BIOC 302 - DNA damage, repair, and genome editing
1/73
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
74 Terms
is DNA damage a mutation?
DNA damage does not become a mutation until its an heritable problem, which may take 1 round of replication or more until the damage becomes heritable.
what is DNA damage?
when there are chemical changes to the nitrogen rich bases in DNA. This can lead to changes in base pairing potential and therefore mutations. It can also lead to double strand breaks thus genome instability
how is DNA damage dealt with?
can be repaired or cells will undergo programmed cell death (apoptosis)
what are the different types of endogenous DNA damage?
polymerase errors
spontaneous deamination reactions
spontaneous hydrolysis of base
oxidative damage
what are the different types of exogenous DNA damage?
UV photo-damage
alkylating agents
DNA polymerase has ___ fidelity. Why?
high fidelity due to multiple mechanisms:
exonuclease proof-reading site
shape discrimination
mis-match repair pathways
However, some mutations sneak by from time to time (3 Billion base pairs ~60 mistakes/cell division → Mutation)
what is spontaneous deamination?
when there is non-catalyzed (spontaneous deamination of cytosine, which causes for cytosine to be converted into uracil. Since uracil is very chemically similar to thymidine, can base pair with adenine and if left unrepaired, a A-T mutation will occur
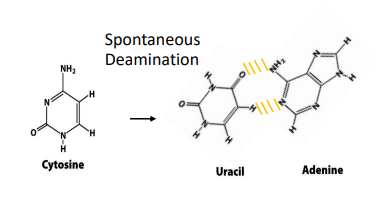
what is deamination?
the spontaneous loss of exocyclic amino groups
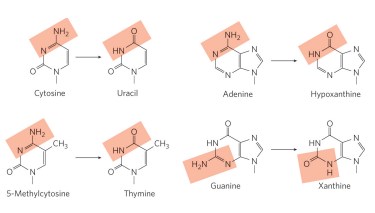
how often does deamination of cytosine to uracil occur?
around 100 events/day. However, uracil is recognized as foreign in DNA and is removed (likely why DNA contains thymine instead of uracil)
what deamination reaction can cause a problem in DNA?
5-Methylcytosine (a very important gene regulator) → thymine. This is a problem bc thymine is found in DNA therefore this deamination is not easily detected and a thymine may end up where its not supposed to

B
C→U so we’re gonna have U-G
U-G is split for replication, U pairs with A making A-U, and G pairs with C making C-G
what are depurination reactions?
a hydrolysis of the N-beta-glycosyl bond between the base and the pentose of a nucleotide. This reaction creates an AP (apurinic, apyrimidinic) site or abasic site (no base).
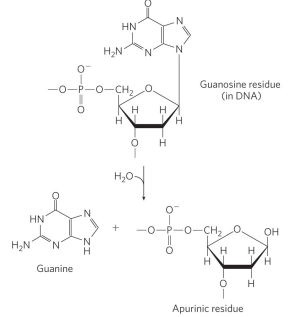
depurination reactions are more common with ___
purines
what can depurination reactions cause?
stalling or very error prone rxn. When unzipped by helicase, no information for polymerase to read (bc the base is the information portion of a nucleotide)
how can DNA be damaged by oxidative damage?
reactive oxygen species such as hydrogen peroxide, hydroxyl radicals, and superoxide radical damage DNA. Hydroxyl radicals are responsible for most oxidative DNA damage.
How is oxidative damage to DNA minimized?
cells have an elaborate defense system to destroy reactive oxygen species
What is an example of how oxidative damage can cause DNA damage?
ROS can cause guanine to become 8-oxo-guanine. While the new carbonyl group doesn’t actually interfere with base-pairing, it causes problems in a different way. Usually, the anti conformation around the glycosyl bond is favored - but the new steric clash between carbonyl and ribose oxygen causes for 8-oxo-guanine to favor the syn conformation. This causes for an increase in Hoogsteen base pairing between G and A. After replication, this will generate an A-T.
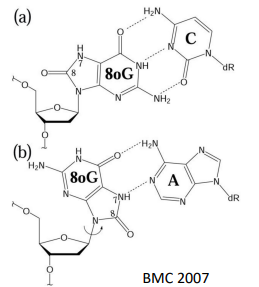
how does UV light cause damage to DNA?
Causes formation of cyclobutane pyrimidine dimers and 6-4 photoproduct.
