BIOL 265 - Final
1/192
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
193 Terms
The antiparallel nature of the two strands of DNA
When DNA is replicated, one strand of DNA is replicated continuously whereas the other strand of DNA is replicated discontinuously. The continuous and discontinuous replication of the respective strands of DNA is due to which characteristic of DNA?
Leading strand, lagging strand, Okazaki fragment, RNA primer
Match A-D
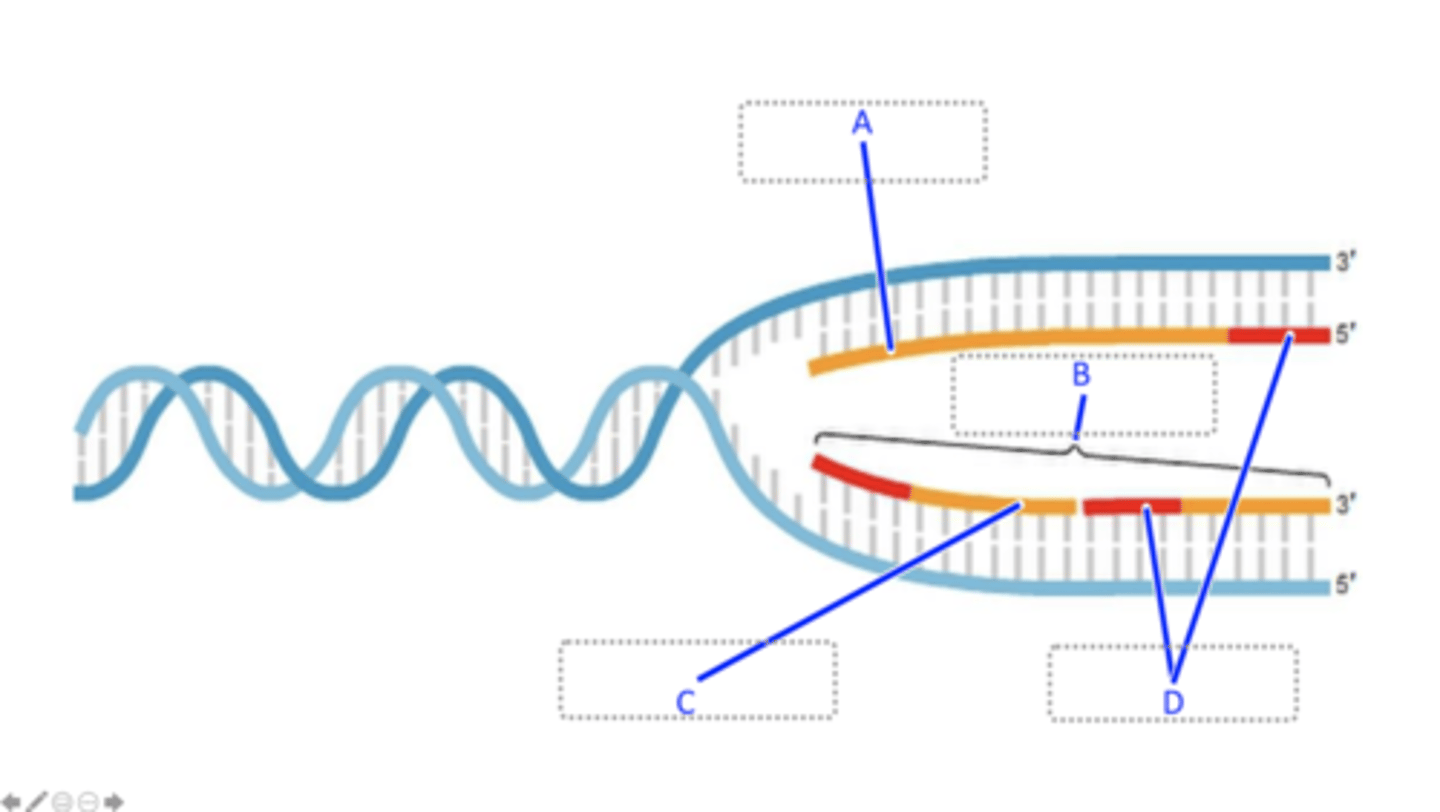
Helicase unwinds the DNA double helix, Single strand DNA binding proteins bind to each template strand, RNA primers are laid down, DNA polymerase synthesizes DNA, RNA primers are removed, DNA ligase joins DNA fragments together
6 steps in the process of DNA replication
Conservative mode of DNA replication, Dispersive mode of DNA replication, Semi-conservative mode of DNA replication
In 1958, Meselson and Stahl conducted an experiment to determine which of the three proposed models of DNA replication was correct. Below are three images labelled A, B and C, each of which illustrate one of the three models of DNA replication. Match the label of each image on the left-hand side with the correct model of DNA replication from the dropdown options provided on the right-hand side.
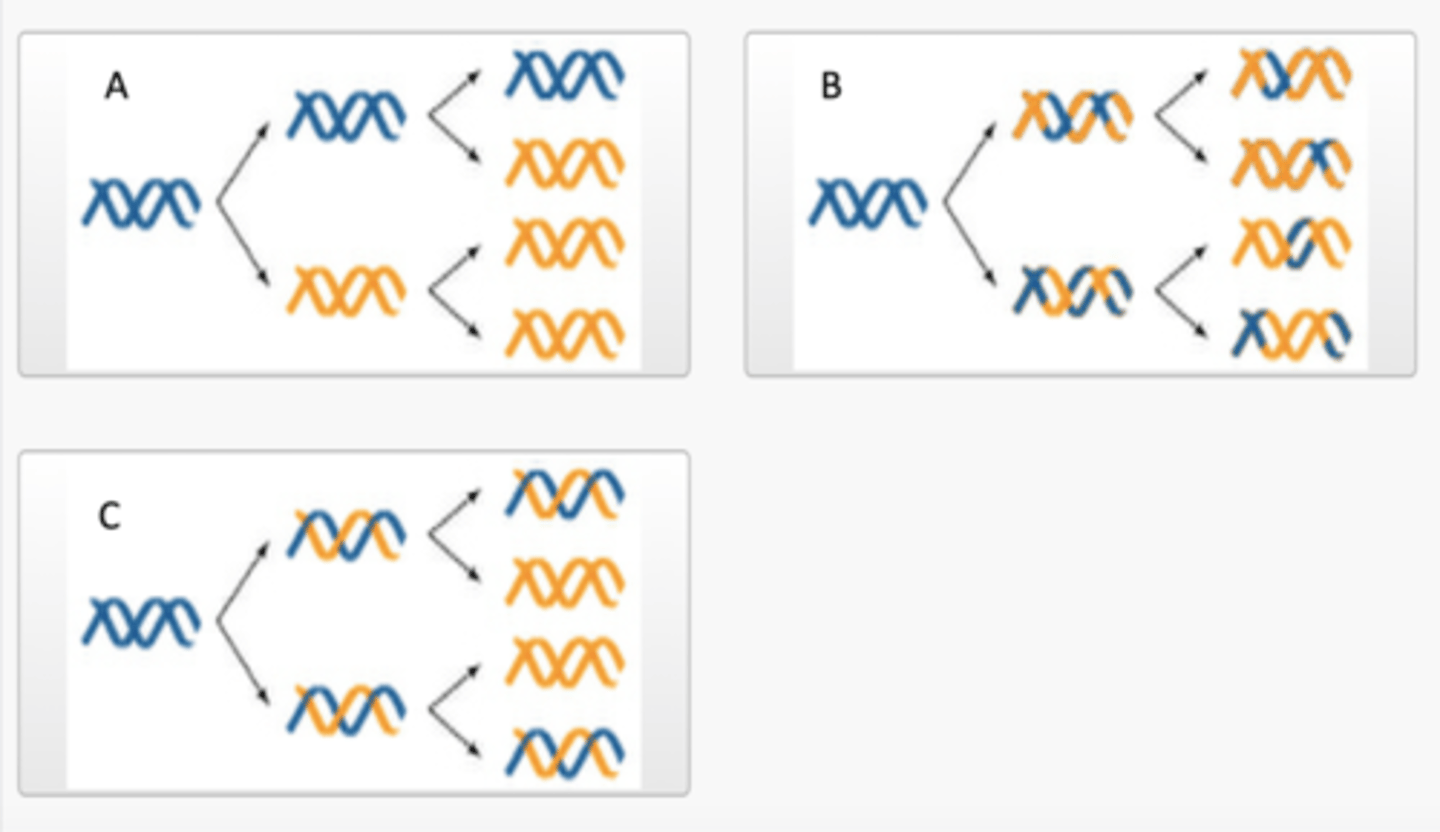
The first round of DNA replication resulted in hybrid 15N/14N DNA and the second round of DNA replication resulted in both hybrid 15N/14N DNA and 14N/14N DNA. No 15N/15N DNA was produced as a result of the first or second round of DNA replication.
The Meselson and Stahl experiment starts with E. coli containing 15N/15N labeled DNA grown in 14N media (first round of DNA replication). The cells from the first round of replication were then further grown in 14N media for the second round of DNA replication. What result did Meselson and Stahl observe by density centrifugation that provided strong evidence for the semiconservative model of DNA replication?
Immediately after transfer of the bacterial cells into radioactive medium; no radioactive phosphorous is incorporated into the DNA. At the end of one round of DNA replication; one strand of each DNA molecule has radioactive phosphorus incorporated. At the end of two rounds of DNA replication; 50% of DNA molecules have radioactive phosphorous in one strand and 50% of DNA molecules have radioactive phosphorous incorporated into both strands.
Phosphorous is required to synthesize the deoxyribonucleoside triphosphates used in DNA replication. A geneticist grows some E. coli in a medium containing nonradioactive phosphorous for many generations. A sample of the bacteria is then transferred to a medium that contains a radioactive isotope of phosphorus (32P). Samples of the bacteria are removed immediately after the transfer and after one and two rounds of DNA replication. Under these experimental conditions, newly synthesized DNA strands contain 32P and the original DNA strands contain nonradioactive phosphorous. Match the timing of analysis of the bacterial cells given on the left-hand hand side with the corresponding distribution of radioactivity in the DNA of the bacteria from the dropdown options provided on the right-hand side below.
-DNA topisomerase: relaxes the supercoiled DNA
-Helicase: unwinds the DNA double helix
-Single strand binding protein: prevents the reanneling of the DNA strand
-DNA polymerase I: replaces RNA primers with DNA
-primase: synthesizes RNA primers
-DNA ligase: catalyzes formation of phosphodiester bonds between adjacent DNA fragments resulting in a continuous DNA strand
50% of DNA molecules have radioactive phosphorous in one strand and 50% of DNA molecules have radioactive phosphorous incorporated into both strands.
-Loss of 5' -> 3' polymerase activity: no DNA synthesis to fill gaps caused by removing RNA primers
-Loss of 5' -> 3' exonuclease activity: no RNA primer removal during DNA replication
-Loss of 3' -> 5' exonuclease activity: decreased polymerase fidelity
DNA polymerase I has 5′-> 3′polymerase activity, 5′-> 3′ exonuclease activity, and 3′-> 5′ exonuclease activity necessary for DNA replication. Mutations in the specific portions of the gene that encodes DNA polymerase I can cause the enzyme to lose one of these activities. Match the type of loss‑of‑function mutation in DNA polymerase I provided on the left-hand side with the correct corresponding consequence of the lost activity from the dropdown options provided on the right-hand side below.
DNA polymerase synthesizes DNA in the 5' to 3' direction, and DNA polymerase requires a primer for DNA synthesis
The "end‑replication problem" (telomere problem) exists in eukaryotic chromosomes and is characterized by the chromosomes shortening with each round of DNA replication in somatic cells. Explain why the "end-replication problem" occurs with eukaryotic chromosomes.
synthesis of a short RNA sequence that initiates DNA synthesis
DNA replication involves multiple steps that require different enzymes. Which process does primase catalyze in DNA replication?
