BIO 4
1/110
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
111 Terms
DNA is inside
the nucleus
Chromosomes
human genome containing 46 chromosomes arranged in 23 pairs
Chromosomes are made of
DNA proteins (histones) packed together
genomes
complete set of genetic information
double-helix
two polynucleotide strands, twist about one another
strands are antiparallel
sugar phosphate backbone
Bases extend towards
middle of the helix
complementary base pairing
base hydrogen bond with another
A pairs with T (2 hydrogen bonds)
G pairs with C (3 hydrogen bonds
A with T has how many hydrogen bonds
2
G with C has how many hydrogen bonds
3
one molecule of DNA corresponds to
1 chromosome (two strands together)
5 prime carbon
where a phosphate ends
DNA is the same
throughout the body, it only depends on which parts of the DNA is getting expressed
DNA replication (simple)
making a copy of the DNA in a cell
DNA replication (complex)
connecting nucleotides to form a long chain of DNA according to the template
each strand can serve as a template for the formation of a new, complementary strand
complementary nature of DNA double helix is important in replication
semi conservation of DNA replication
unwind and unzip DNA
old strand serves as a template
complementary nucleotides are added to form a new strand
result of replication of DNA
parent (old) strand serves as template
two daughter helices that are identical to parent helix and to each other
each daughter helix contains one old strand and one new strand
what catalyzes the polymerization of nucleotides
DNA polymerase
catalyzes the addition of nucleotides to existing 3 OH groups-requires primers
DNA synthesis proceeds
in the 5 to 3 direction ONLY
primers
short RNA starters are synthesized by primase (an RNA polymerase)
prokaryotes
single origin of replication
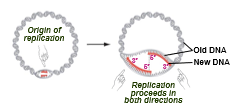
eukaryote
multiple points of origin
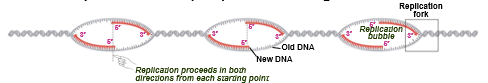
leading strand synthesis
continuous (5)
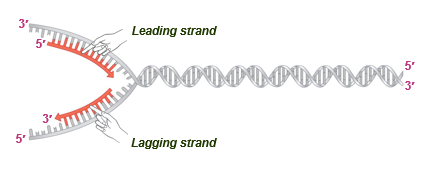
lagging strand synthesis
discontinuous (3)
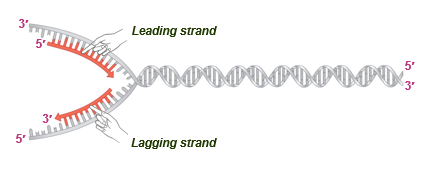
topoisomerase
relieves twisting forces
the target of antibiotics and anticancer agent
helicase
opens double helix
DNA is opened, unwound, and primed by
primase, topoisomerase, and helicase
primase
synthesizes RNA primer
DNA polymerase 3, in 5 to 3 directions, synthesizing leading strands
DNA lagging strand and DNA polymerase 3
DNA polymerase synthesizes first an Okazaki fragment and then another fragment as more template DNA is exposed
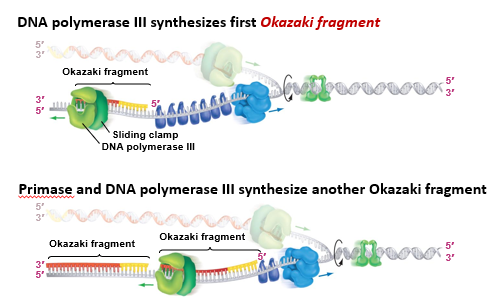
DNA polymerase 1
removes ribonucleotides of primer, replaces them with deoxyribonucleotides

DNA ligase
closes gap in sugar-phosphate backbone after DNA polymerase 1
Chromosome shortening (simple)
ends (telomeres) could shorten because of a problem with lagging strand synthesis
mismatched bases
errors may be made during synthesis
what can damage bases
UV light/ radiation damages nucleotides ‘light damage’
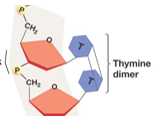
UV light induces covalent bonds between adjacent thymine’s forming a dimer
telomeres
ends of linear chromosomes
chromosome shortening (complex)
lagging strand is missing the end (cannot copy the end)
telomere shortening is a sign of aging (both cells and individual)
telomerase (made of RNA and protein) replicates the end chromosome
special cells turn on telomerase to keep dividing (stem cells, immune cells, cancer cells)
Copy mistakes and proofreading
DNA polymerase can repair mismatched bases after replication (mismatch repair)
felt through a ‘bump’ from mismatched pair
light repair
photolyase uses light energy to break covalent bonds between thymine dimers (reverse of above reaction)
new sunscreens/skin creams claim that they can do this
nucleotide excision repair
remove and fix damaged or wrong nucleotides (bases)
xeroderma pigmentosum- forms skin cancers from being unable to repair
genes
units of genetic information (DNA) that carry instructions for building polypeptides (proteins) or functional RNA molecules along with regulatory sequences
Gene Expression
process of converting archived information (DNA sequences) into molecules (proteins) that actually do things in cells
protein production
messenger RNA serves as
intermediary between genes and proteins
central dogma of molecular biology
by Francis Crick ‘the central dogma of molecular biology’
summary of the flow of information in cells
codes for RNA molecules that do not function as mRNA and are not translated into proteins
transfer RNA (tRNA)
ribosomal RNA (rRNA)
transfer RNA
interpreter molecules, transfers amino acids to the ribosomes
ribosomal RNA
components of ribosomes
information is not always in one direction
information flows from RNA back to DNA (retroviruses)
transcription
DNA to mRNA
complementary base pairing rules
process by which messenger RNA (mRNA) is made from a DNA template
translation
mRNA to protein
uses genetic code
Process by which proteins and peptides are synthesized from mRNA
genetic code
rules that specify the relationship between nucleotide sequence and an amino acid sequence
4 nucleotide bases specify into
20 amino acids
there is a 3 base code to ensure that there is enough
triplet code
each amino acids is coded for by a group of three bases
codon
group of three bases that specifies a particular amino acid
start codon
identifies the site at which protein synthesis should start
there is only one
stop codons
