BCH 5413 - Exam 2, Module 2
1/78
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
79 Terms
How is eukaryotic transcription different from prokaryotic transcription?
Eukaryotes have multiple RNA pol complexes containing more subunits, transcription occurs within the nucleus while translation occurs in the cytoplasm, pre-mRNA is processed to become mRNA, and DNA must be unwound from the chromatin structure before transcription occurs.
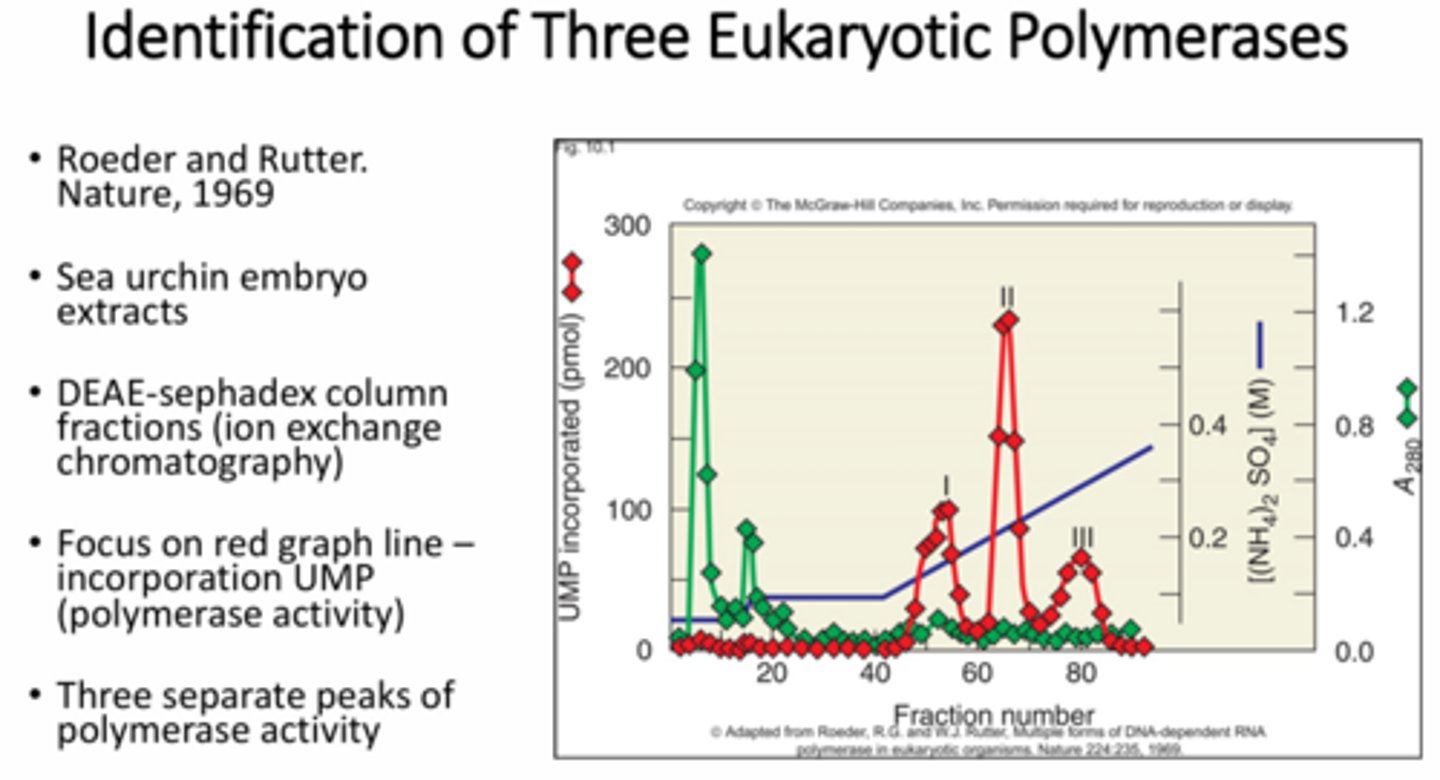
How were researchers able to determine there were 3 eukaryotic RNA pols?
Sea urchin embryo nuclear extracts were fractionated using DEAE-Sephadex ion columns and assayed using UMP for RNA incorporation. Three peaks of polymerase activity were seen. Further study illustrated RNA pol I nucleolar localization and RNA II and II nucleoplasm localization
What is the main function for each eukaryotic RNA pol?
I - rRNA
II - mRNA
III - tRNA
How were researchers able to use alpha-amanitin to assist in understanding the function of each RNA polymerase?
Alpha-amanitin inhibits RNA pol activity - each RNA pol was inhibited one at a time and observed what RNA was absent and assigned that to the inhibited RNA pol
Briefly describe how researchers determined that RNA Pol II has 12 subunits.
All the proteins in the cell were labeled with 35-sulfur (for visualization). One subunit was labelled with an epitope tag (via genetic alteration using an expression vector containing the gene for an epitope tag) and immunoprecipitated using that flag and an anti-epitope antibody. After isolation, a detergent (SDS) was used to separate all the protein subunits within the RNA Pol II complex. These proteins were run on a gel using electrophoresis to isolate them.
How might ChIP be used to study the functions of RNA pol subunits?
Used to identify protein-DNA interactions. Once the protein is bound to its target sequence, it can be fixed and isolated. After isolation, the protein can be separated from the DNA, and the DNA sequence can be analyzed. This can be used to identify what subunits are acting with DNA directly, where along a gene RNA pol binds, and which gene mutations lead to changes in RNA pol interaction.
How might Immunoprecipitation and Western Blotting be used to study the functions of RNA pol subunits?
Immunoprecip. - studies protein-protein interactions, can be used to identify which subunits are interacting to understand how the RNA pol complex is formed
Blot - isolating proteins by size, can be used to identify which proteins are present and how that can change depending on experimental conditions
How might Northern Blotting be used to study the functions of RNA pol subunits?
Used to study RNA, can be used to study how changes in the RNA pol complex can influence transcription
What is the function of RPB1 in the RNA pol complex?
This is the largest subunit in the RNA pol complex. It binds to the DNA to position the complex to select the transcription start site. Additionally, there are heptad repeats that can be phosphorylated in different patterns in order to attract different processing factors.
*Beta prime ortholog
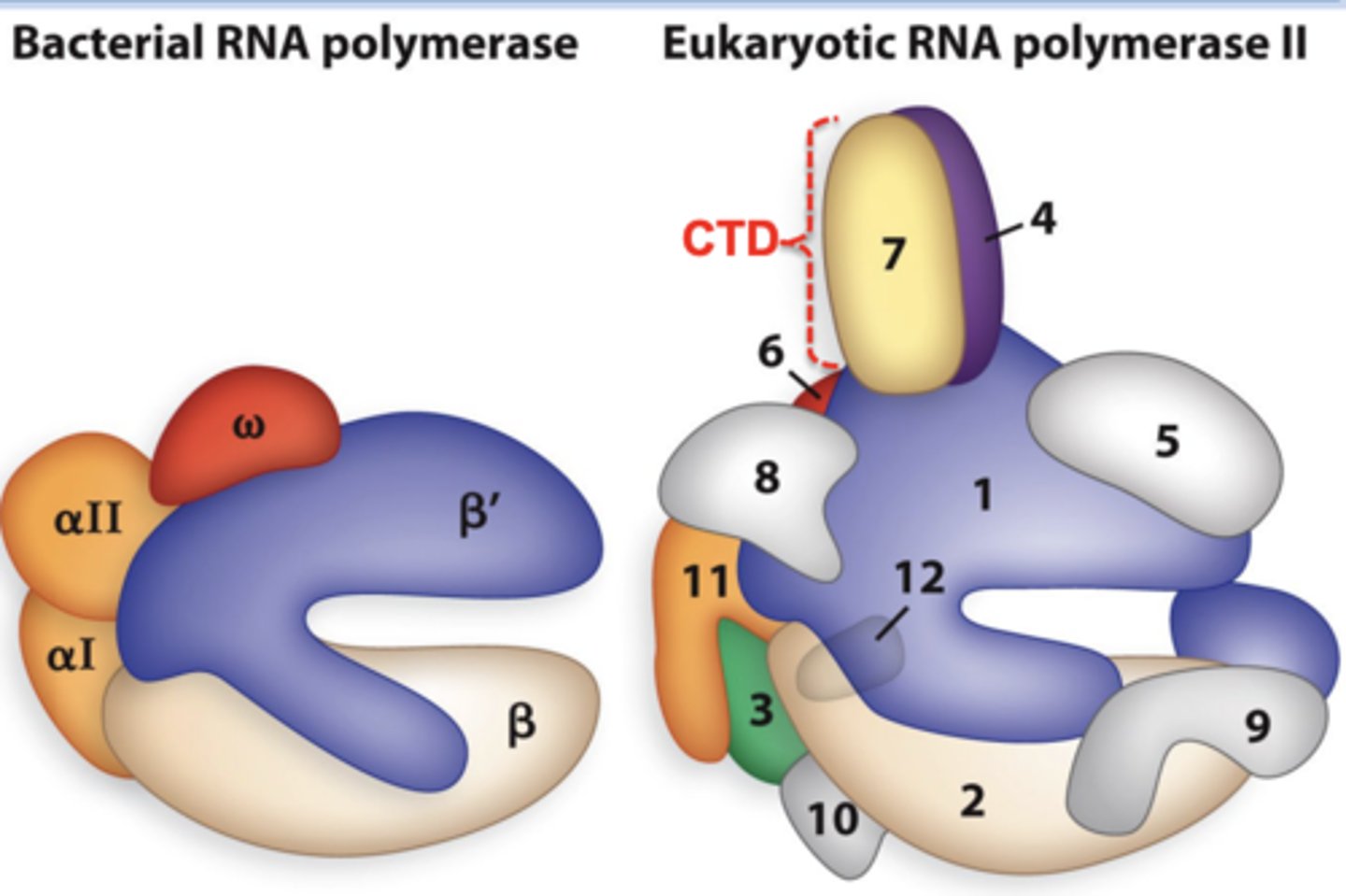
What is the function of RPB2 in the RNA pol complex?
This is the second largest subunit in the RNA pol complex. It's involved in start site selection, contains the transcription active site, and influences the elongation rate during transcription.
*Beta ortholog
What are the functions of the eukaryotic core promoter?
1 - attracts transcription factors and RNA pol to the start site
2 - sets start site and orientation
3 - initiates base level transcription
4 - consists of elements (located) about 40bp from the start site (in either direction)
Briefly describe how the S1 nuclease protection assay works AND describe what it shows regarding the function of TATA box in a eukaryotic promoter.
This assay measures the abundance of specific RNA transcripts by hybridizing a labeled ssDNA probe to target RNA. Then it digests any unpaired regions with S1 nuclease so that just DNA-RNA hybrids are left. Using this assay in eukaryotic promoters showed that the TATA box is required to accurately position RNA pol II at the transcription start site.
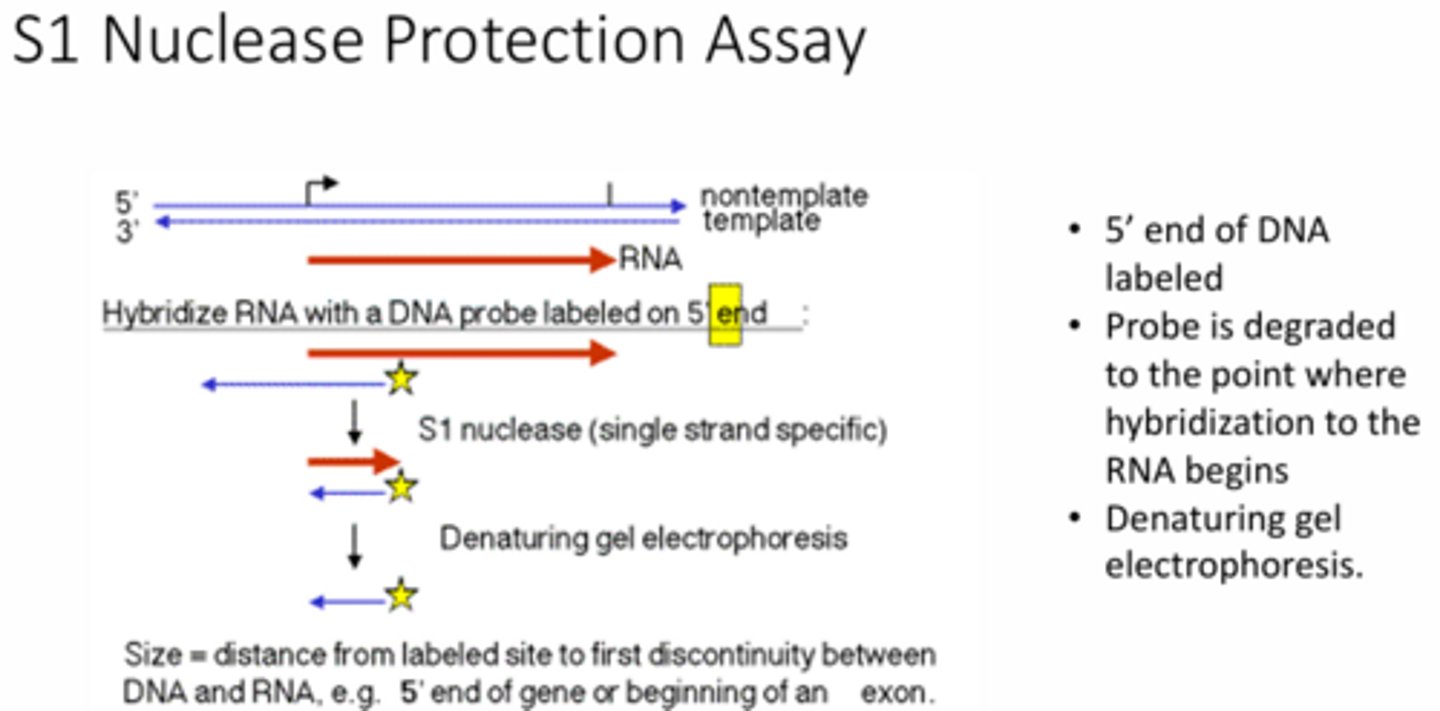
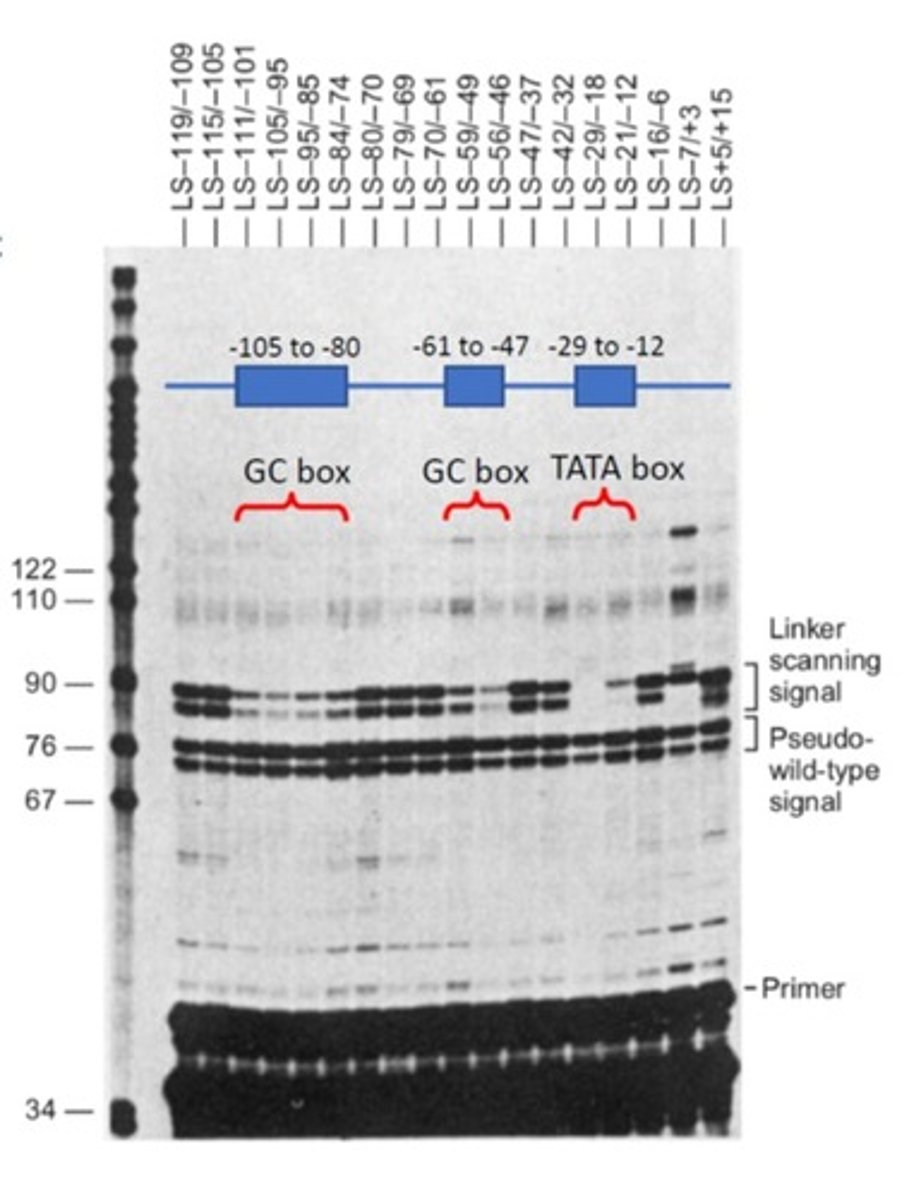
Using the data from the linker scanning experiment shown in the figure, explain why the conclusion was that the TATA box is required for transcription while the GC boxes were not.
When both the GC box sequences were altered/deleted the banding decreased which means that the transcription decreased. However, when the same was done to the TATA box, no banding occurred which means no transcription occurs. Therefore, the TATA box is required in order for transcription to occur.
What is a core promoter element?
The TATA box which is required for transcription. Other elements are included in the core promoter region but they were not discussed (e.g., BRE and DCE)
What are proximal promoter elements?
Elements located upstream of TATA that enhance levels of transcription. They bind to gene specific transcription factors, are relatively position dependent, and they can be orientation independent. The loss of these elements leads to a decrease in transcription only.
What are enhancers?
They are found upstream, downstream, or even far away from the target gene (distal regulatory element). They bind to an activator protein that is associated with RNA pol II to increase stabilization for increased transcription activity (folding over the back). They are tissue specific and can be position and orientation independent.
Briefly describe how a researcher might investigate whether a transcription element is an enhancer.
Regions that are suspected to be enhancers can be deleted or altered in order to compare gene expression. The resultant proteins have an added radioactive amino acid label so that when they are run on a gel the intensity of the bands can be used to compare the wild type gene expression to the altered element expression. This can also be confirmed using a Northern blot by comparing the transcriptional activity between the wild type gene and the altered element genes.
What is the function of the transcription factor TFIID?
This is the first transcription factor to bind (to the TATA box) and it contains TAFs that facilitate interactions with transcriptional activators (forms the pre-initiation complex or PIC)
What is the function of the transcription factor TBP?
This is the subunit of TFIID that binds to the TATA box
During PIC formation, how does TBP bind and carry out its function if TATA box is present?
TBP binds and brings in the rest of TFIID and other TAFs in order to start recruitment for transcription.
During PIC formation, how does TBP bind and carry out its function if Inr and DPE are present but the TATA box is not?
When associated with the TAFs, prior to sequence binding, it is able to bind to the initiator element (Inr) and downstream promoter element (DPE) so that it can recruit the rest to start transcription
During PIC formation, how does TBP bind and carry out its function if GC boxes are present but the TATA box is not?
The GC box binds to the Sp1 which can associate TAF4 which is associated with the other TAFs and the TBP which is now able to start recruitment for transcription
What is the function of the transcription factor TFIIB?
This subunit stabilizes TFIID at the promoter, recruits RNA Pol II and TFIIF, and determines the position and direction of transcription
What are the two domains of TFIIB AND how do they assist in determining the position and direction of transcription?
C term – binds to TBP and DNA
N term – binds to RNA pol and positions within 26-31bp downstream of the TATA box
The asymmetrical binding, along with the contacts inside the open DNA helix, determines the direction of transcription. Just know that because each end makes specific interactions they are able to determine the position and orientation of transcription.
What is the function of the transcription factor TFIIH?
This subunit contains kinase and helicase activity to assist in transcription initiation and facilitates the transcription from initiation to elongation (it is also necessary for promoter opening and clearance). By using ATP, it is able to phosphorylate the CTD tail of RNA pol and unwind the DNA double helix.