Recruiting low fidelity polymerases allows replication to continue when distortions in the DNA template strand stall synthesis by high fidelity polymerases.
Why do cells maintain low fidelity, error-prone DNA polymerases for certain tasks even though more accurate, high fidelity DNA polymerases are available?
An endonuclease cleaves one of the two DNA strands at a specific site creating a free 3' hydroxyl end and a free 5' phosphate end, 3' end of one DNA strand begins covalent extension using the intact DNA strand as a template, template strand rolls as extension of the 3' end of one of the DNA strands continues, DNA strand that is displaced is cleaved off and the cleaved DNA strand then undergoes circularization
4 steps in the process of rolling-circle DNA replication
DNA polymerase can only synthesize DNA in the 5' -> 3' direction and the direction of DNA synthesis is opposite to the direction of unwinding
Lagging strand synthesis occurs discontinuously because
a reaction between the free 3' hydroxyl of the growing strand and the 5' phosphate of a free nucleotide
Free nucleotides are added to a growing daughter DNA strand by
Telomerase would lose the ability to synthesize new telomeric sequences to extend the telomere, telomerase would be unable to correctly associate with telomeres
The enzyme telomerase is part protein and part RNA. What would be the most likely effect of a large deletion in the gene that encodes the RNA part of telomerase, and how would the function of telomerase be affected?
linear eukaryotic replication
Which mode of DNA replication is bidirectional, both continuous and discontinuous, and has multiple replicons?
continuous cell division
What would be the likely direct result of activation of telomerase in somatic cells?
RNA has a 2' OH group in addition to a 3' OH group in the sugar of its nucleotides
RNA is different from DNA in which of the following ways?
pyrimidine (base) deoxyribose (sugar), purine (base) deoxyribose (sugar), and pyrimidine (base) ribose (sugar)
Match the label corresponding to an image of a nucleotide or nucleoside
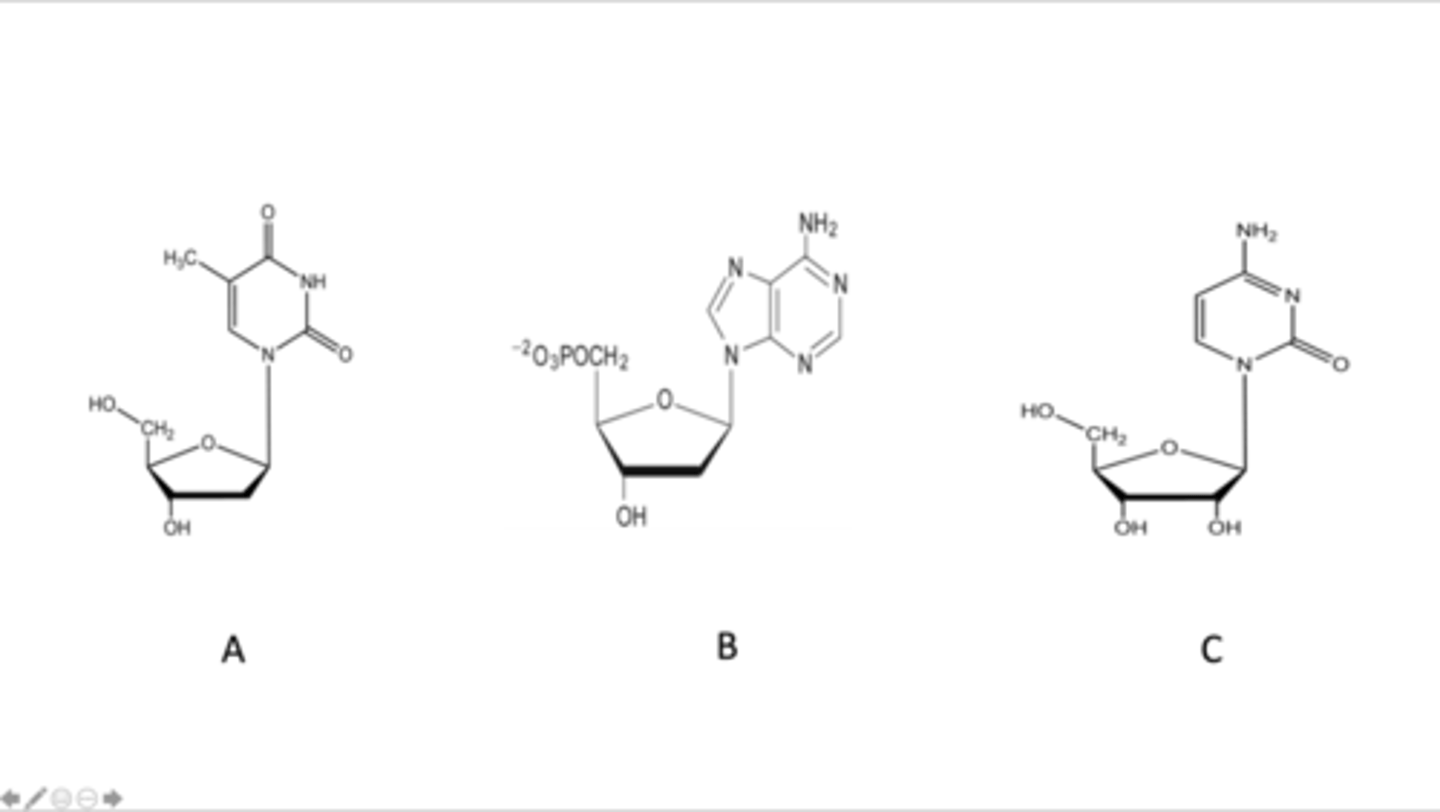
Bottom strand, bottom strand, top strand
RNA synthesis occurs from left to right for gene A and gene B, whereas RNA synthesis occurs from right to left for gene C.
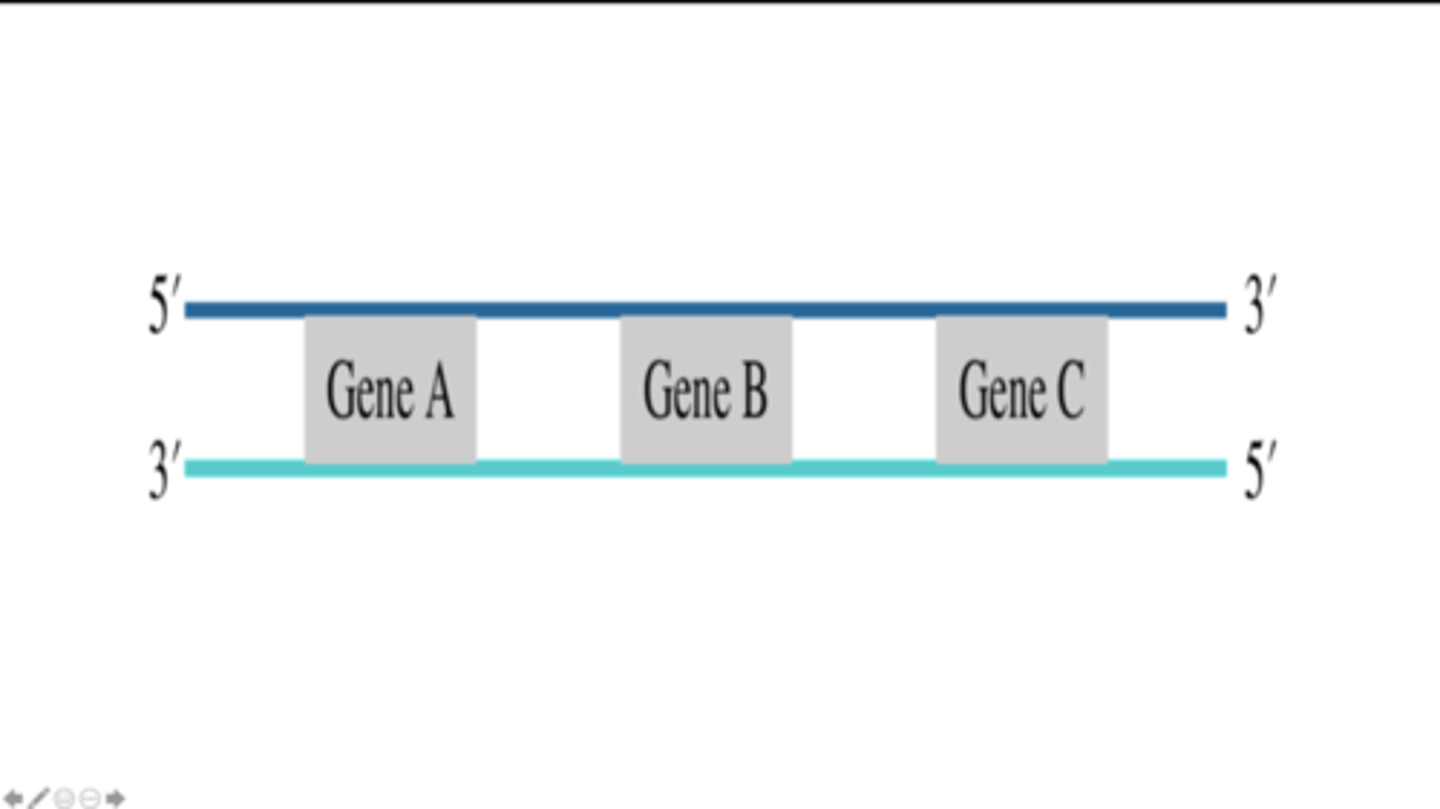
5′-GCAUAUGCGGUAC-3′
Shown below is DNA nucleotide sequence that is part of the RNA-coding sequence of a transcription unit. If the bottom strand is the template strand that is transcribed, then what is the nucleotide sequence of the RNA that is transcribed from this DNA sequence?
5′-GCATATGCGGTAC-3′
3′-CGTATACGCCATG-5′
RNA polymerase is recruited to the promoter region of a gene and the DNA at the promoter unwinds forming a transcription bubble
What happens during the initiation step of transcription?
RNA polymerase leaves the promoter region and moves along the template strand of the DNA creating an mRNA strand
What happens during the elongation step of transcription?
The mRNA detaches from the RNA polymerase as the RNA polymerase leaves the DNA strand
What happens during the termination step of transcription?
T/A G C A A T T
What is the consensus sequence for the following set of nucleotide sequences?
1. AGGAGTT
2. AGCTATT
3. TGCAATA
4. ACGAAAA
5. TCCTAAT
6. TGCAATT
5'ACUACGGAU3'
What is the nucleotide sequence of the mRNA transcribed from the template strand (top strand) of the following DNA segment?
5'ATCCGTAGT3'
3'TAGGCATCA5'
Silencer, core promoter, enhancer, regulatory promoter
Which sequences play a role in transcription of RNA polymerase II-dependent genes?
RNA polymerase moves along the DNA template in a 3' to 5' direction, unwinding the DNA and synthesizing RNA in a 5' to 3' direction
Which of the following statements regarding the direction of movement of RNA polymerase along the template DNA strand and the direction of synthesis of RNA is true?
Eukaryotes use general transcription factors for initiation of transcription, whereas bacteria use sigma factor for initiation of transcription
How does transcription differ between bacteria and eukaryotes?
a hairpin loop in RNA via inverted repeats, and a stretch of A-U bonds between the DNA template and RNA
During the rho-independent termination of transcription in bacteria, the formation of which of the following is critical?
general transcription factors
In eukaryotic cells, the core promoter consensus sequences of genes are recognized by accessory proteins that recruit a specific RNA polymerase. Which of the following types of accessory proteins serve this purpose?
