Genetics Module 3
1/95
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
96 Terms
loss of function mutations
recessive
gain of function mutations
dominant
effects of point mutations at protein level
missense
nonsense
silent
readthrough
missense
changes amino acid (can have neutral effect)
nonsense
changes codon so that it becomes a stop codon
silent (synonymous)
codes for the same amino acid
eg AGG mutates to CGG but both code for Arginine
Readthrough
stop codon is changed to a codon that codes for amino acid resulting in a longer protein
point mutation example
Sickle cell anemia
one amino acid substiution
Spontaneous Point mutations
Depurination
Deamination of cytosine
Wobble base pairing
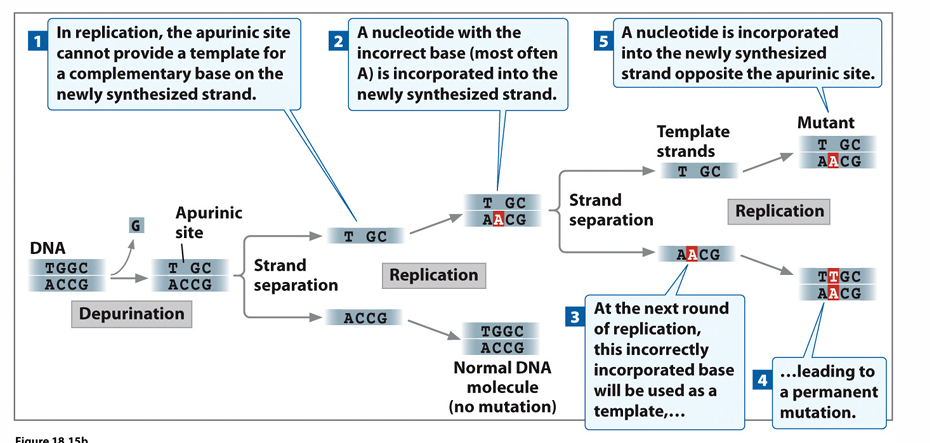
Depurination
Removes glycosidic bond at eithe
r G or A, results in missing purine
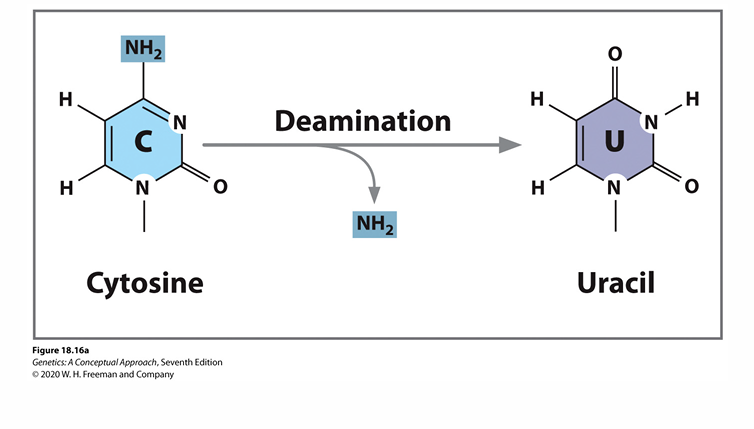
Deamination of cytosine
results in Uracil
causes GC to AT transition
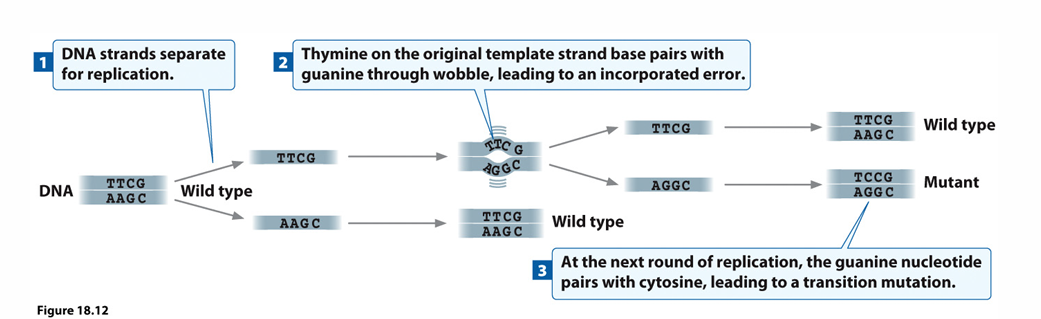
wobble base pairing
mispairing due to flexibility in helix
results in transitions after replication
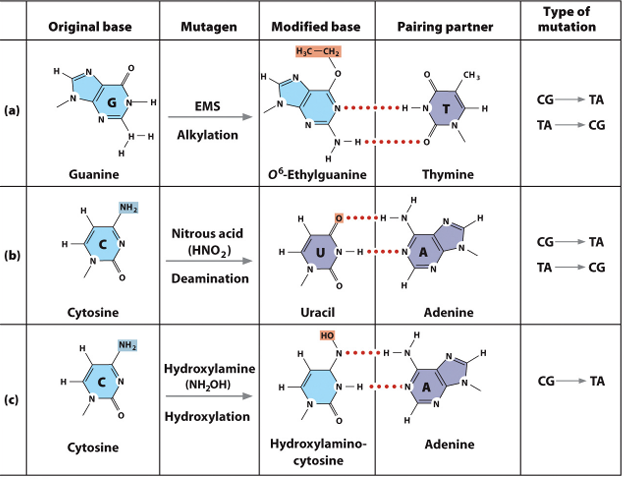
Chemically Induced Mutations
Base Analogs
Alkylating Agents
Deamination
Hydroxylamine
Oxidative Reactions
Intercalating Agents
Base Analogs Cause Transitions After Replication
5-bromouracil normally pairs with A, but can also pair with G.
