BIOC 202 Midterm 3/final
1/53
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
54 Terms
describe the overall rxn of pyruvate dehydrogenase complex (PDC)
conversion of pyruvate to acetyl COA is catalysed by PDC and occurs in the mitochondira matrix
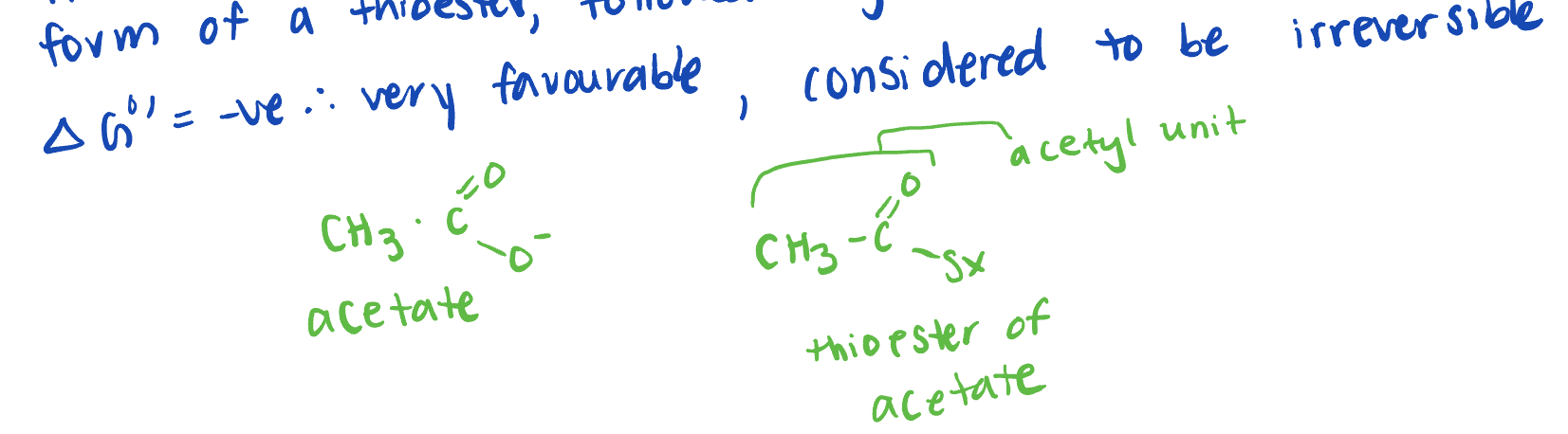
invovles a decarboxylation/oxidiation of pyruvate to acetate in the form of a thioester, followed by the formation of acetyl COA
Pyruvate + NAD+ + COA-SH → Acteyl COA + NADH + CO2 + H+
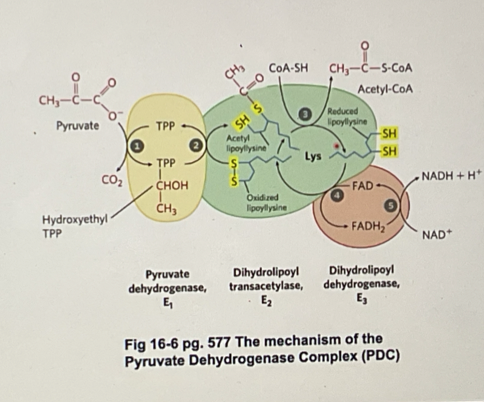
what enzymes is PDC composed of?
E1: pyruvate dehydrogen
E2: dihydrolopyl transacetylase
E3: Dihydrolopal dehydronase
what are the 5 co-factors that compose PDC? and what enzyme subunit are they binded to?
Thiamine pyrophosphate (TPP) → bound to E1
Lipoamine → bound to E2
NAD+ → bound to E3
COA-SH (coenzyme A)→ free
describe coenyme A /COA, COA-SH in PDC
nothing bound
composed of ADP, pantothenate (vit. B5) and B-mercaptoethyl amine
carrier group of acyl groups
form higher energy thioester bonds
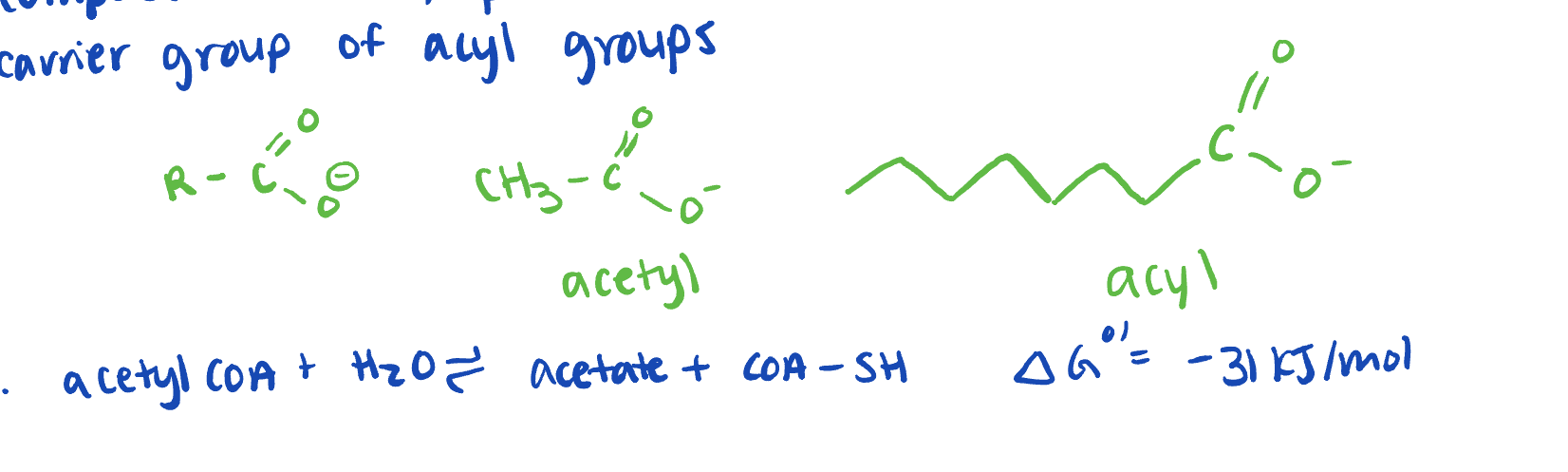
describe thiamine pyrophosphate (TPP) in PDC *** do we have to recongize the structure?
derived from Vit. B1
forms a reactive carbonion easily
carries aldehydes
promotes decarboxylation
describe lipoic acid (lipoamine)
lipoicacid (black) can be attached to alysine on E2 forming a lipoamide
Lipoamide oxidises aldehyde to acyl groups resulting in the acyl group being bound to the disulfide group
acting as a “robotic arm”
Describe the enzyme mechanism of PDC
pyruvate enters E1, binds to TPP and is decarboxylated (forming O2) to form the intermediate hydroyethl-TPP
Lipoamide arm (oxidised) enter E1
The hydrozyethyl group is oxidsed to an acteyl group and is bound to the now reduced lipoamide arm (which is now called dihydrolipoyl group)
the reduced arm carrying the acetyl unit moves in E2 and the acetyl group is transerred to COA, forming acetyl COA (acetyl COA leaces the enzyme) is transferred to COA, forming acetyl COA (acetyl COA leaves the enzyme)
the reduced dihydrolopyl lipoamide arm moves into E3 where it is oxidsed by FAD (FAD is reduced to FADH2)
NAD+ enters E3 and reoxidises FADH2 back to the FAD (NAD+ is reduced to NADH, which leaves E# and now back at step 1)
**note: there are many copies of E1, E2, E3 enzymes and cofactors in the complex
Describe Regulation of PDC? what inhibits/activates E1, 2, 3? what enzyme?