Promoter only
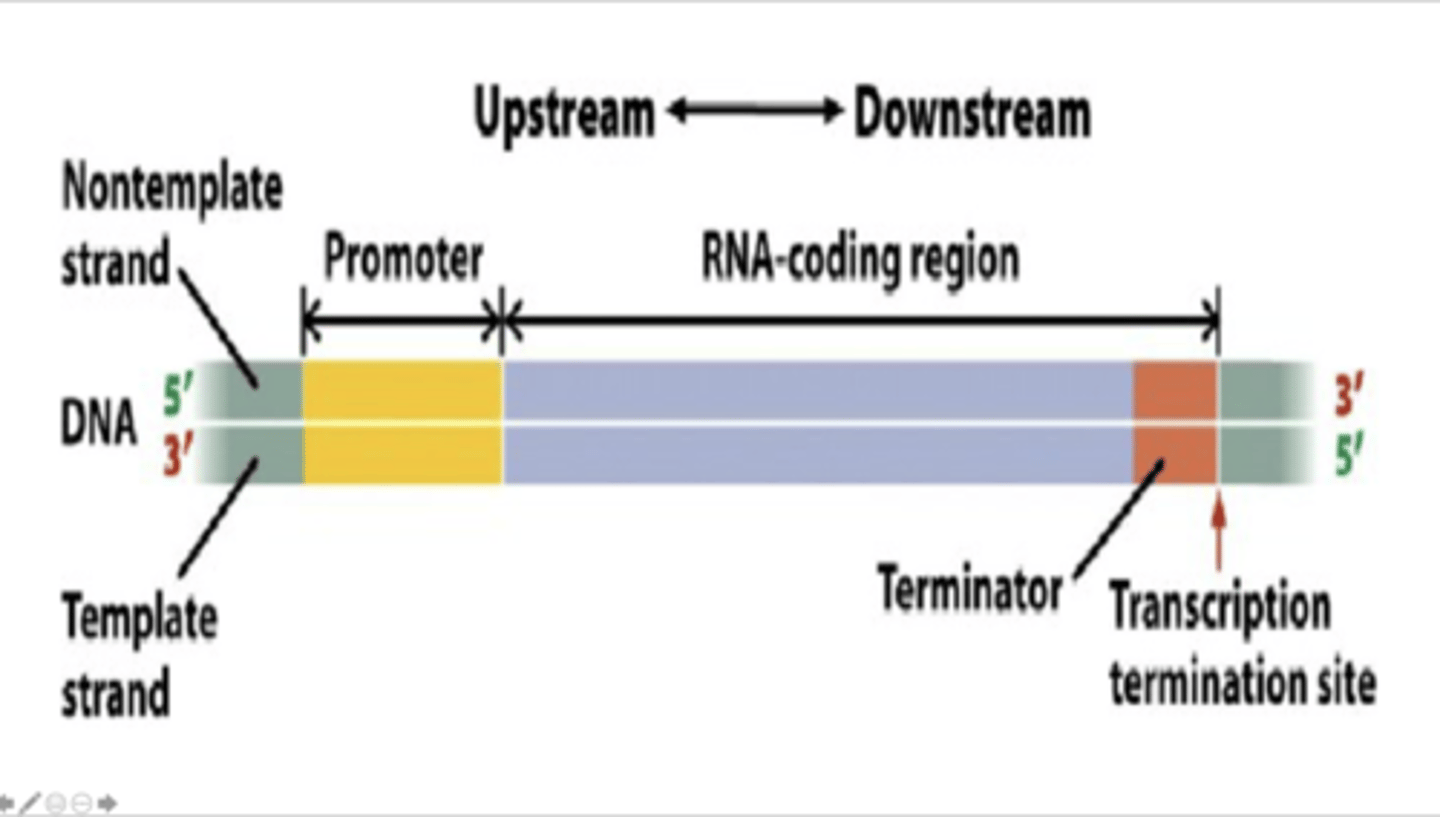
The diagram below represents a bacterial transcription unit comprised of three critical regions: the promoter, the coding region, and the terminator. Which region(s) would not be present in the final RNA transcript?
transcriptional activators
What type of proteins recognize and bind to eukaryotic upstream regulatory promoter consensus sequences?
-code for the amino acid sequence of a protein: exons only
-generally absent from bacterial genomes: introns only
-removed from the primary RNA transcript prior to translation: introns only
-present in the DNA strand used as a template for transcription: both introns and exons
-part of a eukaryotic mRNA strand: exons only
-present in eukaryotic genomes: both introns and exons
Match each description on the left-hand side with whether the description is true of either introns only, exons only or both introns and exons from the dropdown options provided on the right-hand side.
transcription start site, 5' UTR, start codon, splice branch point, stop codon, 3' UTR
Suppose that RNA polymerase was transcribing a eukaryotic gene with several introns all contained within the coding region. In what order would the RNA polymerase encounter the following elements?
exon, intron, 5' cap, poly-A tail
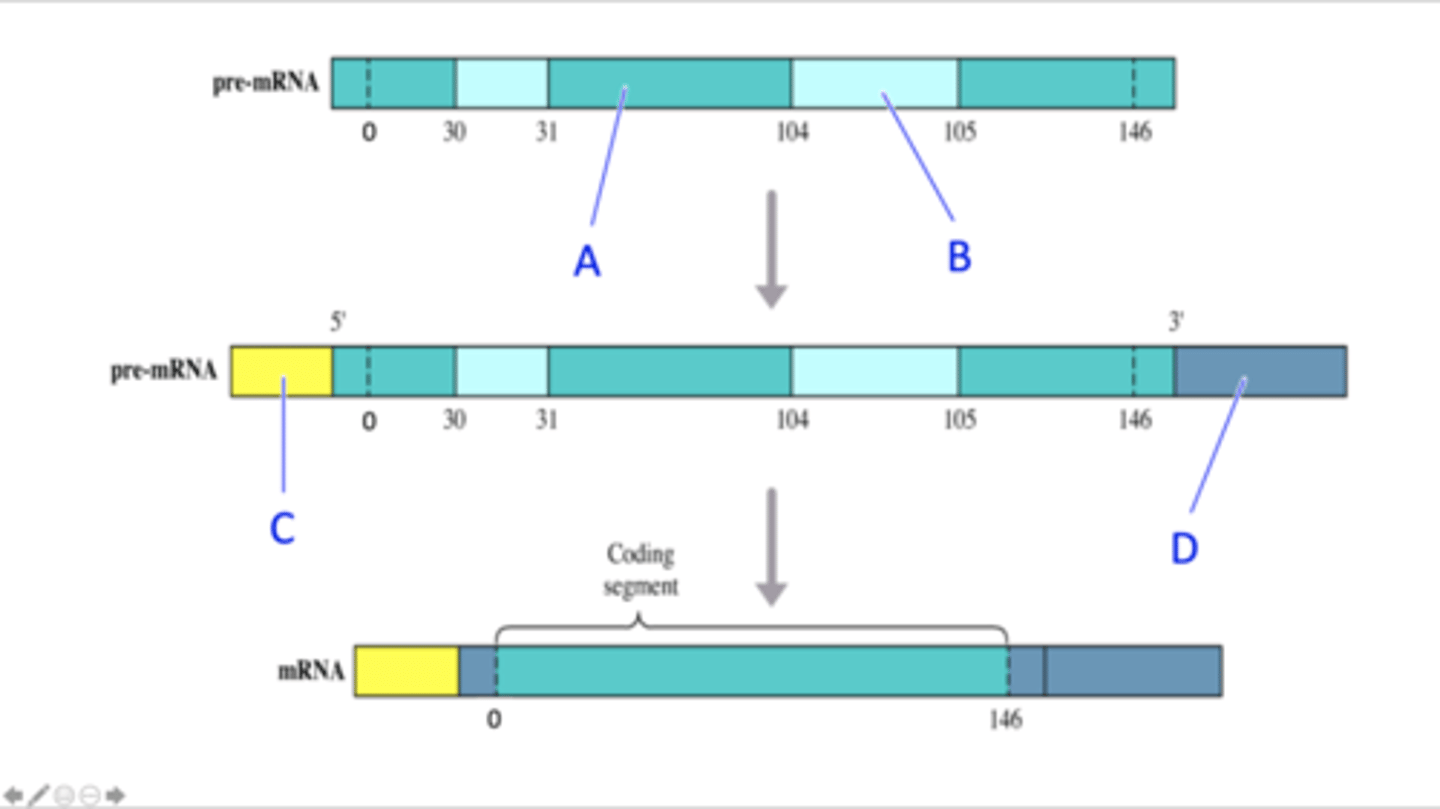
The diagram below depicts the steps in the processing of pre-mRNA into mRNA in eukaryotes. The different parts of the pre-mRNA and the different modifications that occur during processing of the pre-mRNA are indicated by different colours in the diagram. The numbers in the diagram represent the codon numbers (the first codon of the coding region is assigned the number 0, the second codon is assigned the number 2 etc.) and the positions of codons 0, 30, 31, 104, 105 and 146 are indicated in the diagram). What part of the pre-mRNA or modification of the pre-mRNA during processing are indicated by the labels A, B, C and D in the diagram?
6 exons and 5 introns
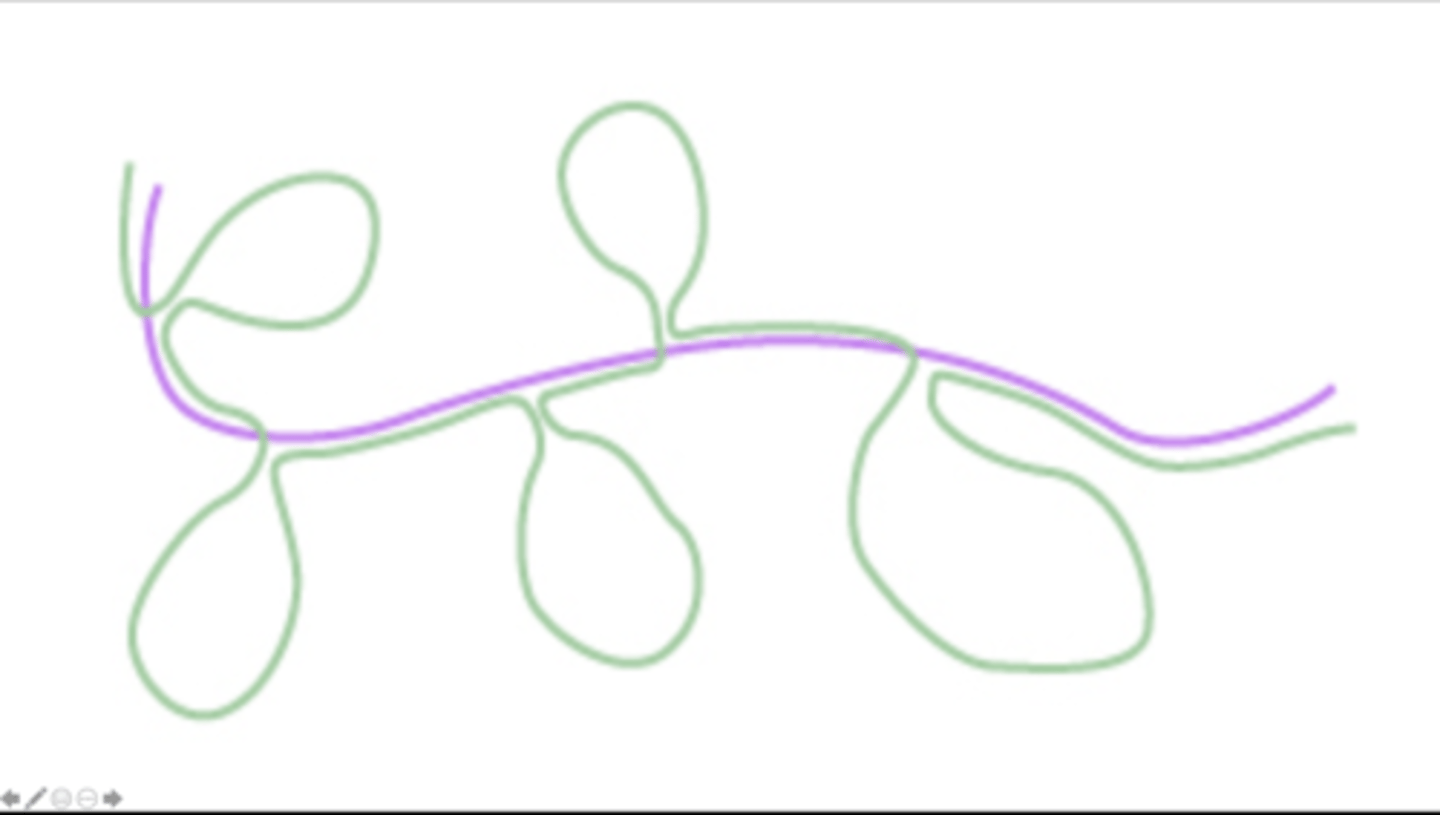
Below is a diagram of the results from a DNA-RNA hybridization (hybridization refers to the formation of hydrogen bonds between the complementary bases of the nucleotides of two nucleic acid strands) experiment for an unknown gene. For this experiment, a mixture of DNA and mRNA corresponding to the gene has been heated to a point of denaturation (denaturation which separates the two DNA strands was carried out by heating to a high temperature that breaks the hydrogen bonds between the complementary bases of the two strands of DNA) and cooled such that the mRNA (depicted in pink colour in the diagram) can hybridize with the template DNA strand (depicted in green colour in the diagram). Based on the diagram, what is the total number of exons and introns contained in the unknown gene?