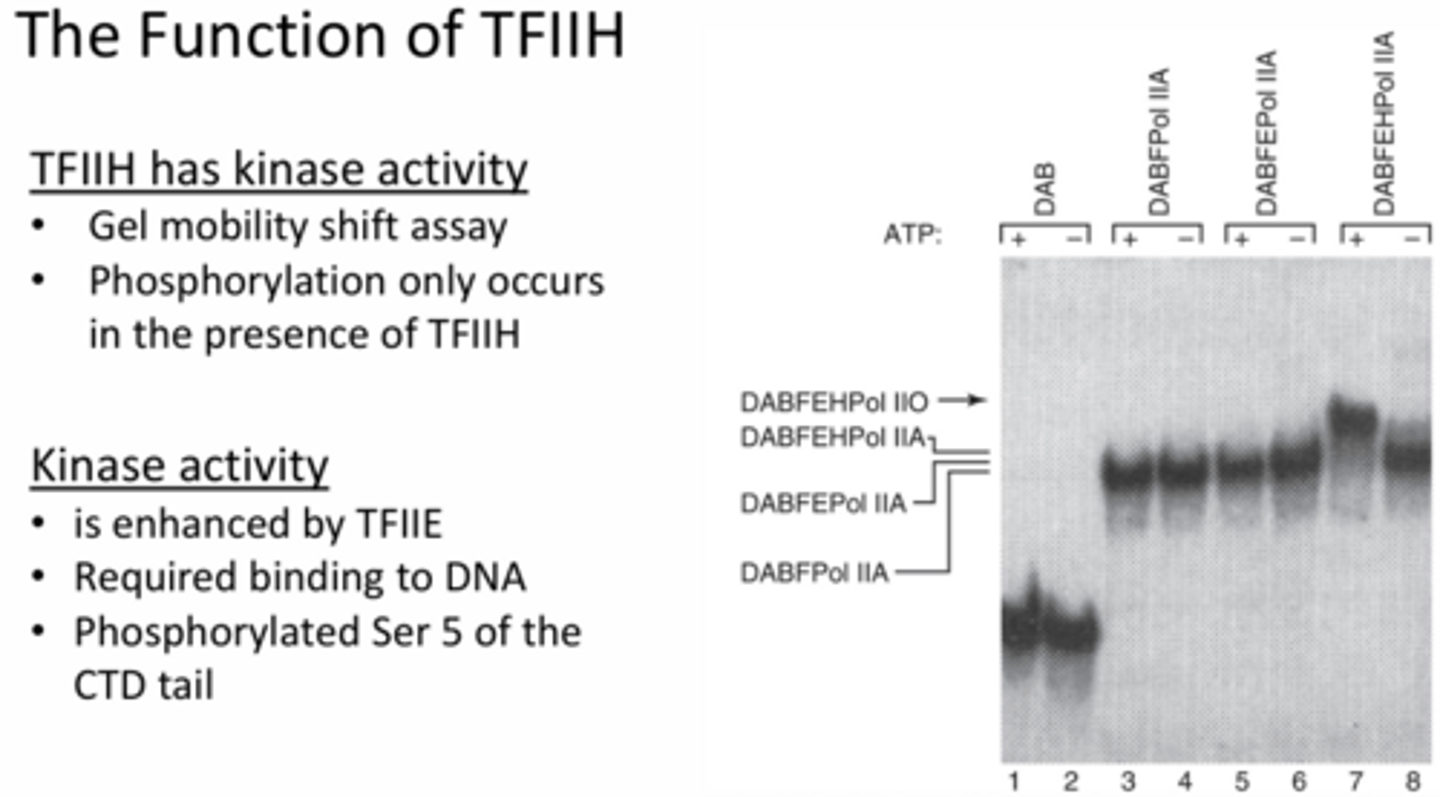
In greater depth, what kinase activity does TFIIH perform and how was this proven?
Phosphorylation of RNA pol II complex - c terminal of RNA pol is phosphorylated (at serine 5) which only happens when bound to DNA (does not happen without DNA being present). This is also enhanced by TFIIE.
The gel shown in the slide illustrates a shift when each new protein is added but the key section is the ATP addition with the DABFEHPol IIa. This shift indicates that phosphorylation is only occurring when TFIIH and ATP are added, which results in a shift that is not seen when the other proteins are added with or without ATP.
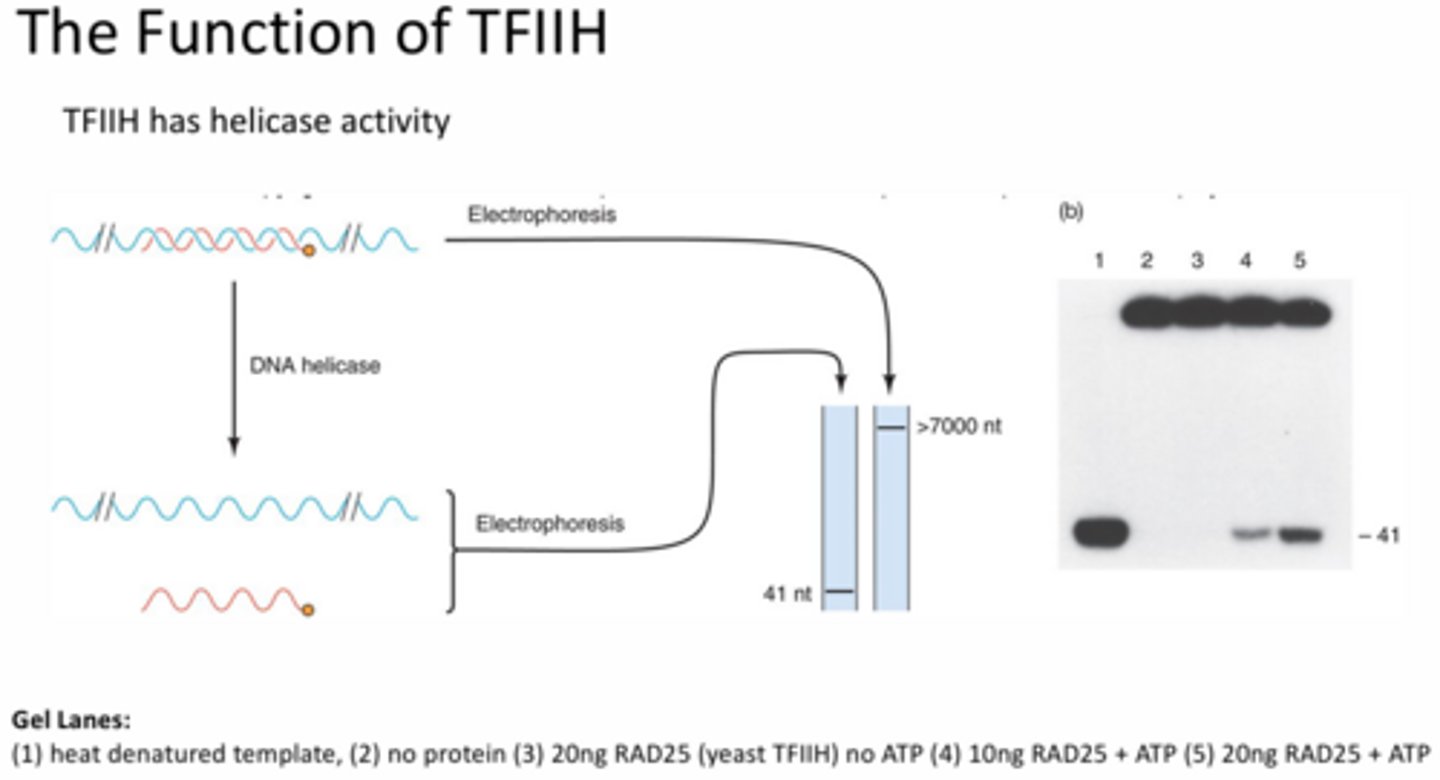
In greater depth, what helicase activity does TFIIH perform and how was this proven?
Unwinding of double stranded DNA. This was illustrated using a helicase experiment in which a complementary probe containing a tag was added to the template and allowed to bind to form a double helix. After various treatments, it is shown that the addition of the TFIIH and ATP resulted in the tagged probe being released from the template, which indicates helicase activity was occurring. This is supported by the heat treated control showing only the probe, the untreated/no protein control only showing the larger double stranded band, and the protein only (no ATP) band being the same as the untreated/no protein control.
In addition to promoter recognition, what are some of the other functions of TAFs?
1 - Mediate promoter recognition of the TFIID complex by interacting with the initiator and the DPE
2 - Histone acetyl transferase activity which may play a role in chromatin structure modification
3 - TFIID recruitment to the promoter
4 - Communicate with transcriptional regulatory proteins and possibly serving as co-activators
Describe the steps necessary for transition from the Pre-Initiation Complex (PIC) to the Early Elongation Complex (EEC).
TBP is bound to the TATA box, TFIIB is bound to the TBP and DNA with a fixed distance to the RNA pol where the transcription bubble is formed and 2-3 nucleotide RNAs are formed (aka promoter opening and abortive initiation) >> eventually the switch 2 domain of the RNA pol is able to stabilize the RNA nucleotides (aka escape commitment) >> as the RNA grows it collides with the B-finger of the TFIIB and causes stress within the transcription bubble (aka clash) >> the stress leads to TFIIB ejection and tightening of the DNA-RNA hybrid (aka promoter escape)
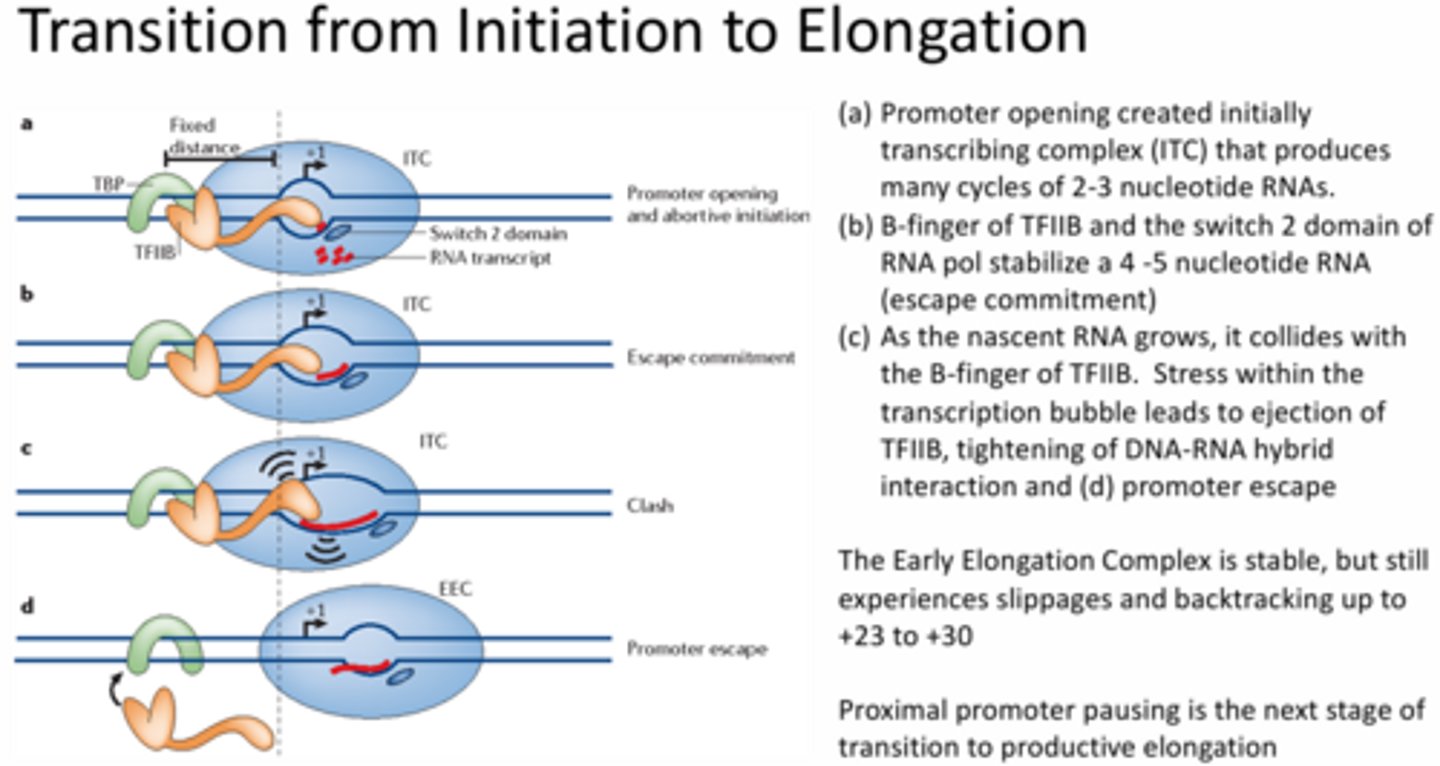
Describe the steps necessary for transition from the Early Elongation Complex (EEC) to productive elongation.
Downstream NELF (negative elongation factor) and DSIF help to pause the RNA pol and prevent elongation. Once the RNA is long enough to exit out of the transcription tunnel the capping factors are added >> the P-TEFbeta (positive transcription elongation factor) kinase will phosphorylate NELF and DSIF >> NELF dissociation and DSIF is converted from a negative factor to a positive factor >> conversion of DSIF results in recruitment of Elongin and ELL (elongation factors). This new complex is now complete and productive elongation can occur.
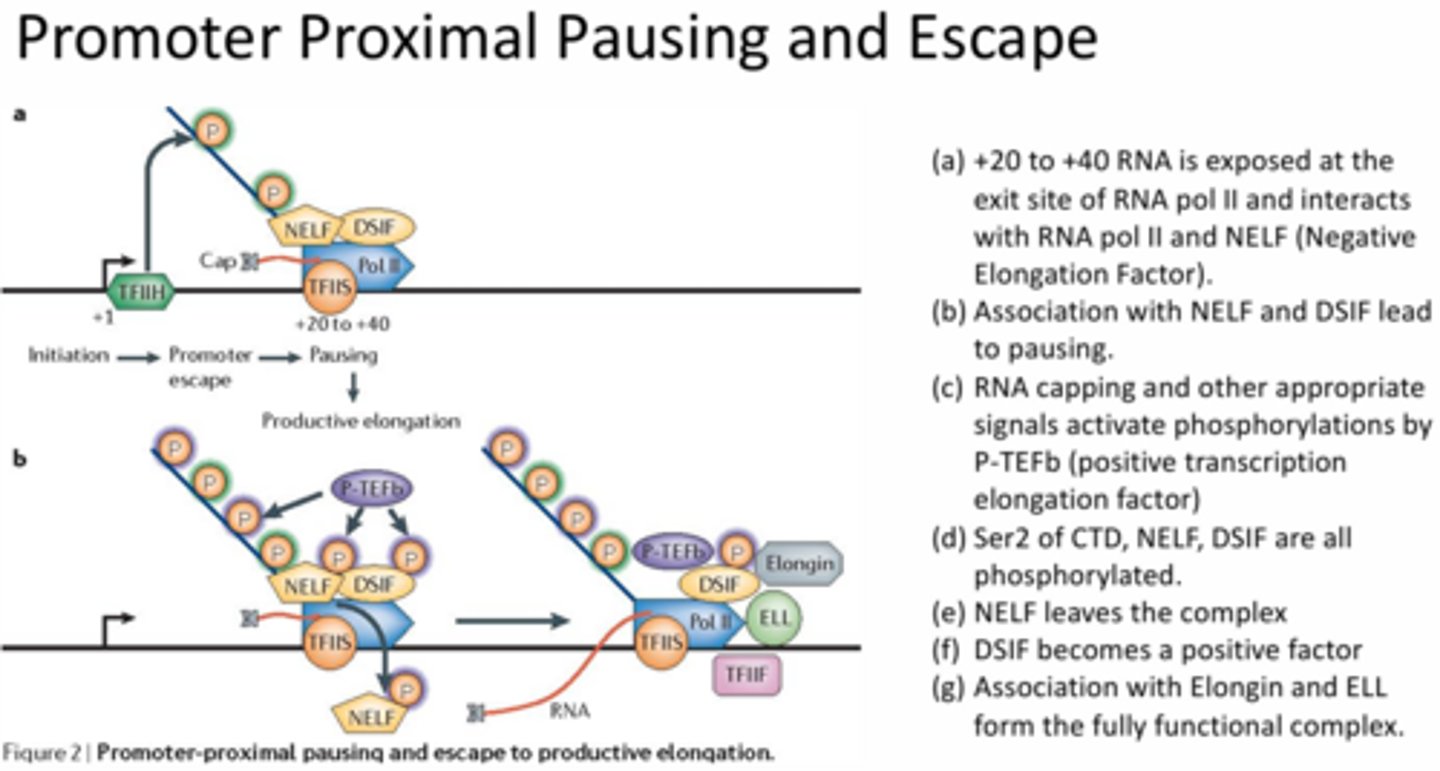
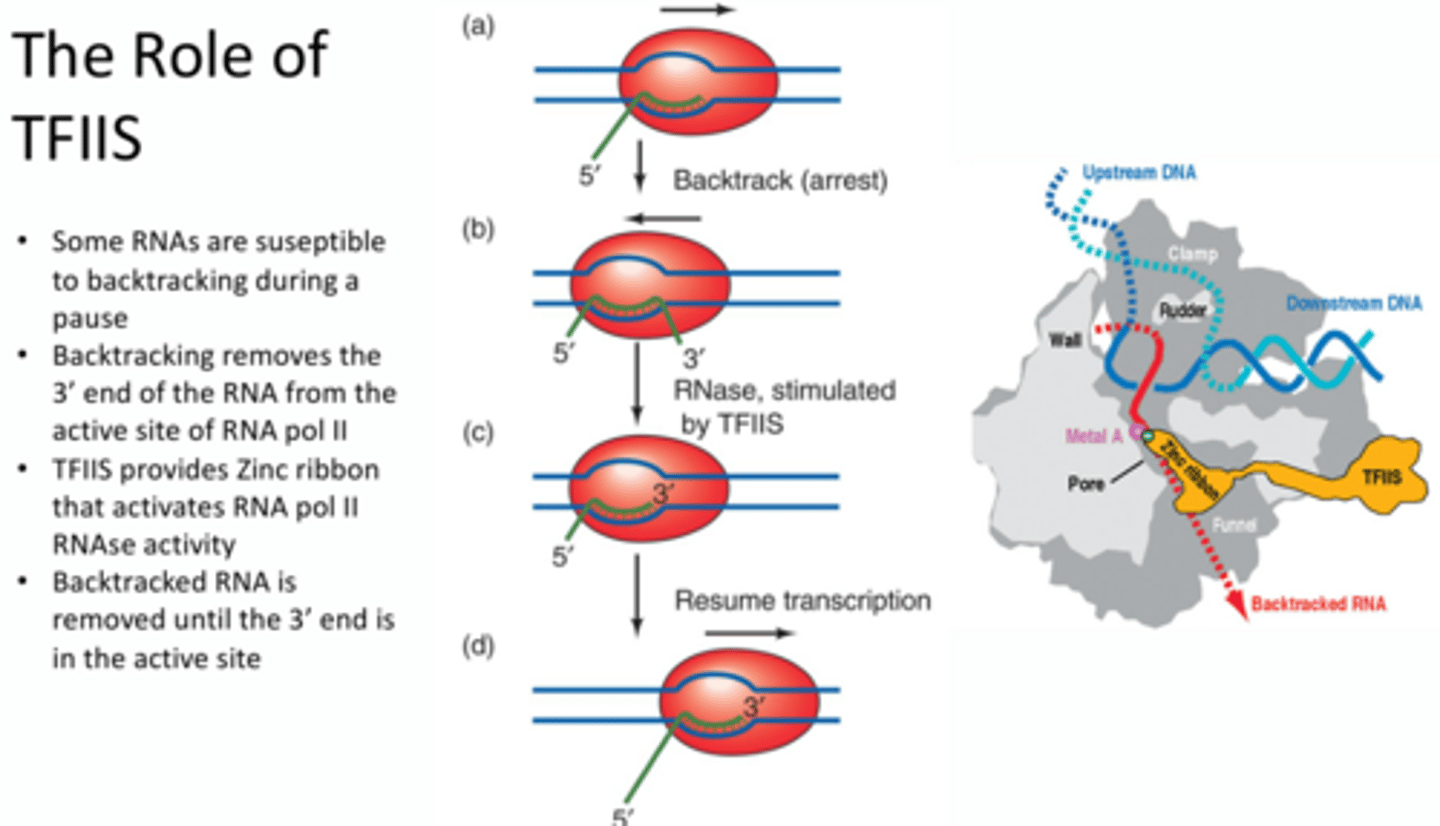
When backtracking occurs, how is the RNA polymerase activity replaced by RNAse activity?