increase of [acetyl COA] allosterically inhibit E2
increase o [NADH] allosterically inhibit E3
BUT the main control is on E1, where the phosphorylation of a serine by a kinase leads to the inhibition of E2 and this the entire complex
this kinase is called: PDC associated kinase
Acetyl COA, NADH + ATP all stimulate the kinase, causing PDC to slow
Pyruvate, NAD+, + ADP all inhibit the kinase resulting in the PDC gradually becoming active again
general phosphatases will gradually (i.e. slowly) dephosphorphorylate E1, resulting in it becoming more active again
cell signalling such as incerase in [Ca2+] +insulin activate PDC assoicated phosphatase (PDCAP) which rapidly dephosphorylates E!, leading to a rapid decrease in activity
describe kreb’s cycle
aka the citric acid cycle, the bicarboxylic acid cycle (TCA)
is the metabolic hub of the cell
it completely oxidizes acetyl COA to CO2 and in the process, generates high energy e- ) in the form of NADH and FADH2) and GTP
these e- are used in oxidative phosphorylation to generate ATP
the kreb’s cycle is also a source of many biological precursores
occurs in the mitochondrial matrix
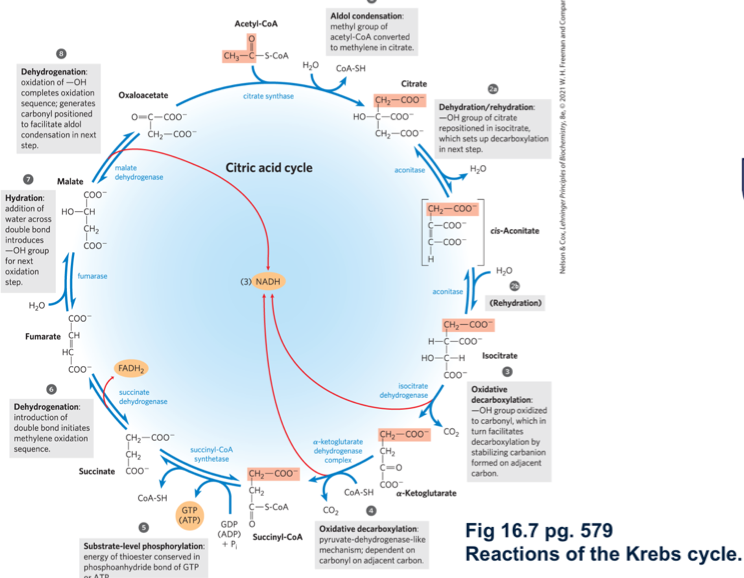
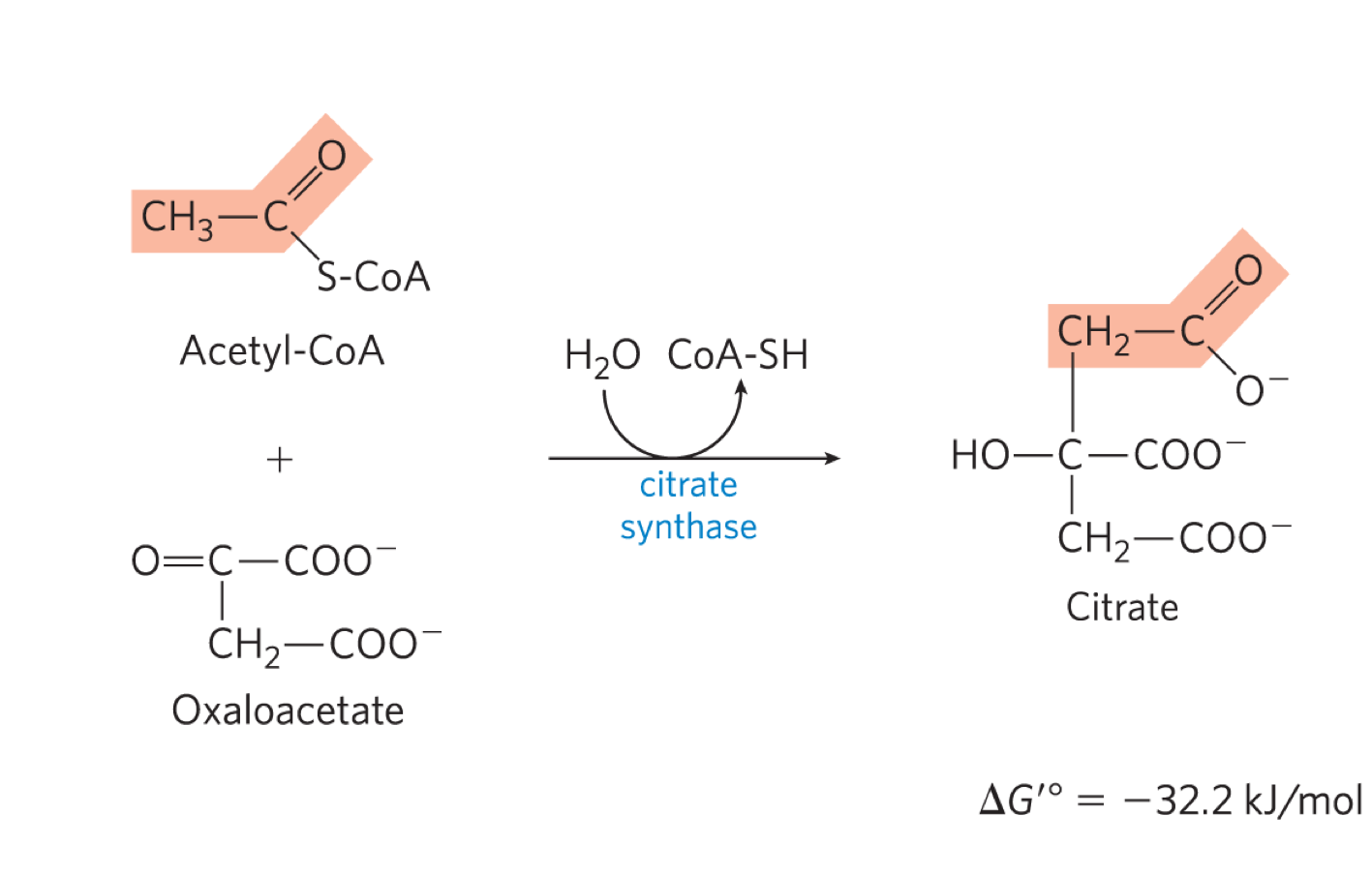
describe rxn 1 of kreb’s cycle
citrate synthase forms citrate by binding oxaloacetate to acetyl COA (going from C4 to C3)
2 parts:
1. aldol condensation to form citryl COA
hydrolysis of ciryl COA to citrate and COA-SH
this rxn drives the entire rxn forward
irreverisble
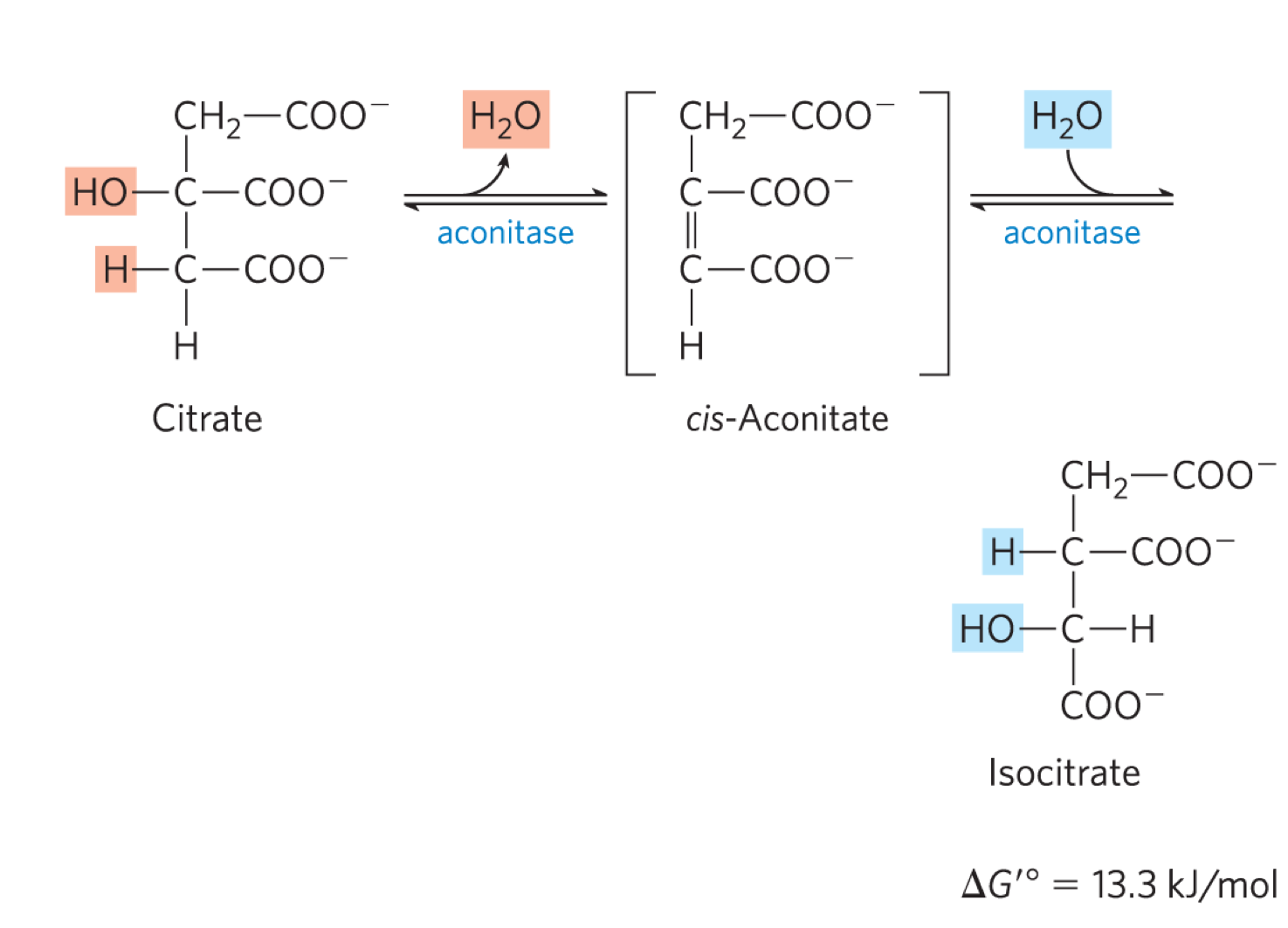
describe rxn 2 of kreb’s cycle
aconitase converts citrate to isocitrate (moving the OH group)
dehydration step to form cis-aconitate, followed by hydration step to generate isocitrate
the delta G knot is +ve but it is drive by the -ve delta G knot of rxn 1 and rxn 2
the OH group is moved onto the CH2 that came from oxaloactetate
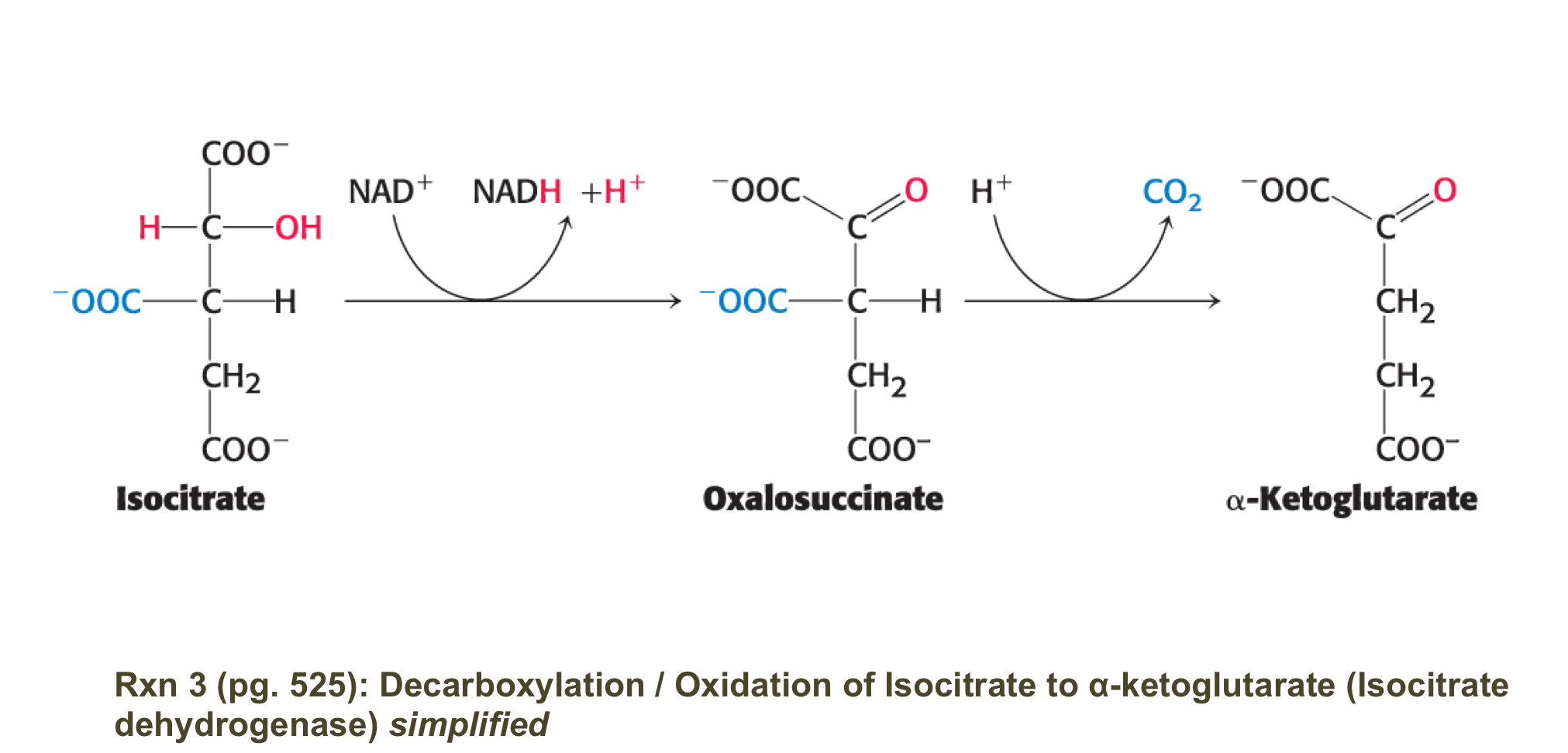
describe rxn 3 of kreb’s cycle
isocitrate is oxidised/decarboyxlated to alpha-ketoglutarate by isocitrate dehydrogenase