These types of damage occurs in a single DNA strand
causes for formation of covalent bonds between bases in the same strand (usually cross-linking of 2 adjacent Ts, but can also happen with C)
causes for kinks in the DNA
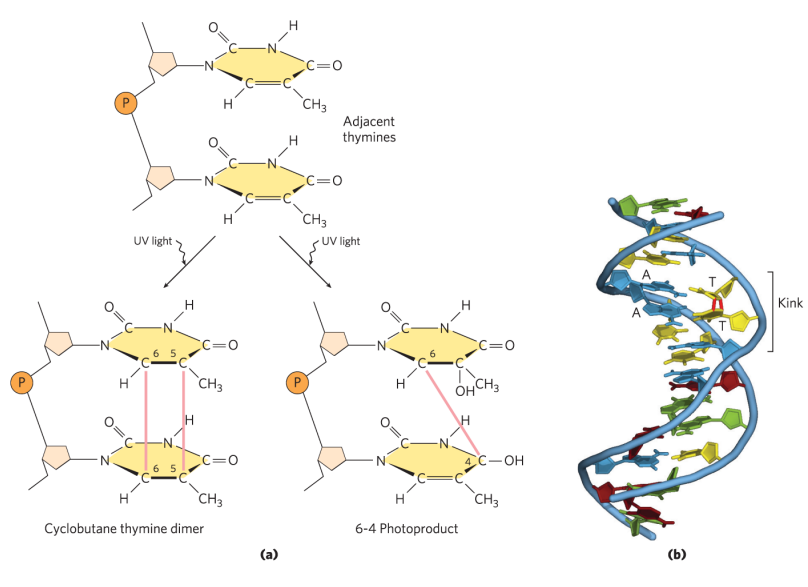

WC interactions won’t be able to occur at active sites bc H-bond donors and acceptors would be at new angles. Therefore, DNA polymerase will start to fall off and there will be stalling of replication
how does ionizing radiation (x-rays and gamma rays) cause DNA damage?
ring opening
base fragmentation
breaks in the covalent backbone of nucleic acids
what does translesion polymerase η Eta do?
Translesion polymerase η bypasses DNA damage caused by UV-induced thymine or cytosine dimers. It has a larger, flexible active site that can accommodate the distorted, cross-linked bases. During bypass of UV dimers, it usually inserts two adenines (dA). If it encounters oxidative damage such as 8-oxo-guanine, it typically inserts a cytosine (dC). This allows DNA replication to continue even when the normal replicative polymerase stalls.
What is an example of alkylation damage in DNA?
Guanine gets modified into O6-methylguanine in the presence of alkylating agents, adding a methyl group. This is a common and highly mutagenic lesion and causes for pairing with thymine rather than cytosine. This pairing occurs bc the carbonyl group on guanine is now blocked by a methyl group so there is no longer the repulsion that would normally occur between guanine and thymine.
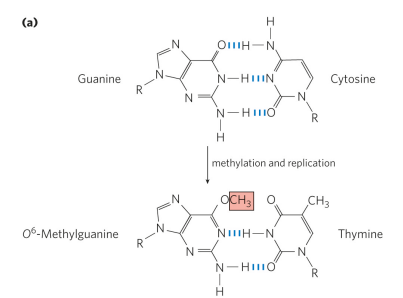
what causes alkylation damage in DNA?
chemical exposure to (alkylation), i.e., cigarette smoke, moldy peanuts, burnt meat
when does DNA dmg lead to heritable mutations?
after replication or single allele variation in post-mitotic cells
Why is DNA repaired instead of degraded like all other biological molecules?
DNA stability is essential for encoding the proteins/RNA that the organism needs (e.g., if we lose an organelle, it can be rebuilt if we have the code for it) and because DNA is easily damaged. We want DNA code to stay the same, and if was broken down and rebuilt, this would not be very possible
what is a mutation?
a permanent change in the nucleotide sequence
what are the different types of mutations?
substitution mutation
insertion mutation
deletion mutation
silent mutation
what is a substitution mutation?
a mutation where one base pair is replaced with another
what is an insertion mutation?
a mutation where there is an addition of 1+ base pairs
what is a deletion mutation?
a mutation where there is a deletion of 1+ base pairs
what is a silent mutation?
a mutation that affects nonessential DNA (most of our genome is not gene, its not-coding regions. Mutations here are not going to change the amino acid but can have a profound effect on regulation) or has a negligible effect on gene function. This type of mutation doesn’t change the amino acid at that position but changes the codon
Some mutations are caused by ____, and some mutations are caused by ____ that then favours new ____ interactions
Some mutations are caused by polymerase errors, and some mutations are caused by DNA damage that then favours new base-pair interactions
what is the Ames test and how does it work?
The Ames test is used to determine if a chemical causes mutations in DNA (is mutagenic). It uses Salmonella typhimurium with a mutation in the histidine synthesis pathway, so the bacteria cannot make histidine and cannot grow on histidine-free media. A small circle of Whatman filter paper soaked in a potential mutagen is placed on the plate. If colonies grow, it means mutations occurred that restored histidine synthesis. More colonies indicate higher mutagenicity, and colonies farther from the center indicate the chemical is mutagenic at lower concentrations. This test can be applied to humans because DNA is chemically similar in all organisms, so chemicals that cause mutations in bacteria will usually also cause mutations in human DNA. The restored histidine synthesis in bacteria is just a reporter for DNA mutations—humans don’t gain the ability to make histidine, but the chemical could still cause mutations in human DNA.