signify that protein synthesis is complete
there are multiple
the codon table
(type of dictionary) to translate mRNA sequences to amino acid sequences
redundant- more than one triplet may specify the same amino acid
unambiguous- each codon has only one meaning
conservative- first two bases of codons that specify same amino acid are usually identical
universal- same genetic code is used by all living things
mutation
any permanent change in an organism’s DNA
effects depend on location of mutation
point mutation
replacement of one nucleotide with another
silent mutation
does not alter amino acid sequence
missense (substitution) mutation
changes one amino acid to another
nonsense mutation
changes codon for an amino acid to STOP codon- polypeptide chain is too short thus non functional protein
base insertion or deletion (frameshift mutation)
alter reading frame of mRNA triplets
affects all codons positioned after site of insertion or deletion
results in non-functional protein
missense mutation example
red blood cells becoming sickle cell from one amino acid change
transcription only occurs on a
gene (not all DNA sequences are transcribed)
called a transcription unit
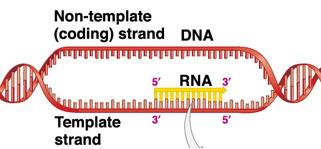
RNA polymerase travels along DNA template strands from
3’ to 5’ (RNA is going 5’ to 3’)
only one serves as a template for RNA, the other one is called coding strand
3 steps in transcription
initiation
elongation
termination
where RNA polymerase begins transcribing
by binding at a promoter
starts copying at +1 site
initiation
promoter
transcription factors or sigma factors
promoter
part of DNA, initiation site of transcription
short sequence of DNA that facilitates binding of RNA polymerase, enabled transcription of downstream genes
acts as ‘initiate transcription here’ signal
transcription factors or sigma factors
proteins
binds to promoter region of DNA
recruit RNA polymerase 2
initiation in prokaryotes
sigma factors (transcription factors in bacteria) binds to promoter
initiation in eukaryotes
basal transcription factor binds to promoter, and regulatory transcription factor binds to enhancer
together they recruit RNA polymerase
RNA polymerase
opens the helix, transcription begins, does not need a primer
copying starts at +1 site
elongation
sigma factor/transcription factor is released
RNA polymerase moves along RNA 3 to 5 synthesizing RNA in the 5 to 3 direction
termination in prokaryotes
transcription stops when RNA polymerase reaches a termination sequence (code for RNA that forms a hairpin)
termination in eukaryotes
transcription termination is triggered at poly(A) signal sequence
a tail of hundreds of A is added to the mRNA
in prokaryotes, transcription and translation are
tightly coupled
what has to happen to RNA transcription before translation
alteration of 5’ and 3’ ends
splicing out of introns
mRNA processing
in eukaryotes
mRNA transcription that is released from RNA polymerase is not ready to be translated it must first be processed
occurs in the nucleus before mRNA is exported to the cytoplasm for translation
addition of the 5’ cap
addition of the 3’ poly(A) tail
removal of introns
5’ cap
composed of modified guanine nucleotide
serves as a recognition signal for translation machinery (ribosome)
3’ poly(A) tail
composed of 100-250 adenine nucleotides
facilitates transport out of nucleus
protects mRNA message from degradation in the cytoplasm
eukaryotic genes contain
noncoding regions
exons
expressive (coding) regions
introns
intervening (noncoding) regions
removal of introns
RNA splicing is catalyzed by small nuclear RNAs and small nuclear ribonucleoproteins snRNAs and snRNPs
forms a multiprotein complex called a spliceosome
products of transcription
messenger RNA (mRNA)
transfer RNA (tRNA)
ribosomal RNA (rRNA)
other non-coding RNAs
other non-coding RNAs
mostly regulate gene expression
translation
process by which proteins and peptides are synthesized from mRNA
done by ribosomes
uses transfer RNA as interpreter molecules
tRNA structure
made of 80 nucleotides from RNA that was transcribed from DNA
all are not alike
differences from tRNAs
anti-codons: pairing with codons
anti-codon sequences
anti-codon sequence determines what amino acid gets attached on top of tRNA
charged tRNA
a tRNA bound to an amino acid
called aminoacyl-tRNA
aminoacyl-tRNA synthetases ‘match maker’ (simple)
matchers correct amino acid to correct tRNA
aminoacyl-tRNA synthetases ‘match maker’ (complex)
each aminoacyl-tRNA synthetase is specific for one amino acid and all the tRNAs that correspond to that amino acid
the enzyme uses ATP to attach (charge) correct amino acid to 3’ end of correct RNA
synthetase reads identity features on the tRNA (anticodon/other recognition sites) to ensure accuracy
process ensures tRNA delivers to the ribosome carries the correct amino acid for its anticodon
ribosome
site of protein synthesis
composed of ribosomal RNA and proteins
ribosomal small subunit
holds mRNA in place
ribosomal large subunit
has 3 binding sites for tRNAs, contains active sites for peptide bond information
Ribosomal A (aminoacyl or arriving) site
holds incoming tRNA with attached amino acids
Ribosomal P (peptide) site
holds the tRNA with growing polypeptide attached
Ribosomal E (exit) site
holds tRNA that will exit, amino acid no longer attached
translation initaitaion
mRNA binds to small subunit of ribosome
initiator aminoacyl tRNA binds to start codon
large subunit of ribosome binds completing ribosome complex