generates CO2 + NADDH
irrverible rxn
occurs in 2 steps:
1. isocitrate is oixdized to oxalosuccinate generating NADH
2. Oxalosuccinate is decarboxylated to alpha-ketoglutorate (spontaneously)
note: the CO2 did not orginiate form the acetyl COA that just entered the cycle
synthase vs synthetase?
synthase: an enzyme catalyzing a synthetic in which 2 units are joined without the direct participation of ATP
synthetase: same as above BUT ATP (NTP) is directly required
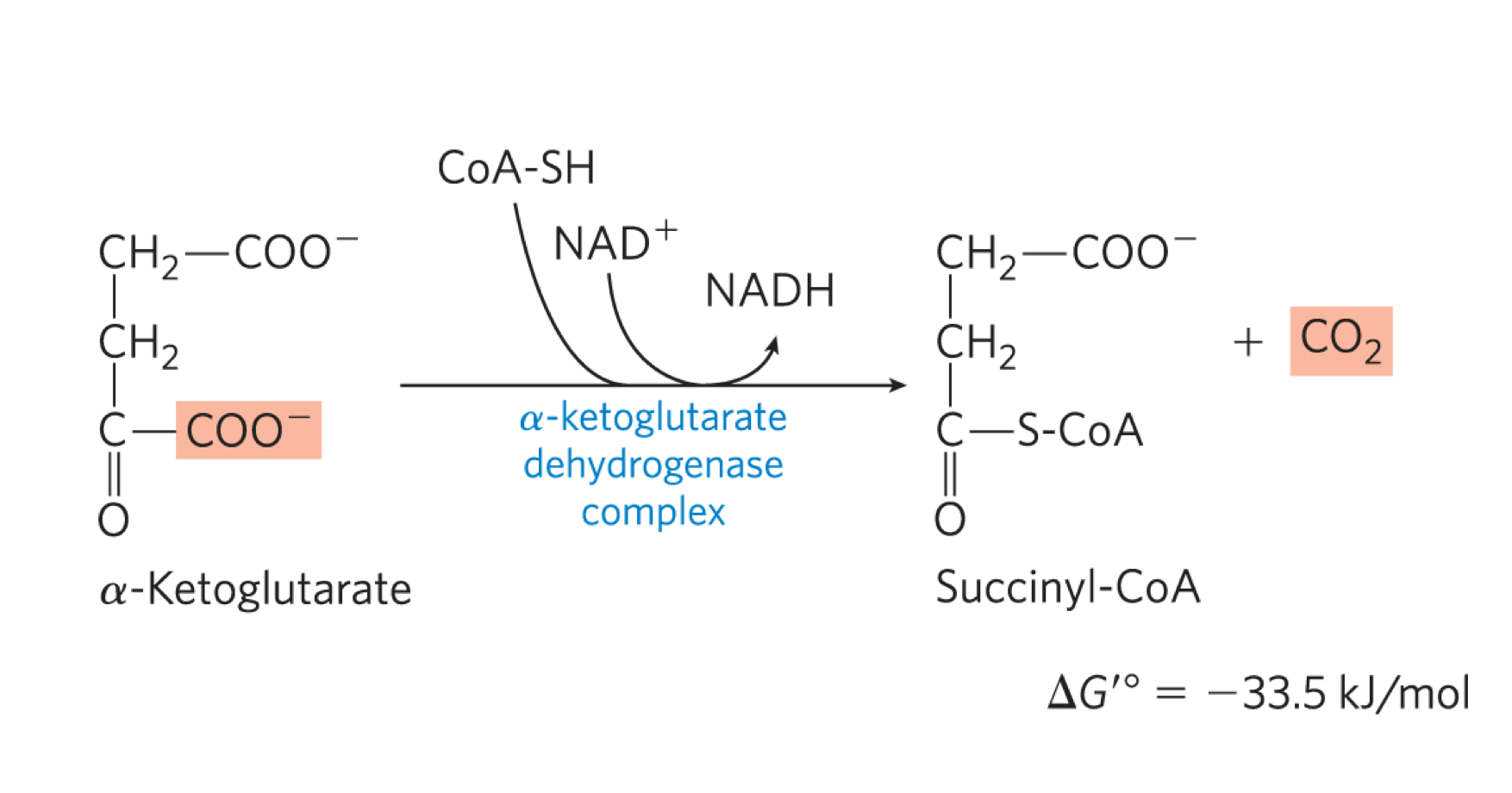
describe rxn 4 of kreb’s cycle
alpha-ketoglutarate is decarboxylated.oxidised and bound to COA by the alpha-ketoglutrate dehydrogenase complex (alpha-KGDH), generating succinyl COA, CO2 + NADH
occurs by the same method as PDC (i.e. same 5 co-factors, similar E1 + E2, identical E3)
back to 4C
irreverisble rxn
high -ve delta G knot
describe rxn 5 of kreb’s cycle
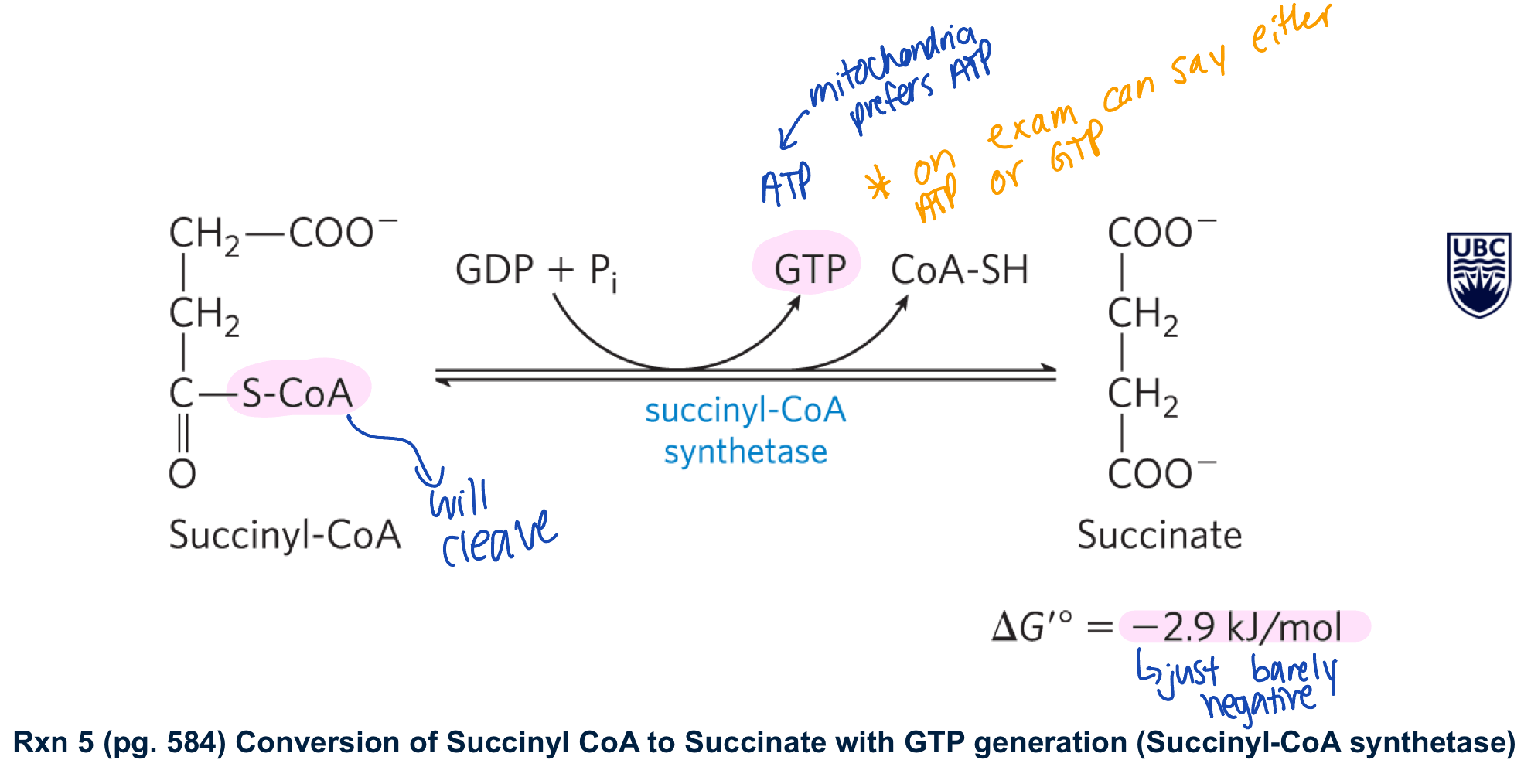
succinyl COA synthetase converts succinyl COA to succinate generate GTP/ATP and COA-SH