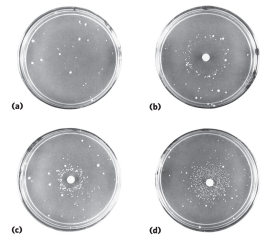
How does DNA repair affect cancer?
8% of human cancers can trace their pathogenic driver to mutations in DNA repair processes. Therefore, DNA repair is tumor protective. Proteins like p53 maintain genome stability (elephants have many copies, so they rarely get cancer). Loss of p53 or other repair defects allows mutations to accumulate, leading to aggressive tumors. Paradoxically, severe DNA damage (e.g., radiation therapy) can trigger apoptosis in cancer cells, killing them. DNA polymerase is highly accurate but occasional mistakes still occur, so repair is essential.
what are the different mechanisms of single-strand base repair?
mismatch repair
base-excision repair
nucleotide excision repair
direct repair
what enzyme is needed for mismatch repair for DNA in E. coli? what does it do?
Dam methylase methylates GATC sites on the adenine residues (binds to the whole DNA sequence and flips adenine into an active site to methylate it) using SAM as the methyl donor. After replication, the template strand is methylated, but the new strand is temporarily unemthylated, creating hemimethylated DNA (this is where we will have post-replication mismatch identification)
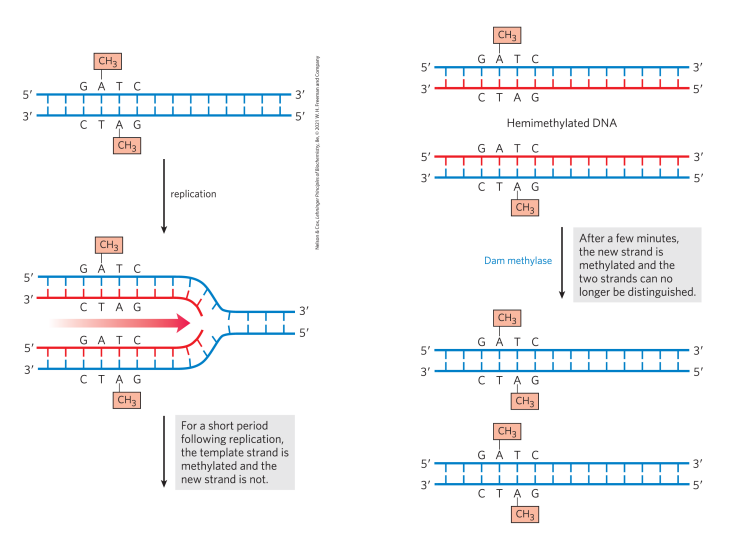
Explain how post-replication mismatch is identified
Mismatched base pair can actually be a far distance away from the methylated sites. The mismatch is caught bc it has a slightly different diameter (no damage to nucleotides), so MutS and MutL recognize the mismatch and then slide along to the nearest flag (methylated adenine in a GATC site) in an ATP-dependent manner. The non-methylated strand gets cleaved, generating a single-strand nick (no prefernce for direction, can go either way of the mismatch, but always happens at the site where the methylation is).
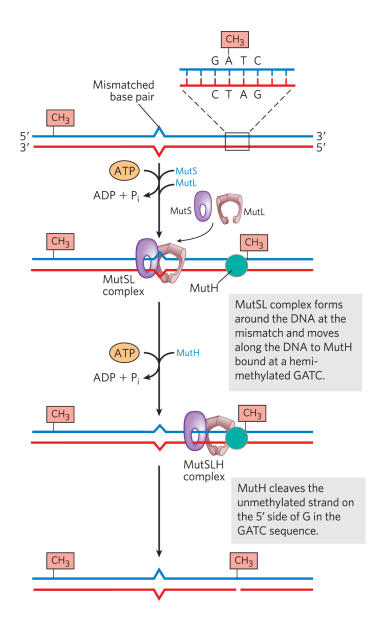
After identification, how is the mismatch repaired in DNA?