The spliceosome produces a mature mRNA by connecting two exons and releasing the intron
What is the function of the spliceosome?
processing of exons in mRNA that results in a single gene coding for multiple proteins.
Which description below applies to alternative mRNA splicing?
In the nucleus, an mRNA copy of a gene is produced, which ribosomes use as instructions to synthesize a specific protein
How do the nucleus and ribosomes work together to generate a protein?
the action of microRNAs that inhibit translation of target mRNA molecules
In eukaryotic gene regulation, RNA interference can occur through
to degrade target mRNA molecules, and to downregulate gene expression
What is the function of siRNA?
The protein translated from the edited mRNA has a sequence different from that specified by the corresponding gene
What is the outcome of RNA editing?
mRNAs of different lengths from the same pre‑mRNA
Alternative splicing produces
the Shine-Dalgarno sequence
Which RNA sequence does not play a role in intron splicing?
the consensus sequence, AAUAAA
What is necessary for the addition of the poly(A) tail to the pre-mRNA?
alpha-carbon atom, R group, amino group, and carboxyl group
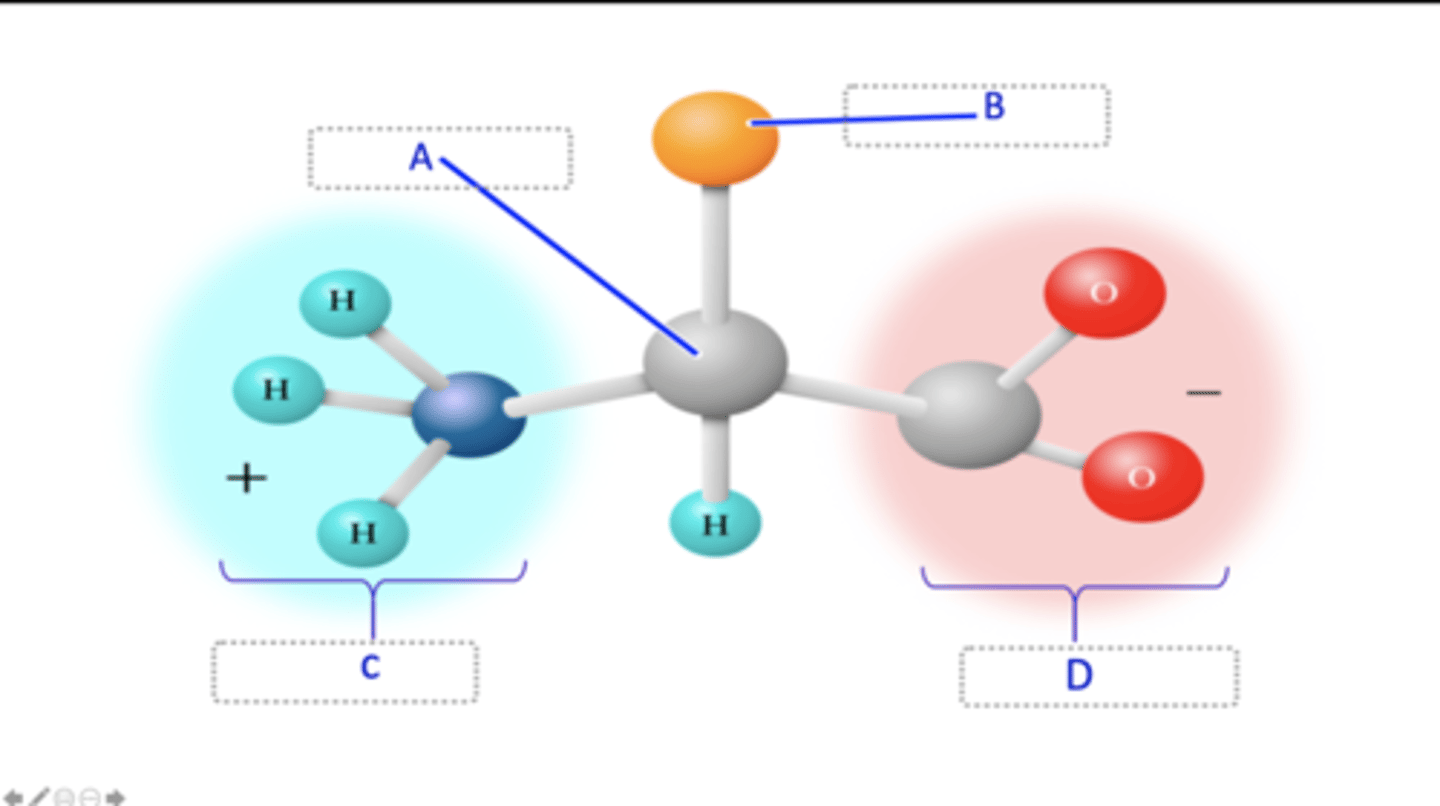
The diagram below is of the structure of a prototypical amino acid. The different parts of this structure are labelled A - D. Match each label (A - D) of the parts of the structure of an amino acid shown on the left-hand side below with the correct name of the part from the dropdown options on the right-hand side below.
R group
Which part of the structure of an amino acid is responsible for the unique properties of an amino acid?
UGC
What is the sequence of the anticodon, from the 3' to 5' end, of the tRNA in the A site?
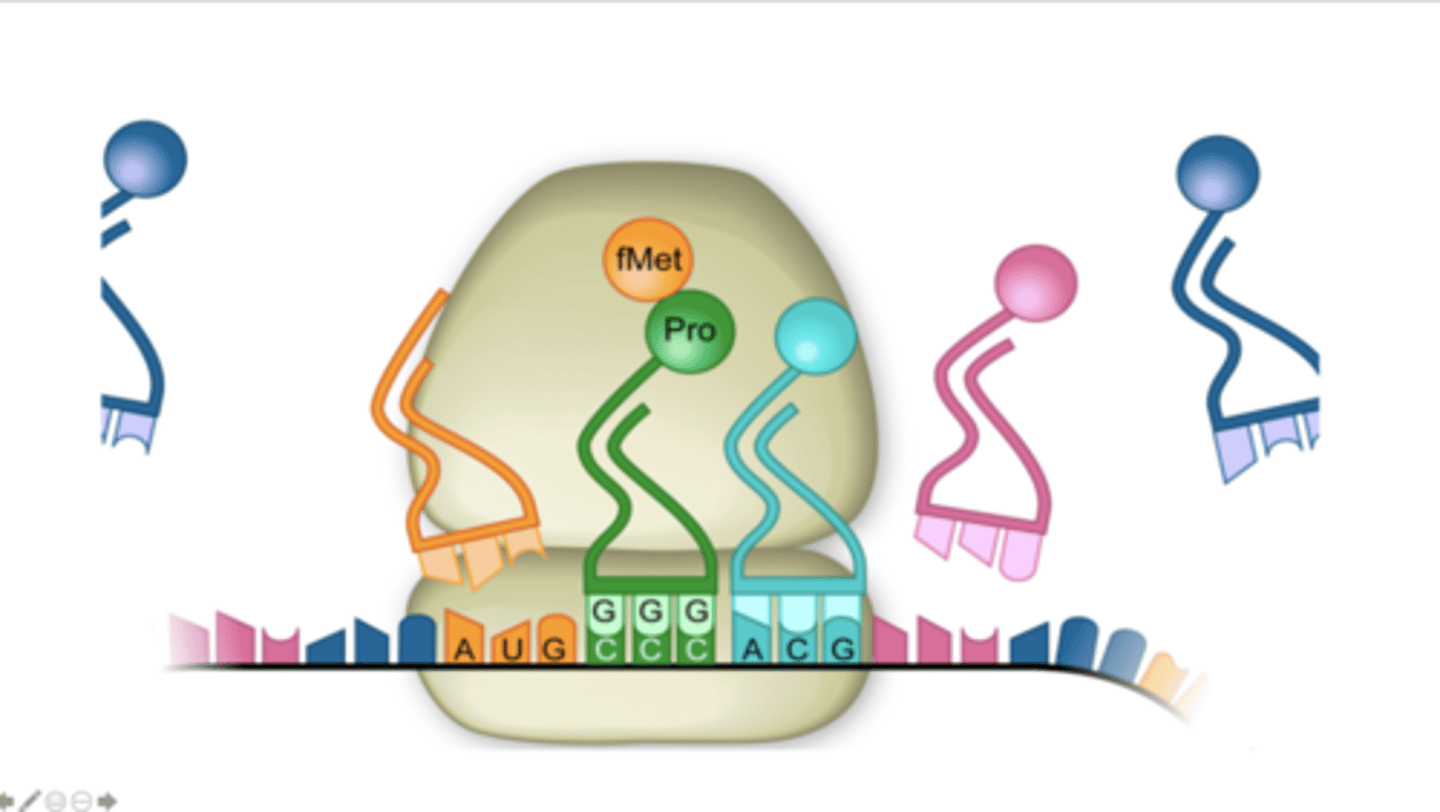
Thr
What is the next amino acid added to the growing polypeptide chain (the growing polypeptide is attached to the tRNA in the P site)?
The small ribosomal subunit complexes with a specific tRNA and specific proteins, and the complex then binds the mRNA and scans for a start codon
Which of the events given in the options below occur during eukaryotic translation initiation?
Specific tRNAs bind specific codons in turn and the ribosome adds amino acids from each tRNA to the growing polypeptide chain
Which of the events given in the options below occur during eukaryotic translation elongation?
The ribosome dissociates from the mRNA after the stop codon is recognized by a Release Factor
Which of the events given in the options below occur during eukaryotic translation termination?
