molecular biology - test
1/14
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
15 Terms
why is phenotype expression from a genotype, not clear-cut
give an example
there may be mutations to other parts of the gene expression process (the gene may be unmutated itself, but the machinery creating the protein may be mutated, changing how the gene is expressed in phenotype)
but even with these mutations, the phenotype cannot be predicted - they are found to increase the probability of a phenotype, but not make it certain
other factors, influences, and outliers still exist - random chance, environmental factors
e.g. hair colour, MC1R genotype variant (mutation in the receptor of melanin production), is thought to give red hair, but not all red hairs have this variant (loss of function mutation to stop melanin), and some non red hairs have this too
wheras, loss of function in mice causes red hair
describe epigentics
how does this work
(description) heritable changes in gene activity / expression, that DO NOT involve changes to the DNA sequence
(how?) commonly due to methylation of DNA (acting as a marker), which is added near a gene to slow down RNA polymerase by slowing the binding of transcription factors
this results in the gene expressed less, therefore acting as a genetic switch - the gene is still included within the genome, it just isnt expressed
this allows cells to maintain their identity within tissues and through successive cell divisons - they remain in DNA replication
this is heritable, some epigentic markers are cleared in fertilisation, while some remain
what are transcription factors
what is their function
proteins that bind to promotor regions on DNA (TATA box, initiation sequences before the gene starts, to signfiy a gene)
this allows RNA polymerase to bind to them and the promotor, to allow it to begin transcription (creating mRNA so that the gene sequence can be used to code a protein)
therefore they control what genes are expressed, and what proteins are created, in a cell - based on cell type, cell location, and extracellular stimuli
this is because, if they dont bind to a promoter region, RNA polymerase then cannot bind to the DNA to begin transcription, and the protein cannot be created (the gene cannot be expressed)
this ensures that not all cells express all genes within the DNA, instead only those required for their function within the organism, overall and at that particular time
name the 3 general types of DNA sequencing
sanger sequencing (e.g. PCR)
next generation sequencing (e.g. Illumina)
nanopore sequencing
explain the process of Sanger Sequencing
(denaturing) the DNA sample of interest is denatured, seperating it into 2 individual strands
(annealing) a specific primer designed to be complementary to the gene region of interest, is added and binds to the start of the region
(elongation) polymerase and nucleotides are added, allowing the DNA strand to be replicated from the added primer
most of these nucleotides are normal, but some have a dye / fluroescence attached (=dideoxynucleotides), coloured depending on the base
this dideoxynucleotide acts as a chain terminator, as they have had a hydroxy group removed, so the polymerase cannot add anymore nucleotides to the DNA backbone afterwards
this creates numerous varying lengths of DNA fragments, stopping on different bases, which are seperated on a gel, where an electric current is applied, causing shorter fragments to travel further across the gel to the positive end (DNA is negative)
a laser is applied, which excites the dye, and as light passes through it produces a wavelength onto a light sensor, which detects and translates it to the appropriate base and computes it
with enough fragments stopping at different bases, enough are computed to produce the DNA sequence
explain the process of Next Generation Sequencing (NGS)
similar to PCR (Sanger), is PCR-based, but with new features and aspects for efficiency
(Fragmentation) sample DNA of interest is broken into fragments via bacterial or physical methods
(Addition) primer binding sites (to allow the generic primer to add, so we dont need to know the exact DNA seq) & capture sequences (to allow immobilisation on gel) are added to the sample DNA
(Denaturation) the DNA sample is denatured, seperating the strand into two
these next steps are automated by the machine (e.g. Illumina)
(Immobilisation) short pieces of DNA to match the added capture sequences, are attached to a surface (flow cell), so when the sample is added, these bind to the DNA
(Amplification) PCR cycles (annealing → elongation → denaturation) are repeated, forming the complementary DNA strand (replication), then washing away the other piece (via denaturation)
these replicants anneal to other capture sites on the flow cell, forming bridges that replication occurs over (as they are the compliments) - then denature to seperate from the bridge
after numerous cycles, one type of capture sequence (Added at one end of the base strands) are cleaved, so one pair of replicant remains captured, then the complementary primer site is added to the free end
(Sequencing) fluroescent dye nucleotides are added, which bind to the added primer (on one type of replicant pair strand, amplified, that remains immobilised)
another chemical can actually add the OH group back, so these nucleotides are NOT chain terminating, and the chain can continue to replicate using these dye nucleotides
this allows the base to be determined at each position of the sequence, as lazer is passed through (4 waves of light) and thoes absorbed are measured, computing the base at each position sequentially, along the entire strand
this is done with millions of fragments (amplified), so sequencing is done accurately and quickly (machine automated)
name the 6 steps of Next Generation Sequencing
DNA Fragmentation
Addition of primer binding sites & capture sequences
DNA Denaturation
DNA Immobilisation
DNA Amplification
DNA Sequencing
=Fadias
what are the limitations of NGS (next generation sequencing)?