1. Helicase unwinds the DNA at the nick site and moves towards the lesion
2. Nucleotides between the nick and the lesion are removed by an exonuclease
3. Nick or exonuclease activity generates a free 3’ OH
4. Non-damaged strand is used as a repair template for DNA polymerase
5. Gap at the end of the repair is sealed with a ligase
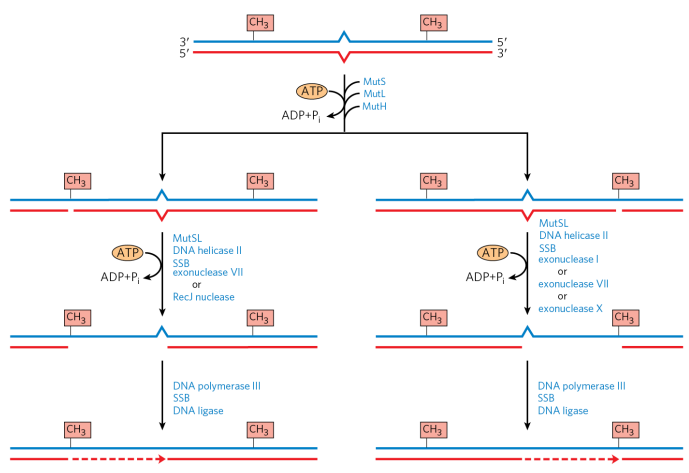
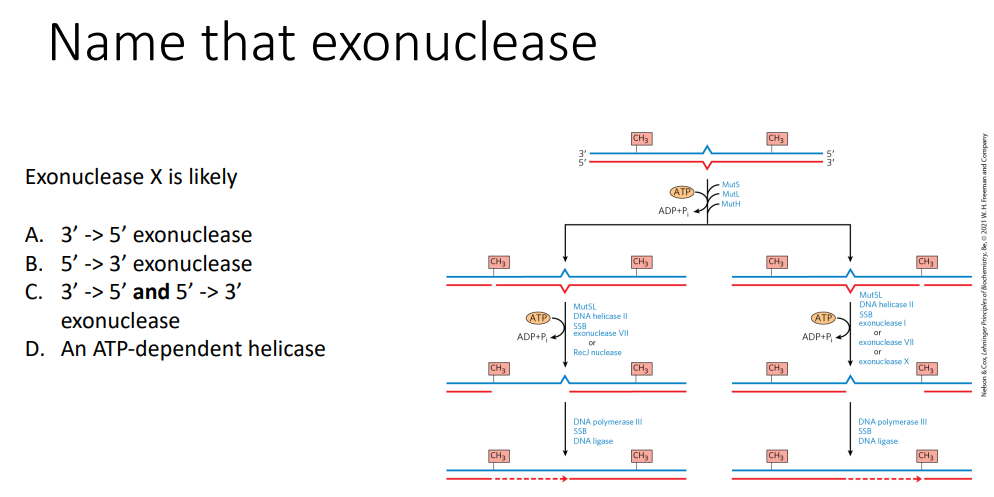
A
what is base-excision repair (BER)?
repair of small, non-helix distorting DNA lesions. DNA glycosylases recognize common DNA lesions and remove the affected base by cleaving the N-glycosyl bond in the process of base-excision repair, generally specific for one lesion type. This forms an AP site or basic site (an apurinic or apyrimidinic site in the DNA resulting from the removal of a base by DNA glycosylase).
What do uracil DNA glycosylases do?
they specifically remove from DNA the uracil that results from spontaneous deamination of cytosine. However, does not remove uracil residues from RNA or thymine residues from DNA (deamination is a 100-fold faster in ssDNA)
Other DNA glycosylases recognize:
formamidopyrimidine (from purine oxidation)
8-hydroxyguanine (from purine oxidation)
hypoxanthine (from adenine deamination)
alkylated bases
pyrimidine dimers
Specific damage is recognized by a specific enzyme that removes the damaged base and generates an abasic site
What are the steps of base excision repair?
1. Damaged base is recognized by a specific glycosylase that removes the base (generates an abasic site)
2. An endonuclease cleaves the phosphodiester backbone at the abasic site, leaving a 5’ phosphate
3. DNA Pol I cleaves off the nucleotides one by one from 5’ → 3’, then replaces missing base (and short extension)
4. Gap generated by the polymerase extension is filled with DNA ligase
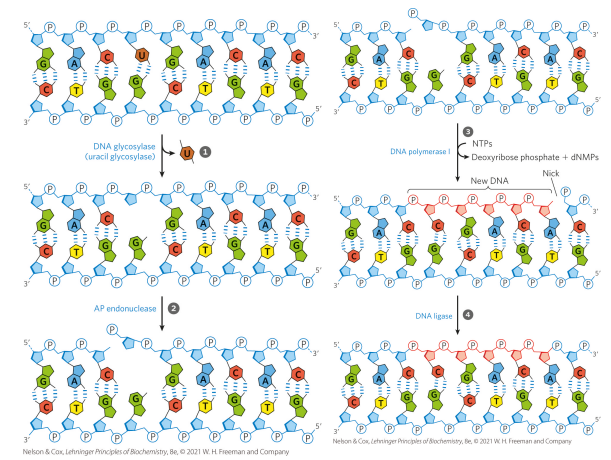
what does the nucleotide-excision repair system do?
repairs DNA lesions that cause large distortions in the DNA helical structure
How does nucleotide-excision repair work?
uses excinuclease which is a multisubunit enzyme that hydrolyzes 2 phosphodiester bonds, one on either side of the distortion. DNA helicase then unzips double stranded DNA between the 2 nicks. Then, DNA pol I (E. coli) or DNA pol ε (epsilon bc we use 2 polymerases, one for lagging and one for leading) (humans) fills the gap and DNA ligase seals the nick (energy needed).