2-aminopurine normally pairs with T, but can also pair with C
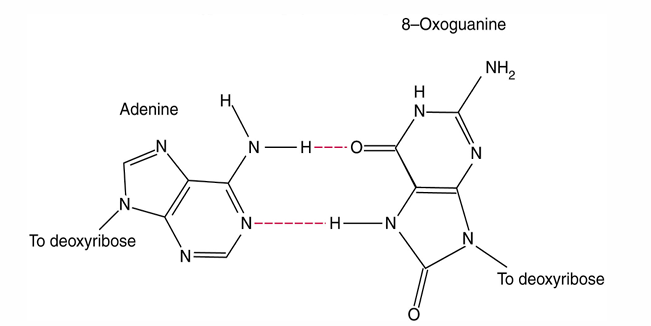
Oxidating Agents Damage DNA
Oxidative reaction converts guanine into 8-oxyguanine.
8-Oxyguanine pairs with A instead of C during replication
Frameshift Mutation example
Cystic Fibrosis
Mutation in a structural protein
Due to a defective CFTR protein- a chloride channel
abnormal salt transport across membranes
Causes of frameshift mutations
Intercalating Agents
Strand Slippage
Unequal Crossing Over
Repeat Regions
Intercalating Agents mutation
Large, planar molecules that slip between base pairs of DNA. This distorts the helix, causing template slippage during replication.
unequal crossing over mutation
Unequal crossing over can cause insertions and deletions
Misalignment of homologous chromosomes during crossing over can lead one product having an insertion and the other having a deletion.
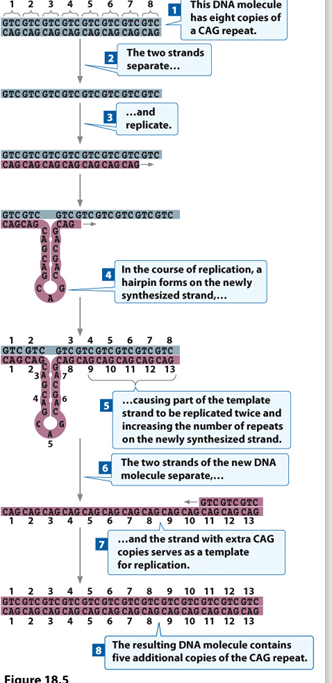
Repeat Regions mutation
repeats may occur via hairpin formation during replication.
direct repair types
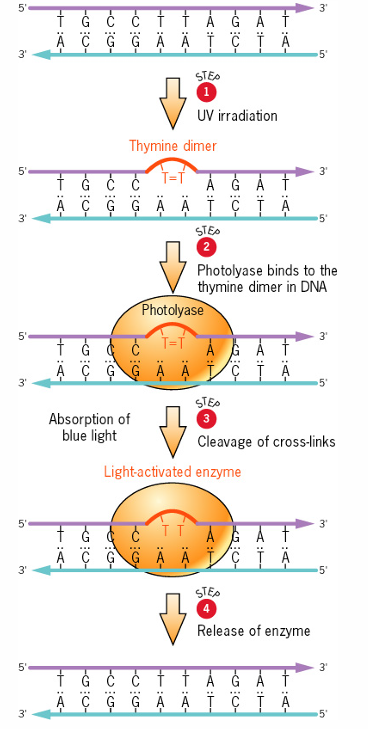
in bacteria- photoreactivation repair of pyrimidine dimers
methyltransferase restores correct form to incorrectly methylated G
mismatch repair
nucleotide excision repair
base excision repair
double strand base repair
direct repair
Corrects structure of abnormal nucleotide without replacing the nucleotide.