this rxn is driven by the -ve delta G knot in the cleavage of the thioester bond
**notes: -GTP can be converted to ATP by a nucleoside diphosphate kinase
-there are isoforms of succinyl COA synthetase that use ADP + generate ATP
reverible rxn
the next steps invovled in the regenrationg of oxaloacetate
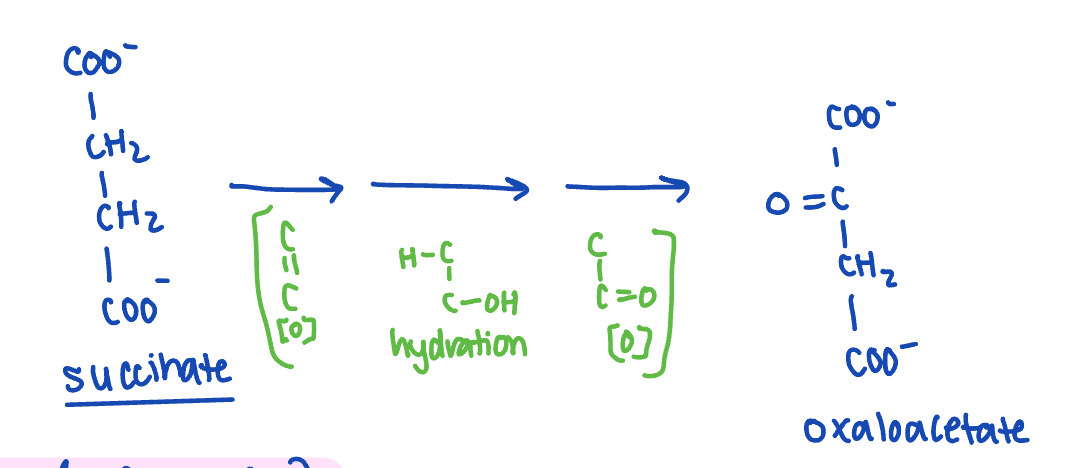
describe rxn 6 of kreb’s cycle
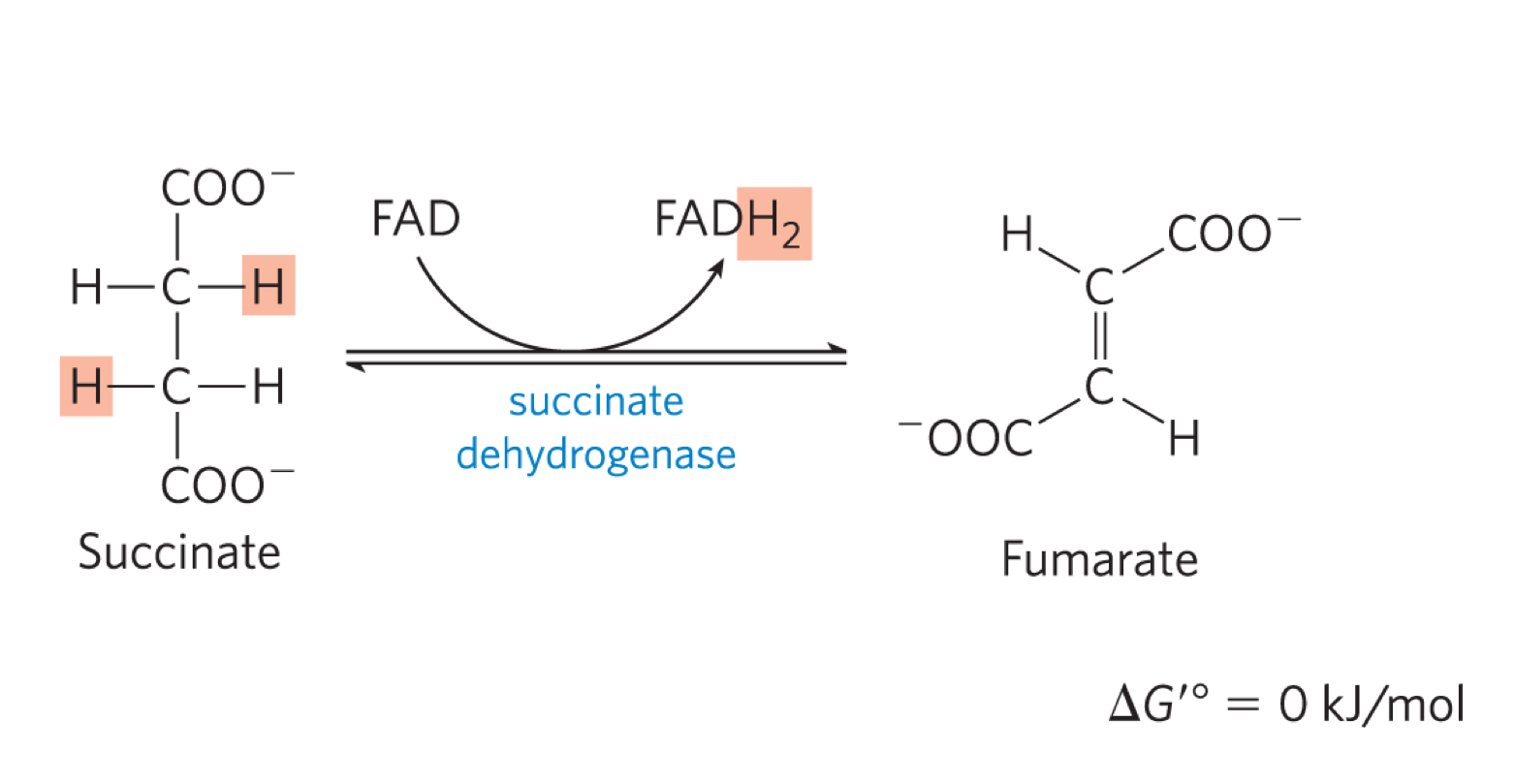
succinate dehydrogenase oxidises succinate, generating FADH2 and fumerate (transt)
the free energy is not high enough to reduce NAD+
succinate dehydronganse is part of complex II
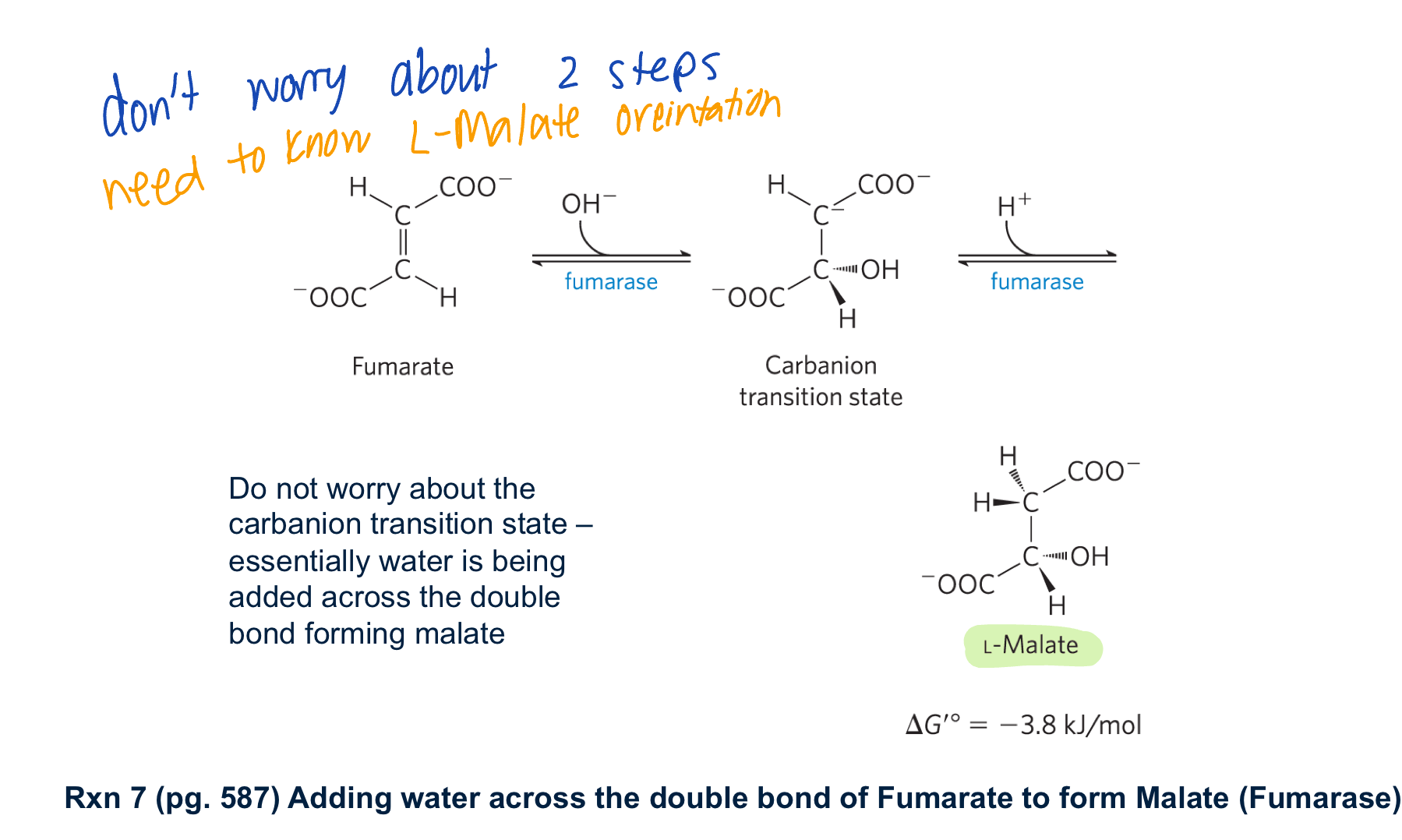
describe rxn 7 of kreb’s cycle
fumerase adds water across the double bond forming L-malate
describe rxn 8 of kreb’s cycle
malate dehydrogenase oxidses malate to oxaloacetate generating NADH
large +ve delta G knot but is driven by the 3 irreverible rxns
also means that the matrix kep low oxaloacetate to keep the rxn moving forward
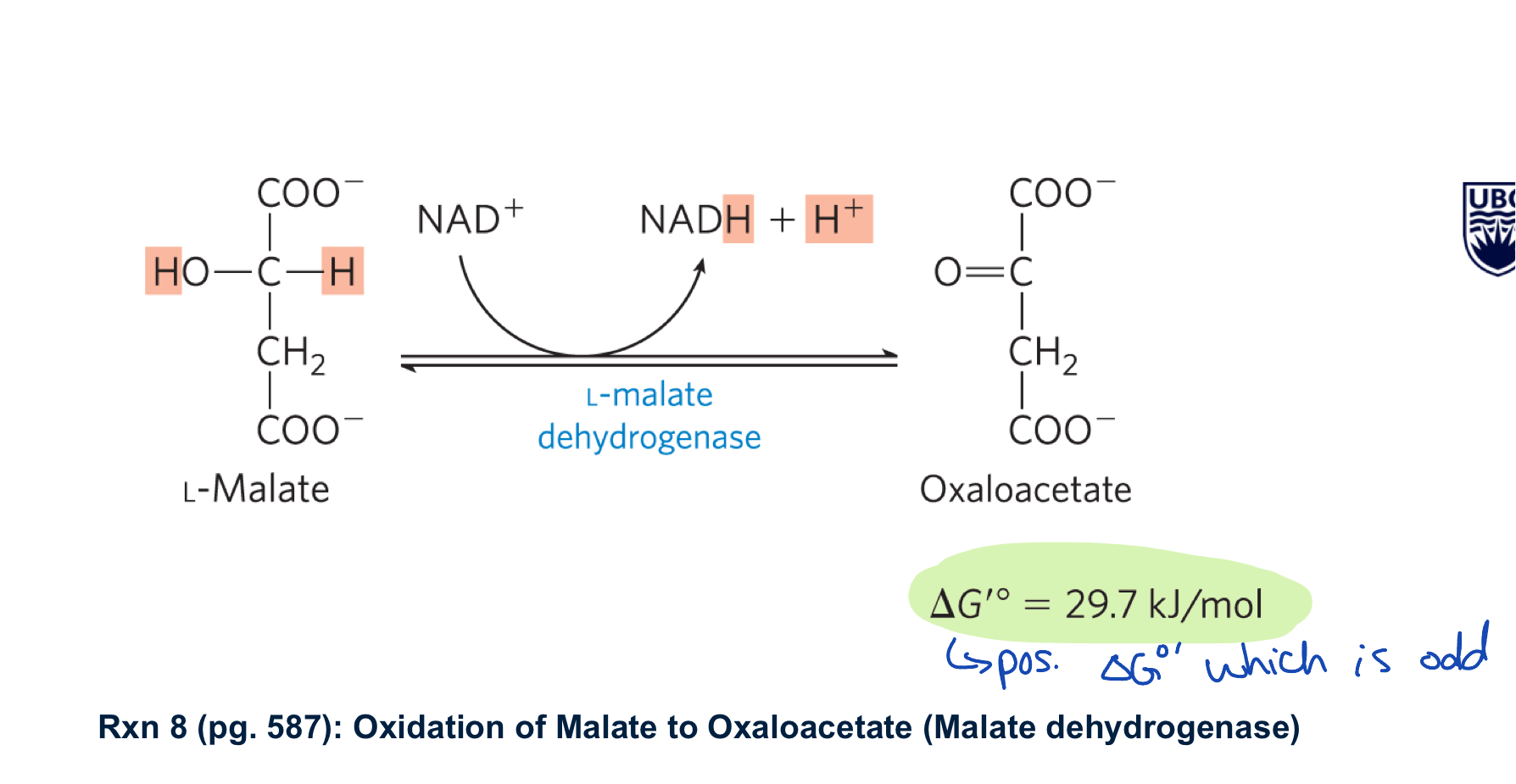
what is the next rxn of kreb’s cycle?
