Quiz 3 Content
1/38
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
39 Terms
5 R requirements of DNA for it to exist
Replicate (make more)
Repair (protect)
Recombine (evolve)
Read Instructions (cellular maintencance)
Regulate the reading (rapid environmental change)
2 fundamentals/ ground truths
nothing makes sense in biology without considering evolution
cells obey the laws of chemistry
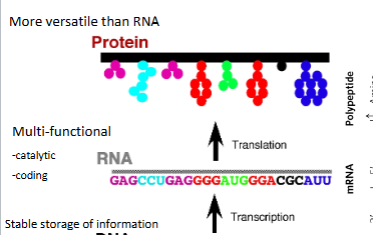
Central Dogma
DNA → RNA → proteins
majority of the chemistry of DNA or RNA occurs through hydroxyl groups
each amino acid is coded for by 3 nucleotides
350 amino acid residues is incodd by 1,050 base pairs of coding DNA!!!
Long-term adaptation of cellular systems
changes in environment
time frame: years
includes DNA replication, DNA repair, DNA recombination
Short-term adaptation (reversible) of cellular systems
changing environments
time frame: seconds/minutes/days
transcription, RNA processing, translation
Main 6 elements of the cell
P, S, C, O, N, H
4 types of non-covalent interactions
hydrophobic interactions
water is 70% of cells, playing an important role in H-bonds
H-bonds
Ionic
Van der Waals
ionic bonds
cohesion between a + charged atom and a - charged
water + salts compete for these interactions
weakens them
salt interactions
ionic bonds DO NOT EQUAL ionic interactions
ex:
protein side chains (E, R)
H-bonds
relatively really weak
H+ w/ a partially positive charge interacts with an electronegative atom (slightly negative)
FON + S sometimes
2 EN atoms can share a h-bond
water can compete w/ H-bonds which weakens them
examples:
protein secondary structure
DNA base pairing
Van der Waals interactions
even weaker than H-bonds
asymmetric electrical charges can lead to 2 atoms in proximity to be attracted to each other
“hand-in-a-glove”
Hydrophobic interactions
water interacts with itself w/ H-bonds
forming surface tension
Nonpolar groups avoid interactions with water, causing nonpolar groups to self-associate
ex:
interior of proteins
double lipid membrane
Covalent Bonds in cells
stronger than noncovalent bonds/interactions
stable chemical link between 2 atoms by sharing 1 or more pairs of electrons
the amount of energy it takes to break covalent bonds depends on atoms and environment
Enzymes are required to break covalent bonds
2 Main functions of DNA
Inheritance
Gene encoding
evolution deep dive
RNA may have been the first genes/catalysts
theory: lipid vesicles containing RNA gave rise to protocells
protocells can self-replicate
eukaryotic cells evolve from 3 major changes
evolution of linear chromosomes
evolution of the nucleus
endosymbiotic theory
eukaryotic cells associating w/ photosynthetic bacterium
mitochondria + chloroplasts contain circular DNA (evidence)
Viruses
came from mobile genetic elements that exist within a cell
bacteriophages
transfer genes among bacteria
bacteria can also take UP DNA from the environment
How does DNA deal with environmental changes?
natural selection
gene transcription
DNA-methylation
can shut down genes temporarily
origin of life
~ 4 Billion Years ago
simple microorganisms extracted en. from chem. compounds + sunlight
utilized en. to make biomolecules from the simplest elements + compounds on the Earth’s surface