-single DNA strand is used to produce mRNA: transcription
-described as semiconservative: DNA replication
-both DNA strands are duplicated: DNA replication
- requires tRNA: translation
- amino acids added to growing polypeptide chain: translation
-requires an RNA primer to be initiated: DNA replication
Match each phrase or term given below on the left-hand side to whether the phrase or term relates to either DNA replication, transcription or translation from the dropdown options on the right-hand side
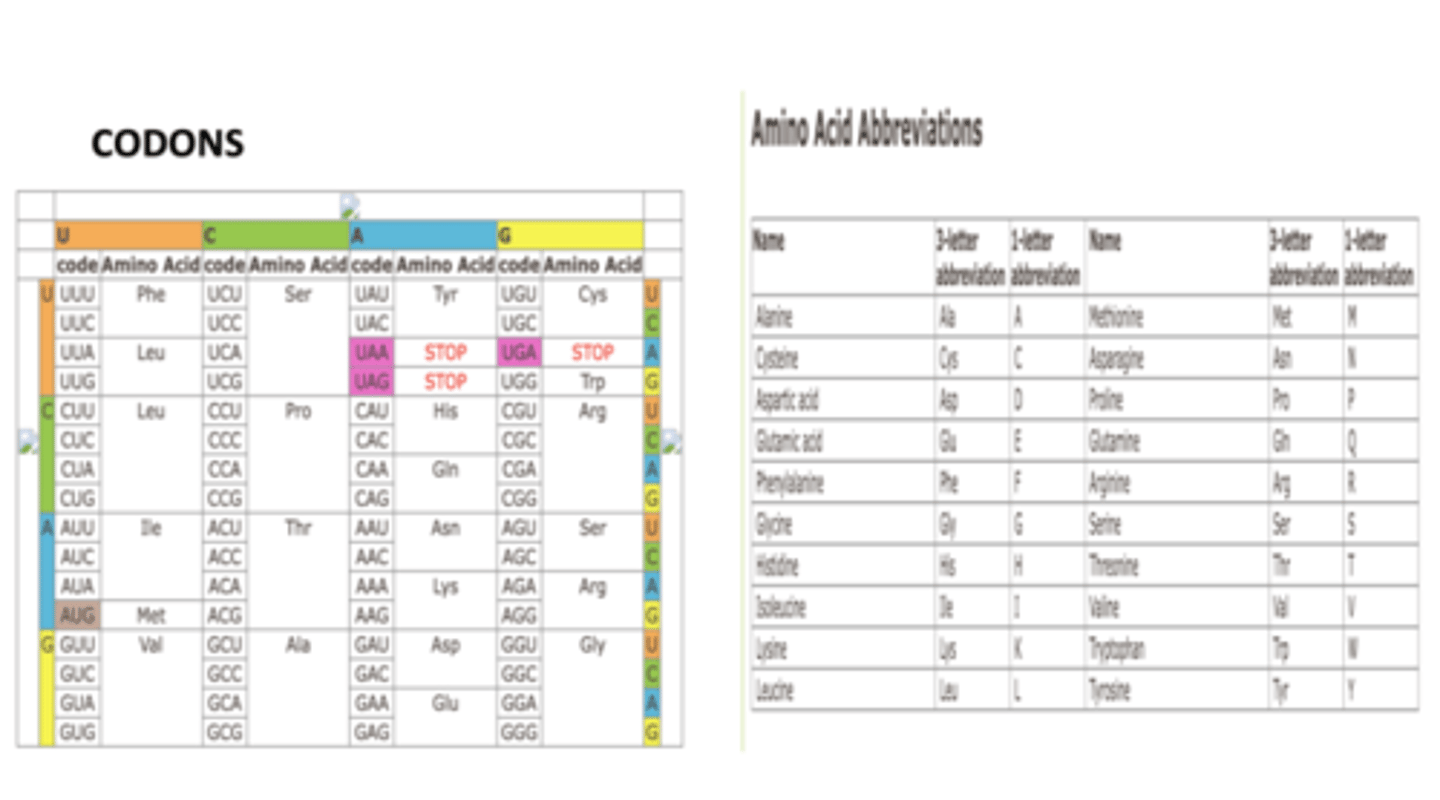
more than one codon can specify an amino acid
The genetic code is said to be degenerate, which means:
The charging of tRNAs by aminoacyl tRNA synthetases occurs within the ribosome
Which of the options provided is false regarding the charging of tRNAs by aminoacyl tRNA synthetases?
the structure formed is called a polyribosome, and they all produce proteins with the same number of amino acids
When multiple ribosomes simultaneously work along a single RNA molecule, translating the RNA molecule
the initiation codon
The correct reading frame of a nucleotide sequence is established by
The 16S rRNA of the small ribosomal subunit binds directly to the Shine‑Dalgarno sequence in prokaryotes
Which of the following statements concerning the Shine‑Dalgarno sequence is true?
EF-Tu
Which of the following elongation factors (EFs) binds to a charged tRNA?
The ribosome encounters a termination codon in both prokaryotes and eukaryotes, and RF1 or RF2 binds the A site of the ribosome in prokaryotes
Which of the following events contribute to the termination of translation?
5'-ACC-3', and 5'-ACU-3'
A tRNA has the following anticodon sequence 5'-GGU-3'. What are the possible codons that can pair with the anticodon of this tRNA?
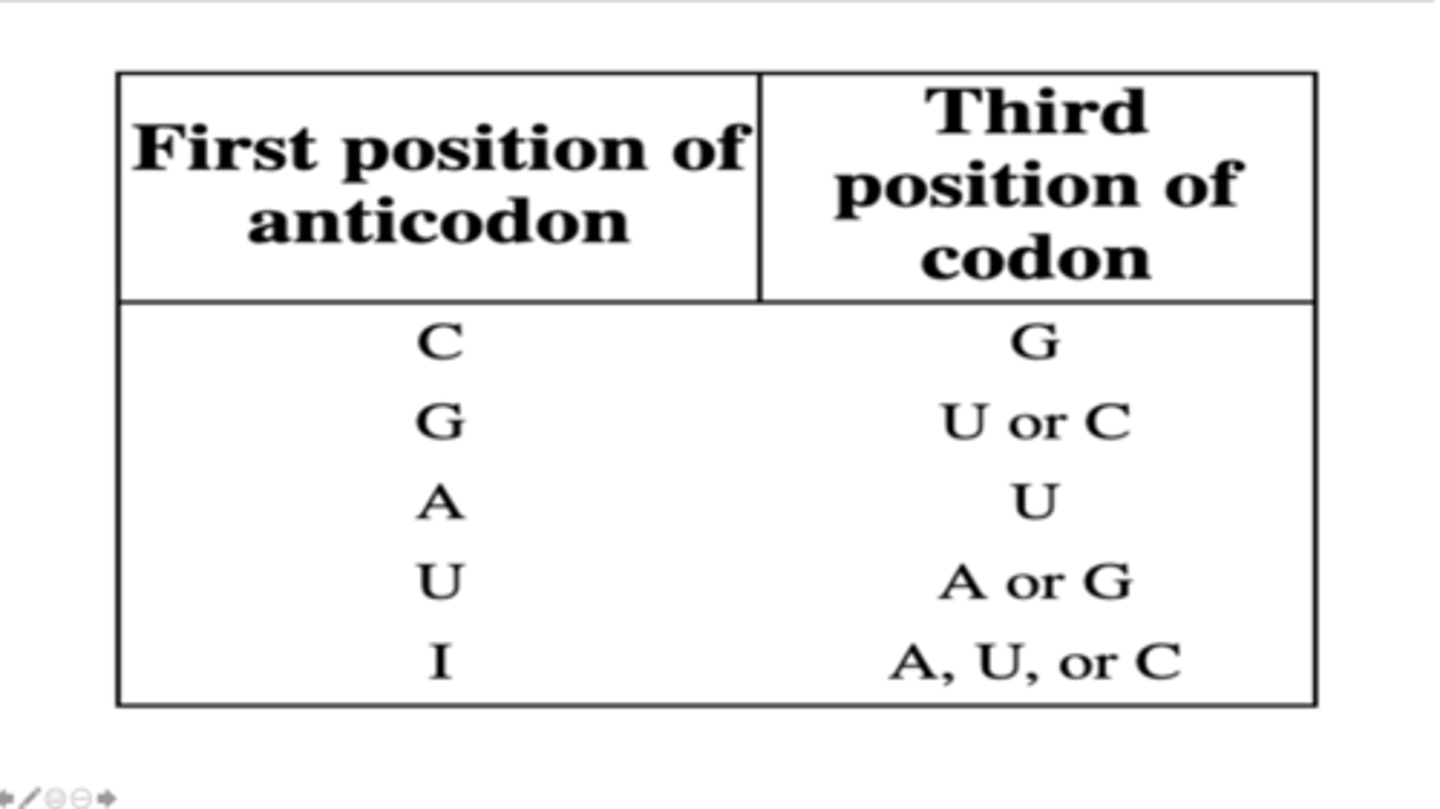
5'-GUC-3', 5'-GUU-3', and 5'-GUA-3'
A tRNA has the following anticodon sequence 5'-IAC-3'. What are the possible codons that can pair with the anticodon of this tRNA?
5'-ACG-3' and 5'-ACA-3'
A tRNA has the following anticodon sequence 5'-UGU-3'. What are the possible codons that can pair with the anticodon of this tRNA?
5
Imagine that you discover a bacterial species on a meteorite. The genetic material of this bacterial species is made up of 4 different types of nitrogenous bases and this species contains 911 different amino acids. What would the minimum number of bases need to be in a codon of this species in order to code for all of the 911 amino acids?
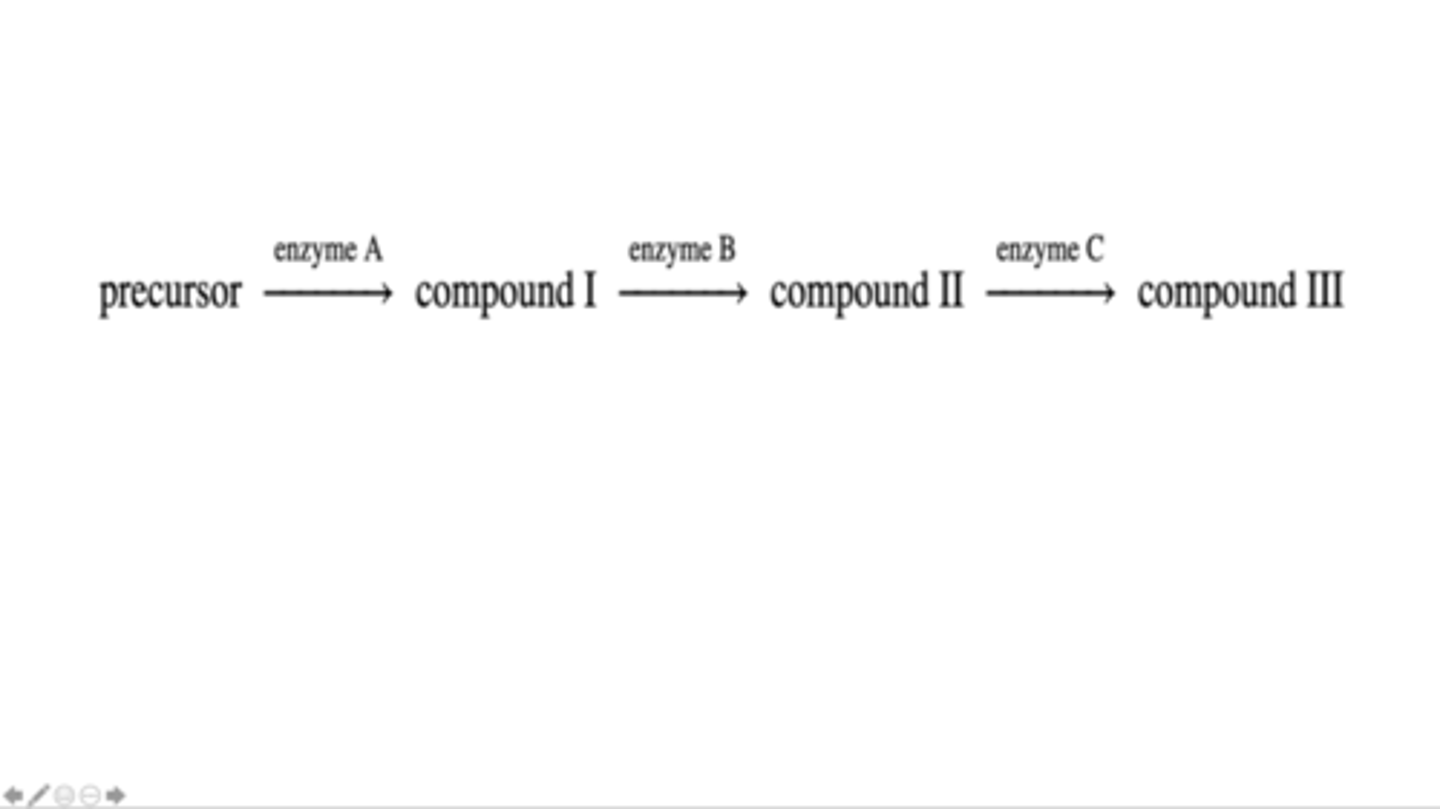
enzymbe B only
Compounds I, II and III are in a biochemical pathway in a particular bacterial species. This biochemical pathway begins with a precursor and is shown below. In this pathway, enzyme A catalyzes the conversion of the precursor into Compound I, enzyme B catalyzes the conversion of Compound I into Compound II and enzyme C catalyzes the conversion of Compound II into Compound III. An auxotrophic mutant of this species was discovered that could survive and grow in minimal medium only when the minimal medium was supplemented with either Compound II or Compound III. This mutant could not survive and grow when the minimal medium was supplemented with Compound I. Which enzyme(s) is (are) likely to be functionally mutated in this mutant?
DNA in a prokaryotic cell is stored in multiple chromosomes.
Which of the following statements about prokaryotic and/or eukaryotic cells is false?
G1 : preparation for DNA replication
Which of the following options represents a correct match between a phase of the cell cycle and an event that occurs in that phase?
transition points during the cell cycle that ensure that cellular components are functioning properly.
What are checkpoints in the cell cycle?
G2/M checkpoint : detection of any DNA damage prior to DNA replication
Which of the following options represents an incorrect match between a cell cycle checkpoint and its function?
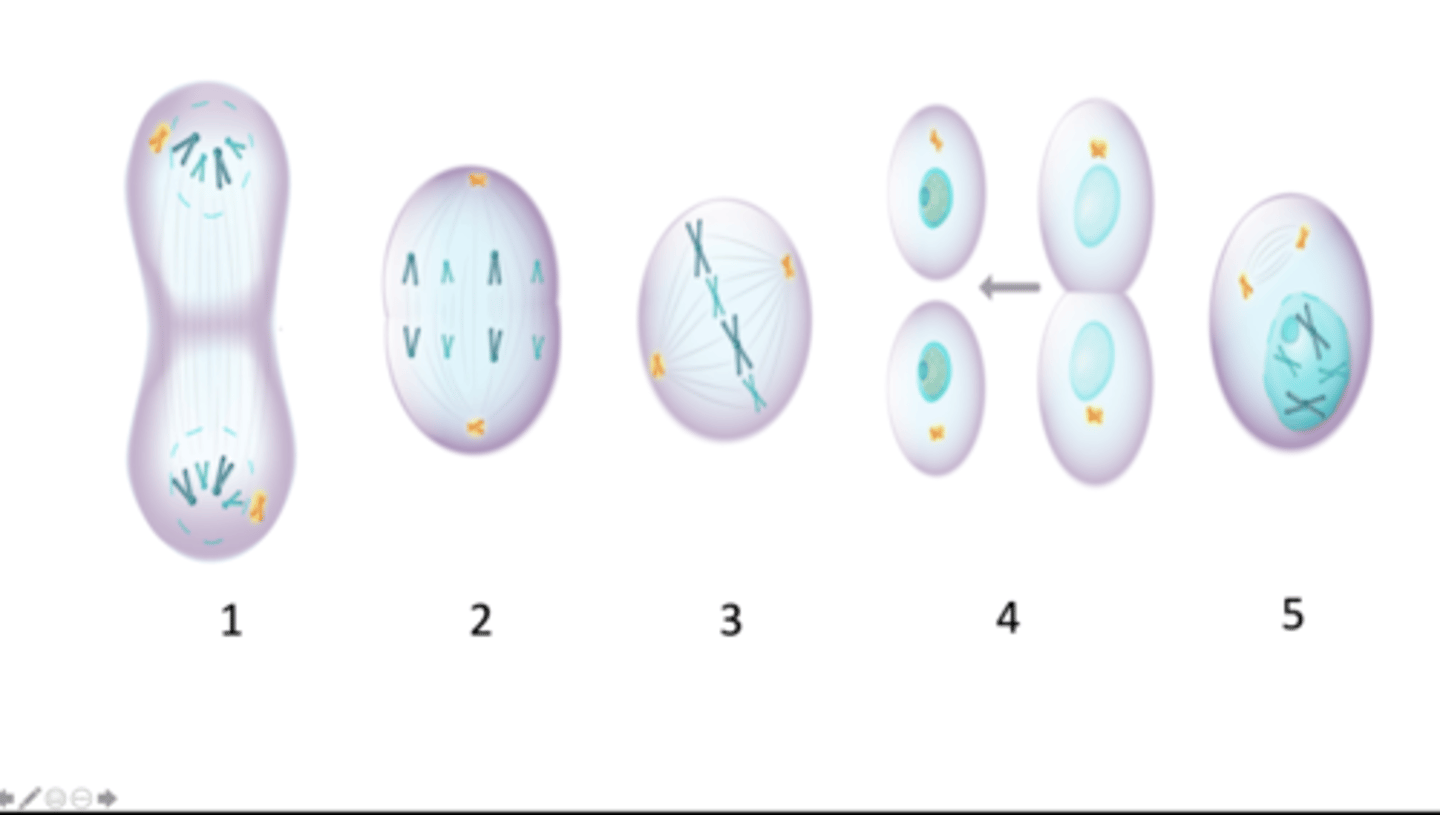
telophase, anaphase, metaphase, cytokinesis, prophase
Below are images (labelled 1 - 5; label is underneath an image) of a cell at different stages of the M phase of the cell cycle. The cell is a somatic cell from a diploid organism containing 4 chromosomes in total. Homologous chromosomes are represented in the same colour. Which of the options represents the correct combination of image (identified by its numeric label) and name of the stage of M phase?
two genetically identical daughter cells
Which is the product of division of a single cell by mitosis?
Sister chromatids of each chromosome separate from each other and migrate towards opposite poles of the cell
What happens during anaphase II of meiosis?
Chromosomes align as homologous pairs in the middle of the cell
What happens during metaphase I of meiosis?
Members of each pair of homologous chromosomes separate from each other and migrate towards opposite poles of the cell
What happens during anaphase I of meiosis?
Individual chromosomes align in the centre of the cell
What happens during metaphase II of meiosis?
14.4 minutes
A biologist examines a population of cells of a particular cell type in a particular tissue of an organism. The biologist counts 160 cells in interphase, 20 cells in prophase, 6 cells in prometaphase, 2 cells in metaphase, 7 cells in anaphase, and 5 cells in telophase. The complete cell cycle requires 24 hours. What is the average duration of metaphase in the cells of this cell type?
21 chromosomes and 42 DNA molecules
Somatic cells of rats (a diploid organism) contain 42 chromosomes. How many chromosomes and how many DNA molecules would the first polar body in rats have?
Sister chromatids of each chromosome separate from each other and migrate towards opposite poles of the cell during mitosis but not during meiosis
Which of the following options is not a difference between mitosis and meiosis?
8 chromosomes and 16 DNA molecules
A diploid cell has 8 chromosomes in G1 of interphase. How many chromosomes and how many DNA molecules are present per cell when this cell just completes metaphase of mitosis (just completes metaphase implies that the cell has completed metaphase but has not yet initiated the following stage of mitosis)?
8 chromosomes and 8 DNA molecules
A diploid cell has 8 chromosomes in G1 of interphase. How many chromosomes and how many DNA molecules are present per cell when this cell just completes anaphase II of meiosis (just completes anaphase II implies that the cell has completed anaphase II but has not yet initiated the following stage of meiosis)?
6
What is the diploid number of chromosomes (2n) in this plant species?