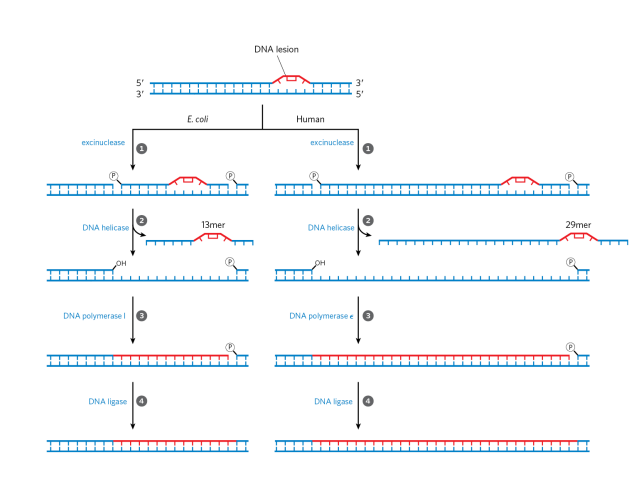
what is direct repair and when is it used?
Direct repair reverses specific DNA damage without cutting the DNA. It’s used for lesions like O6-methylguanine, which would otherwise mispair (thymine instead of cytosine) and cause mutations after replication.
Explain direct repair of O6-methylguanine
O6-methylguanine-DNA methyltransferase (MGMT) catalyzes the transfer of the methyl group on O6-methylguanine to one of its own Cys residues (single methyl transfer event). While the chemistry of the guanine residue is restored, MGMT is not regenerated, thus it is termed a “suicide enzyme.” This type of repair is very expensive by cellular energetic measurements bc protein synthesis isn’t cheap.
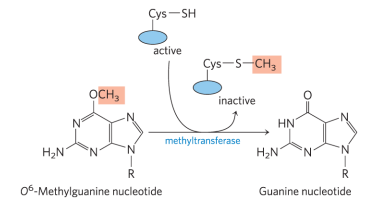
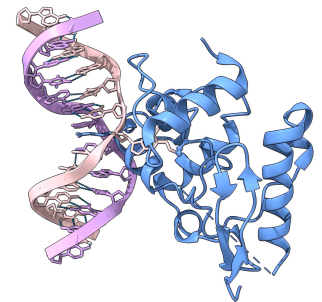
MGMT. What role is Cysteine playing in the repair process (ball and stick side chain)? What role is Arginine playing (stick presentation with tiny label)?
cysteine holds onto the methyl group
Nucleotide is flipped out during repair so can’t do base stacking in the helix so arginine takes over and does base stacking in place of the damaged guanine
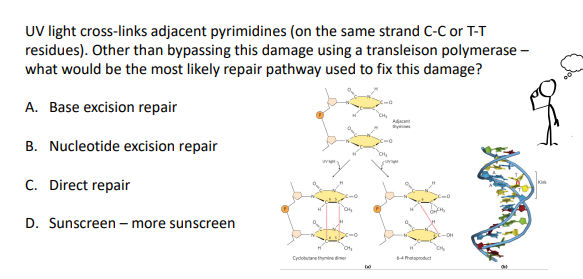
B (bc the lesion is so bulky)
what are the causes of DNA double strand breaks
programmed double-strand breaks
damage that leads to double strand breaks
what are the programmed double-strand breaks?
during meiosis - recombination
V(D)J recombination - our immune system is able to adapt and part of that adaptation is making double strand breaks and then joining 2 pieces of the genome together (no apoptosis)
what damage leads to DNA double-strand breaks?
incomplete NER or BER repair pathways and subsequent DNA replication
UV, IR, or radiation damage
replication stalling
(from image - unrepaired lesion can cause single stranded region in the new daughter strand. Unrepaired break can cause a double strand break)
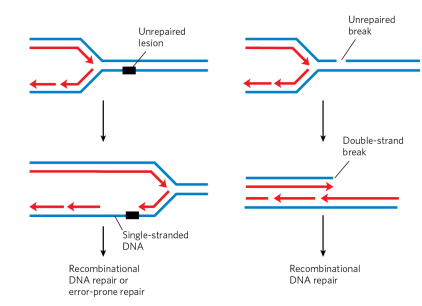
when does DNA double strand breaks occur?
at the S phase of replication, so we need to make sure all our DNA is as fit as possible before S phase. A big part of the pause at the G1 phase is to make sure no dmg is in the genome before S phase
what are DNA repair foci and when do they form?
DNA repair foci are large clusters of macro-molecular bodies at the site of damage (double-strand breaks, e.g., after ionizing radiation) to coordinate DNA repair
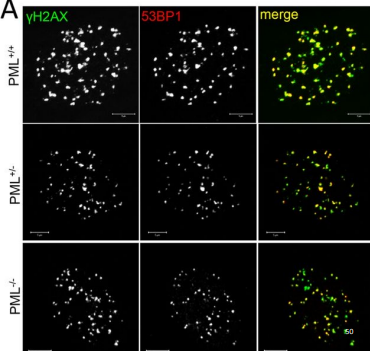
what is γH2AX and what does it do?
its a modified histone (H2Ax) that allows for cell to signal that there’s been a lot of damage that needs repair. Helps recruit repair proteins to the break
what is 53BP1 and what does it do?
53BP1 is a protein that’s recruited when there’s been a DSB. It is the deciding protein that helps the cell determine whether to do NHEJ or homologous recombination
what role does PML play in DNA repair foci?
PML helps organize repair foci (recruitment of yH2AX and 53BP1 is a PML-dependent event) and keeps the 2 broken DNA ends close together, preventing them from drifting apart during repair
what happens when PML is absent or reduced?
While yH2Ax and 53BP1 still appear, with no PML, we lose the perfect correlation that we had when PML was present. Organization of repair foci is disrupted, meaning the DNA ends are less coordinated, and repair becomes less efficient
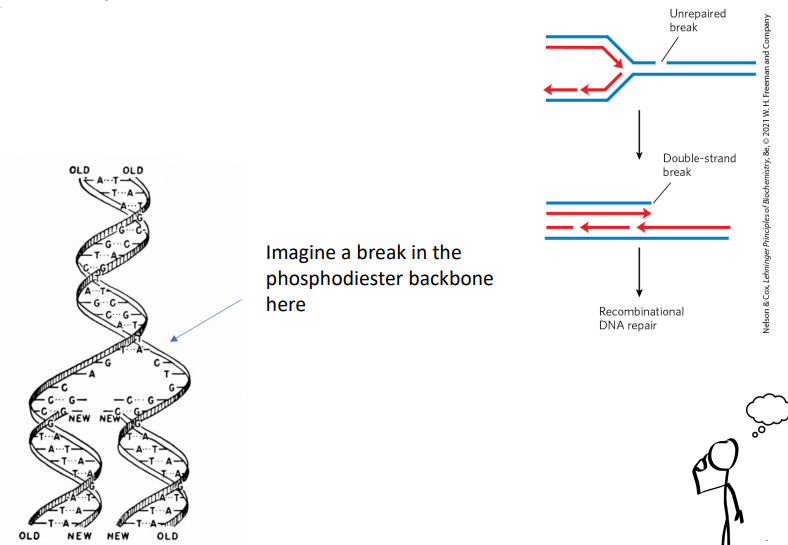
how do single-strand breaks lead to double-strand breaks during S-phase?
if there was a break in the phosphodiester backbone, polymerase will go along the strand and add nucleotides until it reaches the nick and ends there bc it doesn’t realize that there’s more after the nick.
what are the 2 main double-strand break repair pathways?
non-homologous end-joining (NHEJ)
homologous recombination (HR)
what does NHEJ do?
joins 2 broken DNA strands together using very small microhomology - can result in small deletions or insertions.
what is HR?
