Genetics Exam 3
1/154
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
155 Terms
Chromatin condensation pattern
DNA → Chromatin → Metaphase chromosome
Chromatin
DNA + Protein (histone)
Euchromatin
lightly packed chromatin
rich in gene concentration and most often under active transcription
Heterochromatin
tightly packed chromatin consisting mainly of genetically inactive sequences
Constitutive heterochromatin
VERY gene poor
wound tightly most of the time
typically make up centromeres and telomeres
Facultative heterochromatin
can go back and forth from hetero and euchromatin
used in gene silencing and barr bodies (inactive X chromosome)
Folded Fiber Model
No proteins in chromosomes
concluded each chromatid must be a single fiber
random folding
no evidence to support
Nucleosome Model
most commonly accepted model for DNA packaging
DNA wrapped around histones (nucleosomes)
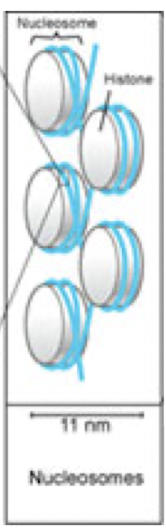
Nucleosome
simplest packaging structure of all eukaryotic chromatin
localized areas of transcription
DNA wrapped around histone
8 proteins
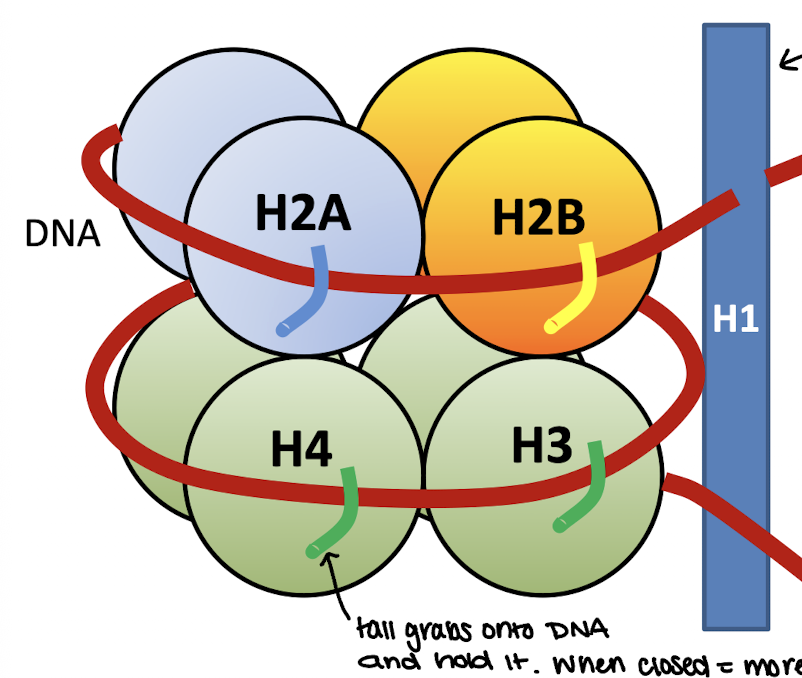
Core histones
H2A, H2B, H3, H4
120 amino acids each
highly conserved during evolution
form the core particle
VERY BASIC in charge (25% lysine and arginine) = hold onto acidic DNA
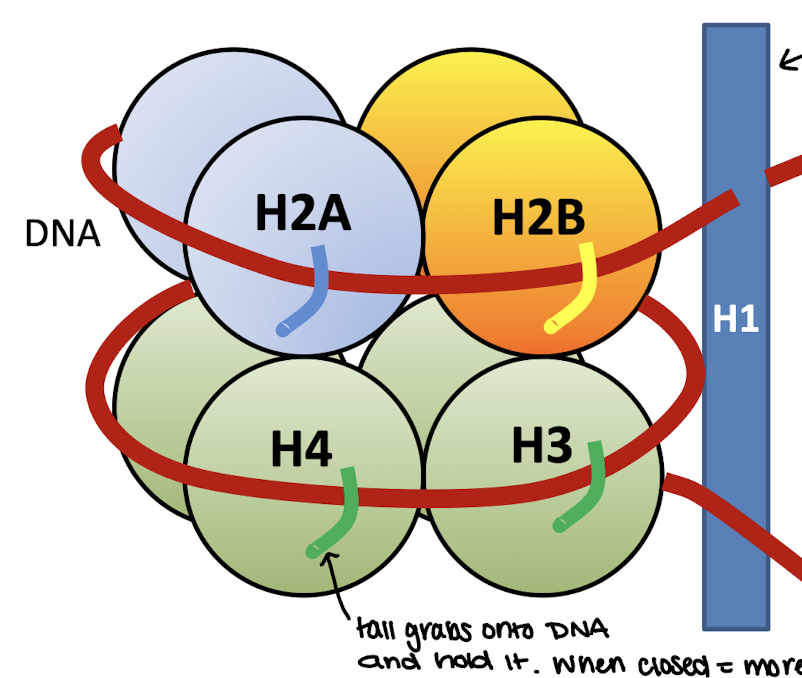
Linker Histone
H1
200 amino acids
tissue specific expression and not highly conserved during evolution
loosely associated with core particle
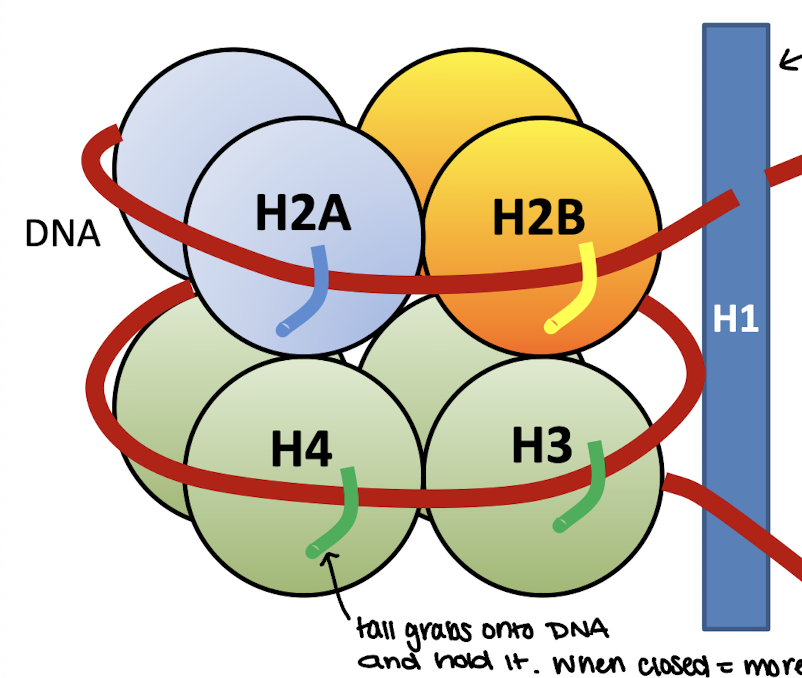
Linker DNA
DNA that connects one nucleosome to the next
Width of DNA
2nm
Width of nucleosome
10nm fiber
how many base pairs per nucleosome
200
How many histones in a nucleosome
9
2 sets of 4 core = 8
1 linker
by what factor do nucleosomes reduce DNA length
7x
arrangement of nucleosomes
linear
“beads on a string”
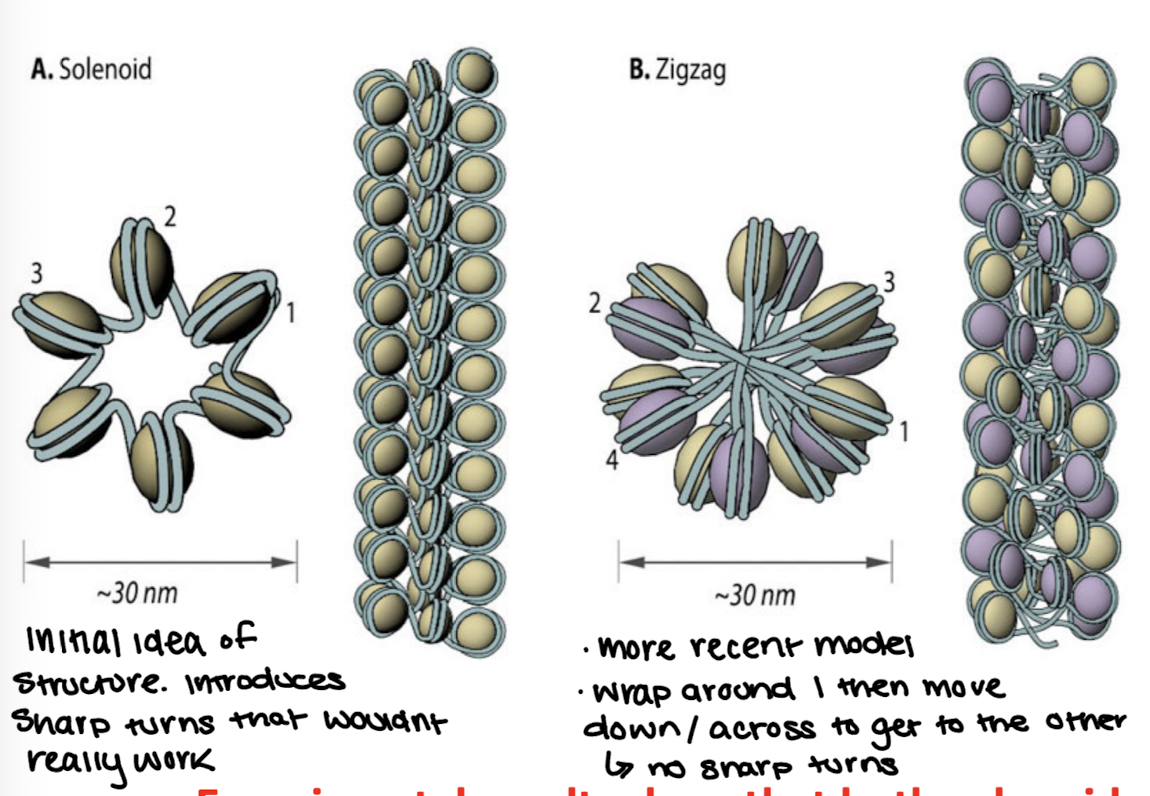
Solenoid
helical coiling of 10nm fibers consisting of 6 nucleosomes