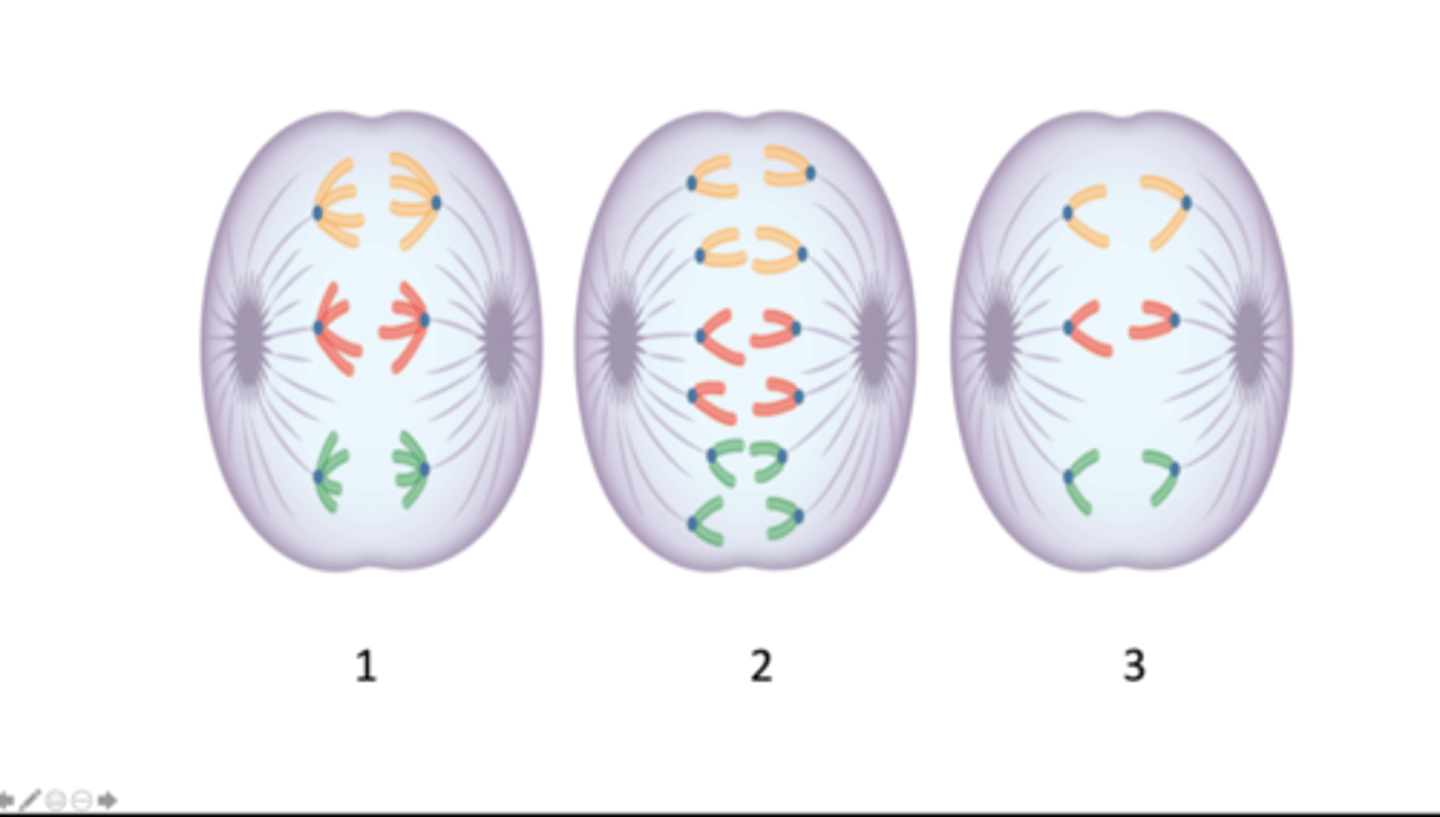
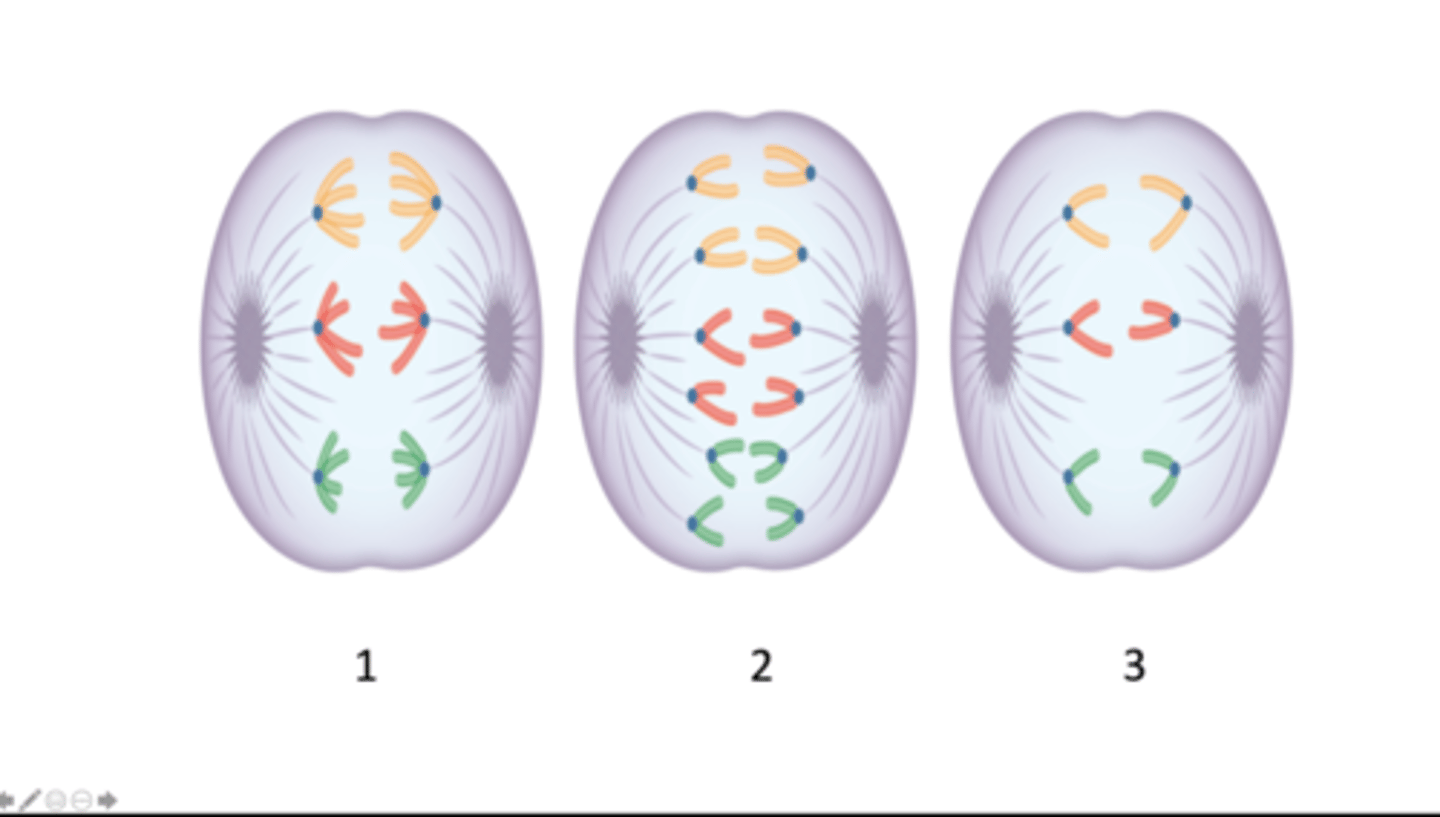
anaphase I of meiosis, anaphase of mitosis, anaphase II of meiosis
Which stage of mitosis or meiosis does each image represent?
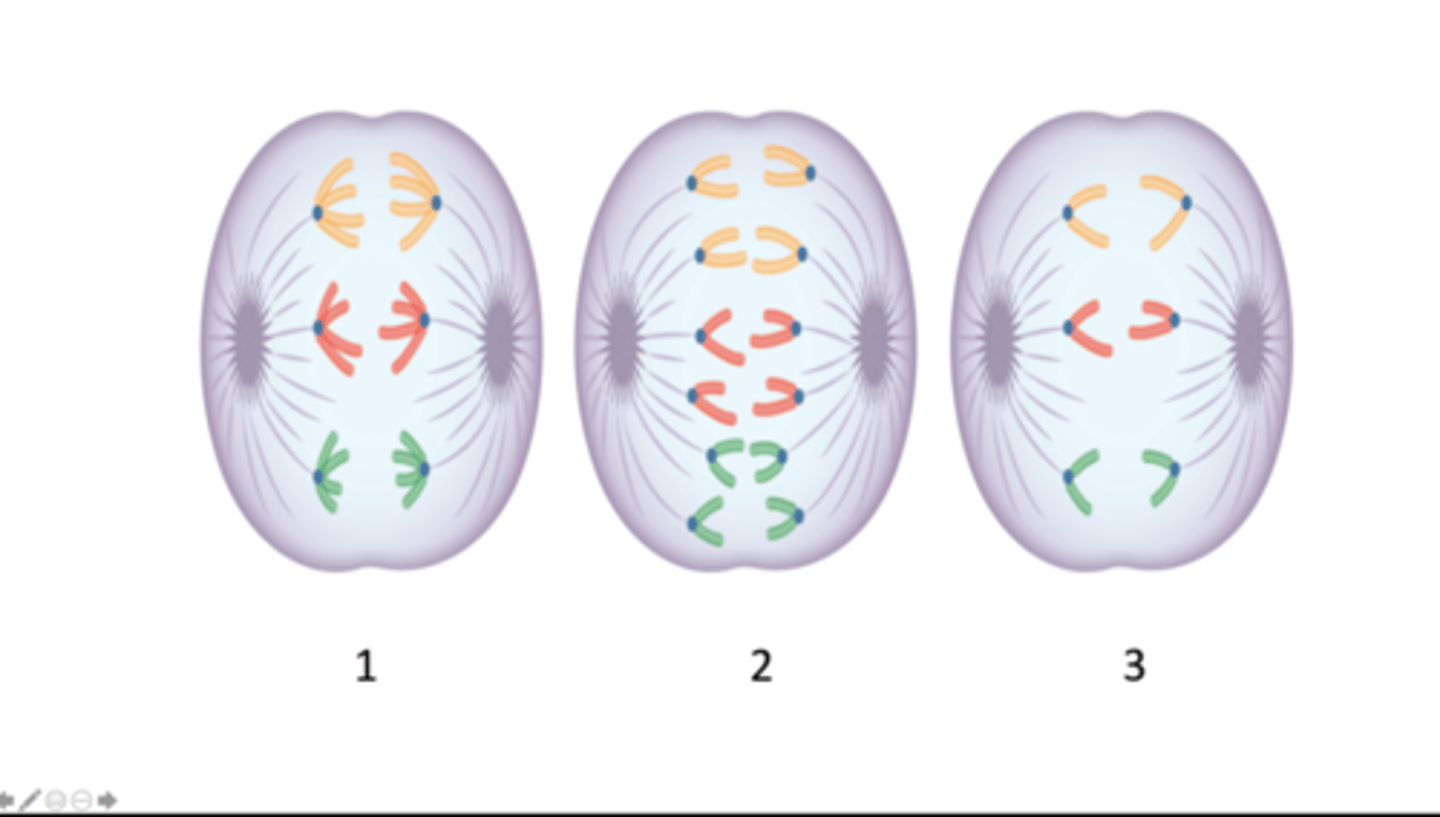
six, twelve, six
How many chromosomes are present in the cell depicted in each image 1 - 3?
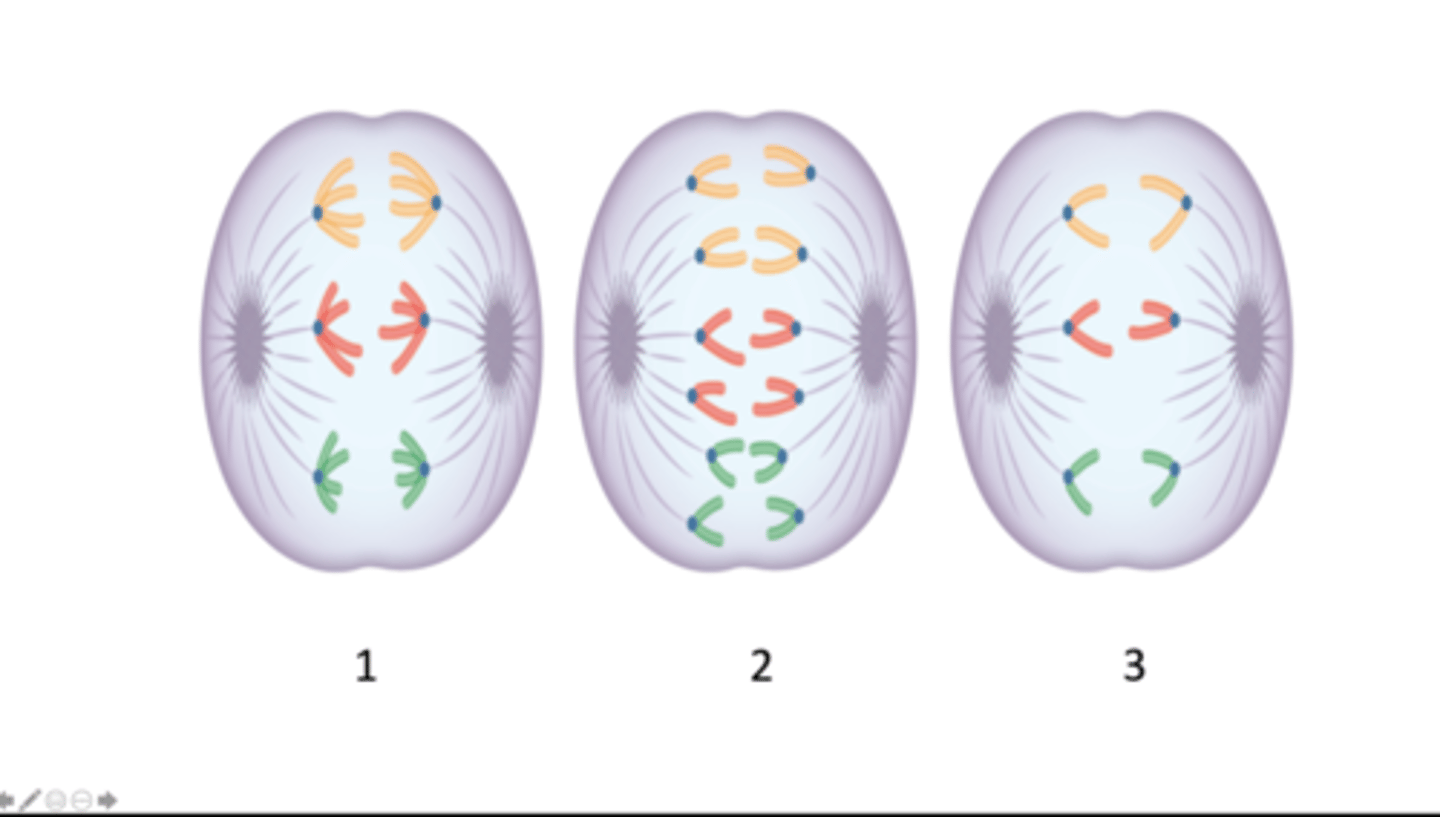
twelve, six, six
How many DNA molecules are present in the cell depicted in each image, 1 - 3?
asexual reproduction, growth of multicellular organisms, and replacement of damaged cells
What are the functions of mitotic cell division?