mismatch repair
Mismatch repair proteins recognize abnormal helical structure and identify the incorrect base. Exonucleases remove an area of the new strand from the methylated sequence to the mismatch. DNA polymerase fills in the gap and ligase seals the nick. • Does not remove lesions (damaged DNA).
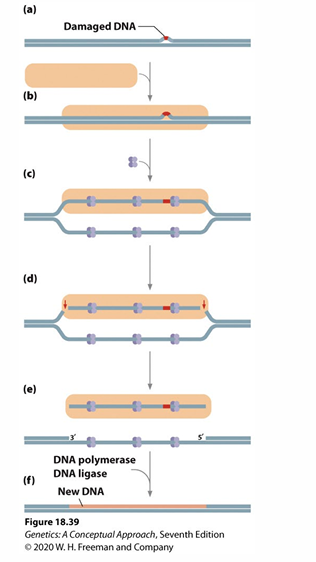
nucleotide excision repair
Removes bulky lesions (damaged DNA)
Enzyme cleaves sugar phosphate bonds on both sides of lesion removing several nucleotides
DNA polymerase fills in gap, DNA ligase seals nick
Can remove lesions unlike MMR.
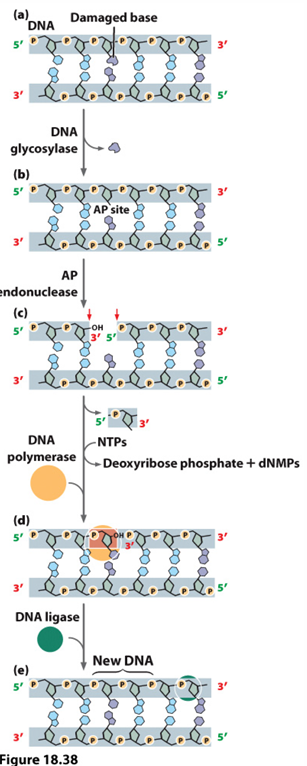
base excision repair
Removes modified bases
Glycosylases bases recognize and remove defective resulting in an AP site
Then AP endonuclease cleaves the phosphodiester bond next to the missing base (causes a nick) and then removes the rest of the nucleotide
DNA polymerase fills in the gap, DNA ligase seals the nick
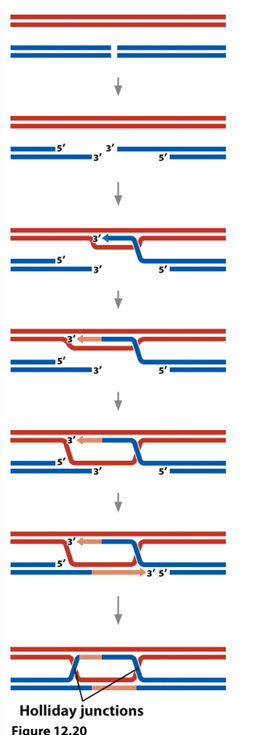
double strand break repair
Homologous Recombination Repair
-Uses the sister chromatid to repair the break
Nonhomologous end joining
-Joins broken ends
-Often leads to translocations, deletions and insertions
translesion DNA polymerases
deletion types
terminal- acentric fragment lost in cell division
interstitial- 2 breaks
cri du chat
missing part of chromosome 5
deletion effects
haploinsufficiency- single copy not enough for wild type phenotype to occur
pseudodominance- expression of normally recessive phenotype because no homologous allele
duplication types

evolutionary significance of inversions
inversions may lead to speciation
translocation types
Reciprocal
Non-Reciprocal
Robertsonian
Reciprocal Translocations
Two nonhomologous chromosomes exchange arms (or parts of arms).
No gain or loss of DNA.
Non-Reciprocal Translocations
A segment from one chromosome is moved to a nonhomologous chromosome.
No gain or loss of DNA.