millions & billions of fragments are read simultaneously in the machine, of their DNA sequences based on the dye nucleotides added, which requires lots of computational power to put it back together into the actual genome (the fragments of sample DNA)
also provides lots and lots of data, so has storage and processing of lots of data
what is a type of Third Generation Sequencing?
how does this work?
(nanopore sequencing)
a nanopore (protein complex w/ a pore) is inserted into a synthetically created membrane, and a potential is applied
when molecules pass through the pore & membrane, the potential is disrupted
(process)
a specific helicase is used, designed to bind onto the nanpore, which unwinds the approaching DNA, so it passes through single-stranded
as each base passes through, the membrane potential is altered slightly, which is computed, and can be differentiated
this allows real-time reading of the DNA base sequence
what are the advantages & disadvantages of Third Generation Sequencing?
(advantages)
real-time output of DNA base sequence
doesnt require primers (not PCR based)
is contained in a portable device, so can be used on the field to easily and quickly access genetic info (vs large machines of PCR & next gen sequencing)
can read a whole piece of DNA (access the whole genome) rather than being limited to fragments it must patch together which is often difficult
(disadvantages)
lower accuracy as it just relies on the reading of a single strand of DNA rather than repeated fragments
costly
name the 3 main ways of determining protein structure
x-ray crystallography
NMR spectroscopy
cryo EM (electron microscopy)
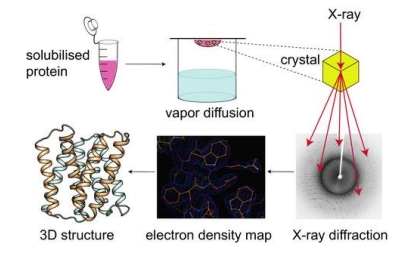
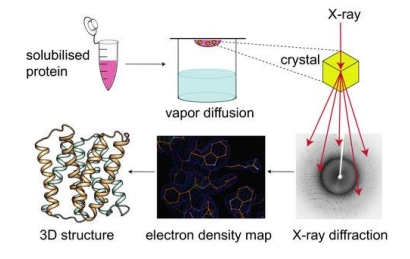
how is xray crystallography used to determine protein structure
what is the main advantage & limitation
the original method used to solve the first structures, still a main method
(method)
firstly grow a crystal (solid structure in an organised lattice) of the protein of interest
then pass xray through this crystal, and measure diffraction
diffraction is then computed to determine the refraction image (where xray was unable to pass through, therefore protein must be here), and therefore the crystal structure
(advantages)
using xray is beneficial because it has small enough wavelengths to pass between protein atoms (vs visible light wavelengths which are longer than the distance between atoms)
this gives it a high resolution (can detect and compute, so therefore see in the structure, individual atoms)
(limitation)
must use protein crystals, as the xray signal is too weak to create refraction when we pass it through a protein solution
however, these crystals are hard to grow, as proteins resist forming this highly ordered structure, and may clump / form inaccurate structures (must change pH etc)
furthermore, some proteins may not be able to crystallise at all
how is NMR spectroscopy used to determine protein structure?
what are its advantages & limitations?
(method)
uses protein in solution
reads the changes in magnetic field of the atoms in the molecule, which change depending on the spin of the electrons, but also the atoms in the chemical environment
this creates a slightly different chemical environment for each atom in the molecule, which is computed with a graph with points at each atom and their changes in magnetic field
this provides information on the structure of the molecule, based on how known atoms together, create certain changes in chemical environments
(advantages)
uses proteins in solution, removing the struggle of crystallising proteins (xray crystallography main method), and shows the protein in their natural environment (rather than a static crystal)
(disadvantages)
is not high resolution, is best for small proteins as larger ones provide too much data which is difficult to compute and work out the structure from
how is Cryo EM used to determine protein structure?
what are the advantages & limitations?
(method)
alphafold