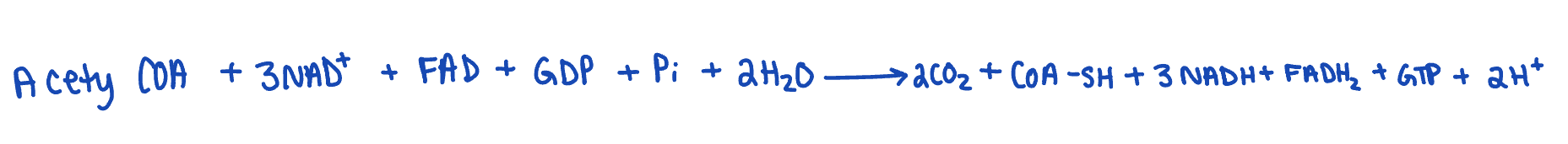
describe regulation of kreb’s cycle
all allosteric:
isocitrate dehydrogenase
activiated by ADP
inhibited by ATP and NADH
alpha-ketogluterate dehydrogenase
inhibited by ATP, NADH, and succinyl COA
optional* citrate synthase (only in bacterica)
inhibited by ATP
what is oxidative phosphorlyation?
the formation of ATP as a result of the transfer of e- from NADH + FADH2 to O2 by electron carriers
what is electron motive force? how can it be harnessed? what does it form?
the e- attached to NADH + FADH2 to have high transfer potential (aka EMF)
this EMF can be harnessed by the electron transport chain (ETC) to transfer protons (H+) out of the mitochondrial matrix, through the inner mitochondrial membrane (IMM) and into the intermembrane space (IMS)
the resulting electrochemical gradient forms a proton motive force (PMF)
PMF can be used by ATP syntase to form ATP (a chemical with high transfer potential)
mitochondrial composition
the ETC and ATP synthase is located on the IMM
the IMM is impermeable to small molecules and ions
the OMM is permeable to small molecules and ions
the IMM requires transporters to move things across it
the OMM has many pores and is considered leaky
hence, the IMS is similar to the cytosol (often referred to as the cytosol)
IMM: inner membrane of mitochondria
OMM: outer membrane of mitochondria
how can electrons be transferred?
electrons can be transferred as:
free e-
H+ atoms
hydride ions (H:-)
standard reductive potential
different molecules can have different tendencies to accept e-
this can be measured as the standard reductive potential (Eo’) in volts (v)
the more positive the Eo’ the higher the molecules affinity for e-
what is the half rxn and coupled rxn of NAD+ and O2
NADH + H+ → NAD+ + 2H+ + 2e-
½ O2 + 2e- +2H+ → H20
NADH+ H+ + ½ O2 → NAD+ H2O → coupled half rxn
O2 has a higher affinity for e- than NAD+
NADH is more likely to donate e-
in order for e- to be transferred, the 2 rxn must be coupled
what is the nerst equation

Describe the ETC
e- are transferred through a series of e- carriers (most of which are embedded in C1-IV) of increasing Eo’ until they reach O2 the final e- acceptor
in the process, H+ are moved into the IMS
the ETC is composed of 4 major complexes, each containing multiple protein subunits and e- carrier
there are also 2 e- carriers that act as shuttles moving e- from complex to complex
describe complex 1 in ETC
NADH-Q oxidoreductase
accepts 2e- from NADH
e- are transferred to FMN and then a series of 4Fe-4S clusters and finally to coenzyme Q (ubiquinone), reducing it to QH2 (ubiquinol)
the results in 4H+ pumped out of the matrix and into the IMS
NET RXN:
NADH (matrix) + 5H+ (matrix) + Q → NAD+ (Matric) + 4H+ (IMS) + QH2
describe complex II in ETC
succinate dehydrogenase is part of this complex
e- going from succinate to furmarate are transferred to FAD (forming FADH2), then to the succinate-Q reductase (i.e rest of CII) flowing through a series of Fe-S complexes and then finally Q (forming QH2)
i.e. these are the e- from FADHs of the Kreb’s cycle
e- from NADH in CI do not pass through CII