TFIIS – provides zinc ribbon that activates RNA pol II RNAse activity. When backtracking occurs the 3’ end sticks out through the entry tunnel so no new nucleotides can be added to the active site. TFIIS binds the zinc ribbon to another metal ion to activate the RNAse activity to degrade this extruding 3’ end until it is within the active site so that transcription can occur.
TLDR - TFIIS assists in the degradation of backtracked RNA
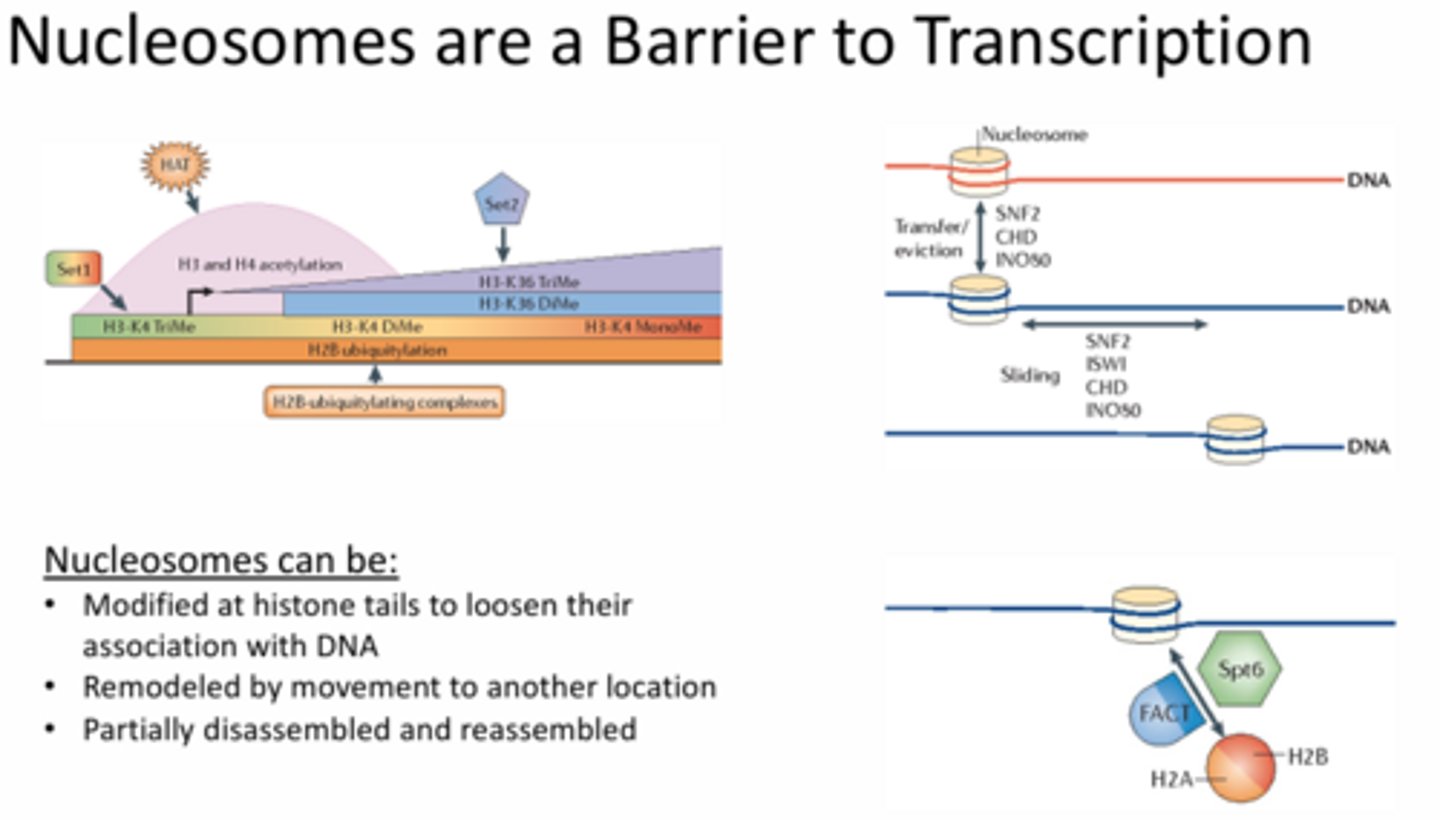
What are the different ways in which nucleosome barriers to transcription can be overcome?
1 - modified at histone tails to loosen their association with DNA
2 - remodeled by movement to another location
3 - partially disassembled and reassembled
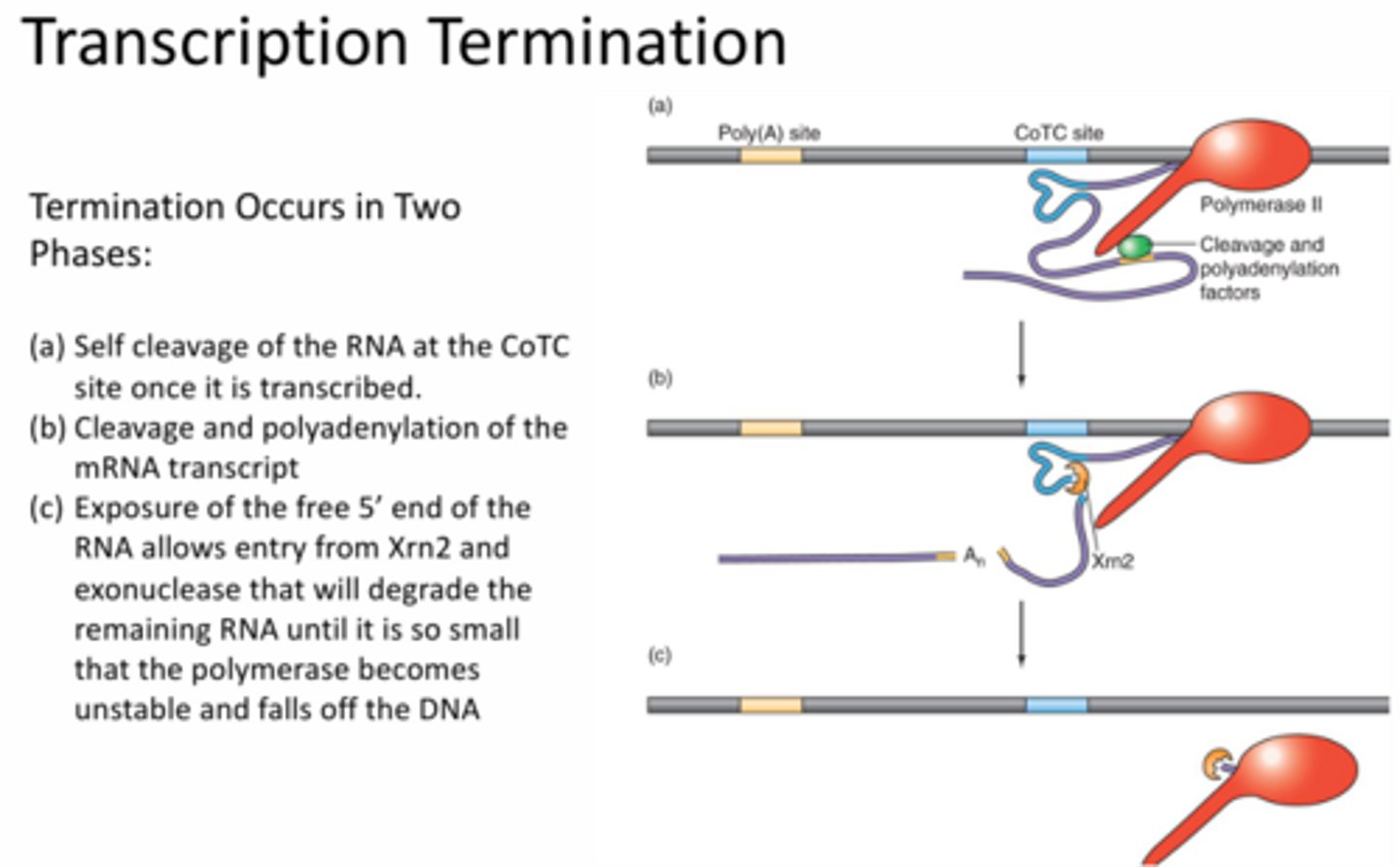
Describe the steps of termination.
Two phases:
1 - RNA self cleaves at the CoTC site by folding in on itself
2 - 5' end of the non-coding RNA is exposed and degraded by Xrn2. Xrn2 eventually catches up to RNApol which will fall off the DNA because it's become unstable.
What are the Transcription Factor domains and their function?
1 - Binding domain - recognition and binding to specific DNA sequences
2 - Activation domain - stabilization of RNA pol for a higher level of transcription (how they work is unclear just remember they come in contact with something at the promoter to stabilize and increase transcription)
3 - Dimerization domain - not all have this domain, but they help create dimers or larger structure formation to increase the affinity of activators and DNA or to allow for flexibility in DNA binding by mixing and matching promoters
What is the significance of the Transcription Factor domains and the yeast two hybrid system?
These domains were found to be able to act independently of each other. This means that it doesn't matter which binding domain the activation domain is bound to, it will still result in transcription of the gene that is associated with the binding domain. This can be used to study protein-protein interaction by attaching one of the target proteins to the binding domain and the other to the activation domain (this interaction is used in place of direct-domain interaction). If the target proteins interact, transcription of the reporter gene will occur.
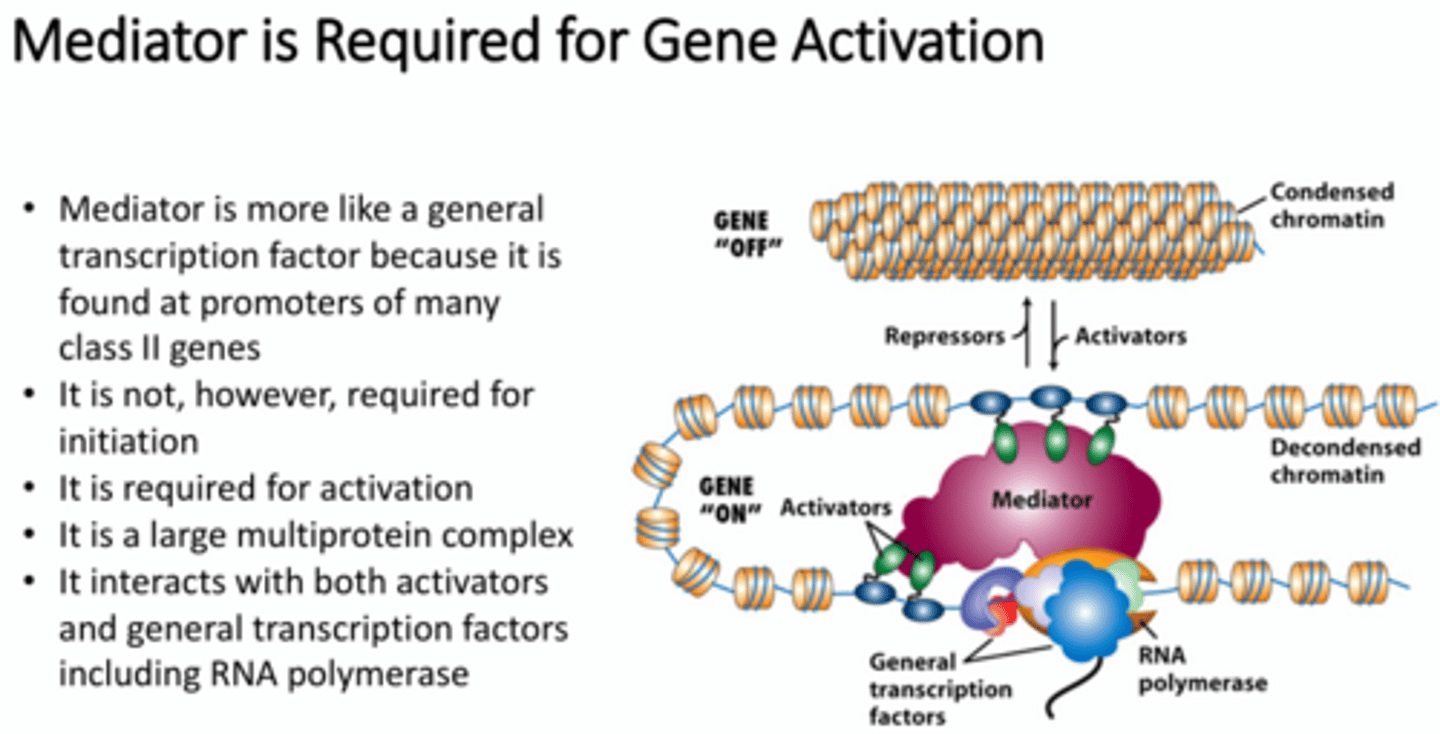
What is a mediator?
A general transcription factor. It is found at promoters of many class II genes. It is a large multiprotein complex that interacts with both activators and general transcription factors and RNA polymerase. It transmits signals from the TFs to the RNA pol
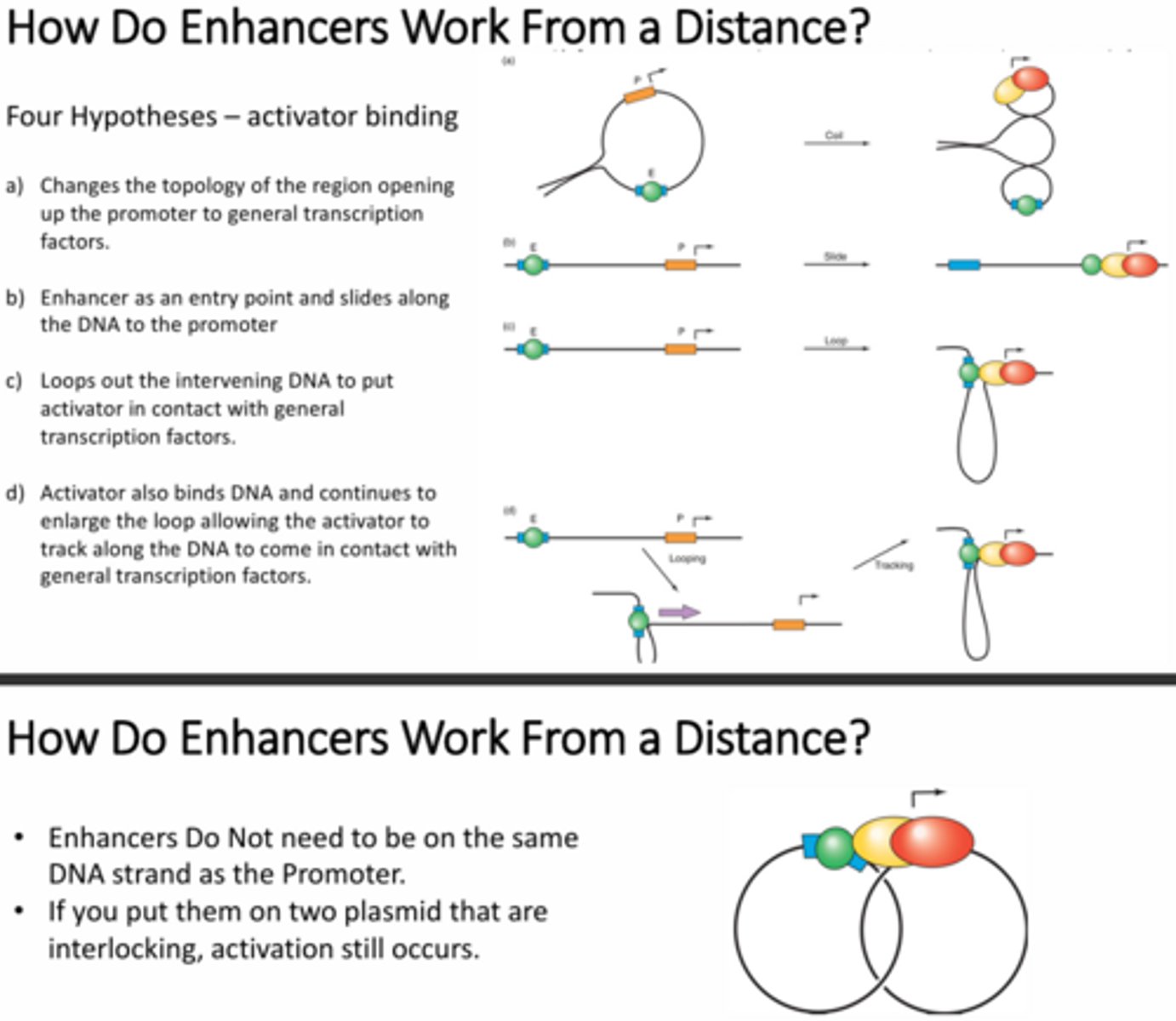
Briefly explain how researchers were able to show that a sliding mechanism for enhancer activation of a promoter is not a likely mechanism?
The sliding mechanism would require that the enhancer and the promoter be on the same strand of DNA because the enhancer would need to slide down the strand of DNA to reach the promoter. They designed an experiment in which there were two interlocked plasmids, one contained the enhancer and the other contained the promoter. They found that transcription did still occur even though the two elements were not on the same strand of DNA, the proteins were just in close proximity due to the linkage of the plasmids.
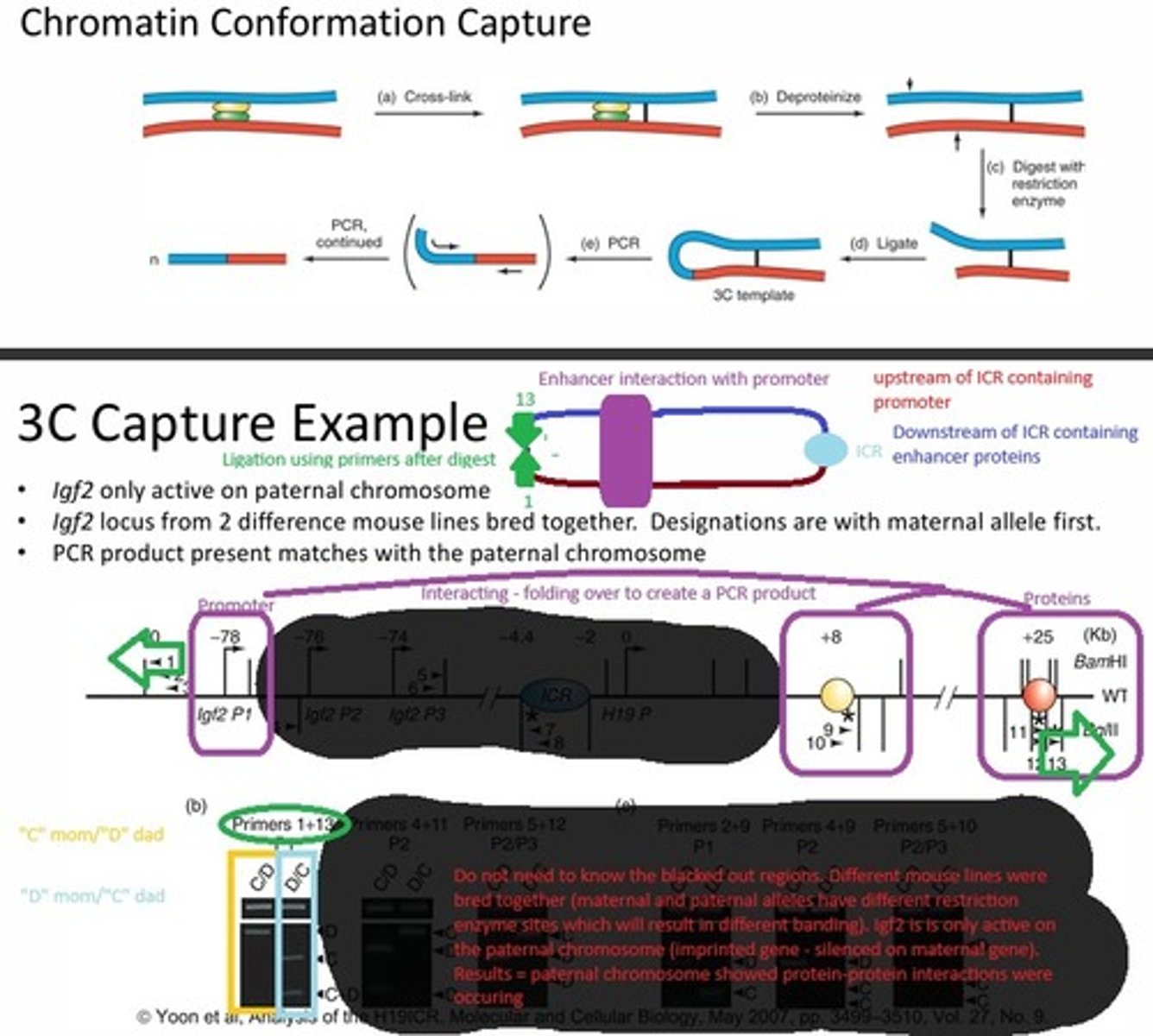
In the 3C chromatin capture experiment, if a PCR product is formed, what does that tell you about an enhancer and a promoter?