template-driven repair of DSB. conservative when sister chromosome is used to repair break
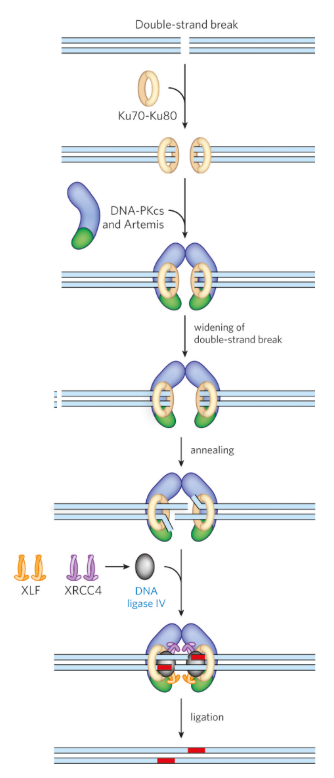
explain and sketch the mechanism of NHEJ
Ku7-Ku80 complex binds to the broken DNA ends, which protects the ends and marks them the fite for repair
DNA-PKcs (kinase) and Artemis (nuclease) are recruited to form the repair complex
synapsis occurs - the 2 DNA ends are brought together and held in place so they don’t drift apart
End processing - Artemis removes single-strand extensions or hairpins, making the strands compatible for joining
Annealing - short regions of microhomology base-pair (W-C only) to align the strands
DNA polymerase fills the missing nucleotides
DNA ligase seals the break by forming phosphodiester bonds
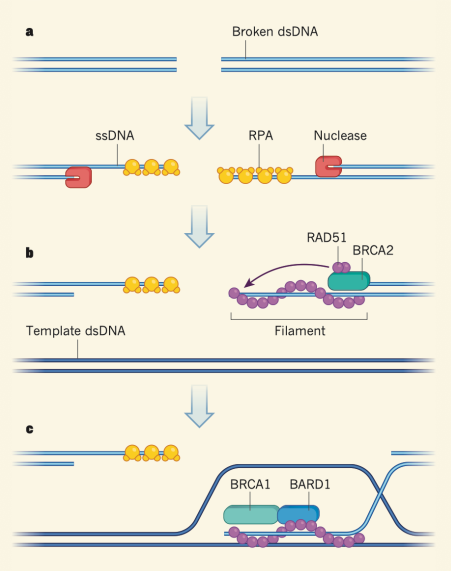
Explain and sketch the mechanism of homologous recombination
MRN-CtIP comes and recognizes broken dsDNA region, recruiting exonuclease MREII
MREII chews away the 5’ end, leaving a 3’ overhang (resection of DNA)
On the other strand, replication protein A (RPA) (in humans) or single-stranded binding proteins binds to the 3’ overhang, protecting it
RPA is replaced by RAD51 (recombinase)
BRCA2 directs RAD51, forming a filament
BRCA1 and BARD1 help direct invasion filament to a region in the genome. RAD51 filament invades a homologous DNA sequence (sister chromatid), forming a D-loop
RAD51 starts to get peeled off one at a time until filament is pulled off
DNA polymerase extends the invading 3′ end using the homologous template.
Second-end capture: The other broken DNA end pairs with the displaced strand in the D-loop and is also extended by DNA polymerase.
resolution of holliday junctions with nucleases
nicks sealed with ligase
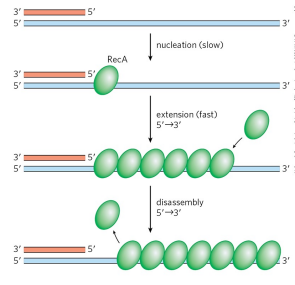
what is RecA?
the version of Rad51 in E. coli (Rad51 is in humans). Upon binding to RecA or Rad51, the 3’ overhand becomes a filament
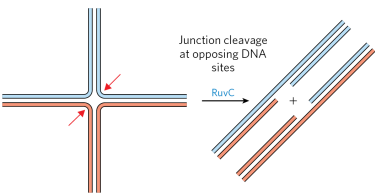
how are Holliday junctions resolved?
nucleases form 2 nicks in the junction. The gap is then sealed with ligase
How did we figure out which repair proteins would bind to what part of the DNA during the repair process? Explain.
A gel shift assay can be used to test the binding of proteins to different DNA structures, such as D-loops, bubbles, replication forks, double-stranded DNA, and single-stranded DNA. In this assay, the DNA is incubated with the protein of interest, such as BRCA1-BARD1. When the protein binds to the DNA, the DNA moves more slowly through the gel, causing a “shifted” band that appears higher up compared to unbound DNA. The higher the band appears on the gel, the stronger the binding affinity between the DNA and the protein. Using this approach, it was observed that BRCA1-BARD1 binds most strongly to D-loop and bubble structures, showing higher bands, while replication fork, double-stranded, and single-stranded DNA showed lower binding.
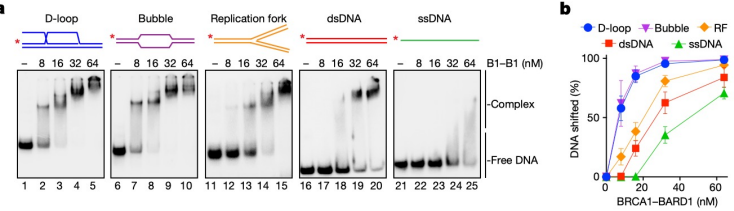

A
why would scientists engineer a genome?