CII is not a proton pump so e- from FADH2 do not move as many H+ across the IMM as NADH
coenzyme Q
Ubiquinine (Q)/ ubiquinol (QH2)
acts as a shuttle moving e- from either CI or CII to CIII
it is a small hydrophobic molecule located in the IMM
contains a repeating isoprensiol tail
the number of repeats varies from species to species
e.g. in humans = 10 (i.e. CoQ10)
ubiquinone can accept 2H+ and 2 e- to be reduced to ubiqunol
i.e. Q+ 2H+ + 2e- ←→ QH2
complex III
Q-cytochrome C oxidoreductase
it contains:
1× 2Fe-2S cluster
2x cytochromes: cytB (contains heme bL (low affinity) + heme bH (high affinitiy))
e- flow: QH2 → 2 Fe-2S → heme c1 → heme c
BUT as QH2 docks, one e- follows the above, the other e- goes to heme b and participates in the Q cycle, before it follows the first e- to the 2Fe-2S cluster
net equation:
QH2 + 2Cyt(ox) + 2H+ (IMM) → Q + 2CytC (red) + 4H+ (IMS)
ox = oxidsed
red = reduced
cytochrome
e- transferring proteins containing 1 or more hemes
Q-cycle
You do not need to know the Q-cycle expect that it is moving e- and pumping H+ through CIII
if both e- flow through CIII, 4H+ are pumped into the IMS:
2H+ come directly from the matrix
2H+ come from QH2 (these H+ orignated in the matrix)
cytochrome c (cyt C)
contains heme c
it is another e- shuttle
water soluble protein containing covalently linked heme
carriers 1 e- from CIII→ CIV
it rolls along the surface of the IMM on the IMS side
Fe3+ + 1e- → Fe2+ (heme)
complex IV
cytochrome C
carriers out the final reductin of oxygen to water using e- from cyt. c
requires 4e- to completely reduce O2 → H2O
in the process, 4H+ are pumped into the IMS
contains:
2 cytochromes:
cyt a(hemea)
cyt a3 (hemea3)
2 copper centers:
CuA
CuB
O2 binds to heme a3 and then bridges between heme a3 + CuB
e- flow: heme c → CuA → hemea → hemea3/CuB →O2
net equation: 2ctyc(red) + 4H+ (IMM) + ½ O2 → 2cytc (ox) + 2H+ (IMS) + H2O
reactive oxygen species
CIV is designed to prevent the release of partially reduced O2
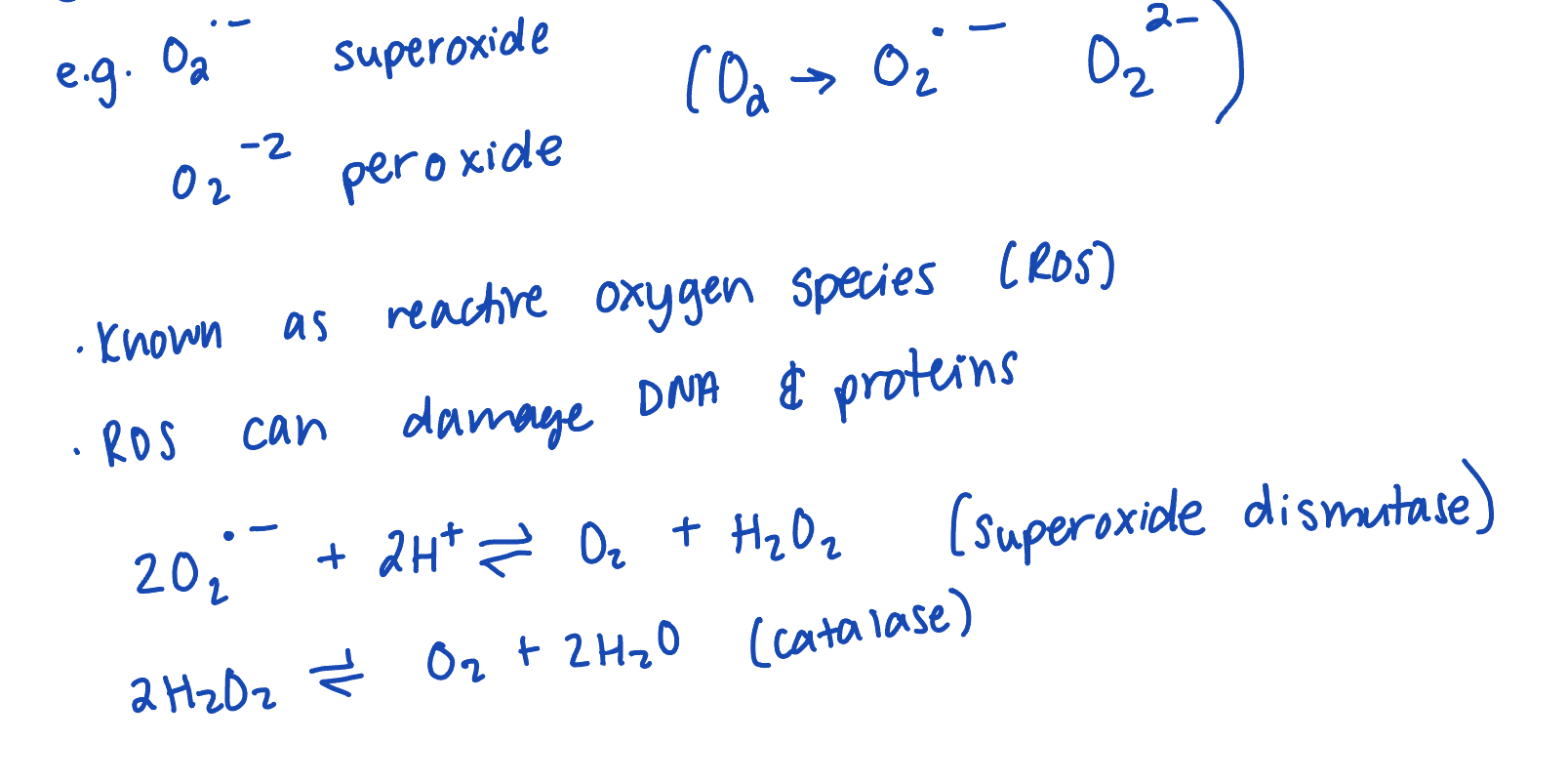
ETC inhibtiors
somes molecules can bind to various e- varriers in the ETC and block e= transfer
e- carriers before the block will become reduced and e- carriers after the block will become oxidsed
therefore, it blocks the ETC, prevents generation of the proton gradient of the H+ gradient, ATP synthase slows/stops, no ATP made

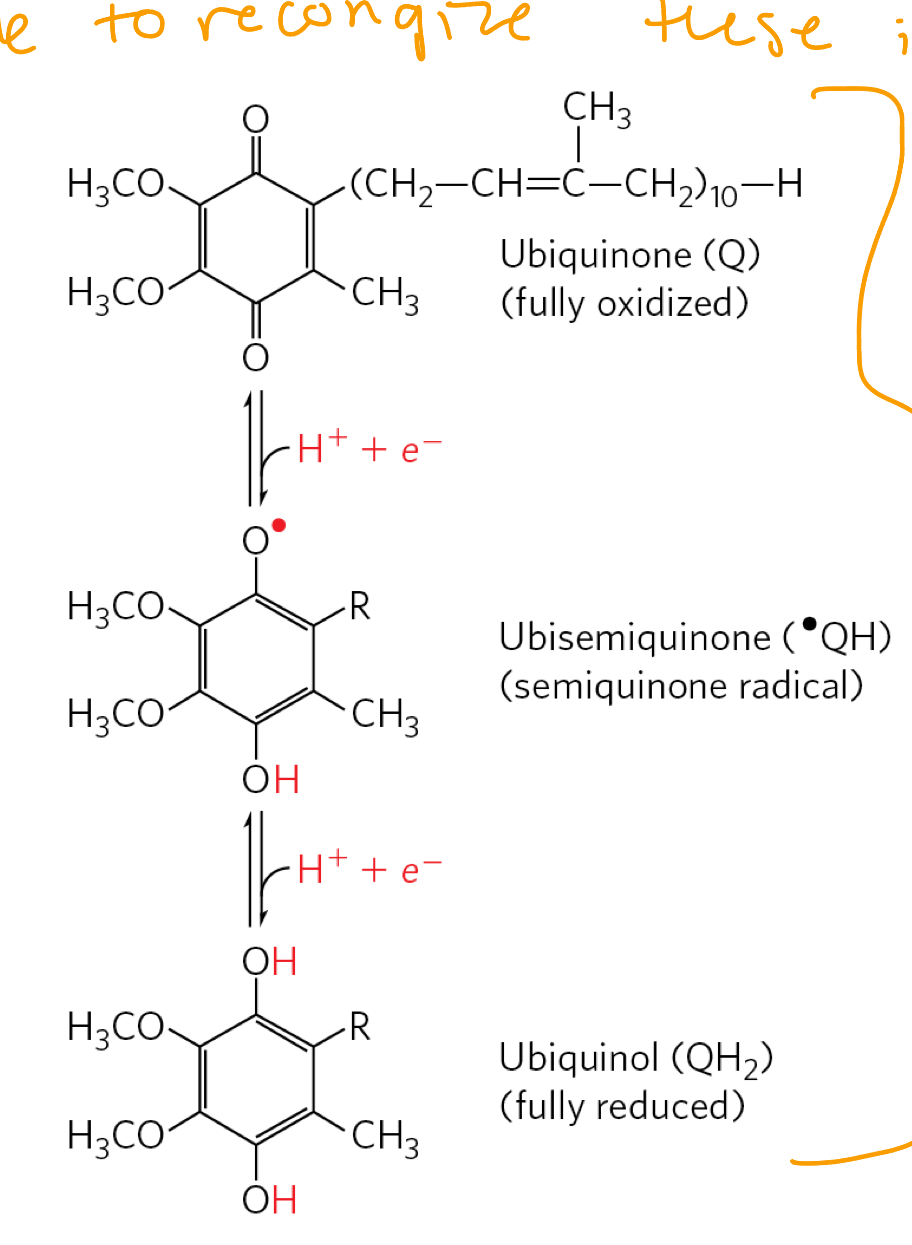
need to recnogize in all forms
ATP synthase and ATP synthesis
need to harness the PMF created by the ETC to generate ATP
the pH difference between the matric and IMS is 1.4 pH units (~25 fold difference)
this is done by ATP synthase
consists of 2 components:
Fo
F1
Describe F1
extends into the matrix + synthesis ATP when coupled to the spin generated by Fo
contain 3 alphabeta subunits arranged in a ball
ATP synthesis occurs in B subunit
in the center of the ball is y (gamma) shaft
the subunit y spins - the alphabeta subunit does not
the y shaft binds to effect alphabeta subunit differently, causing different conformaitons in each of the 3alphabeta subuntis
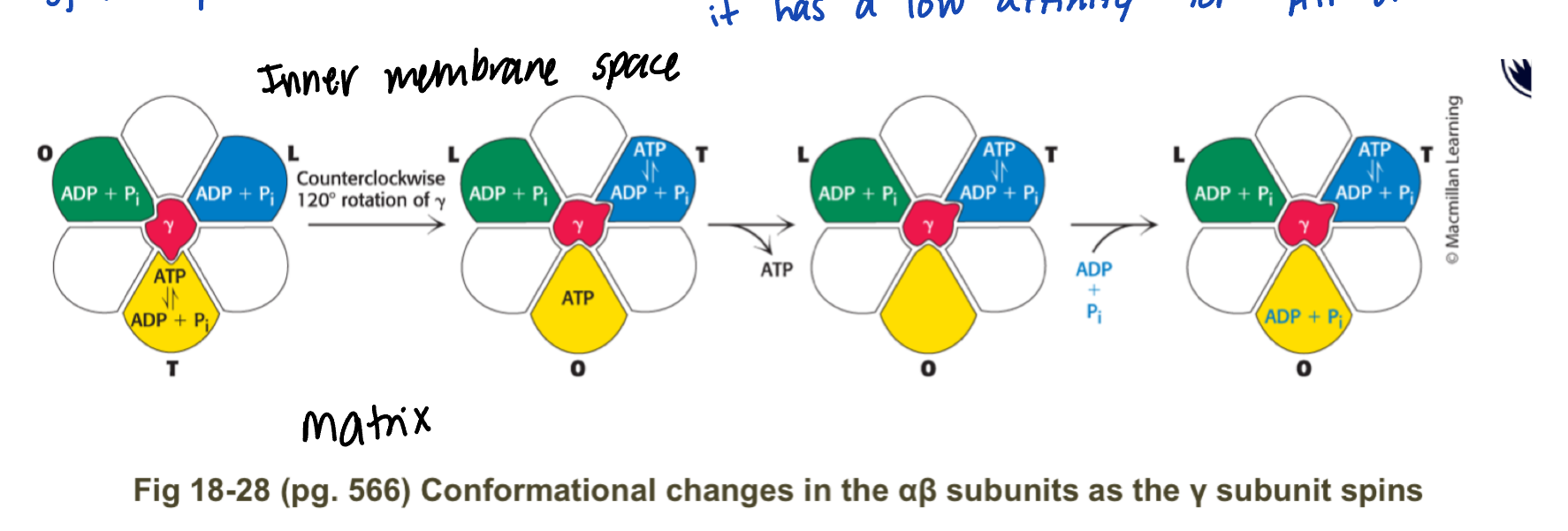
describe the 3alphabeta subunits in F1