The DNA was cross-linked, which would help hold the DNA together where the protein-protein interactions were occurring. Then the DNA was deproteinated and digested but the regions where the proteins were interacting remained connected via the crosslink. Those two sections of DNA were ligated together and PCR was performed. If a product is formed, then the enhancer and promoter are in close proximity and more product means more interactions are occurring
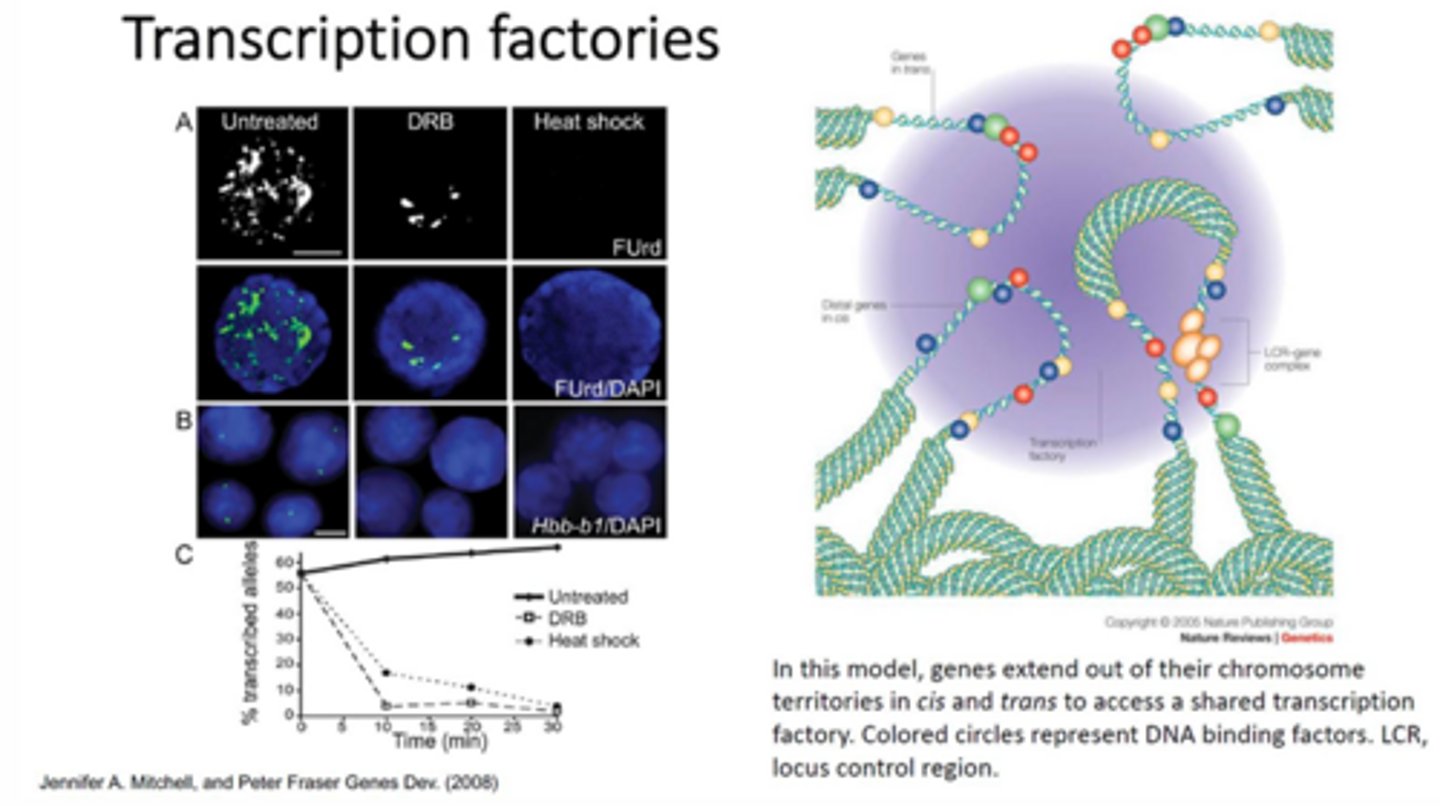
What is a transcription factory? What evidence supports the existence of transcription factories?
Within the nucleus, these are regions where transcription is occurring and the chromosomes/regions that need to be transcribed are moving into these regions and opening up for transcription.
This is supported by labelling newly synthesized RNA so that it can be viewed using fluorescence and viewing the nuclear activity. When this experiment is performed, it is seen that there are many localized regions of RNA rather than it being confluent through the nucleus. After inhibition of RNA pol II, there are some focal points left that illustrate where RNA pol I is transcribing rRNA. After heat shocking the nucleus, there are no focal points of new RNA.
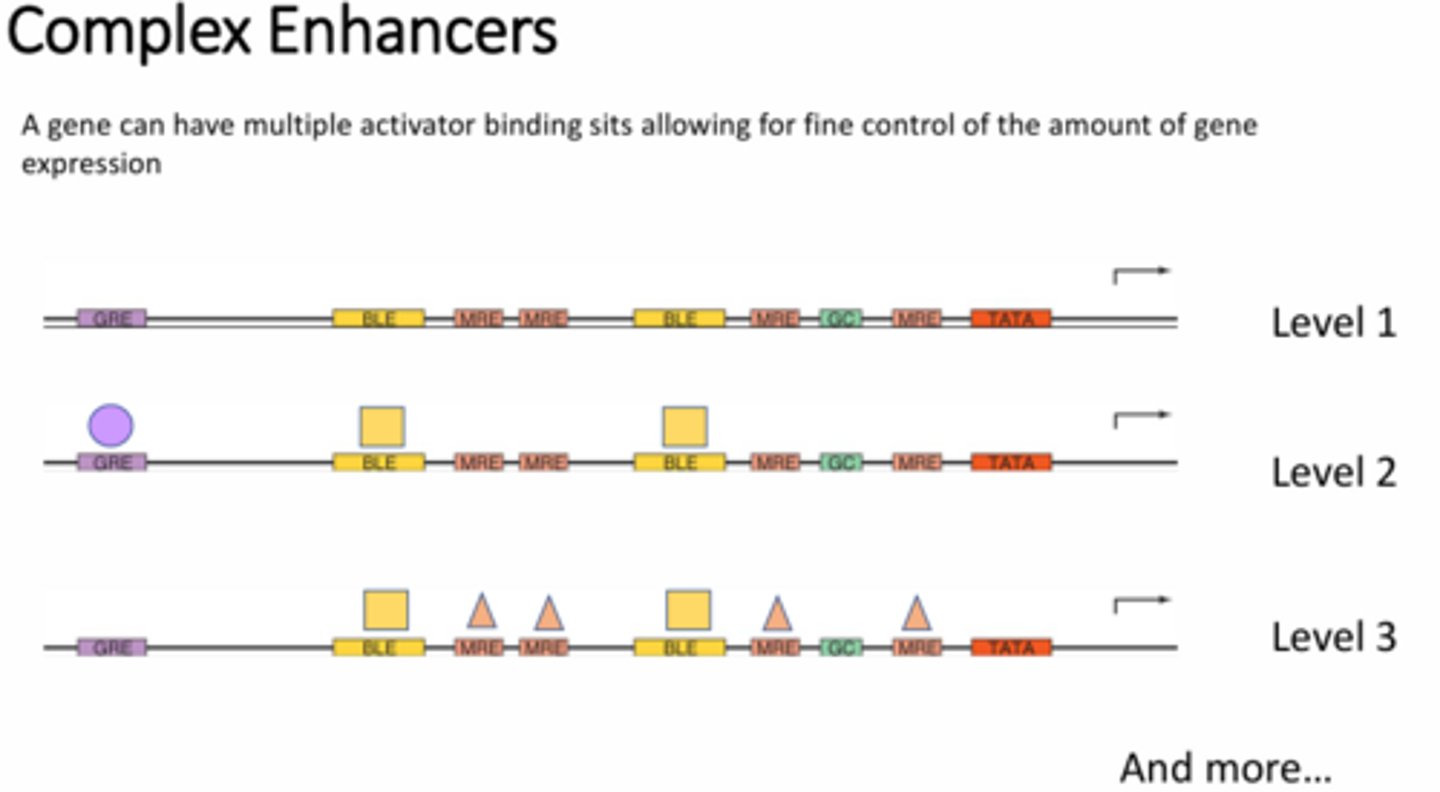
Explain how complex enhancers are able to achieve multiple different levels of transcription.
A single gene can have multiple activator binding sites, which can change depending on tissue/cell type or the conditions surrounding/within the cell. The use of different transcription factors and enhancers interacting with these various sites under different conditions can result in different levels of transcription. In other words, not all enhancers are created equal. Depending on the quantity of enhancers or what other elements are being interacted with, the level of transcription can change.
What is an enhanceosome?
Collection of transcription factors on the DNA that all come together around the TATA Box to form a transcription complex and activate transcription efficiently
What is the advantage of having enhancers act as an enhanceosome?
An enhanceosome is a complex of proteins that is required to be bound at once in other to get a high level of transcription. This results in more of a simple transcription on or off switch rather than different levels depending on what enhancers are present. When the transcription is turned on then the level of transcription is high at the same level every time rather than the expression changing depending on various conditions.
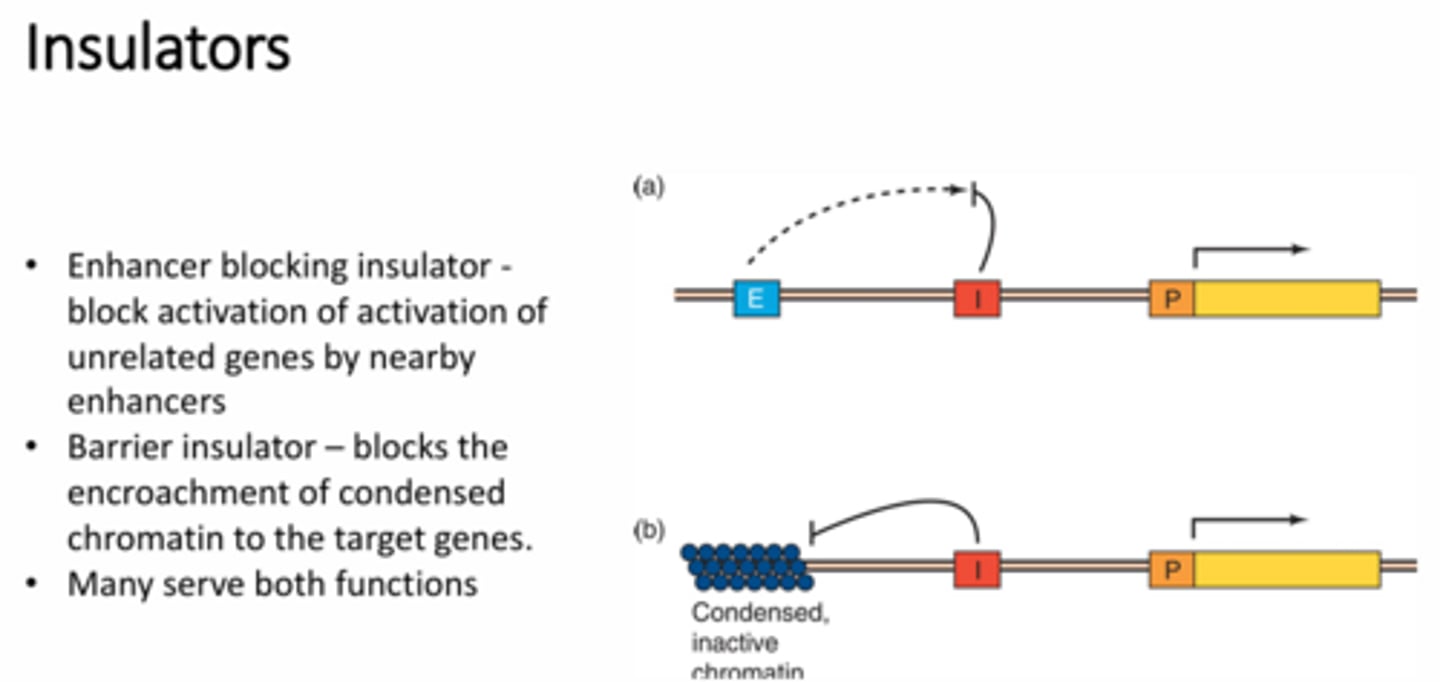
Describe the two mechanisms by which insulators may block the ability of enhancers to stimulate transcription of a gene
1 - enhancer blocking insulator - a protein binds to the insulator region that will block the enhancer from interaction with the nearby promoter region.
2 - barrier insulator - blocking inactive chromatin encroachment into the gene so that the gene can still be transcribed rather than tightly bound up into condensed chromatin
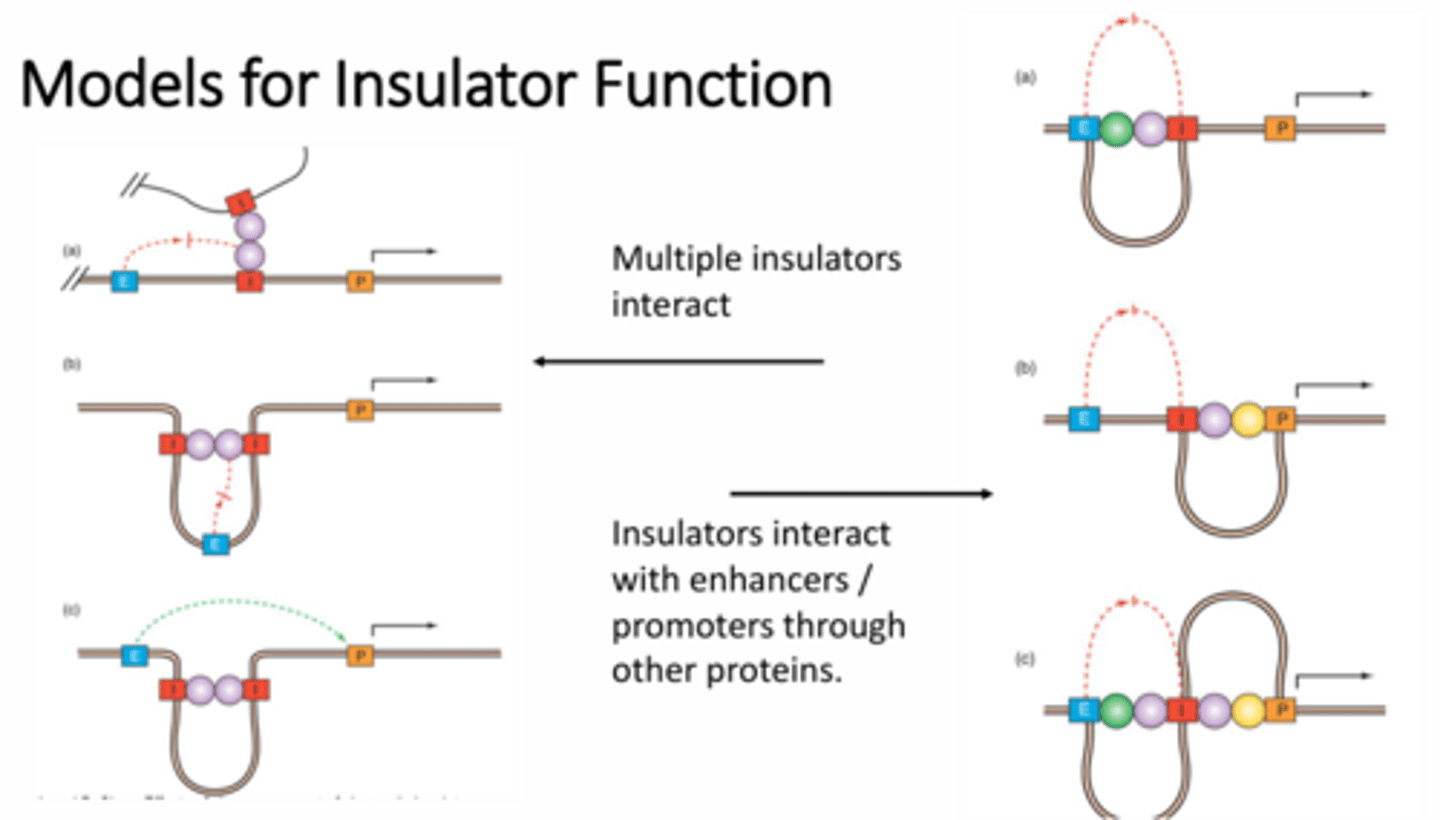
What are the two ways in which the enhancer blocking insulators can occur?
1 - multiple insulators can interact between two different strands of DNA to block the enhancer's ability to fold over and interact with the promoter. If the insulators are on the same strand of DNA they can interact to form loops to prevent the folding of the enhancer or to hide sections of DNA from the enhancer
2 - insulators can also interact with other proteins in order to block enhancer interactions. These extra proteins can result in interactions with the enhancers, the promoters, or both to block enhancement of transcription of a given gene.
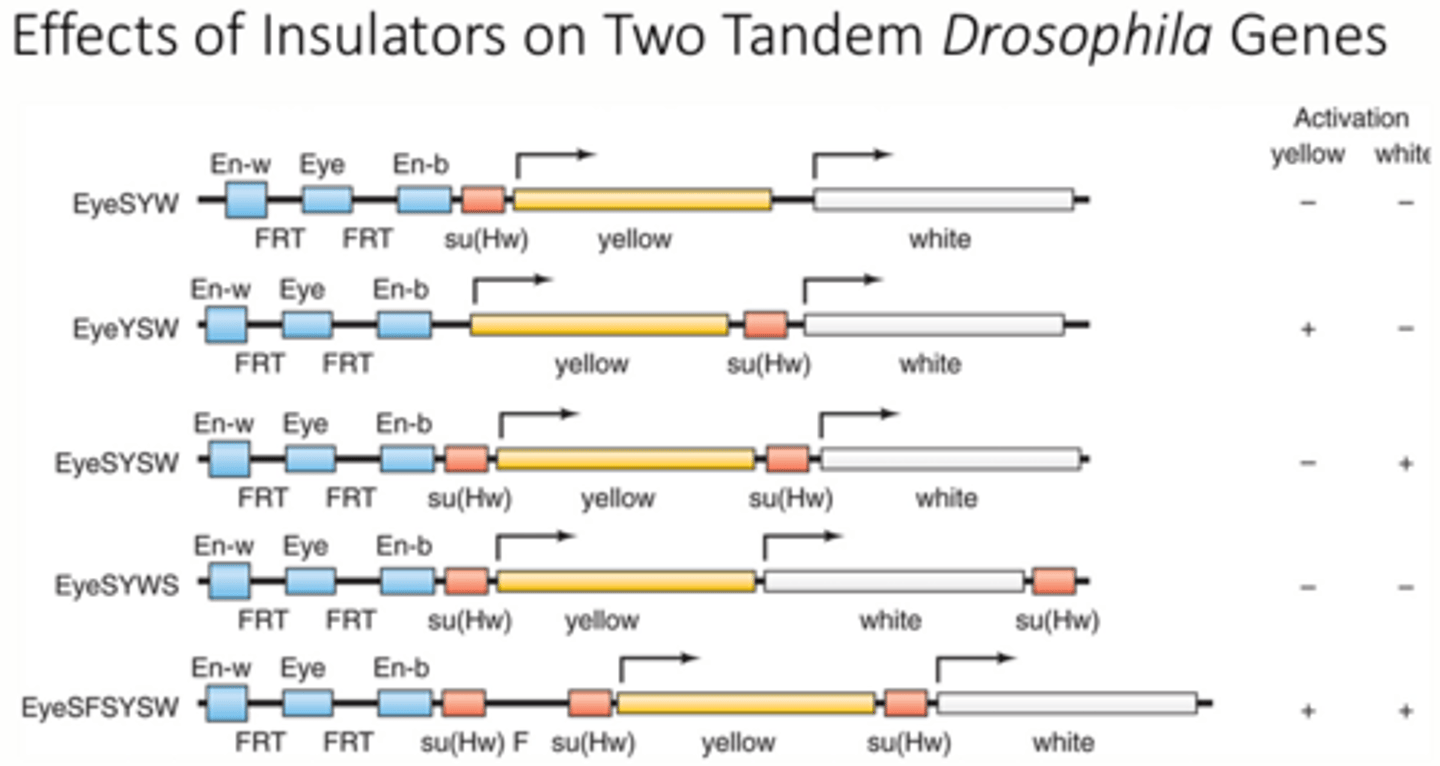
What is an experimental example of insulators in action?