To analyze gene functions
To study genome rearrangements (ie cancer)
To engineer animal models
Engineer disease or insect resistances in crops, animals or humans
To remove unwanted organisms
To treat severe genetic diseases
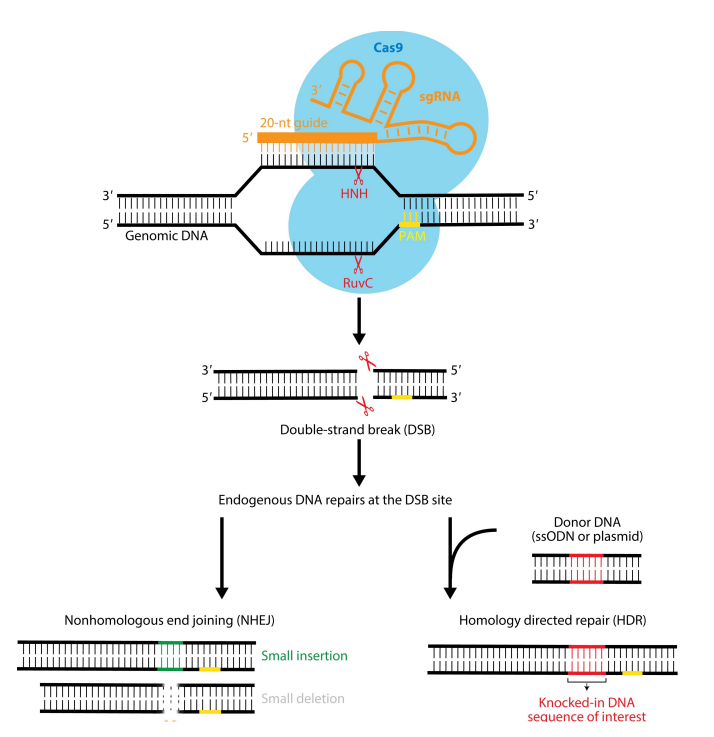
Explain the CRISPR-Cas9 mechanism
Apo protein cas9 (inactive) → no cleaving
binding of the RNA: 2 RNA → crRNA (unique) & tracer RNA (structural). Gets turned into single guide RNA (sgRNA) which is a fusion of the structural RNA and the engineered RNA
re-organization of the PAM recognition region which then enables binding to 5’-NGG-3’ (any nucleotide, 2 guanines)
melts target DNA (target strand and non-target strand)
W-C between sgRNA and target DNA
moves HNH domain over the cute site → allows for cleavage 3 nucleotides away from PAM
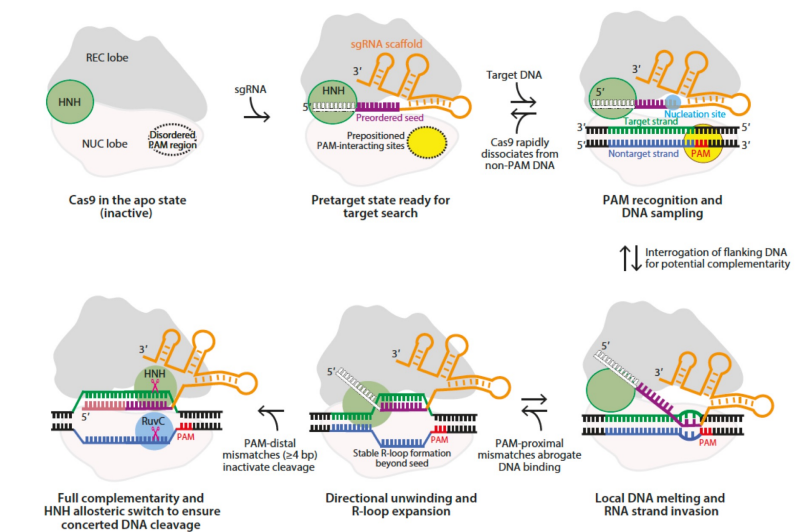
what should you do if asked to find PAM and only one strand of DNA is given?
always write the other strand so you can see both strands and find the PAM (5’NGG3’)
how does Cas9 become active and cleave DNA?
Cas9 undergoes a large-scale structural reorganization during activation, particularly in its HNH nuclease domain. This reorganization occurs when Cas9 binds both the single-guide RNA (sgRNA) and the target DNA. Cleavage of the DNA is only possible after these conformational changes, meaning Cas9 cannot cut DNA unless it is in complex with both sgRNA and its target.
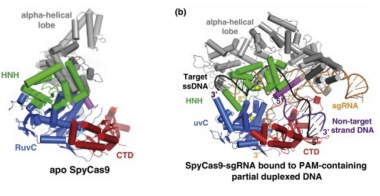
After the CRISPR-Cas9 mechanism, there is a DNA double strand break. Which repair mechanism can be used to add genetic material?
HR repair. If you provide the cell with a repair template - one that is identical to the sequences on either side of the CRISPR DSB you can add genetic material. During HR repair, DNA pol will amplify these regions if the repair template is used for repair