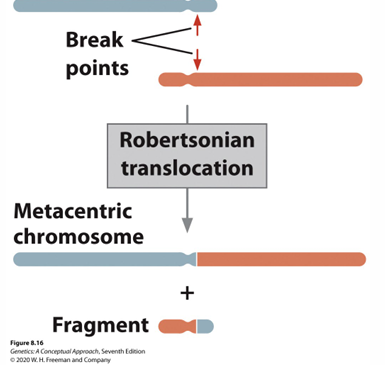
Robertsonian Translocations
• Two telocentric/nearly telocentric chromosomes combine to make one larger, more metacentric chromosome.
• Some small amount of DNA is lost but often not noticeable.
• Isochromosomes – two chromosomes joined are homologs.
Familial Down Syndrome
4% of Down Syndrome cases are familial.
• Chromosome with Robertsonian translocation joining 14 and 21 information
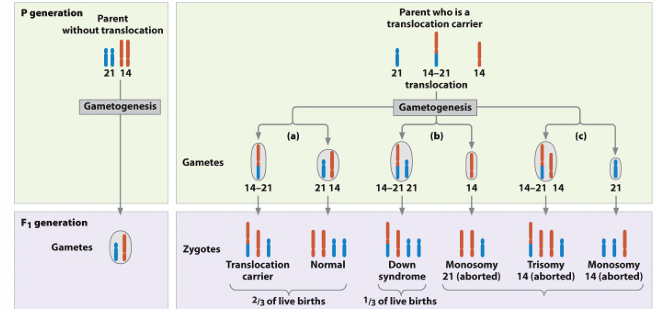
if a person is heterozygous for a reciprocal translocation, what % of their meiotic products will result in nonviable gametes?
50 percent
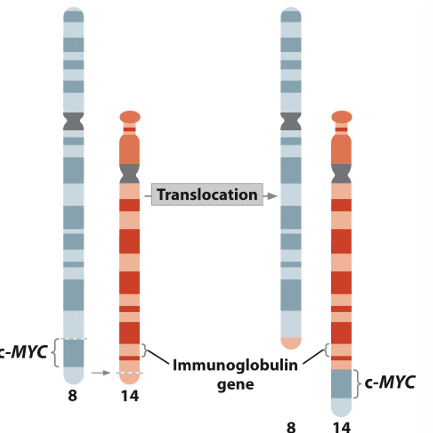
Burkitt’s Lymphoma
Abnormal function of B cells
Reciprocal translocation between chromosomes 8 and 14 places c myc (oncogene that promotes cell division) next to an enhancer which normally stimulates production of antibodies.
Cell division is stimulated in B cells.
Same genes are present, but chromosomal location alters the phenotype
Regulatory Mechanisms for Transcription
rapid turn on and rapid turn off
sequential cascades of gene expression
constitutive expression/housekeeping- always on
lac operon
negative inducible with lactose, positive inducible with cAMP (not glucose)
active repressor binds and turns it off, but when lactose present gene is turned on
I-
repressor cannot bind operator due to repressor’s bad binding site
I^s
super repressor, always on
can’t bind to lactose
lac operon and glucose
high glucose = low cAMP = lac operon is repressed = no transcription
Tryptophan Operon
negative repressible
tryptophan low
no tryptophan so repressor stays inactive, can’t bind to operator so transcription takes place
tryptophan high
tryptophan binds to the repressor and activates it
trp repressor binds to operator and shuts transcription off
Tryptophan Operon Attenuation
Premature termination of transcription
Attenuator – located in the leader sequence and responsible for decreasing transcription when trp is present
When trp high, transcription ends really fast
Puff DNA
loosely coiled and more transcription
how does chromatin change before transcription
decondenses to 11nm fibers beforehand
DNasel
chews up loose DNA
histone modification
acetylation and methylation
what does acetylation do
neutralizes positive charge on histones which loosens DNA and turns on gene
what does deacetylation do
turns off gene
what does DNA methylation do
decreases transcription
changes in histone modification and flowering
FLC gene suppresses flowering
Acetylation of FLC gene turns it on which prevents flowering
Deacetylation of FLC gene turns it off which allows flowering
epigenetic methylation of bees
female bees that eat royal jelly become queens
royal jelly suppresses Dnmt3
Dnmt3 adds methyl groups to DNA
Bees w/ suppressed DNmt3 have less methylation and genes that are normally silenced are expressed
Primary Immune response
Secondary immune response
Antibody structure
2 chains heavy and light
-each has a constant and variable region
Antigen binds to the variable region
antibody diversity- how are millions made
somatic recombination in B cells during B cell differentiation
alternative sites for recombination
somatic mutations
development
regulated growth resulting from interaction of genome, cytoplasm, and environment
programmed sequence of events
not reversible
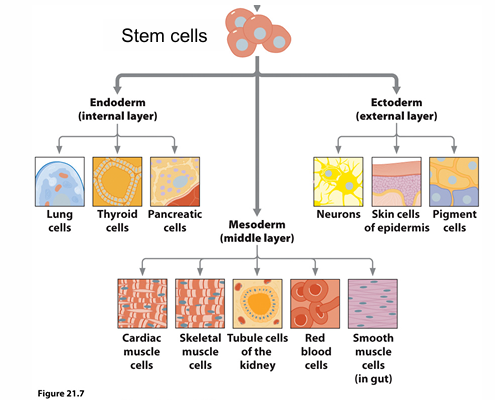
differentiation
aspect of development
forming different types of cells, organs through specific regulation of gene expression
dorsal
up
ventral
down
Maternal drosophila genes
egg polarity genes- establish anterior/posterior polarity and dorsal/ventral polarity
When are Maternal drosophila genes transcribed?