1 - insulator blocks both color genes
2 - insulator blocks only the white gene
3 - insulator blocks only the yellow gene
4 - insulator blocks both color genes
5 - neither color gene is blocked
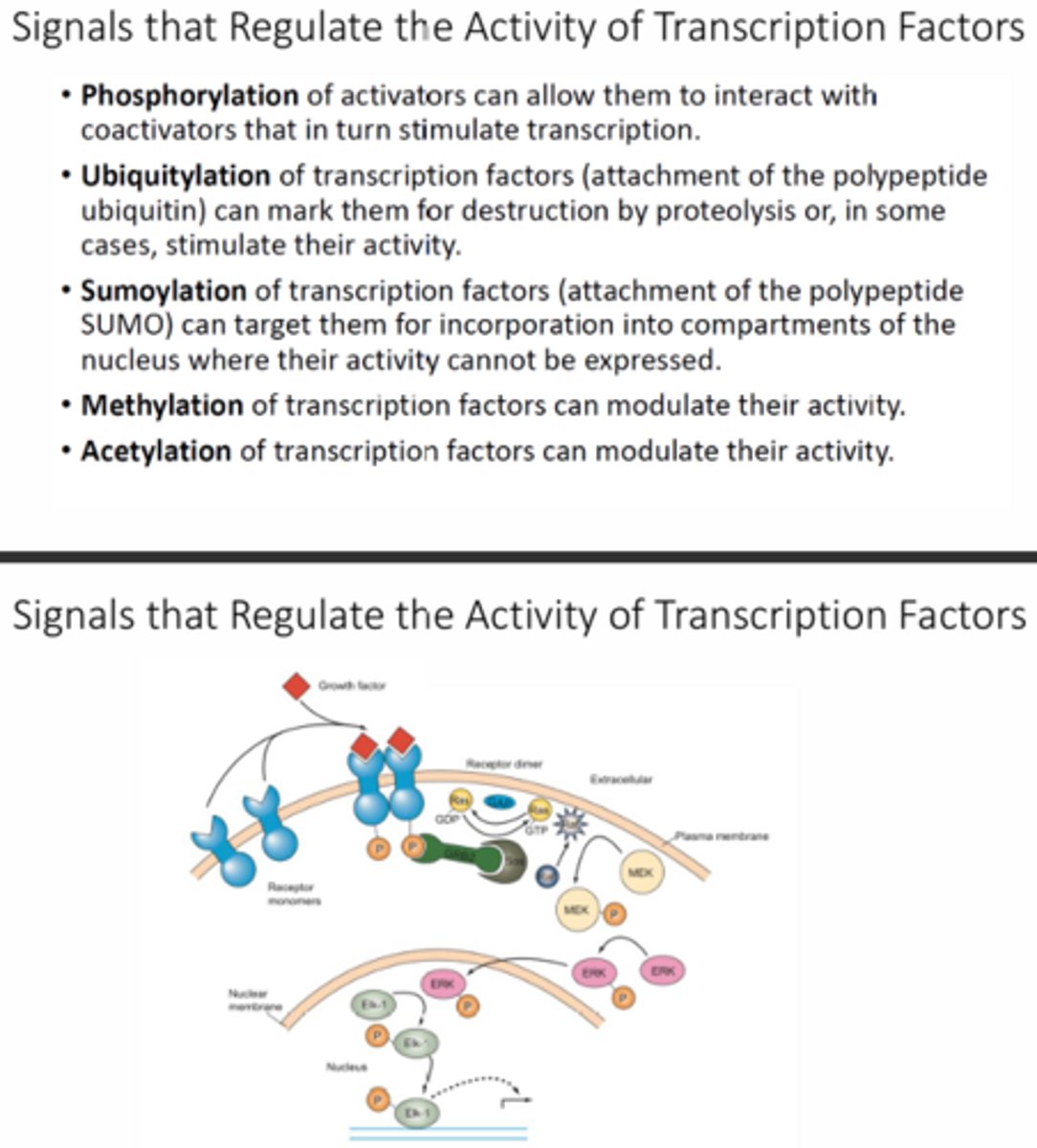
Why is signal transduction an important concept in understanding when are where genes are transcribed?
Signal transduction is key to when genes are transcribed because the cascade can alter when genes are being transcribed based on the conditions outside of the cell or depending on the cell type/location. These cascades can lead to the activation of enhancers/transcription factors to increase transcription or they can inactivate them or they can even lead to the activation of repressors to inhibit transcription.

What are the three processing steps necessary for a pre-mRNA to become a mature mRNA?
1 - 5' capping
2 - splicing
3 - 3' poly-adenylation
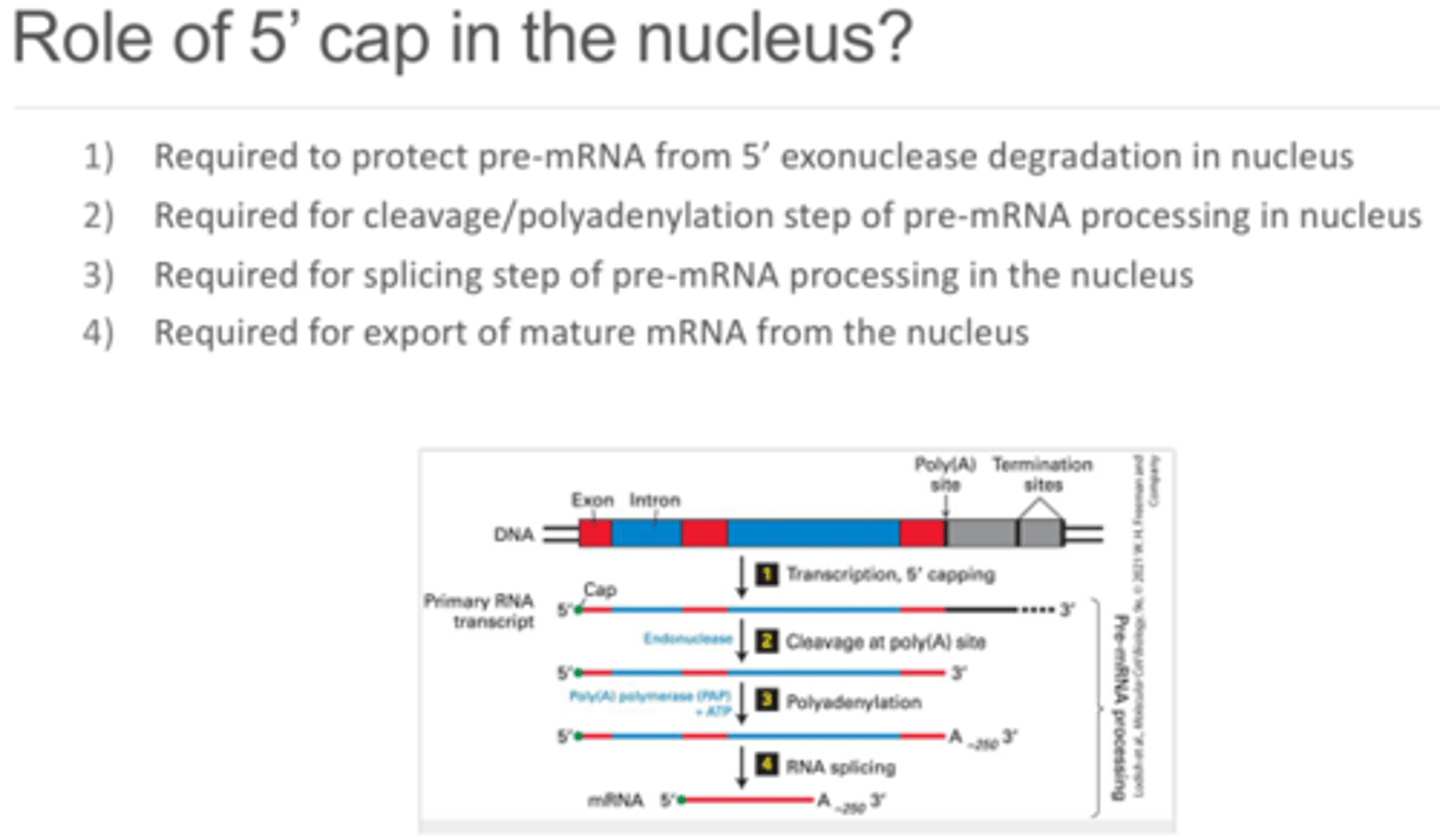
In addition to protecting the 5' from degradation, what are some of the 5' cap's other functions in the nucleus?
In addition to protecting the 5' end from being degraded by exonucleases, it also stabilizes the 5' end of the mRNA. The 5' cap is required for cleavage and polyadenylation of the pre-mRNA processing, the splicing of pre/mRNA, and export of the mature mRNA from the nucleus and into the cytoplasm.
What happens to pre-messenger RNA that is improperly processed?
It is rapidly degraded by the exosome within the nucleus
What does it mean to say the mRNA is never "free"?
Within the nucleus, the mRNA is always coated with nuclear proteins that protect it from degradation and allows for nuclear export. During this export the nuclear proteins are exchanged for cytoplasmic proteins. Therefore, the mRNA is never naked or "free." These proteins and RNA interactions are known as ribonuclear protein (RNP) complexes.
Describe the structure of the RNA polymerase II carboxy-terminal tail domain (CTD). What is its role in capping?
The CTD of RNA pol II is a long tail coming off of the globular portion of RNA pol II and it functions to bind to the enzymes required to make the 5' cap as well as the nascent RNA that is being made. This allows for rapid capping of the pre-mRNA molecules as they are being synthesized.
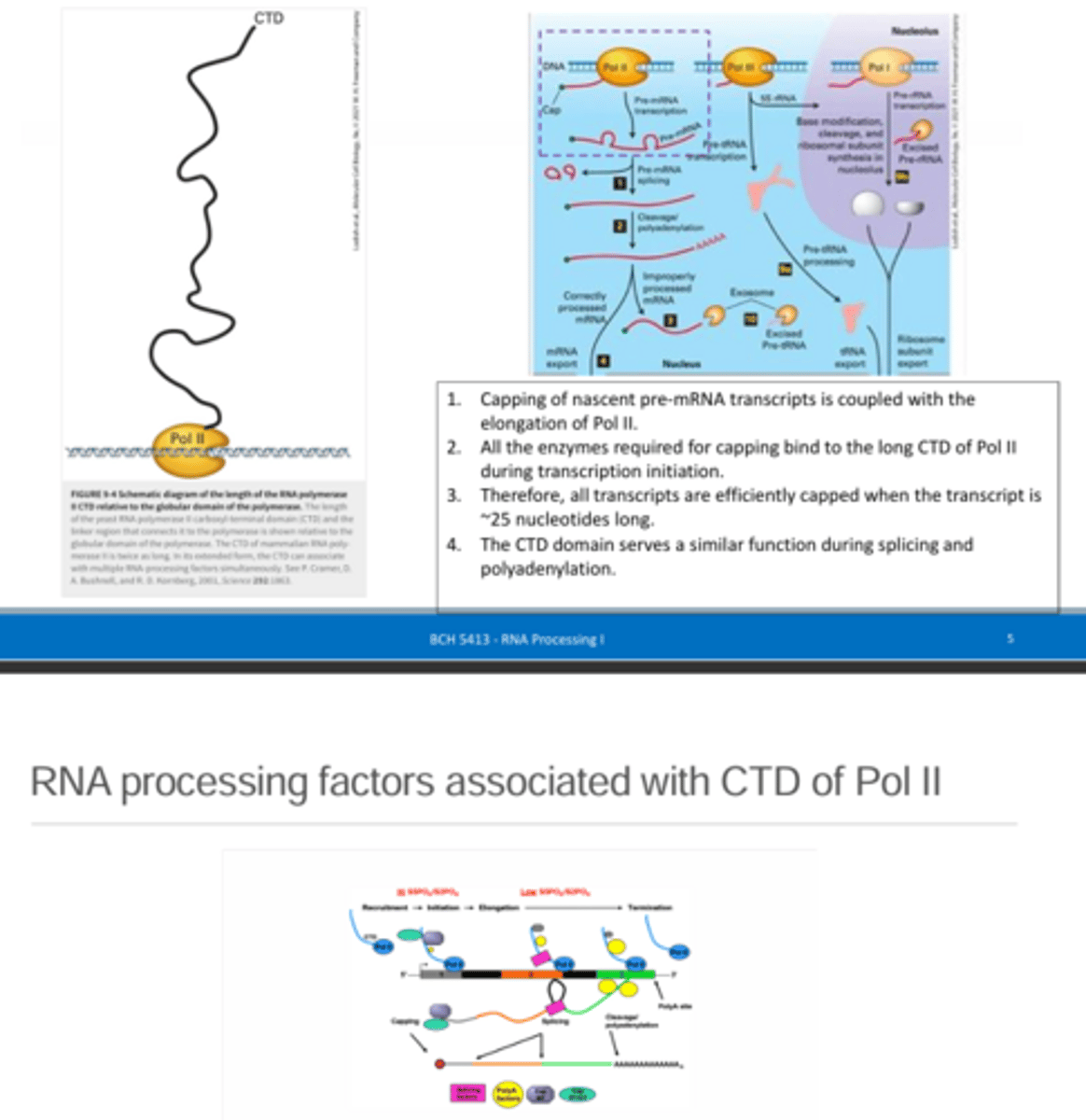
Over time, what happens to the proteins recruited to the CTD of RNA pol II?
They dissociate to allow for other proteins to bind (e.g., proteins associated with splicing or poly-A tail formation depending on the stage of transcription).
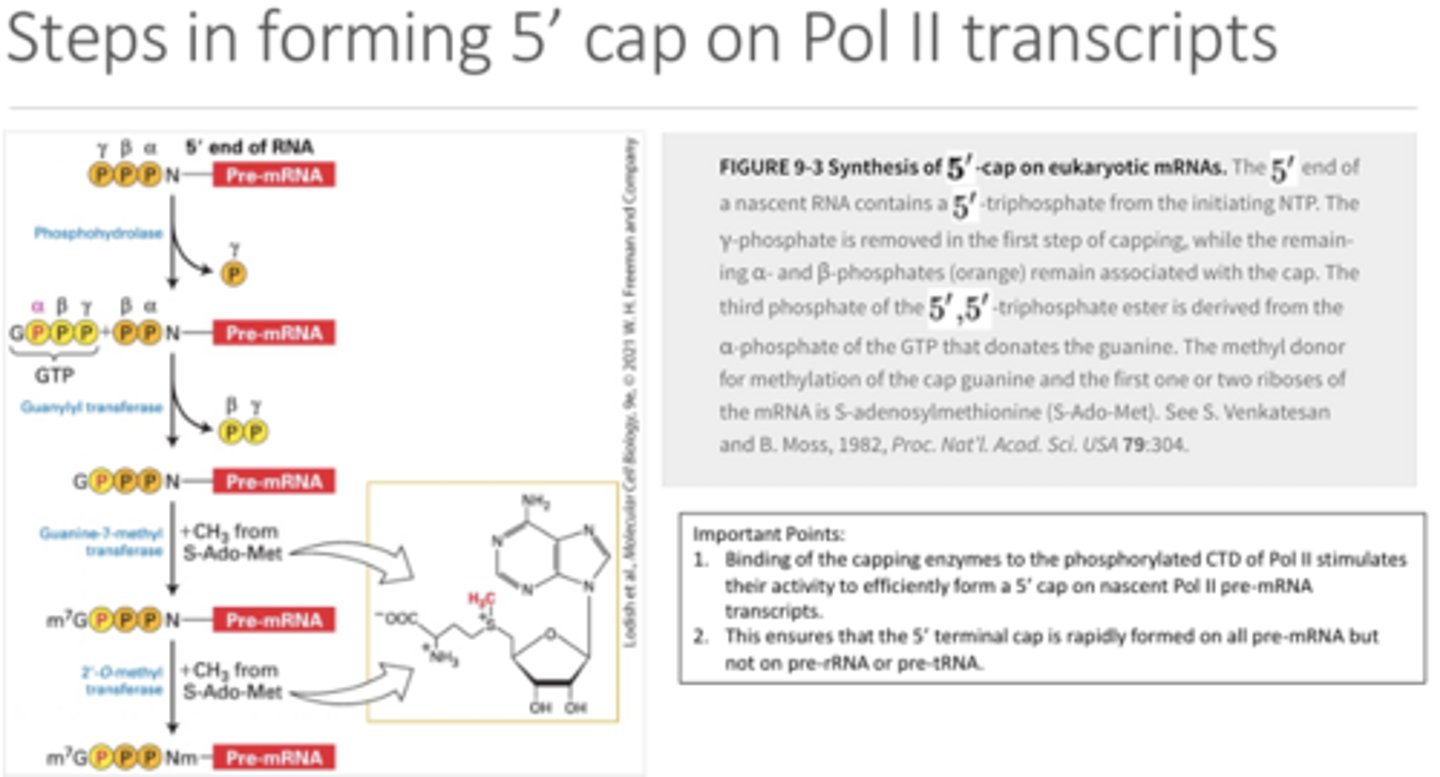
What is the structure of the 5' cap? AND why is it so efficient at protecting the RNA from degradation?
The 5’ most end is the 7-methyl guanine followed by the triphosphate of the first nucleotide via a 5’-5’ linkage. This linkage is very unusual. The 5’ exonuclease is designed to cleave at a 5’-3’ linkage between bases so it is unable to cleave at the 5’-5’ linkage. The triphosphate is then bound to cap 1 nucleotide which is bound to cap 2 nucleotide. Both these cap nucleotides have an extra methyl group added on as well.
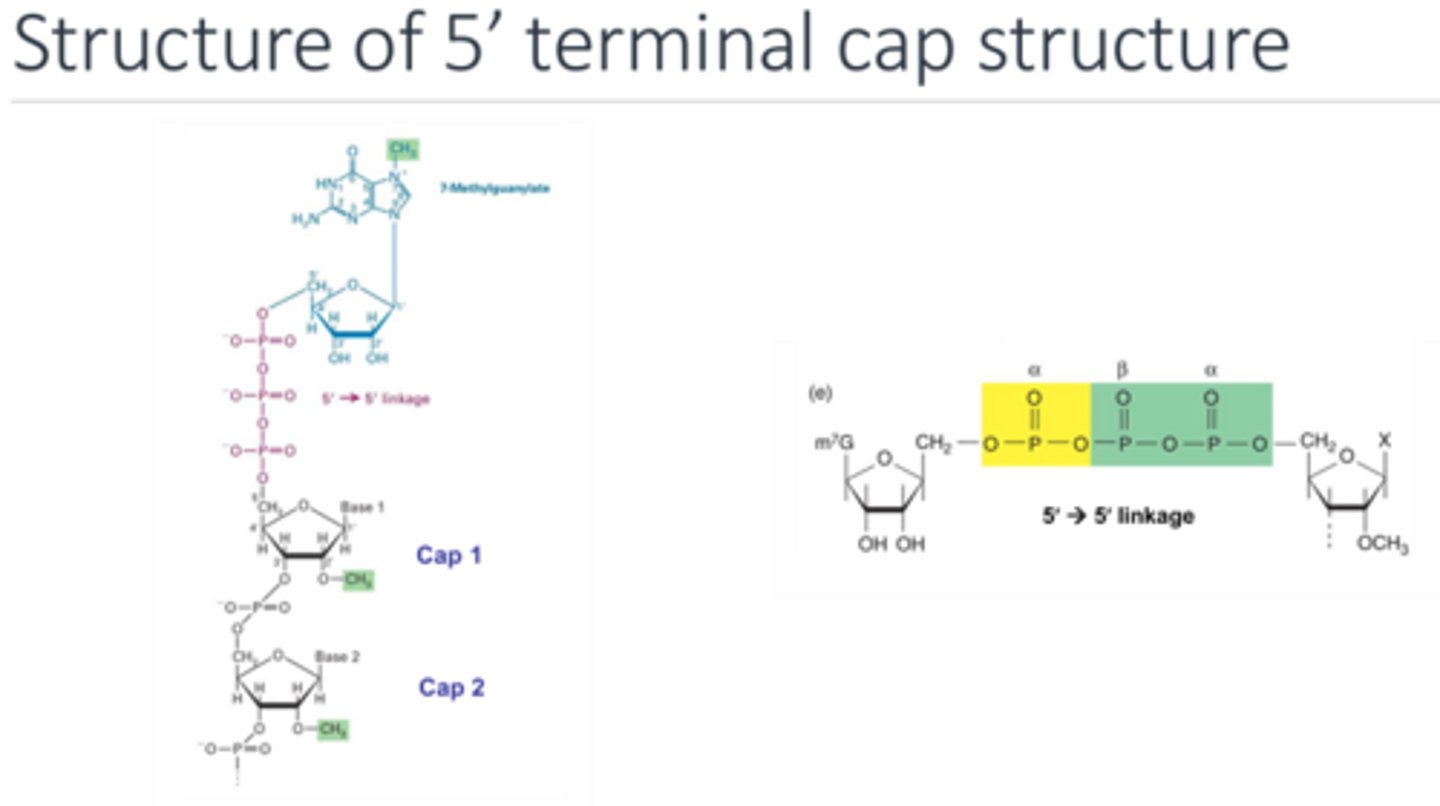
How is the 5' cap formed?
The first 5' nucleotide loses its gamma phosphate and it is replaced by the alpha phosphate (still attached to the donor guanine). Then another methyl group is added to the guanine to make it 7-methyl.
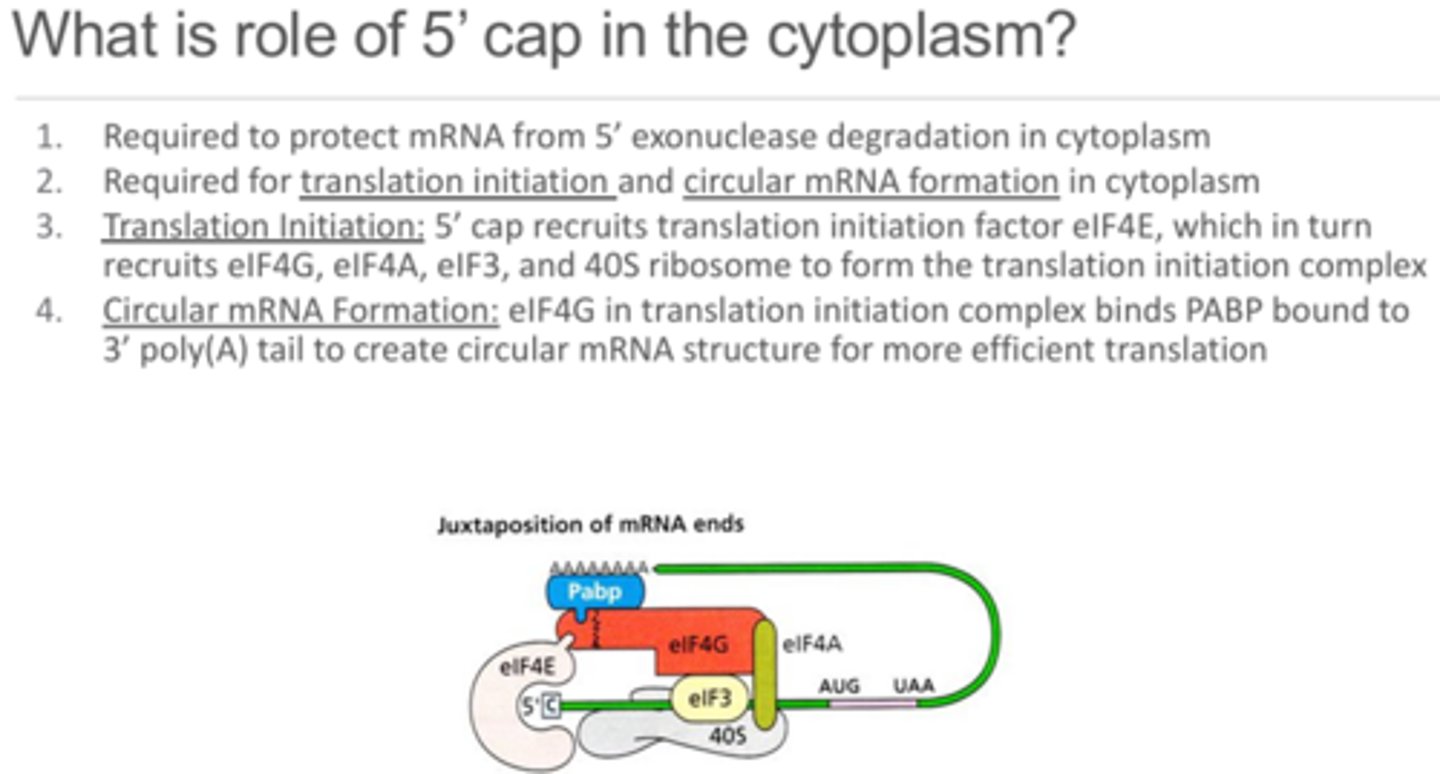
What are the roles of the 5' cap in the cytoplasm?
1 - Protect from 5' exonuclease degradation that is also possible in the cytoplasm.
2 - Translation initiation - 5' cap recruits the translation initiation complex by recruiting eIF4E which recruits the other complex components and the 40S ribosome.
3 - Circular mRNA formation - eIF4E which is bound to the 5' cap binds to eIF4G, which is then able to bind to PABP which is bound to the poly-A tail thus creating a loop or circularization of the mRNA.
Why don't tRNA's and rRNA's get capped?
The CTD is bound to RNA pol II, which transcribes mRNA only.
During pre-mRNA processing, what is the function of the CPSF complex?
Binds upstream of the cleavage site at the AAUAAA sequence in the pre-mRNA, this is the poly-A signal sequence.
During pre-mRNA processing, what is the function of the CFI/II complex?
Stabilizes the interaction between the CPSF and CStF complexes (does not bind directly to the RNA), the main function is also to cleave the RNA and create the 3’OH for the poly-A tail formation.
*Interactions with this complex stabilize the bend in the RNA needed for cleavage
During pre-mRNA processing, what is the function of the CStF complex?
Binds to the GU or U rich sequence downstream of the cleavage site as well as the CPSF complex to create an RNA loop
During pre-mRNA processing, what is the function of the Poly-A polymerase (PAP)?
Stimulates the cleavage of the RNA between the CPSF and CStF complexes (aka at the poly-A site), then adds at least 12 A residues after the other cleavage factors are released along with the RNA remaining with the RNA pol complex.
During pre-mRNA processing, what is the function of the Poly-A binding proteins (PABP)?
Binds to the short pol-A tail and accelerates the rate of A residue addition by PAP, after about 200-250 A residues have been added it signals the PAP to stop
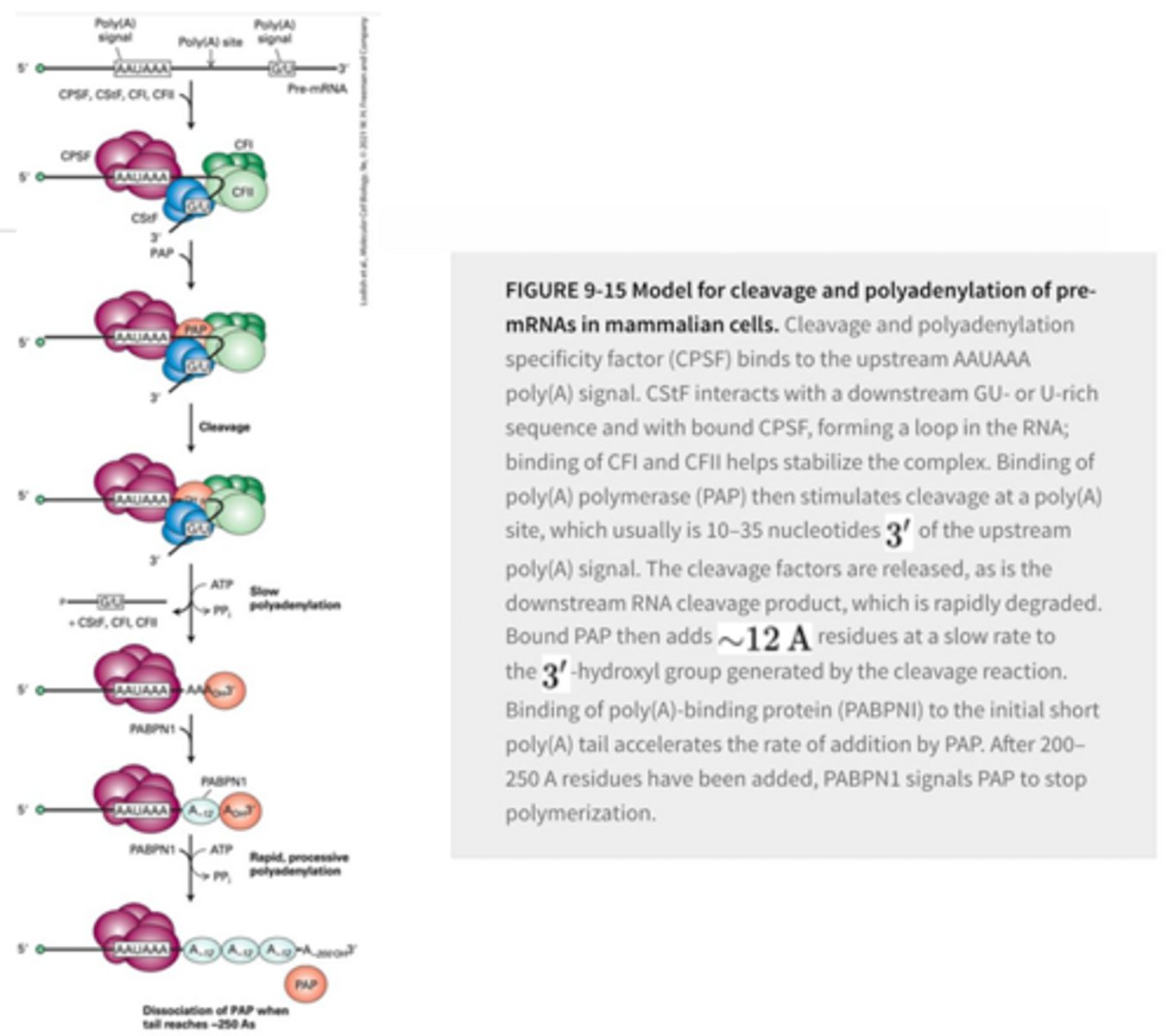
What is the sequence of events for adding the Pol-A tail to the pre-mRNA?
CPSF and CStF binding at their respective Poly-A signal sites and form a bend in the RNA >> CFI/II stabilize the CPSF-CStF interaction and RNA bending >> PAP binding >> PAP cleavage of RNA >> Cleavage product is rapidly degraded and PAP slowly starts to add A residues >> PABP binds to accelerate the A additions until 200-250 are added >> PABP signals PAP to stop
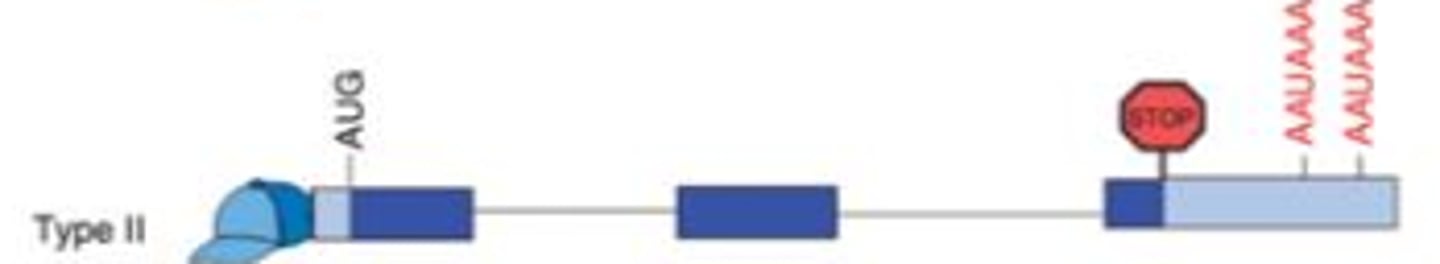
In type II alternative polyadenylation, the protein coding sequence is exactly the same. What, then, is the purpose of the alternative polyadenylation?
This is mainly used when a gene is expressed in different cell types - one will use one of the poly-A sites and the other cell type will use the other site. These sites can result in different mRNA stabilities based on the sequences present in between the two poly-A sites. This can have big effects on how long the mRNA lasts within the cell and thus how much of the protein is made from that mRNA.
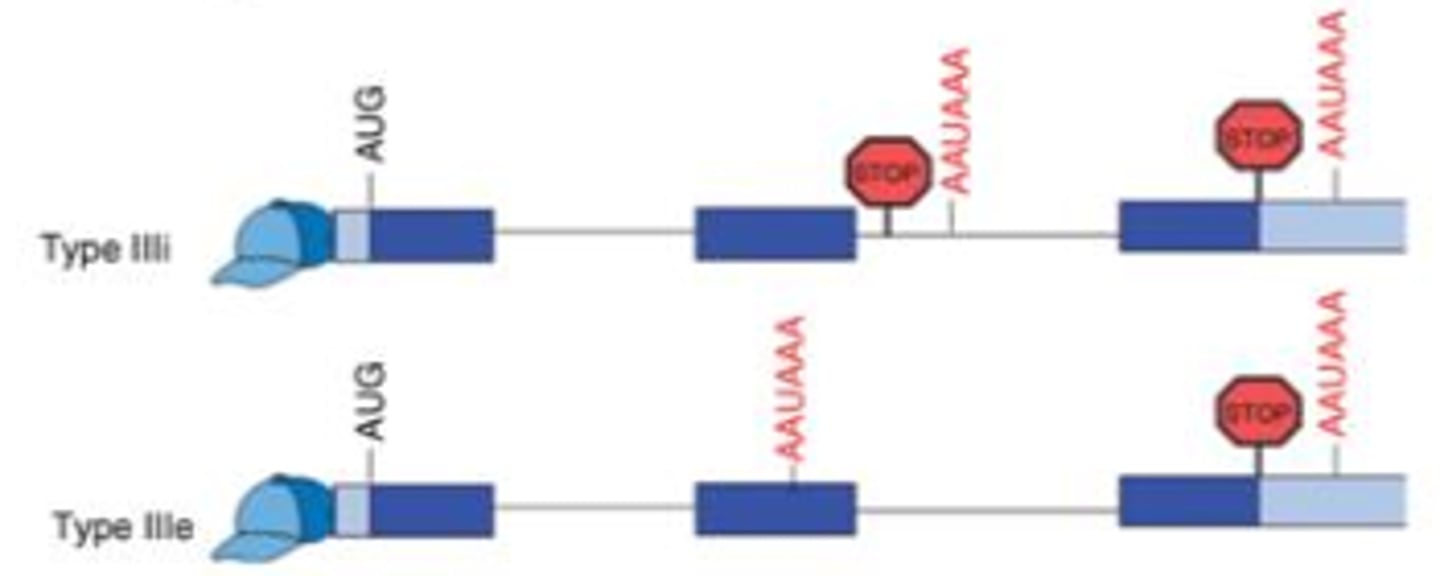
What happens in type III alternative polyadenylation?
The pol-A site within an intron can lead to cleavage and pol-A tail formation with the loss of everything downstream (other exons). While the other site will have cleavage and poly-A tail formation resulting in the mRNA containing all of the exons. In this case different proteins can be created from the same strand of pre-mRNA transcript.
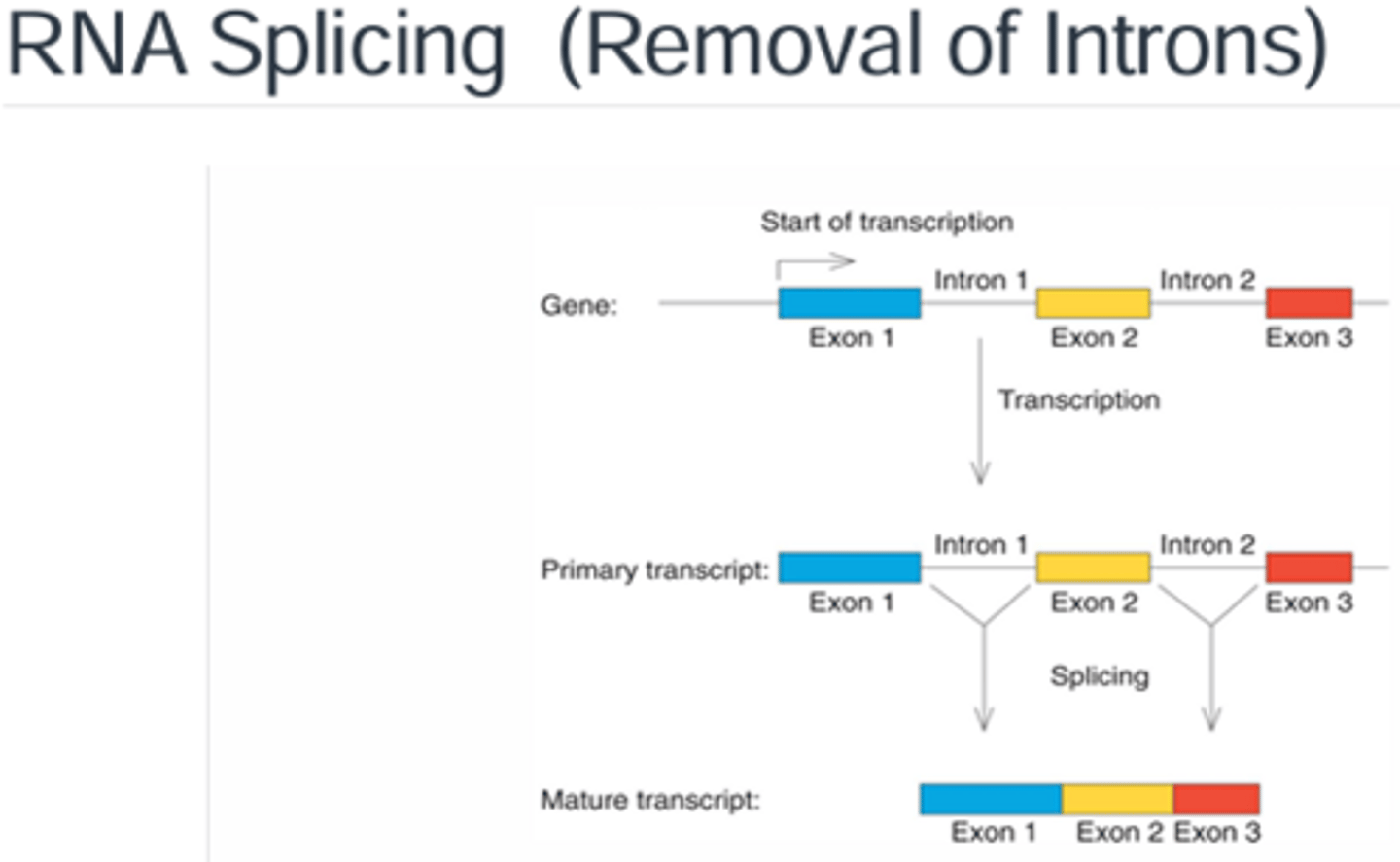
What is the difference between an intron and an exon.