Difference between the 3 branches of the TOL
bacteria + archaea = prokaryotes
lack a nucleus, small size/genome
eukaryotes
have double membrane organelles'
larger in size/genome
Model organisms
E. Coli → bacteria
Arabidopsis, yeast, fruit fly, mouse, zebra fish, C. elegans, human cancer cells → eukaryotes
DNA polymers (aka chromosomes)
pack DNA into chromatids
we have 22 pairs + an X and a Y chromosome (XX for females)
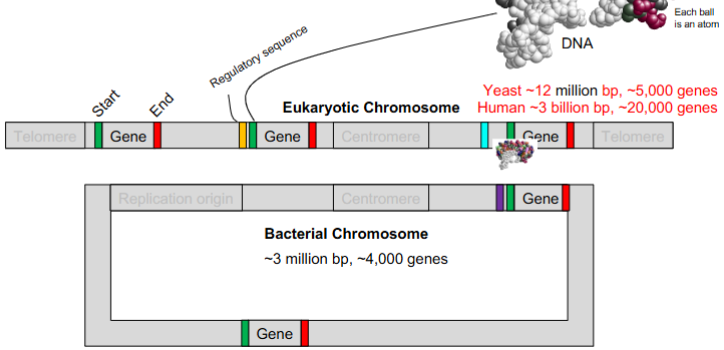
characteristics of eukaryotic chromosomes
linear
contain genes
replicaiton origin
centromeres
telomeres
have introns and extrons
introns: transcribed but NOT TRANSLATED
extrons: transcribed and translated (to a protein)
characteristics of bacterial chromosomes
circular
contain genes
replication origins
centromeres
NO telomeres, NO introns
changes in the hereditary instructions allow evolution to occur
mutation - changes int he nucleotide seq. of DNA
changes the instructions for a cellular component
can be rarely beneficial, mostly neutral, occasionally detrimental
wild type - unmutated cells (reference genotype)
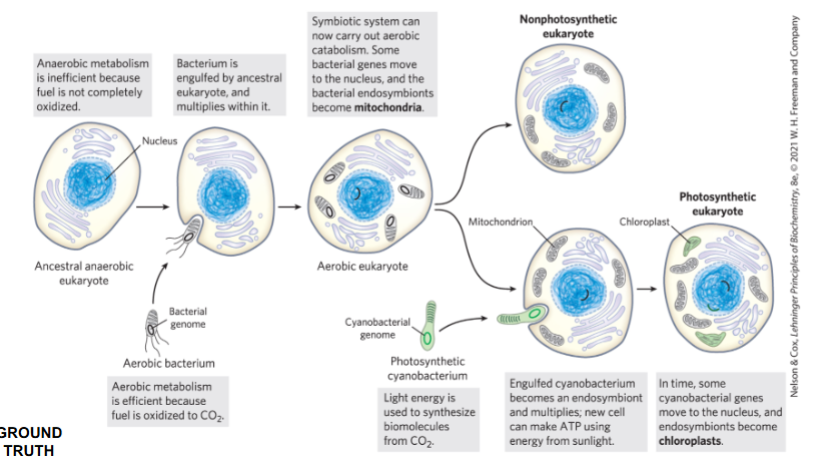
endosymbiosis summary
cyanobacterium engulfed by a eukaryotic cell → chloroplast
proteobacterium engulfed by a eukaryotic cell → mitochondria
mitochondria have their own circular DNA that can code for their own organelle-specific translation machinery
plasmids
small circular DNA
likely mini-chromosomes w/ a few genes
can move among bacteria when they have sex
fungi make antibiotics to kill bacteria, so there is an arms-race between plasmids and the archaea
bacteria - highly evolved, but how?
bacterial viruses (phages) can transfer genes among bacteria
Bacteria can also take up DNA from their environment
they replicate rapidly
3 main functions of gene regulatory sequences
where to start transcription
where to end
how often to transcribe
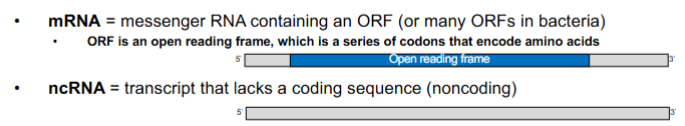
different types of RNA (4 main)
mRNA
messenger RNA
contain an ORF
ORF - open reading frame
series of codons that encode amino acids
ncRNA
transcript that lacks a coding sequence (NONCODING)
tRNA
transfer RNA
translates codon to amino acids on a ribosome
rRNA
ribosomal RNA
make up a ribosome
Homologs
proteins encoded by genes that share readily detectable sequence similarities
similarities in genes/protein sequences can help determine phylogenetic relationships
genes can be conserved across deep lineages
multicellularity development
egg → stem cells → specialized cells
specialized cells are the product of expressing specific genes
cell signaling can trigger the expression of particular sets of genes
What are some features that distinguish a eukaryotic genome from a bacterial genome?
linear DNA vs. circular DNA
eukaryotic DNA have introns, bacterial genomes don’t
eukaryotic DNA have histones
eukaryotic genome is much larger
eukaryotic DNA is located in the nucleus, which bacteria don’t have (in cytosol)
Why does Eukaryotic DNA compact itself?
neutralize the negative charges of the phosphoryl groups in the DNA backbone
w/ cations, histones, and polyamines
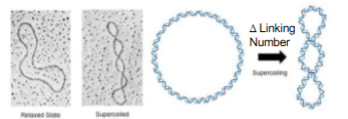
DNA has supercoiling
positive supercoiling tightens it
negative supercoiling unwinds → for DNA replication/transcription!