during egg development
When are Maternal drosophila genes translated?
after fertilization
drosophila segmentation genes
affect number and polarity of segments
-gap genes
-pair rule genes
-segment polarity genes
homeotic genes
determine identity of each segment
bicoid maternal gene
bicoid mRNA anchored at ANTERIOR end
nanos maternal gene
nanos mRNA anchored at POSTERIOR end
egg from bcd+
normal larva
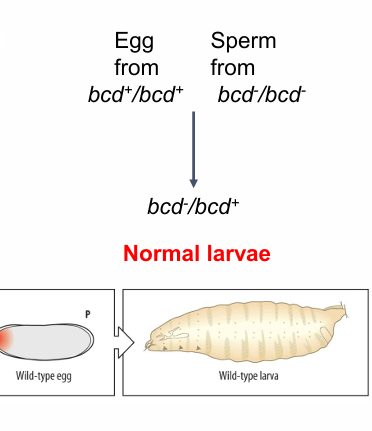
egg from bcd-
no anterior- headless
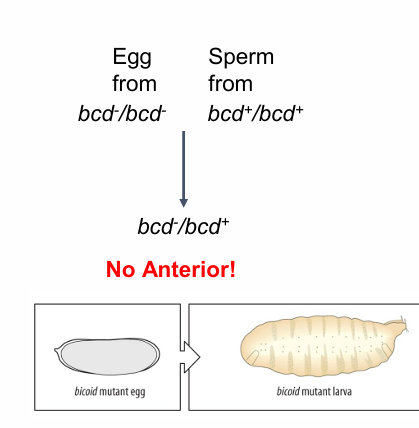
where is bicoid protein highest?
In anterior
Where is nanos protein highest?
In posterior
bicoid and caudal
bicoid represses translation of caudal mRNA
affects POSTERIOR
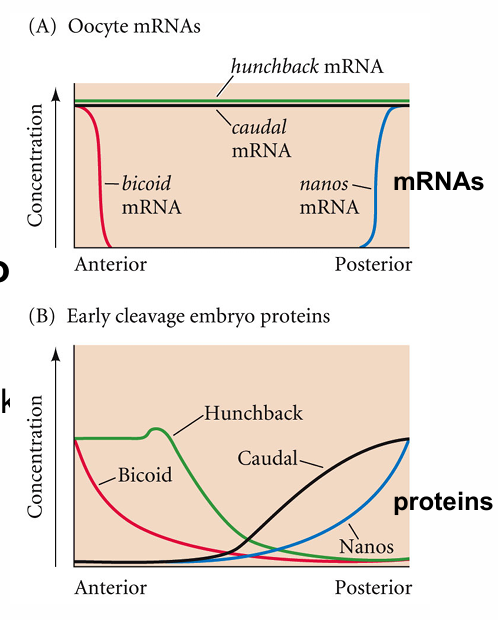
bicoid and hunchback
bicoid stimulates hunchback expression
in ANTERIOR
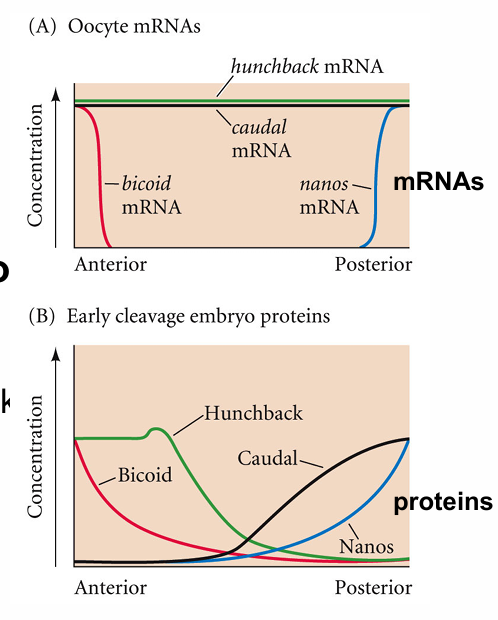
Nanos and hunchback
Nanos inhibits hunchback translation in ANTERIOR
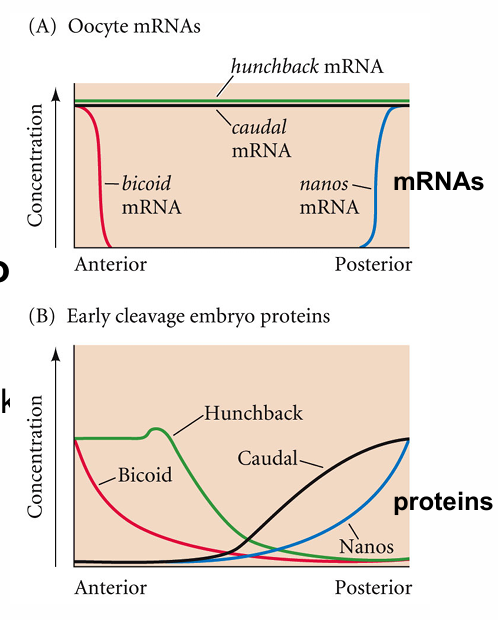
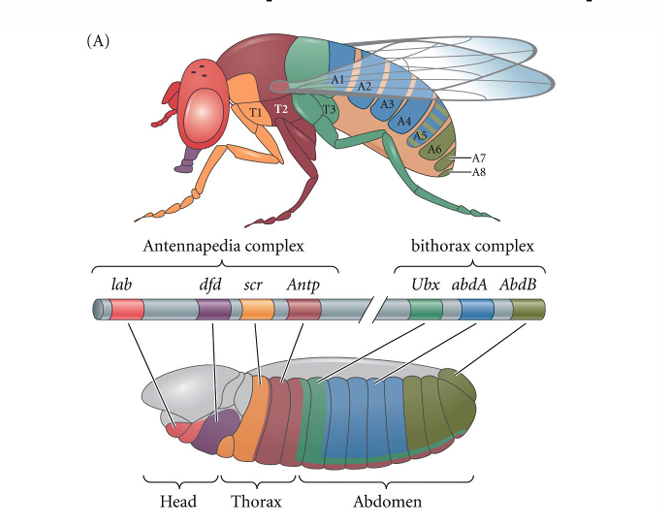
homeotic genes
give specific identity to each segment
genes in order from anterior to posterior
each gene is on only in specific segments based on concentration of earlier gene products
homeotic drosophila mutation example