Exons are sequences in the mRNA that are the coding sequences for the proteins while the introns are the in-between sequences that are not translated into proteins
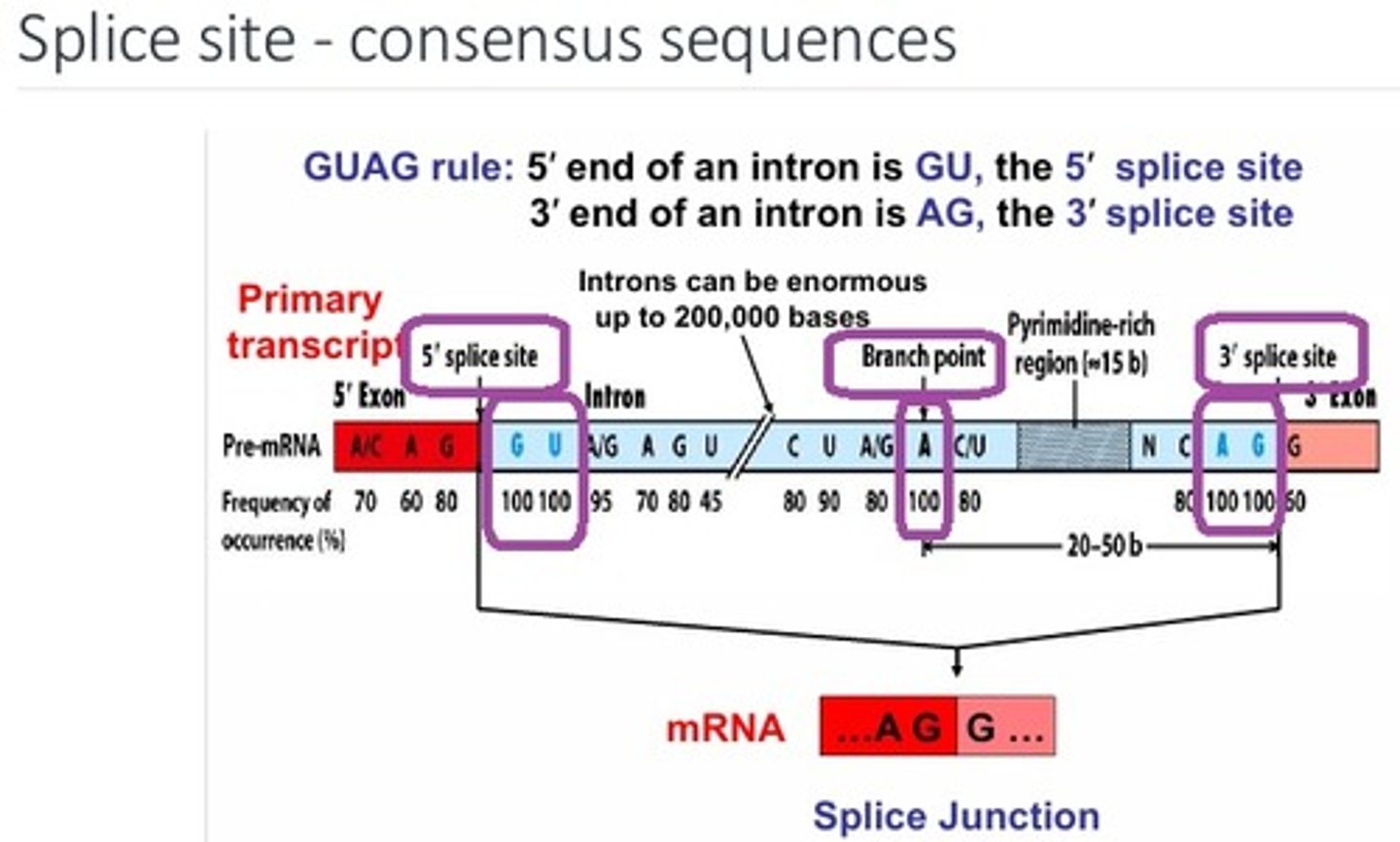
What are the three sequences that are 100% conserved for a splice site and where are they located?
1 - 5' splice site = GU
2 - 3' splice site = AG
3 - branch point 20-50bp from 3' splice site = A
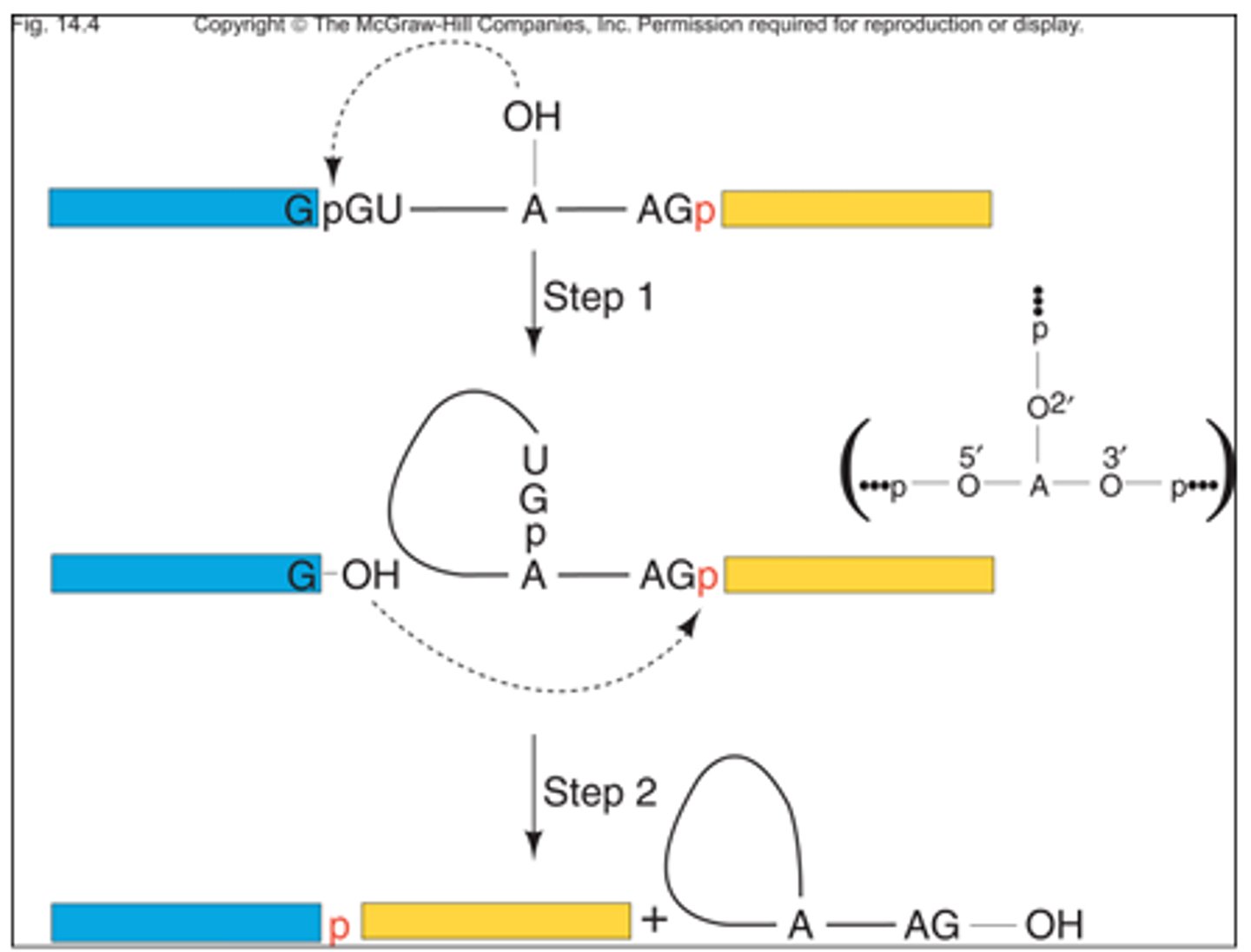
What is the meaning of the term transesterification?
The displacement of alcohol from an ester by another alcohol, similar to hydrolysis (aka the conversion of a carboxylic acid ester into a different carboxylic acid ester)

What is the meaning of the term phosphodiester bond?
A covalent bond in which a sugar is bound to a phosphate like the bonds found between nucleic acids along a single strand (phosphate and 3rd Carbon OH reaction)
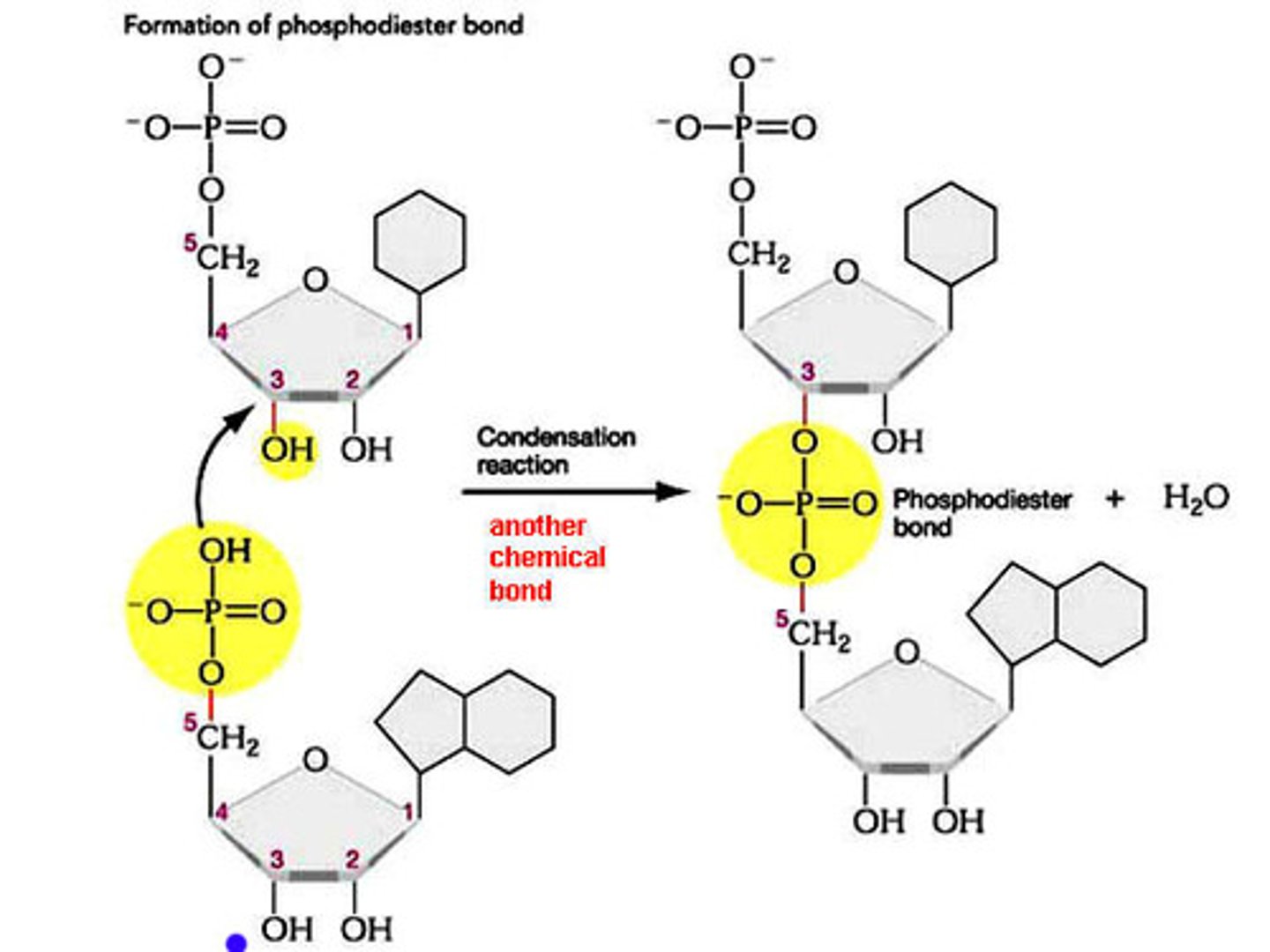
What specific atoms from what specific nucleotides are involved in the first transesterification reaction?
The 2'-OH on the A from the branch site attacks the 3'-phosphodiester bond at the 5' splice site, essentially swapping the OH for the phosphate
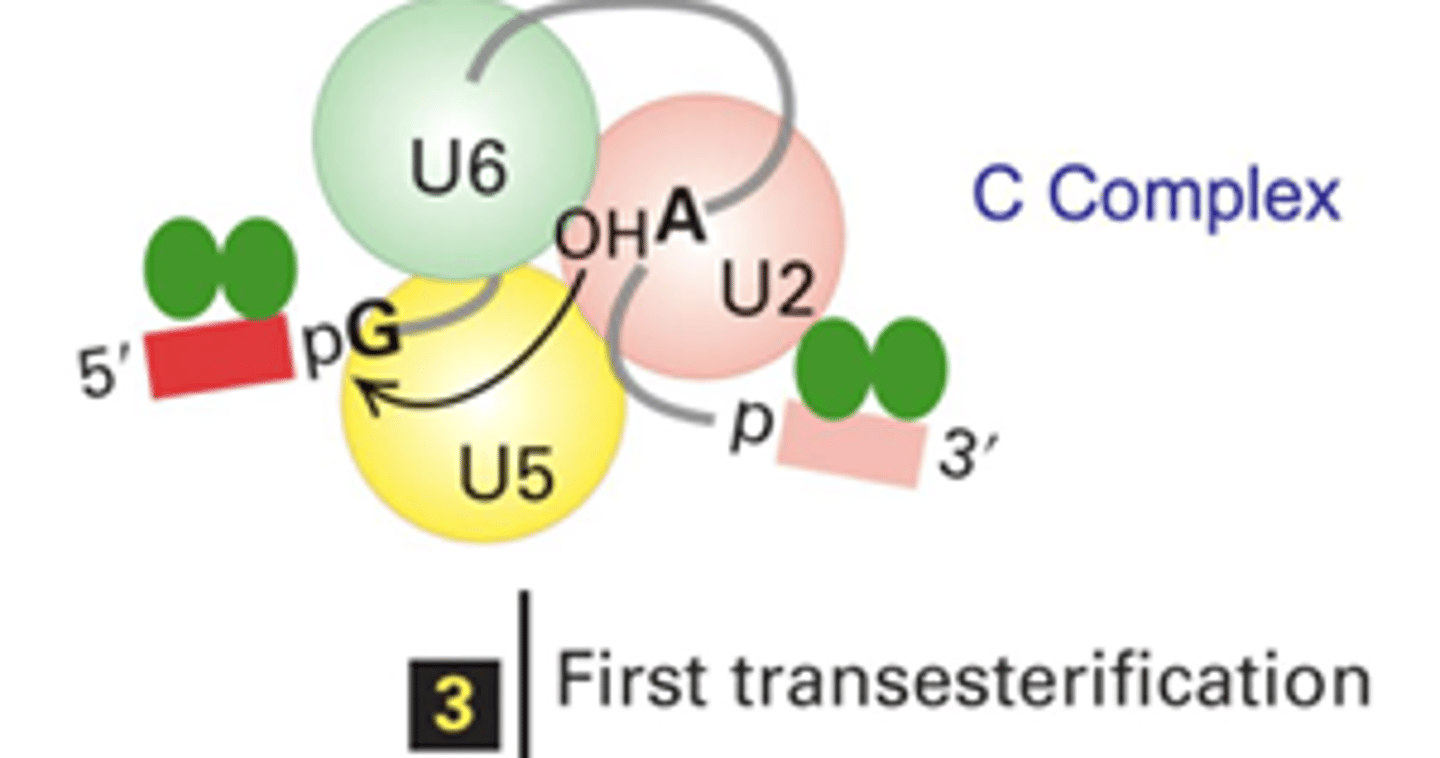
What specific atoms from what specific nucleotides are involved in the second transesterification reaction?
The 3'-OH of the 5' splice site attacks the 3' splice site phosphodiester bond (returning the OH to the removed intron and combining the two exons)
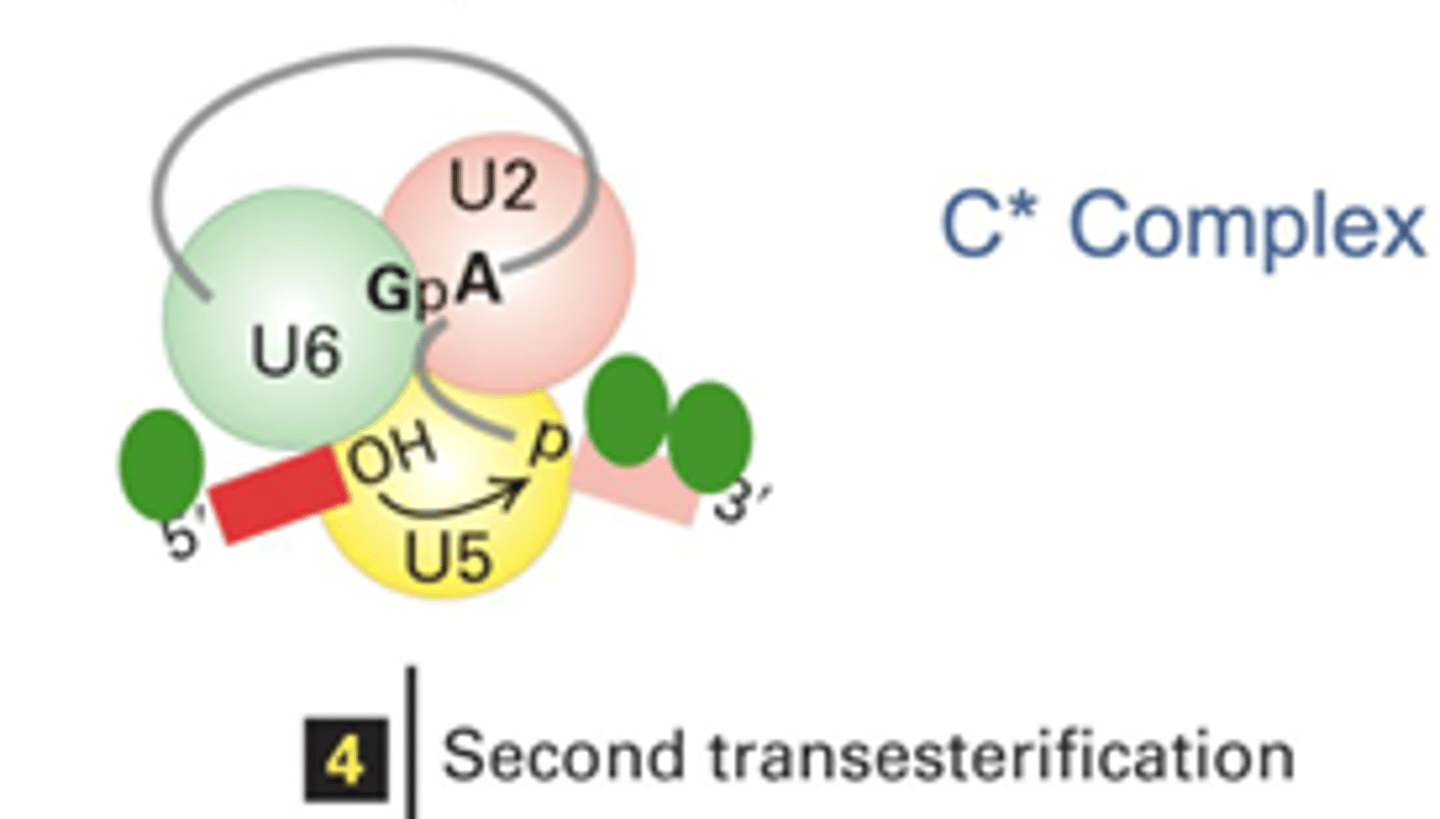
Explain why steps c and d are transesterification reactions and why the reactions break and or form phosphodiester bonds
They are transesterification reactions because of the movement of the OH group (alcohol group) and result in the movement of the phosphodiester bonds that occur between the bases' sugars and the phosphate group in order to create a complete RNA strand.