Topoisomerase: a type 1 mechanism
Topiosomers
form of circular DNA that differ only in a topological property
ex: linking numbers
Topoisomerase
enzyme that changes DNA supercoiling
resolves distortion from replication/transcription
type 2 breaks both strands and then both are resealed
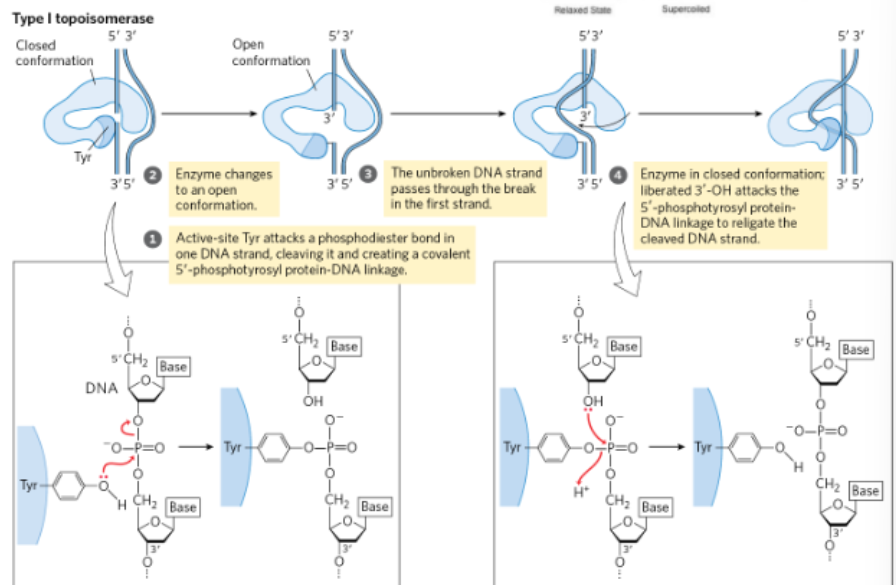
topoisomerase mechanism
KNOW THE ARROW PUSHING
Active site Tyrosine attacks a phosphodiester bond (breaks it)
enzyme changes to an open conformation
the unbroken strand swivels through the break in the first strand
The enzyme in closing position
liberate 3’ hydroxyl attacks the phosphotyrosine linkage to re-ligate
reform the phosphodiester bond
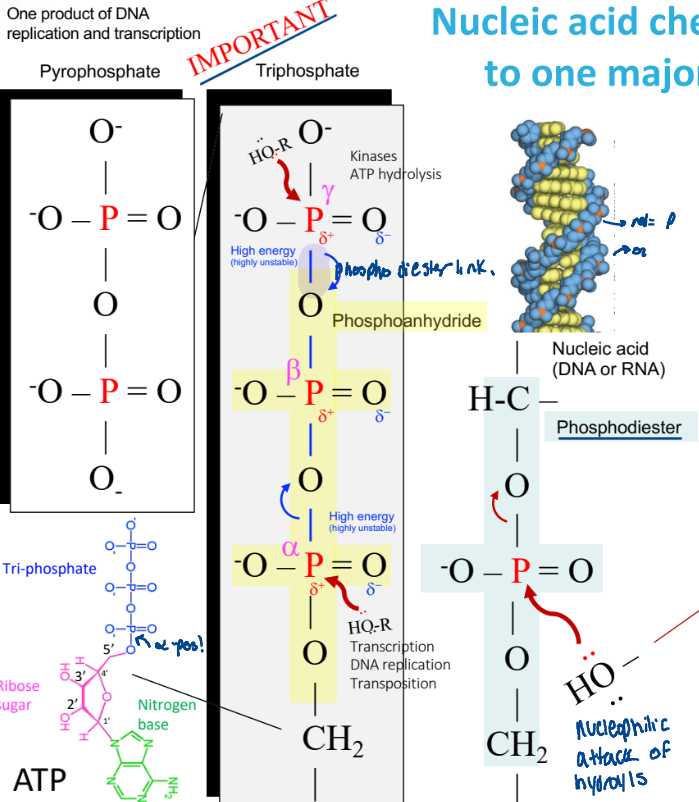
Nulcei acid chemistry => nucleophilic attack of hydroxyls
a hydroxyl group attacks the p, breaking a phosphodiester bond
ex:
alpha, beta, gamma numbering
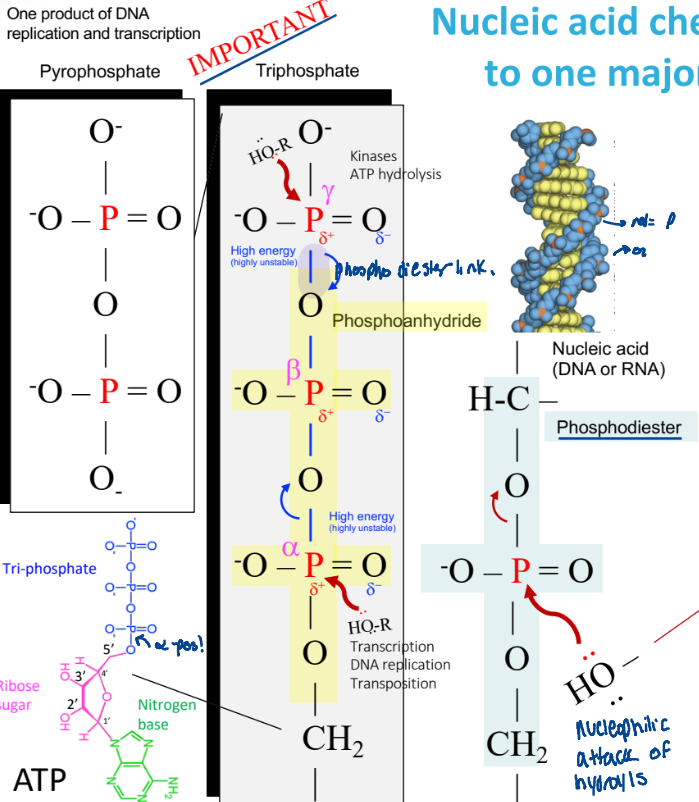
Why can phosphoanydride bonds break easily?
highly unstable
weak electron sharing
release the phosphate
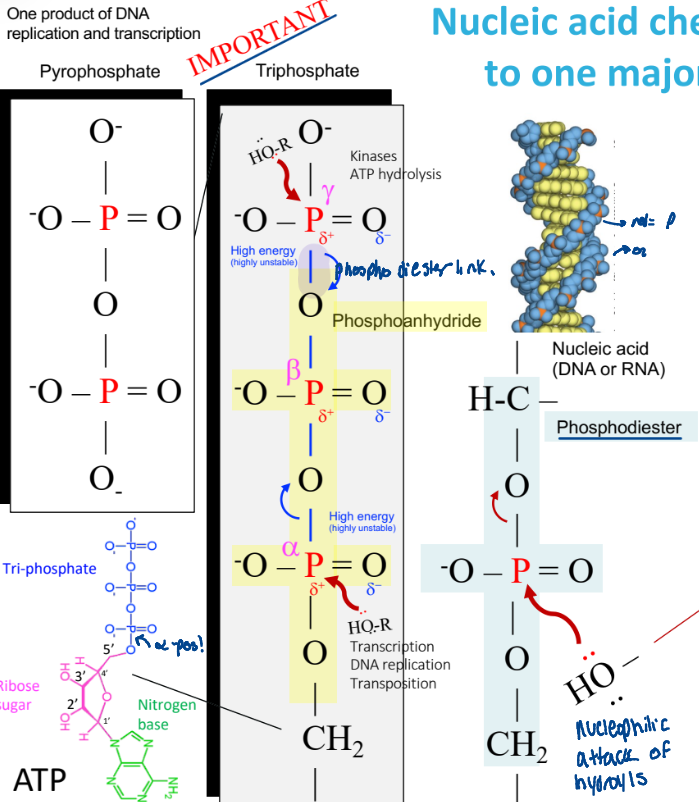
reactions at the alpha position
transcription
DNA replication
transportation
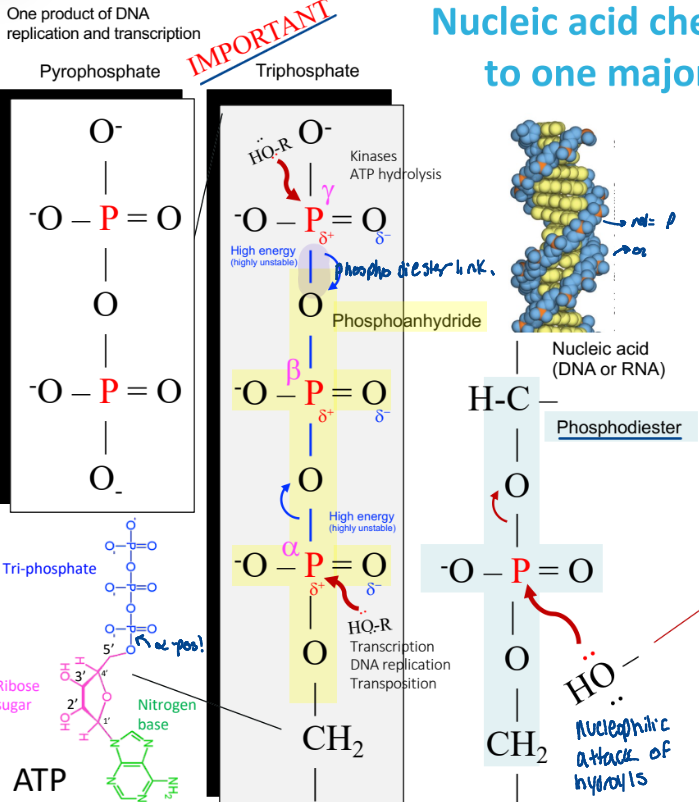
reactions at the gamma position
kinases
ATP hydrolysis
for phosphorylation
constituents of nucleic acids
deoxyribonucleic acid (DNA)
hydroxyl group at the 2’ position
ribonucleic acid (RNA)
hydrogen at the 2’ position