H1 histone helps pack into circle formation
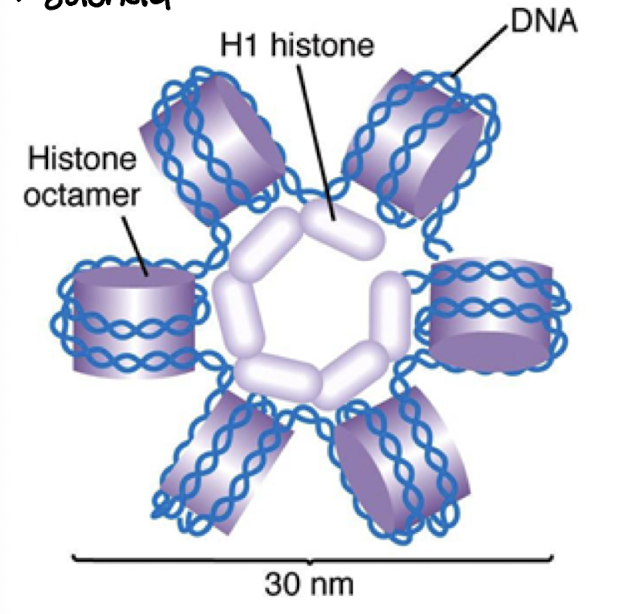
width of solenoid
30nm
by what factor does supercoiling reduce the length of DNA
7x
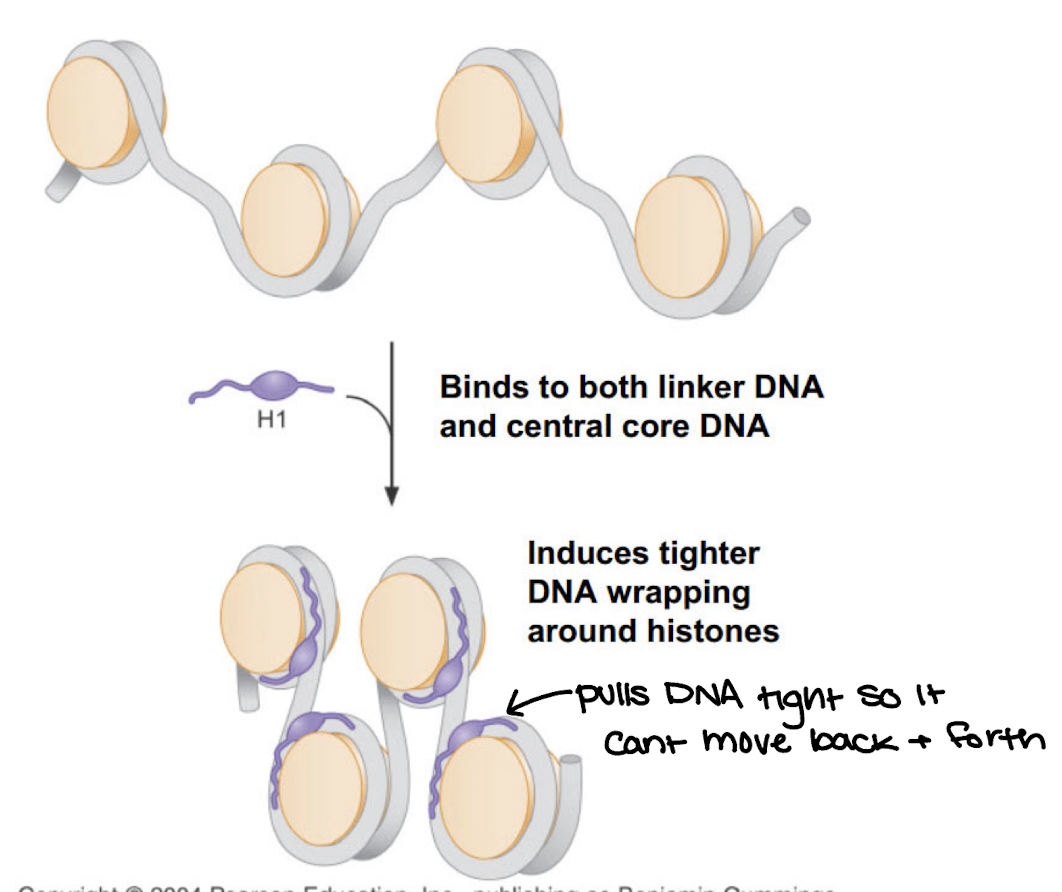
How does Histone H1 work
Binds linker DNA and portion of 146 BP core histone
induces compaction of DNA
“alternate model” of DNA super coiling
Zigzag model
DNA backbone is not flexible enough to bend between nucleosomes so straight linker DNA connects opposite nucleosomes
more recent model
Both kinds will exist, just at different parts of the chromatin fiber
Higher order coiling
Built around a scaffold of topoisomerase II
“chromatin loops”

Width of higher order coiling
300nm
what is the compaction level of euchromatin
higher order coiling
300nm
Final condensation
spiral scaffold composed of topoisomerase II and about 15 non histone proteins
right before metaphase chromosome

Width of final condensation
700nm
compaction level of heterochromatin
final condensation
700nm
when is DNA the most accessible
interphase
between G1 and G2
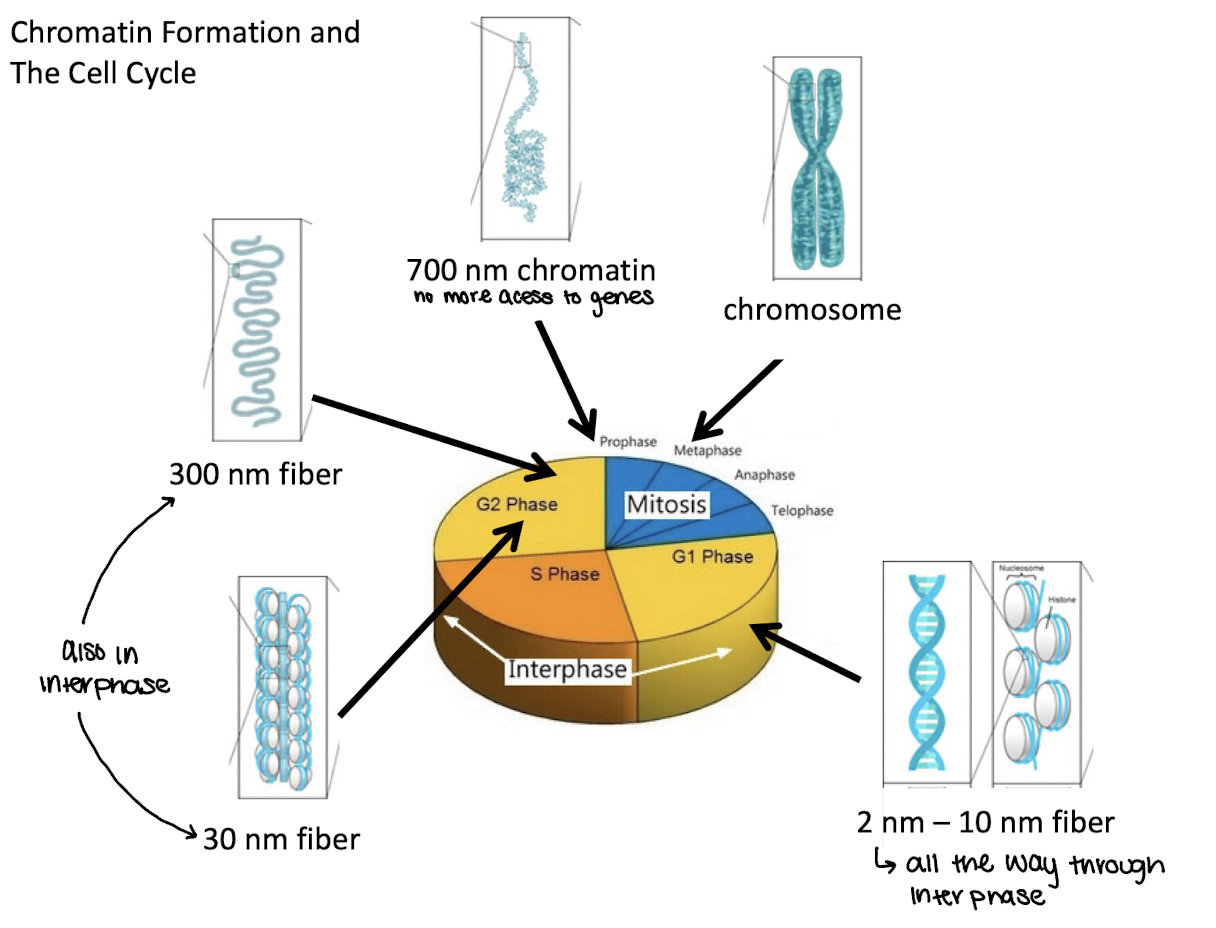
what phase of the cell cycle are 2nm-10nm fibers in
G1
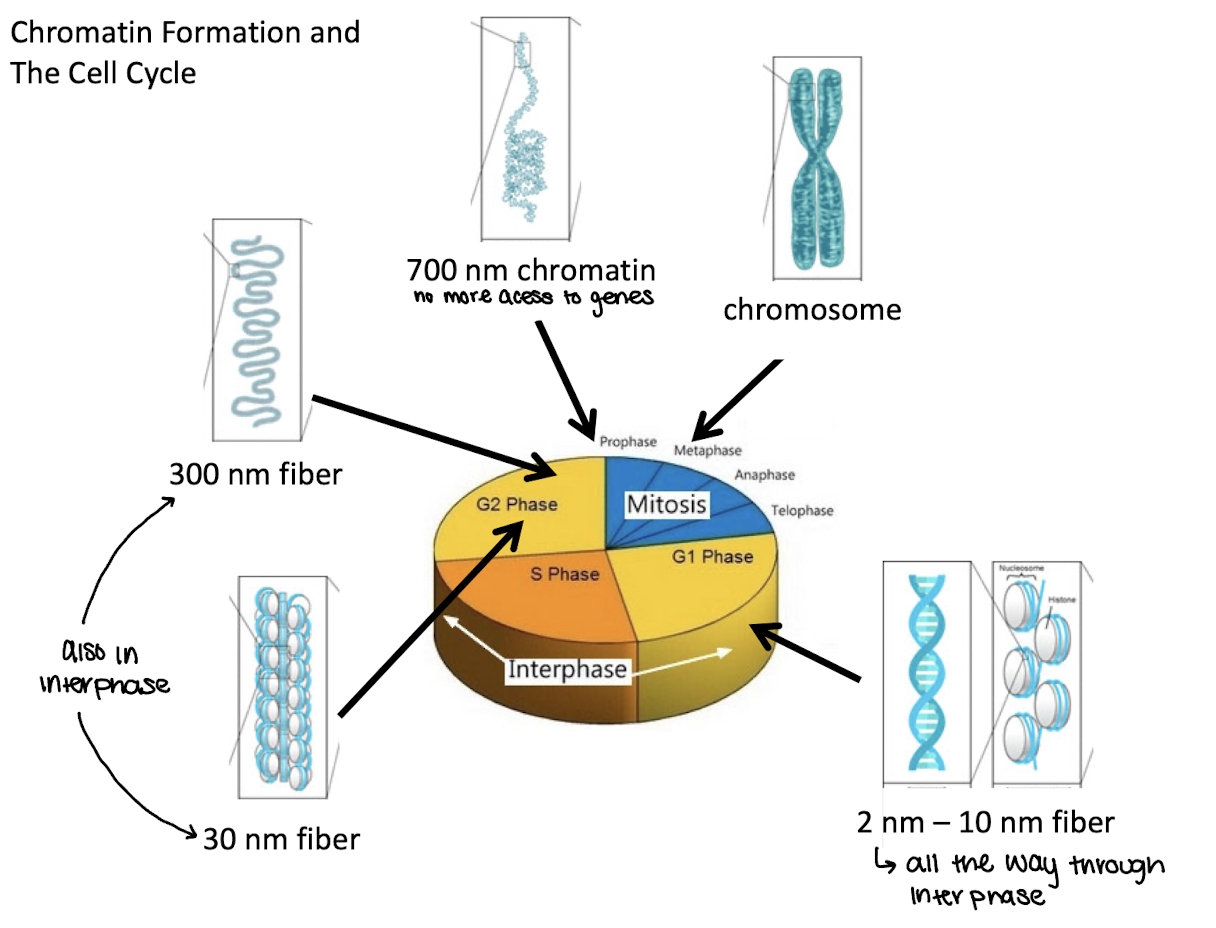
what phase of the cell cycle are 30 and 300nm fibers in
G2 and interphase
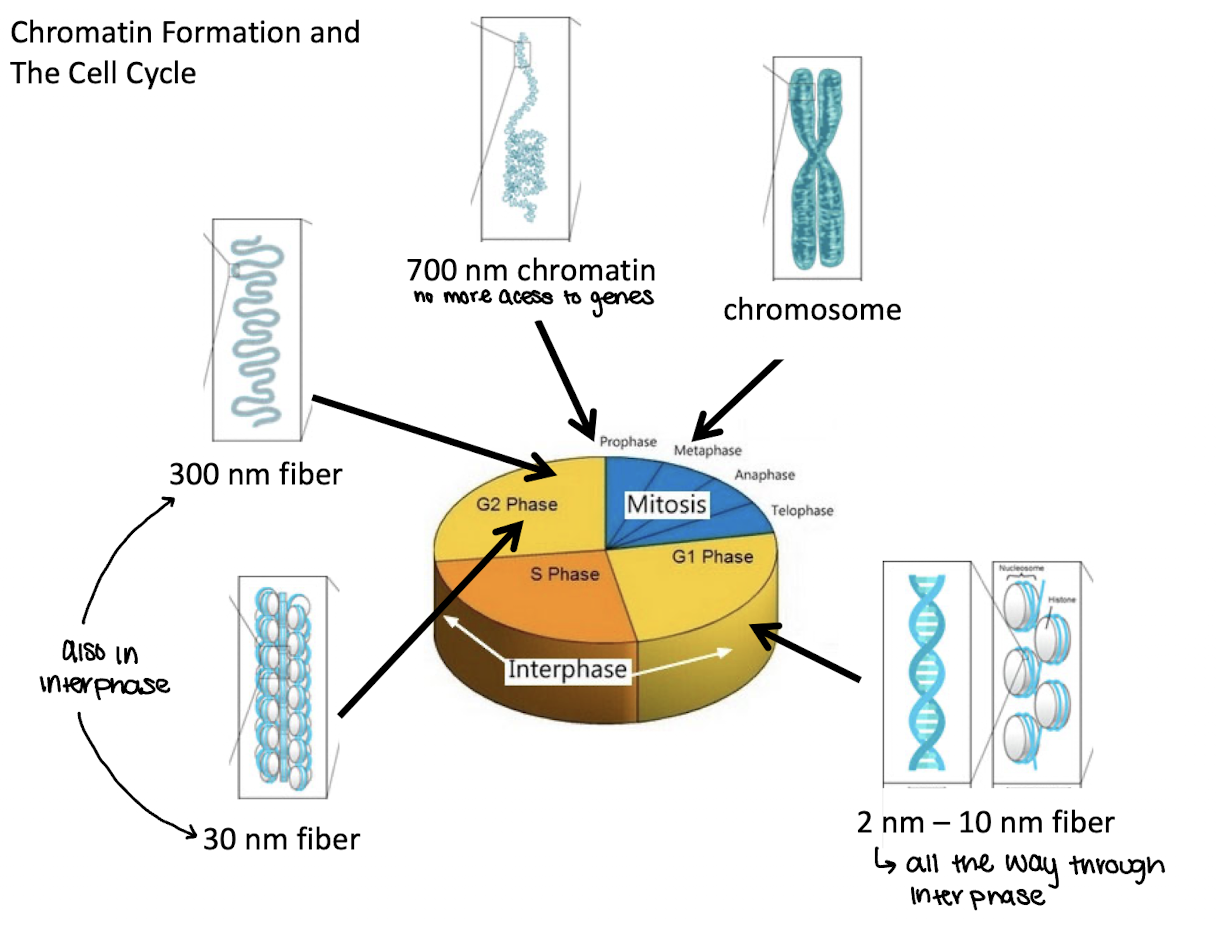
what phase of the cell cycle are 700nm chromatin in
End of G2/prophase
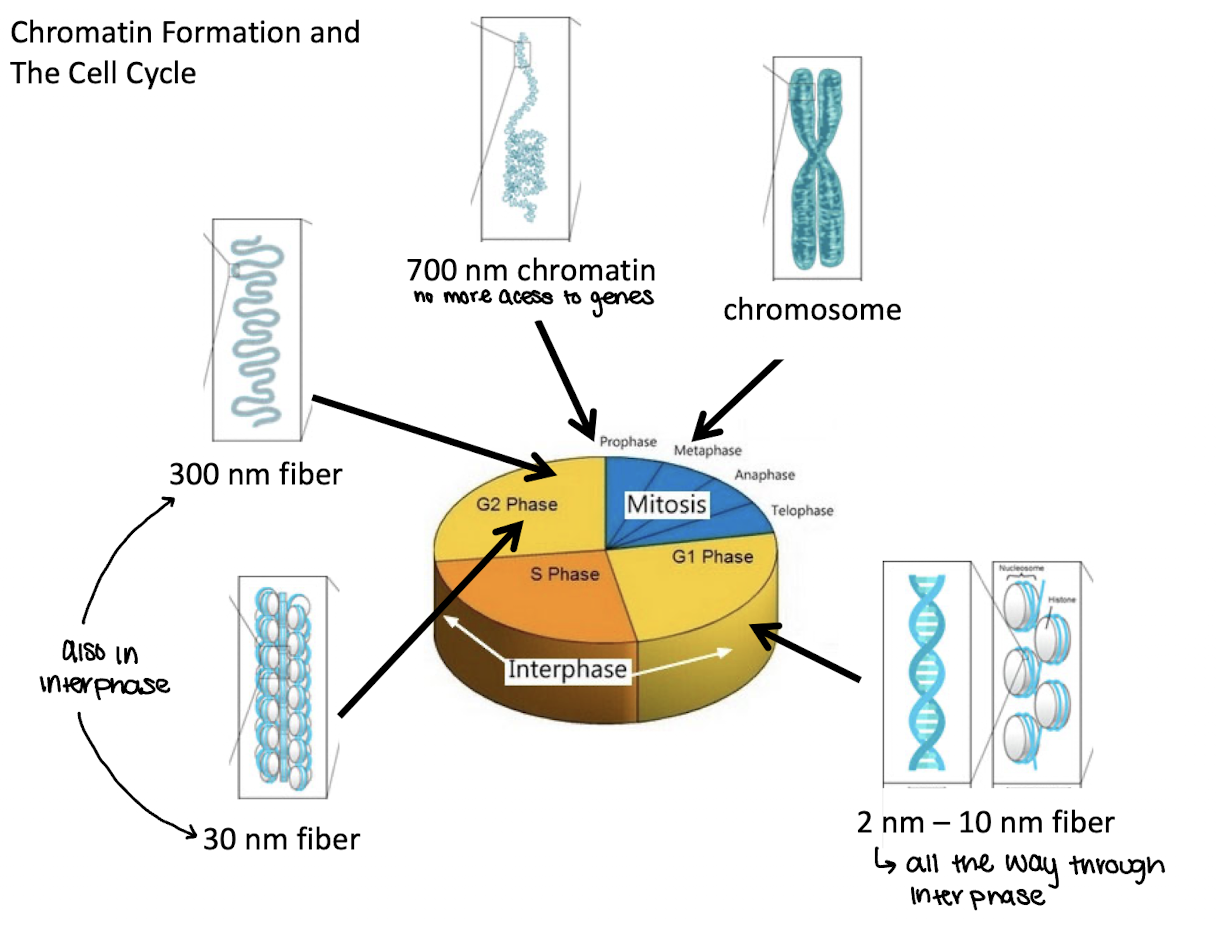
what phase of the cell cycle are chromosomes in
metaphase
Biological complexity
the result of the hierarchical organization of nested levels of cells, tissues, and higher order part types interacting together
C-value paraxon
genome size does not correlate with organismal complexity
G-value paradox
the number of genes does not correlate with organismal complexity
not just size of genome but actual coding genes
Classes of nucleotide sequence
highly repetitive (HR)
moderately repetitive (MR)
single copy (unique)
Highly repetitive DNA sequence (HR)
mostly located in heterochromatic regions around centromere/telomere → no coding DNA
comprises 10% of human genome
function is structural and organizational
occurs at variable lengths
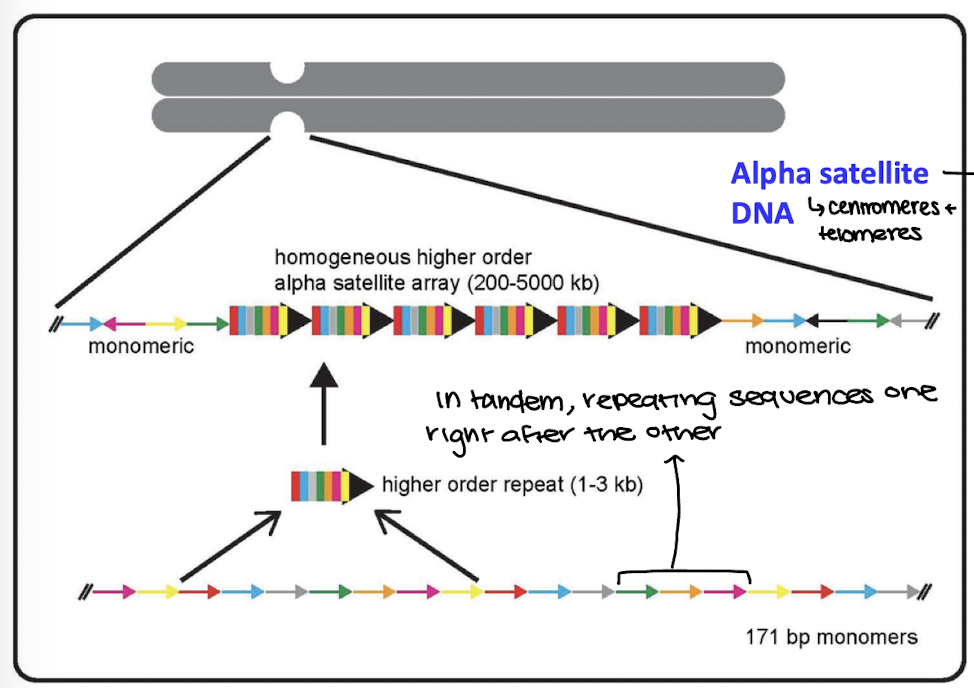
ex: alpha satellite DNA
Alpha Satellite DNA
highly repetitive DNA sequence
in tandem repeats
structural function (centromeres, telomeres)
Moderately repetitive DNA sequence (MR)
found mostly in euchromatin or facultative heterochromatin
comprises about 30% of the human genome
average 300bp in size
function: transcription factor binding, spacing between promoter elements, cytosine methylation, alternative splicing, mRNA stability, transcription start and termination sites
includes ‘redundant’ genes for histones, rRNA, and proteins (gene-products present in cell in large numbers)
ex: microsatellite DNA
Microsatellite DNA
variable number of tandem repeats typically occurring in non-coding regions of the genome
occurs through a mutation process known as “slippage recognition”
useful genetic markers as they tend to be highly polymorphic
used to sequence genome, markers for certain diseases, testing in forensics

Single copy DNA sequence (unique)
found throughout euchromatin
comprises 1-5% of human genome
single or low copy number
“coding DNA regions = genes”
20,000 protein coding genes
what is the other 45% that is not HR, MR, or unique DNA
noncoding DNA
Gene
basic physical and functional unit of heredity
sequence of unique nucleotides (genotype) that carry the genetic information which is to be expressed (phenotype)
how many copies of each gene do we have?
2
1 from mom
1 from dad
% of DNA that is the same in all people
99.5%
molecular level definition of gene
transcriptional unit
DNA sequence that gives rise to an RNA molecule
Exon
coding sequence
phenotype
intron
intervening sequences
areas of genes that don’t typically code for phenotype
transcription
DNA → RNA
Translation
RNA → Protein
Does DNA have codons?
no, only RNA
Flanking regions
areas on either side of the gene
5’ untranslated region
mRNA that is directly upstream from the initiation codon
regulates translation
recruits ribosomal subunits
3’ untranslated region
mRNA directly following the translation termination codon
post transcriptional influences on gene expression
Promoter
DNA sequence where the transcription machinery binds and initiates transcription
on/off switch
ex: TATA box
TATA box
type of promoter
on off switch for transcription
found in DNA
5’ TATAAA 3’
Enhancer regions
recruit proteins to regulate transcription
dimmer switch
ex: CAAT box and GC box
CAAT box
enhancer region
upstream 60-100 bases to initial transcription site
required for inducible genes
GC box
enhancer region
region of DNA that can be bound with proteins (activators) to activate transcription of a gene
Termination
end of the gene
DNA recruits protein to stop transcription
Terminator
section of nucleic acid sequence that marks the end of a gene or operon in genomic DNA during transcription
Solitary genes (unique)
single copy of a gene in haploid and two copies in diploid
makes up most of euchromatin
Duplicated genes
portion of chromosome is duplicated resulting in an additional copy of a gene
copy is called paralog gene
either original or copy may mutate and change function of the gene
usually occurs during an error in meiosis
Multigene families
set of several similar genes, formed by duplication of a single original gene
usually located in similar regions
used or synthesized at different times
Pseudosomes
dysfunctional relatives of genes that have lost their protein-coding ability
result of multiple mutations within a gene
Repeated genes
multiple copies of small genes clustered throughout the genome at specific sites
often back to back
ex: genes for tRNA or rRNA
why do cells divide
outside of the cell is unable to keep up with the inside because the inside grows at a faster rate
DNA replication
process by which genetic information is duplicated
ensure that each cell in an organism has a complete copy of the genome
G0
resting phase
G1
growth phase, cell increases in size and prepares for DNA synthesis
S
synthesis phase
DNA replication
G2
growth phase, cell increases in size and prepares for mitosis and cell division
M
cell growth stops and cell divides into two daughter cells
Semi-conservative model
one parental and one daughter strand
Conservative model
both parental strands stay together after replication and daughter strands go together
Dispersive model
parental and daughter DNA are interspersed in both stands
randomly break apart and come back together
High-fidelity
accurate replication, few errors due to proofreading
instability of mis-matched base pairs
proofreading/ exonuclease activity of DNA pol III
Origin of replication
AT rich region (looser and easier to break strands apart)
GATC methylation sites (precise timing of replication)
Initiator protein
DnaA protein
replication bubbles
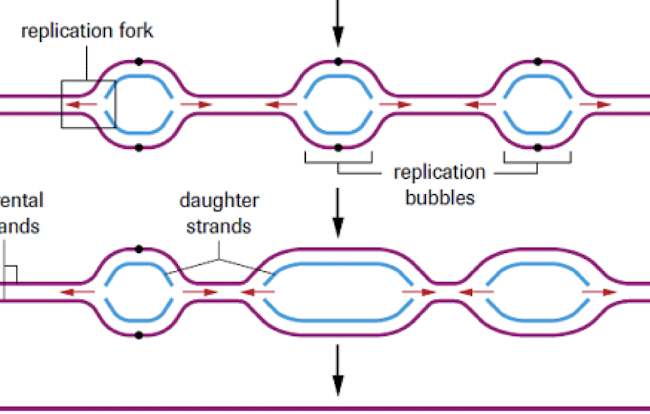
Bi-directionality of replication
replicate both strands at the same time in opposite directions
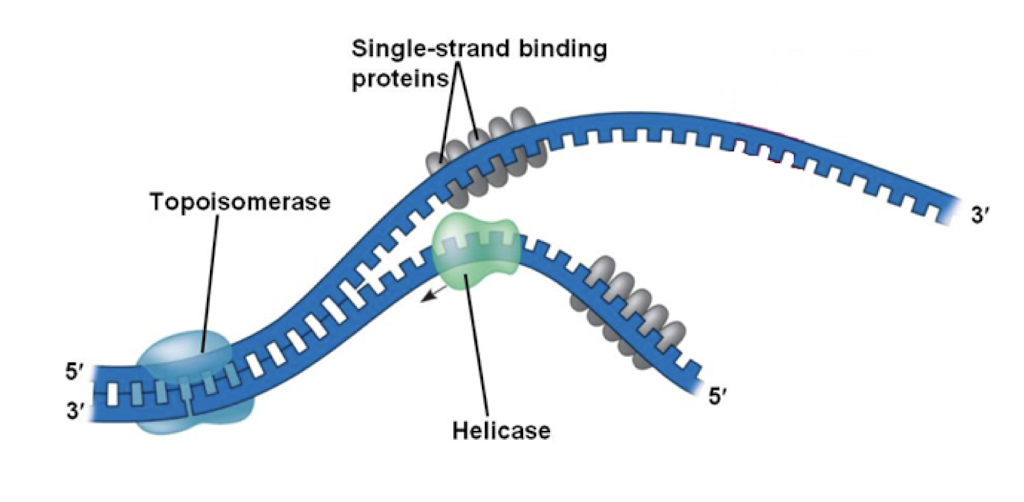
Topoisomerase (gyrase)
reduce torsional strain, unwinds double helix
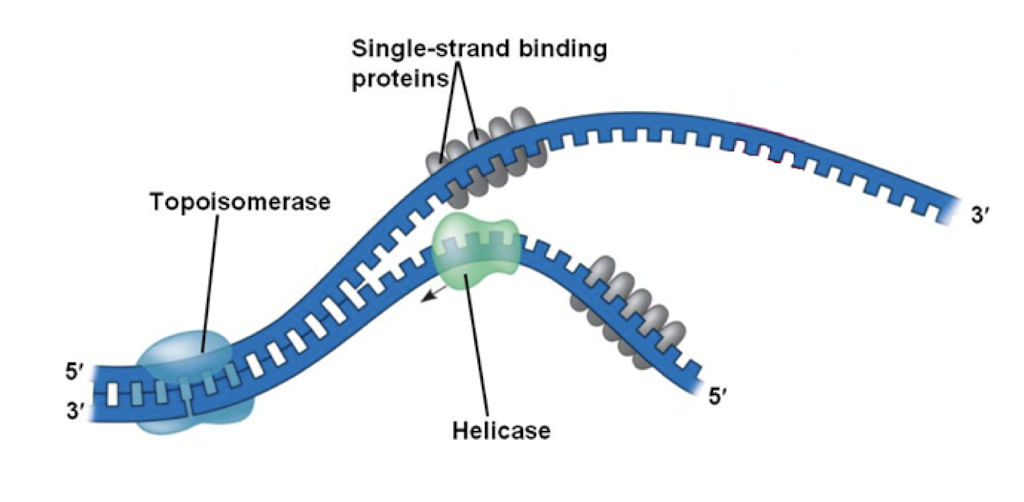
Helicase
breaks hydrogen bonds between complimentary nucleotides
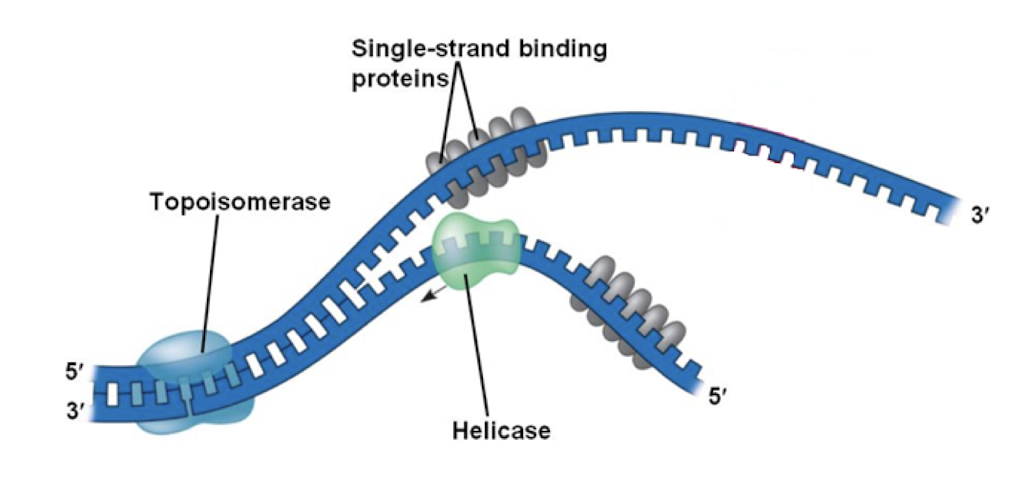
Single-strand binding proteins
stabilize ssDNA until elongation begins
prevents DNA from coming back together
Primase
RNA polymerase that adds a ribonucelotide primer to ssDNA
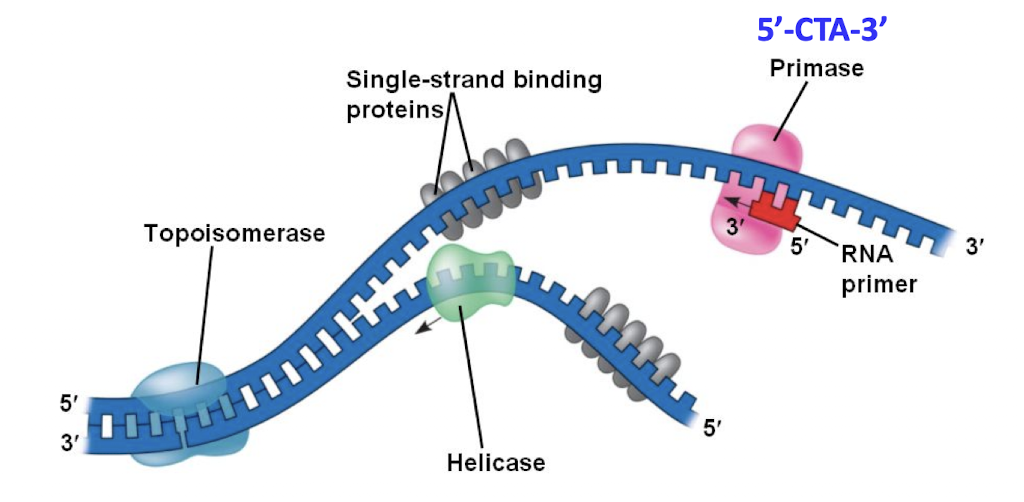
Primers
10-12 bases in length
binds to CTA region
creates a fake “double stranded DNA” so DNA polymerase can bind
uses RNA because its easier to remove later
removed after elongation and replaced with DNA nucleotides
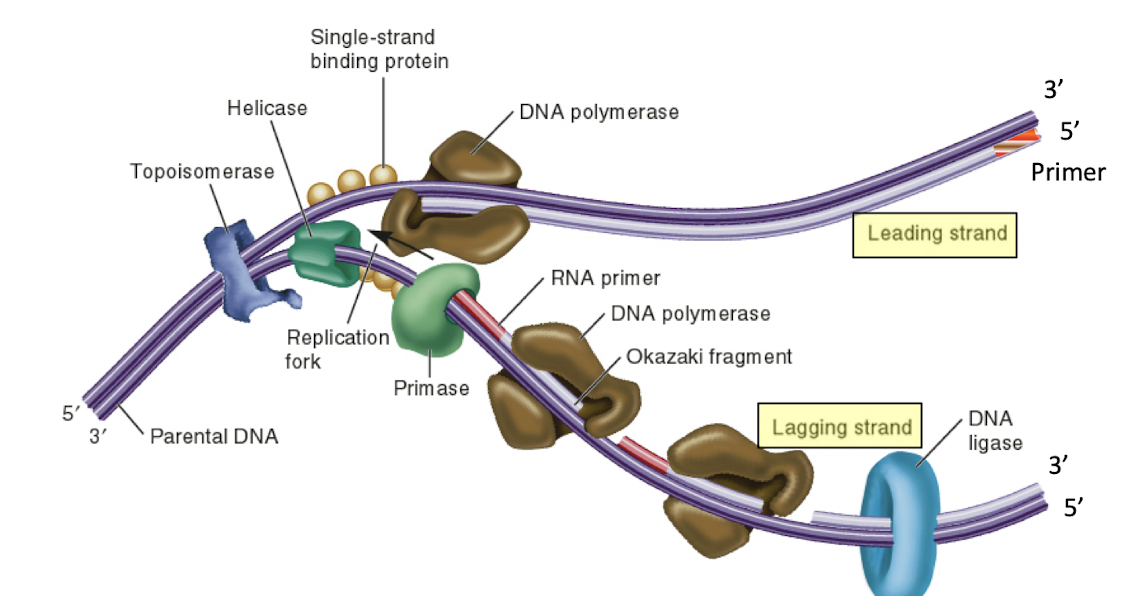
Steps of DNA replication
initiation
Unwinding
Priming
elongation
Initiation of DNA replication
initiator proteins bind
replication bubbles form
replication forks
Unwinding of dsDNA strand
topiosomerase unwinds DNA
helicase separates strands
ss binding proteins stabilizes ss DNA
priming of DNA strand
primase adds ribonucleotide primer to ssDNA which allows polymerase to bind
DNA polymerase III
responsible for most of the replication process
enzyme that catalyzes attachment of nucleotides to make new DNA during replication
prokaryotic DNA polymerases
DNA pol I → remove RNA primer and start synthesis
DNA pol III → most of replication process
DNA pol II, IV, V → repair DNA
Eukaryotic DNA polymerases

can DNA polymerase initiate DNA synthesis
not by itself, it requires an RNA primer (needs to bind to double strand)
what direction does DNA polymerase synthesize in
5’ → 3’
what direction is the template strand read in DNA replication
3’ → 5’
how do nucleotides join in the DNA backbone
3’ hydroxyl to 5’ carbon
phosphodiester bond forms, phosphate ions released
leading strand
DNA pol reads 3’ → 5’ into the replication fork, no breaks