Ubx green in T3
When deleted, Antp red extends into T3 causing extra wings
homeotic homeoboxes
Region of the protein formed from the homeobox DNA is called homeodomain
Proteins containing homeodomain are DNA binding proteins
Homeodomain binds to specific DNA sequences and regulates transcription
apoptosis
removal of tissue between fingers
creation of joints
neural pruning
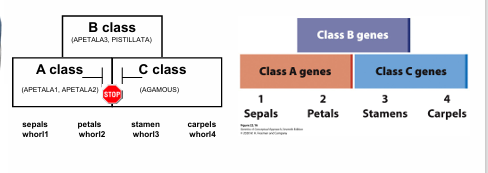
plant homeotic genes
Class A= whorl 1 sepal and whorl 2 petal
Class B= whorl 2 petal and whorl 3 stamen
Class C= whorl 3 stamen and whorl 4 carpel
normal chromosomes
2n=8
nullisomic
2n-2=6
The correct sequence of segmentation gene action in the development of the Drosophila embryo is
Gap
Pair rule
Segment polarity
whorl 1
sepal
whorl 2
petal
whorl 3
stamen
whorl 4
carpel
gap genes
divide embryo into broad segment
are a type of segmentation gene
eurkaryotic regulation
Changes in Chromatin
initiation of transcription
RNA processing and stability
Protein modification
In eukaryotes, repressors can function by:
A plant species is described as 2n=20.
How many chromosomes present in a triploid cell from this plant?
n=10
nx3= 30
In eukaryotes, repressors can function by: