transfer, carries amino acids.
rRNA → ribosomal, structural component of ribosomes.
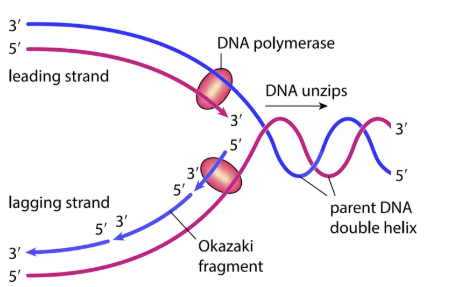
DNA Replication
Occurs during S phase of cell cycle.
Steps:
DNA unzips (hydrogen bonds broken).
DNA polymerase attaches to each strand, adds complementary nucleotides (5′ → 3′ direction).
Leading strand → continuous replication.
Lagging strand → discontinuous replication → short Okazaki fragments.
DNA ligase joins nucleotides and Okazaki fragments with phosphodiester bonds.
Semi-conservative replication → each new DNA has one original strand + one new strand.
The Genetic Code
Sequence of bases = code for sequence of amino acids in proteins.
Features:
Triplet code (3 bases = 1 amino acid).
Universal (same in all organisms).
Punctuated → start (e.g., TAC for methionine) & stop codons.
Redundant/degenerate → amino acids have more than one triplet.
Protein Synthesis
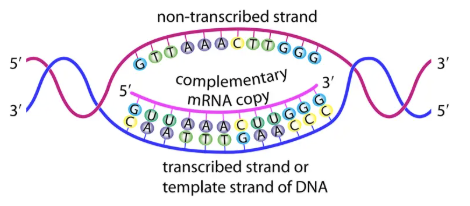
Stage 1: Transcription (in nucleus)
DNA → mRNA.
Enzyme: RNA polymerase.
Only template strand copied.
Base pairing: A → U, G → C, T → A, C → G.
Primary transcript → modified by RNA processing.
Introns removed, exons joined (splicing).
Alternative splicing → different proteins from same gene.
mRNA leaves nucleus via nuclear pore.
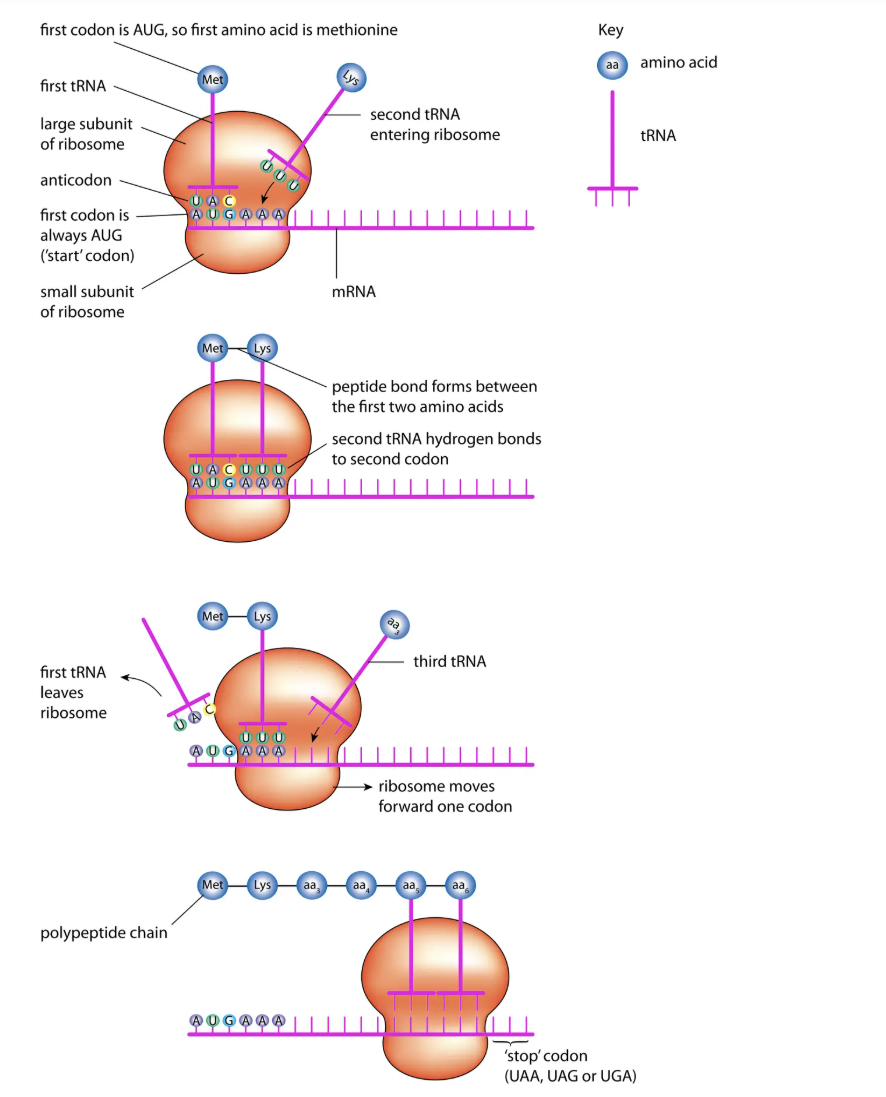
Stage 2: Translation (at ribosomes)
mRNA → polypeptide.
tRNA:
Carries amino acid at one end.
Has anticodon (3 bases complementary to mRNA codon).
Steps:
tRNA binds to codon on mRNA.
Ribosome holds two tRNAs side by side → peptide bond forms.
Ribosome moves along mRNA, process repeats.
Stops at stop codon → polypeptide released and folded.
Gene Mutations
Mutation = random change in DNA base sequence.
Caused by copying errors, radiation, carcinogens (mutagens).
Types:
Substitution → one base replaced.
May change amino acid (missense), no change (silent), or stop codon (nonsense).
Example: Sickle cell anaemia (Glu → Val substitution in haemoglobin β-chain).
Deletion → base removed.
Insertion → base added.
Deletion/insertion → frame-shift mutations → big effect on protein.