Gametic chromosomes have a different combination of alleles for different genes than parental chromosomes as a result of independent assortment that occurs during meiosis. Mitosis produces two genetically identical diploid daughter cells. The sister chromatids of a chromosome are not genetically identical as a result of crossing over during meiosis.
Meiosis and mitosis are both forms of cell division. However, the outcomes of these processes differ. Consider a diploid organism with two sexes. From the options below, select reason(s) why meiosis typically produces genetic variation or reason(s) why mitosis does not produce such genetic variation.
They have the same phenotype but different genotypes.
How is a true breeding round‑seeded pea plant different from a hybrid round‑seeded pea plant?
Bb × bb
Which of the crosses shown in the options below would likely produce a genotype ratio of 1:1? B and b are alleles at a single gene locus and allele B is dominant over allele b.
25%
In pea plants, at the seed shape gene locus, the allele for round seed shape, R, is dominant over the allele for wrinkled seed shape, r. Two pea plants that are heterozygous at the seed shape gene locus are crossed. What proportion of the offspring of this cross would be expected to have wrinkled seeds?
AA only
If an individual that has normal pigmentation mates with an individual that has the albino phenotype, then what is/are the possible genotype(s) that would not be expected in offspring of this mating?

AA, or Aa, or aa
If two individuals that have normal pigmentation mate, then what is a possible genotype in an offspring of this mating?

Orange (both RR and Rr)
In cucumbers, fruit colour is determined by a single gene. At this gene locus, orange fruit colour (allele R) is dominant over cream fruit colour (allele r). A cucumber plant homozygous for orange fruit is crossed with a plant that bears cream fruit. What phenotype(s) and genotype(s) would be expected in the offspring of a backcross between the F1 and the parent that produces orange fruit?
different pairs of homologous chromsosomes will segregate independently during anaphase I
Orange (both RR and Rr)
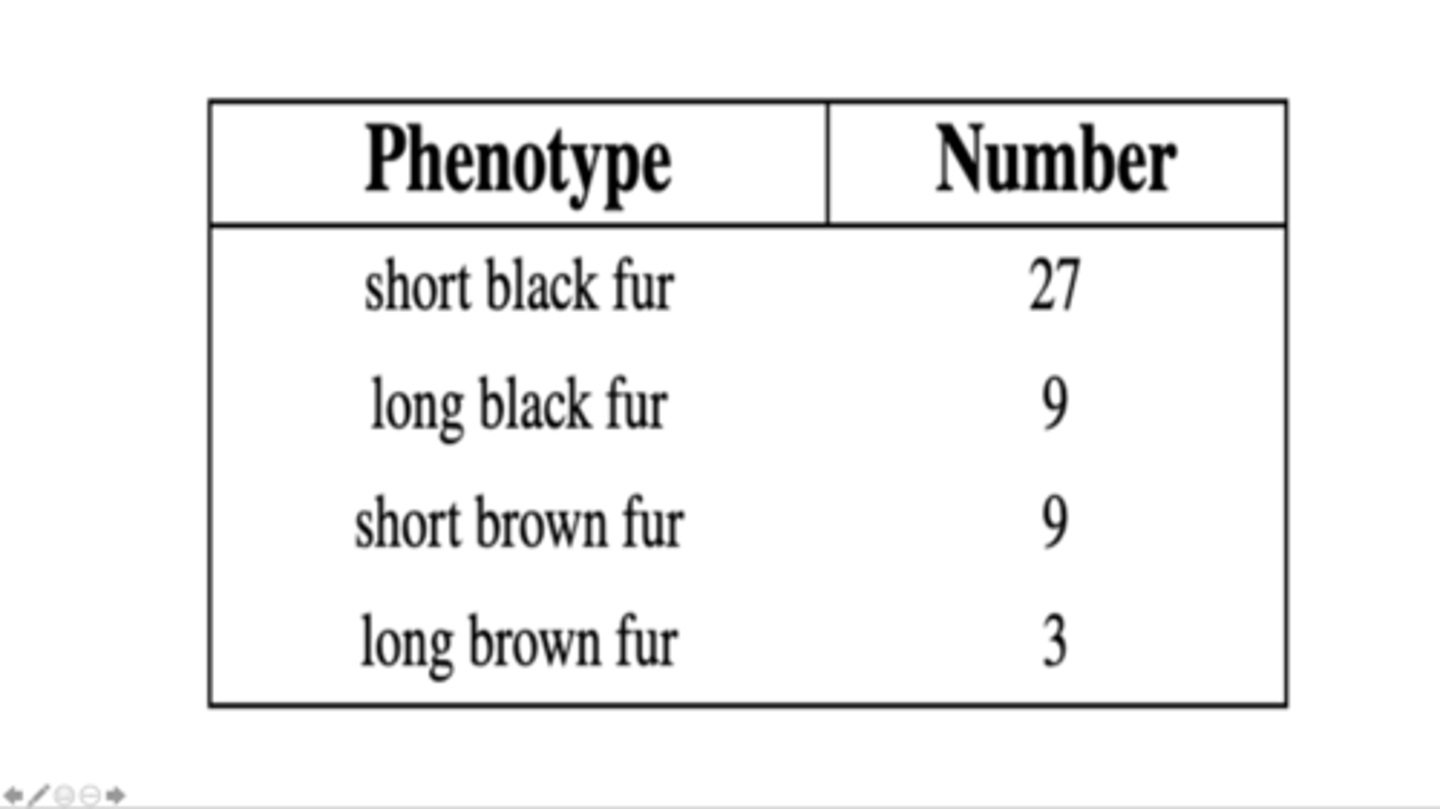
2 genes; 2 alleles for each gene
Determine the number of genes and alleles that control these phenotypes.
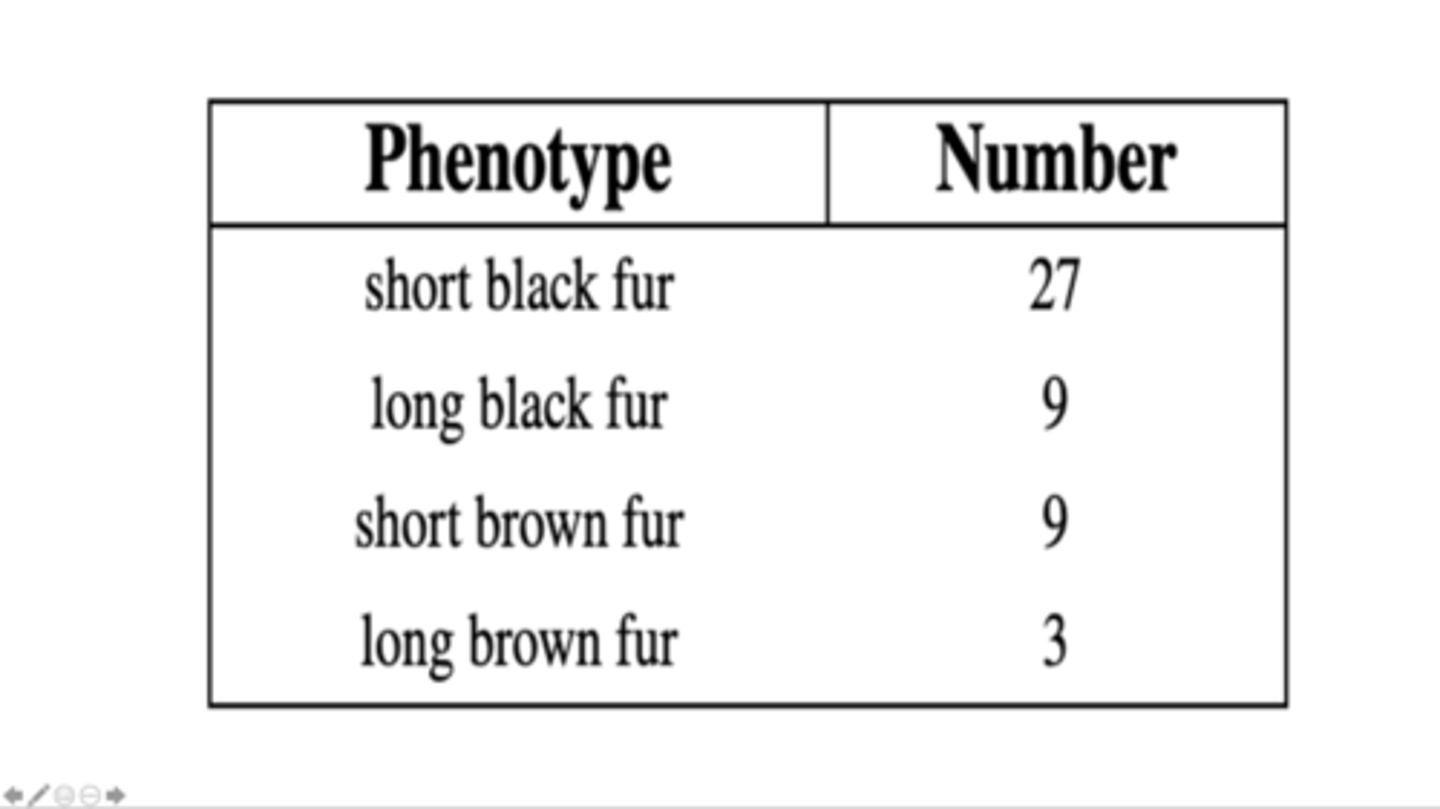
black fur and brown fur are coded by one gene; short fur and long fur are coded by a second gene
Which phenotypes are coded for by the same gene?
WwRr or Wwrr or wwRr or wwrr
In fruit flies, wing length and eye colour are characteristics encoded by distinct genes present on different types of chromosomes. With the wing length characteristic, long wings (allele W) is dominant over short wings (allele w). With the eye colour characteristic, red eyes (allele R) is dominant over orange eyes (alelle r). If a fly that is heterozygous for both traits is crossed with a fly that is homozygous recessive for both traits, then what is a possible genotype of an offspring of this mating?
42
If two pea plants that are heterozygous at both the seed colour and seed shape loci are crossed producing 224 offspring, then what is the expected number of yellow, wrinkled offspring?

25
If a pea plant that is heterozygous at both the seed colour and seed shape gene loci is crossed with a pea plant that is homozygous recessive at both of these gene loci, then what percentage of the offspring of this cross are expected to be yellow and wrinkled seeded?

6 yellow, round : 2 yellow, wrinkled : 6 green, round : 2 green, wrinkled
If a pea plant that is heterozygous at both the seed colour and seed shape gene loci is crossed with a pea plant that is green seeded and heterozygous at the seed shape locus, then what is the phenotypic ratio of the offspring of this cross?