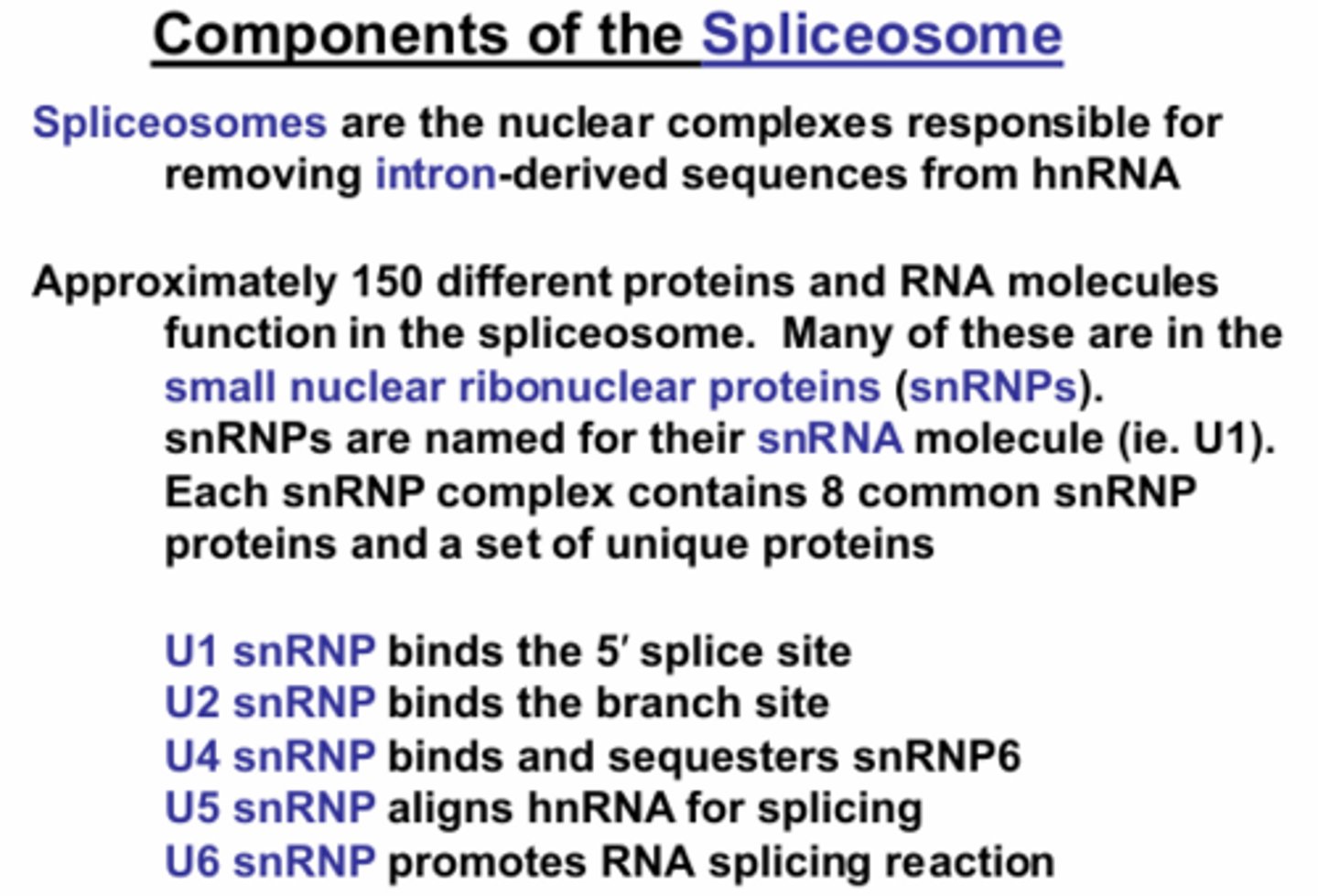
What are the components of the Spliceosome and what are their functions?
U1 snRNP - binds 5' splice site
U2 snRNP - binds branch site
U4 snRNP - binds and sequesters snRNP6
U5 snRNP - aligns RNA for splicing
U6 snRNP - promotes RNA splicing reaction (U6 interacts with the lariat structure and U2)

Briefly describe the steps of splicing.
A complex (U1 binds 5' site, U2AF binds 3' site, to ID branch point, then U2 binds at branch point) >> B complex (U4, U5, U6 recruitment) >> C complex (dynamic rearrangement - U1 and U4 leave, U2 and U6 prepare for 5' attack) >> first transesterification >> second transesterification
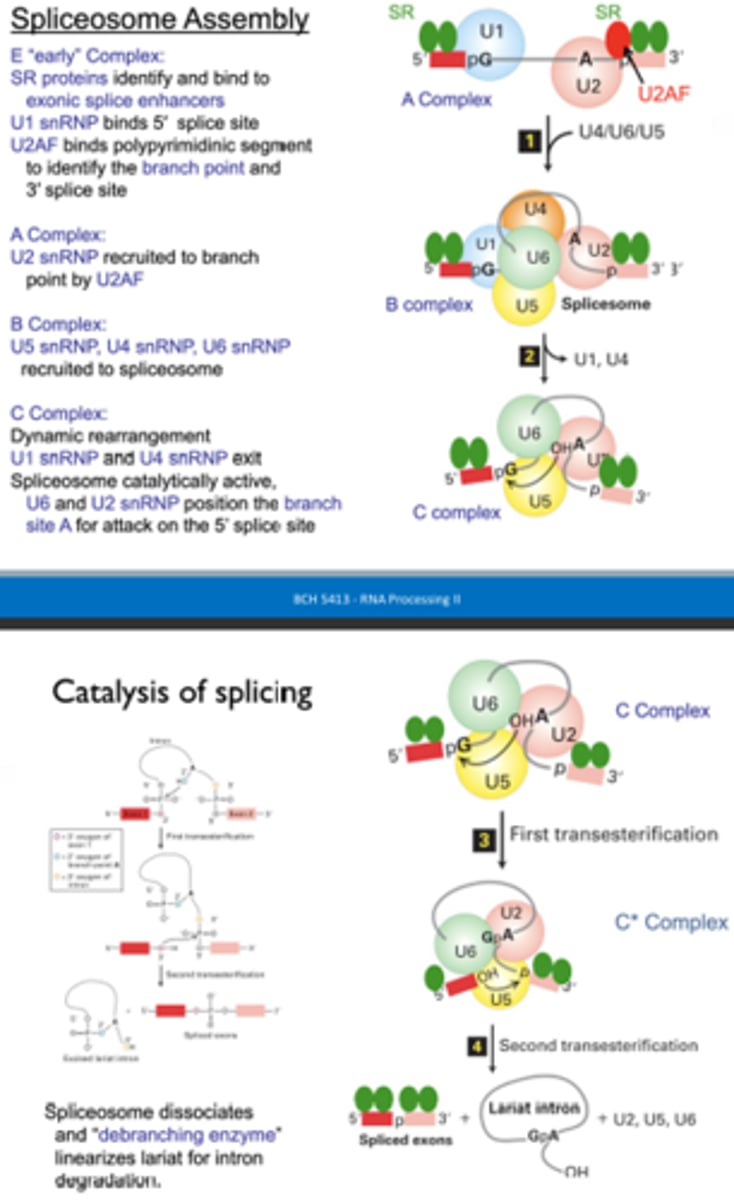
What happens to the lariat intron after it is removed from the pre-mRNA
The debranching enzyme linearizes the lariat intron for degradation.
Explain the complementary base pairing necessary to carry out the first trans esterification reaction.
U2 snRNA binds with the branch point BUT they base pair in such a way that the branch point A is pushed out and not paired with the snRNA. Then the U6 snRNA binds with the intron side of the 5' splice site and the U2 snRNA to hold the reaction sites close together (5' splice site close to the A branch point)
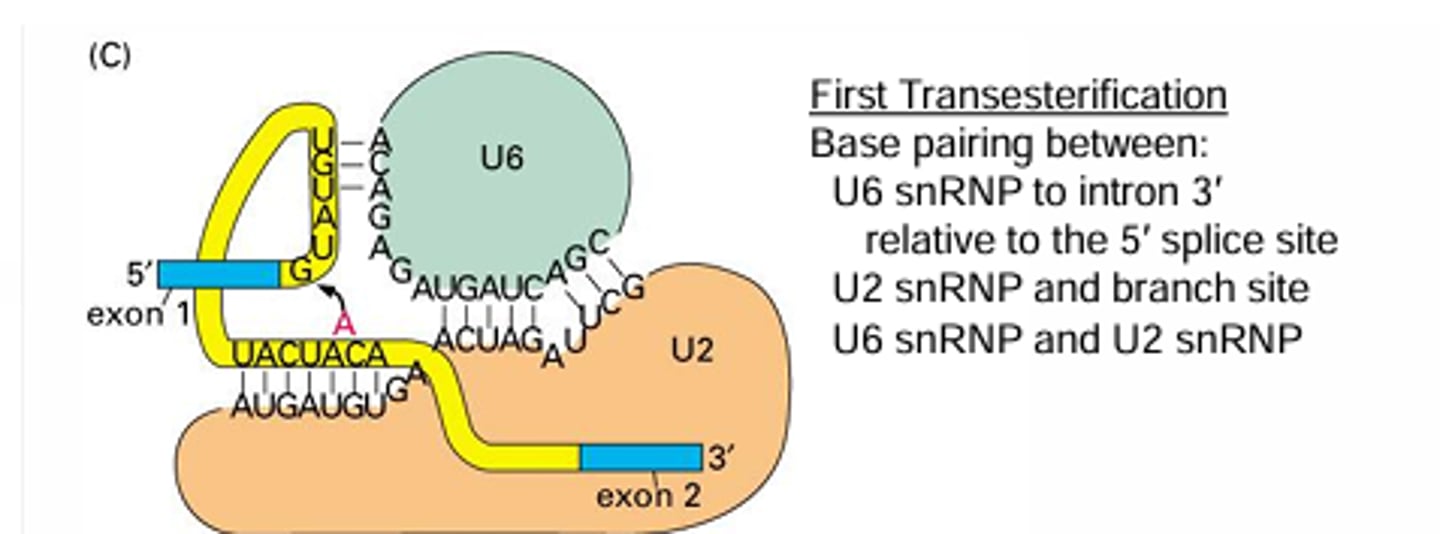
Explain the complementary base pairing necessary to carry out the second trans esterification reaction.
The U6 still holds the same base pairing while U5 binds with two exons in order to hold them close together for the final transesterification to occur.
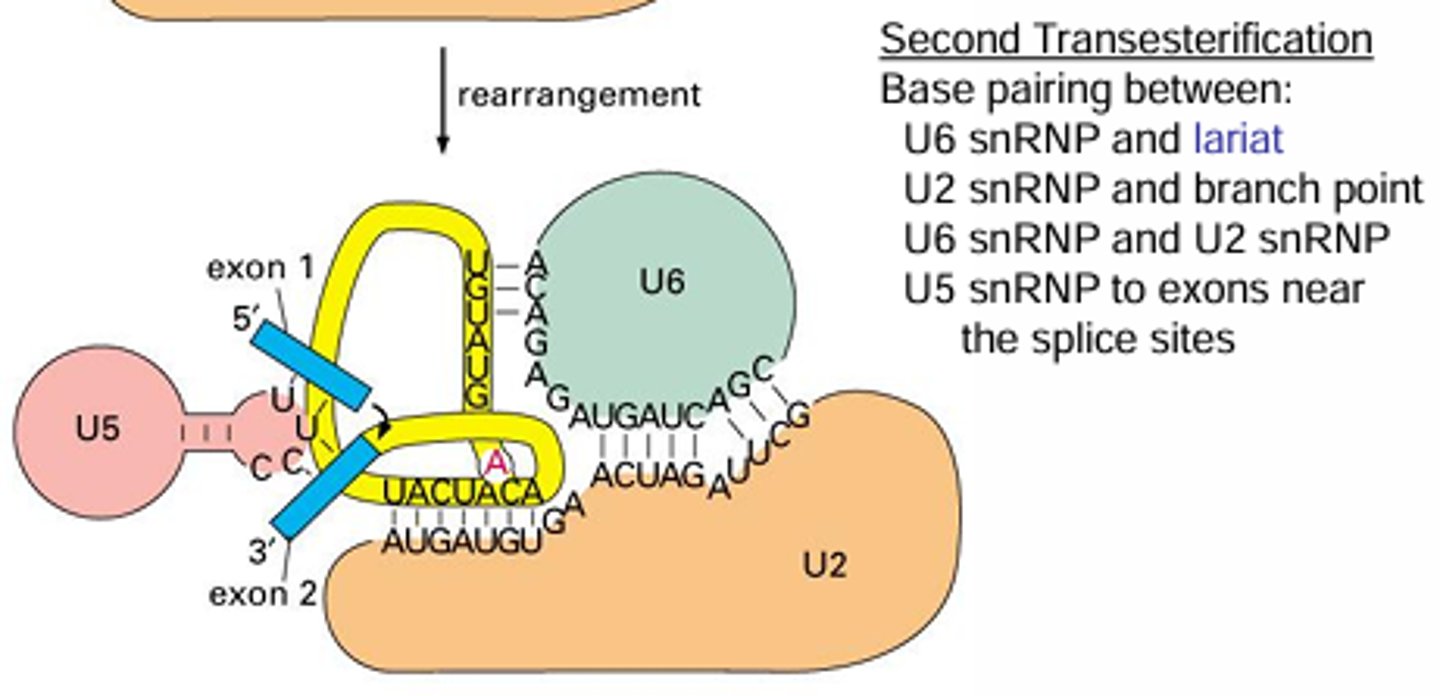
What is alternative splicing and why is it important?
A change in splicing depending on the cell type that results in different proteins being made from the same gene (e.g., fibroblast fibronectin vs hepatocyte fibronectin) - same pre-mRNA but different mature mRNA
This occurs because the snRNPs have different proteins depending on the cell type which will result in different splicing due to the binding sites being slightly different
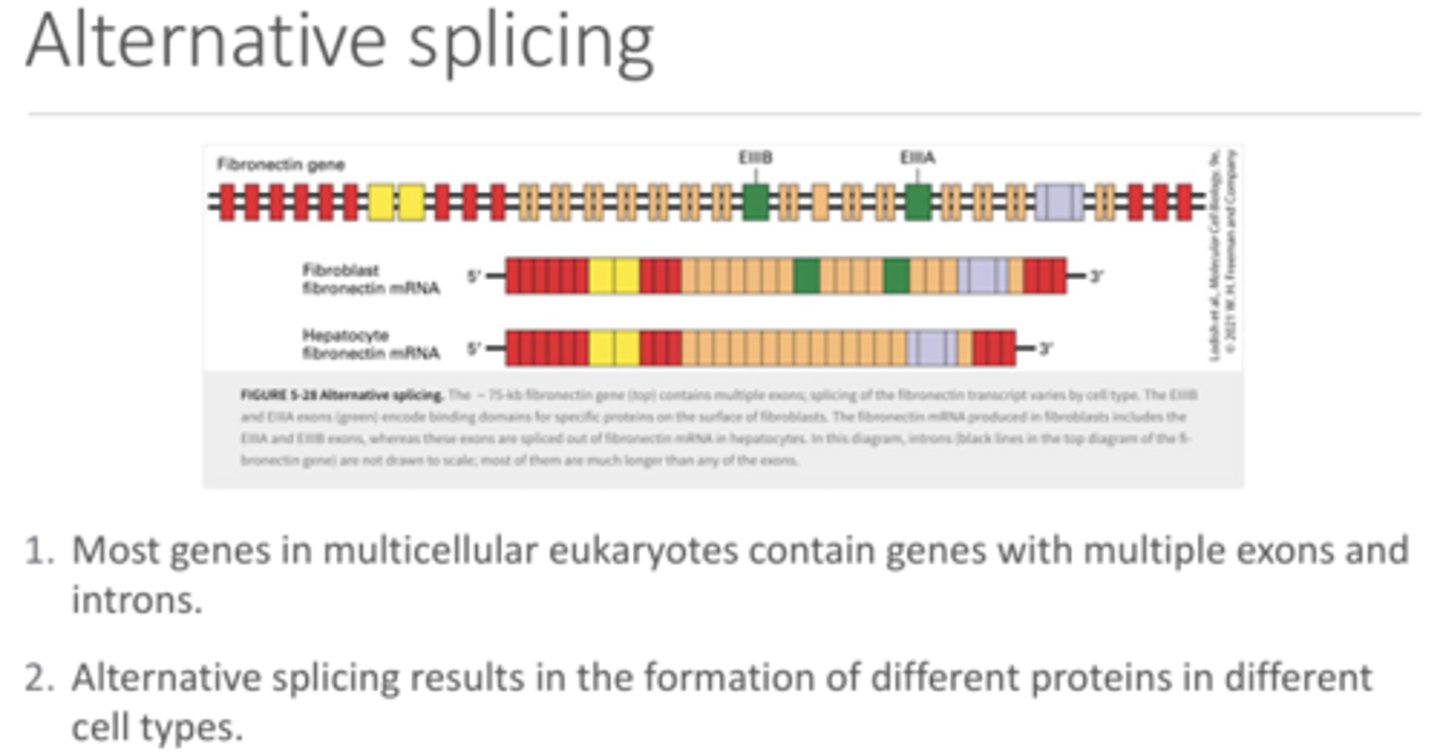
Indicate what nucleic acid(s) would be found at each of the arrows and the significance of the circled region.
A - poly-A tail
B - ssDNA
C - RNA-DNA hybrid
D - ssRNA
Circle - the extra 5' RNA tail that was not hybridized with the DNA due to intro presence remaining in the DNA sequence
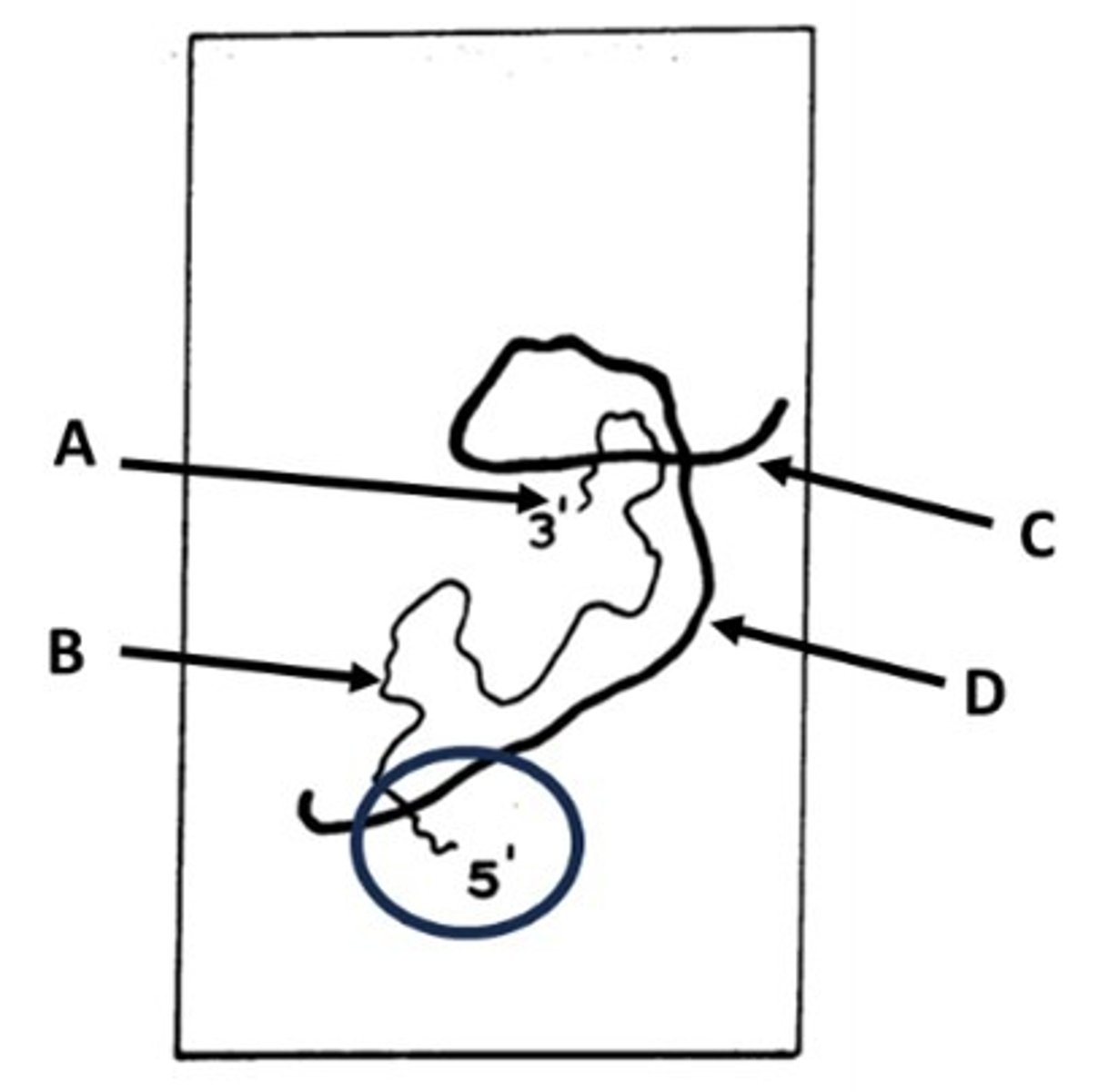
Briefly describe the hybridization photo in relation to the splicing discovery.
Thin line - displaced DNA
Thick line - RNA-DNA hybrid
RNA-DNA binding is stronger than DNA-DNA >> resulting in heteroduplex mapping when the RNA displaces the DNA to hybridize with its complementary sequence. This image proved the existence of splicing due to the 5' RNA tail that was not able to hybridize with the DNA until a much longer strand was used (looping was observed at the intron sites).