alphabeta lose site (L, BADP) = loading conformation
ADP + Pi can bind and become trapped
alphabeta tight site (T, BATP) = ATP synthesis step, where ATP is made and found tightly to the B subunit
alphabeta open site (O, B-empty) = release confromation, it has a low affinity for ATP or ADP
using the sping from Fo, the y shaft spins
this causes each of the alphabeta subunits to cycle through the L,T, and O conformations
ATP generation +release
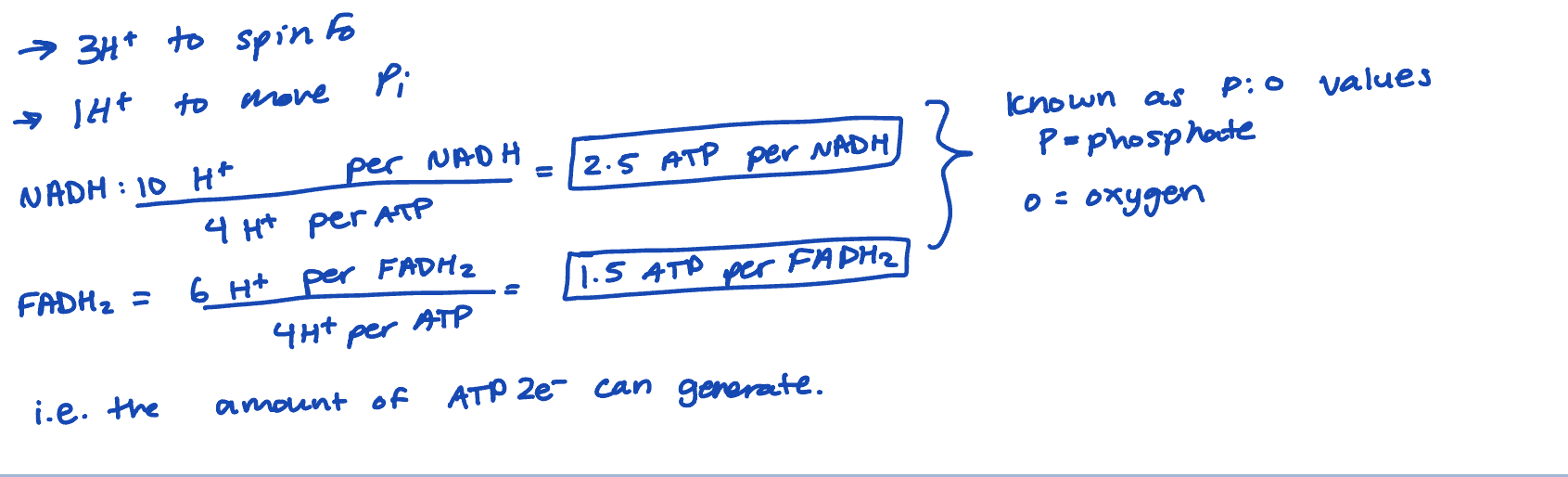
best estimates suggest 4 H+ need to move into the matric per ATP made
describe Fo
embedded in IMM
contains the half channels that H+ flow through
uses H+ to generate spin
how does Fo cause the y shaft to spin? → only describe subunit c
Fo is composed of many subunits, but focus on subunit c and a
subunit c:
composed of 2 alpha-helices that can span the membrane
there are 10-12 (10 in humans) c subunits arranged in cylinder
the entire cylinder will rotate
halfway down one of the alpha-helices is a kep aspartate/aspartic acid residue which can be protonated or deprotonated depending on pH
can only move into or be exposed to the membrane, if it uncharged (i.e. protonated) but it can move when charged if it is covered by the subunit a clamp
the clamp masks the charge
how does Fo cause the y shaft to spin? → only describe subunit a
Fo is composed of many subunits, but focuses on subunit c and a
subunit a:
aka the clamp
the subunit covers 2 c subunits
it is stationary (i.e. does not rotate)
it has 2 half channels
1 is open to the IMS
the other is open to the matrix
How does Fo use the proton gradient to spin?
a charged apartate subunit C is in the IMS half channel, a charged aparate subunit c is in the matric half channel
a proton diffuses from IMS (where [H+] is high) through the IMS half channel and protontes the aspartate to an aparatic acid
now this subunit c is charged
the subunit c complex cylinder can rotate clockwise by unitc and freshly protnated uncharged subunit c move in the membrane
this brings a charged subunit c into the IMS half channel and an uncharged subint into the matrix half channel
a proton diffuses off the apartic acid, down the matrix half channel and into the matric
this subunit c is now charged (back to step 1)
regulation of oxidative phosphorylation
normally ATP synthesis and ETC are coupled
ATP us formed only as fast as it is consumed
ATP synthase cannot spin if ADP + PI are not bound to the loose alpha beta
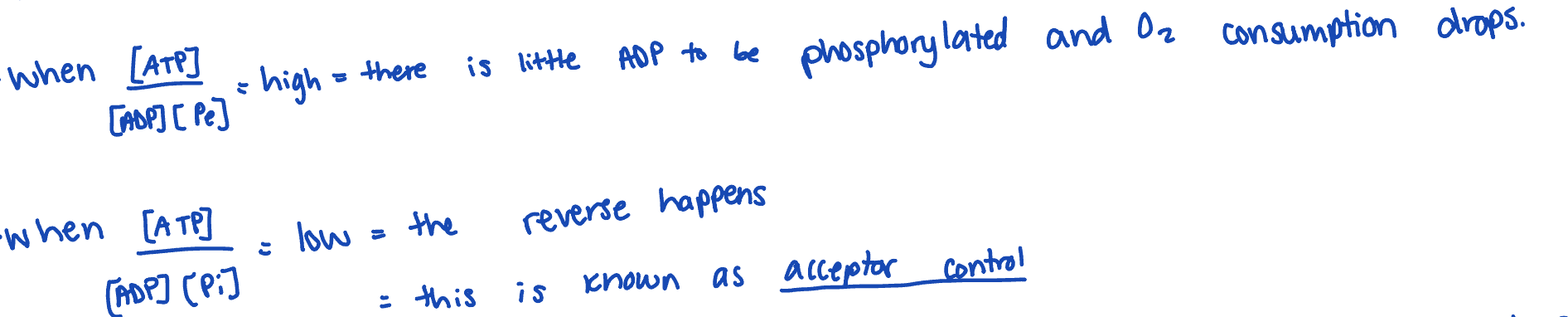
acceptor control
the regulation of cellular respiration by the availability of ADP as a phosphate acceptor
Describe DNP, brown fat, and FCCP
both uncouplers
2,4 dinitrophenol (2,4 DNP) can uncouple the ETC and ATP synthase by carrying H+ across the IMM
this reduces the proton gradient and the ETC speeds up to restore it
BUT the rate of ATP synthase stays the same (if 2,4 DNP is really high is drops)
brown fat carries thermogentics (aka uncoupling protein 1) and this is found in newborn babies and mammals to adapt to the cold
this is a proton channel in the IMM
transport across the IMM
electrons from NADH
e- transport to the ETC
occurs by shuttles (translocases)
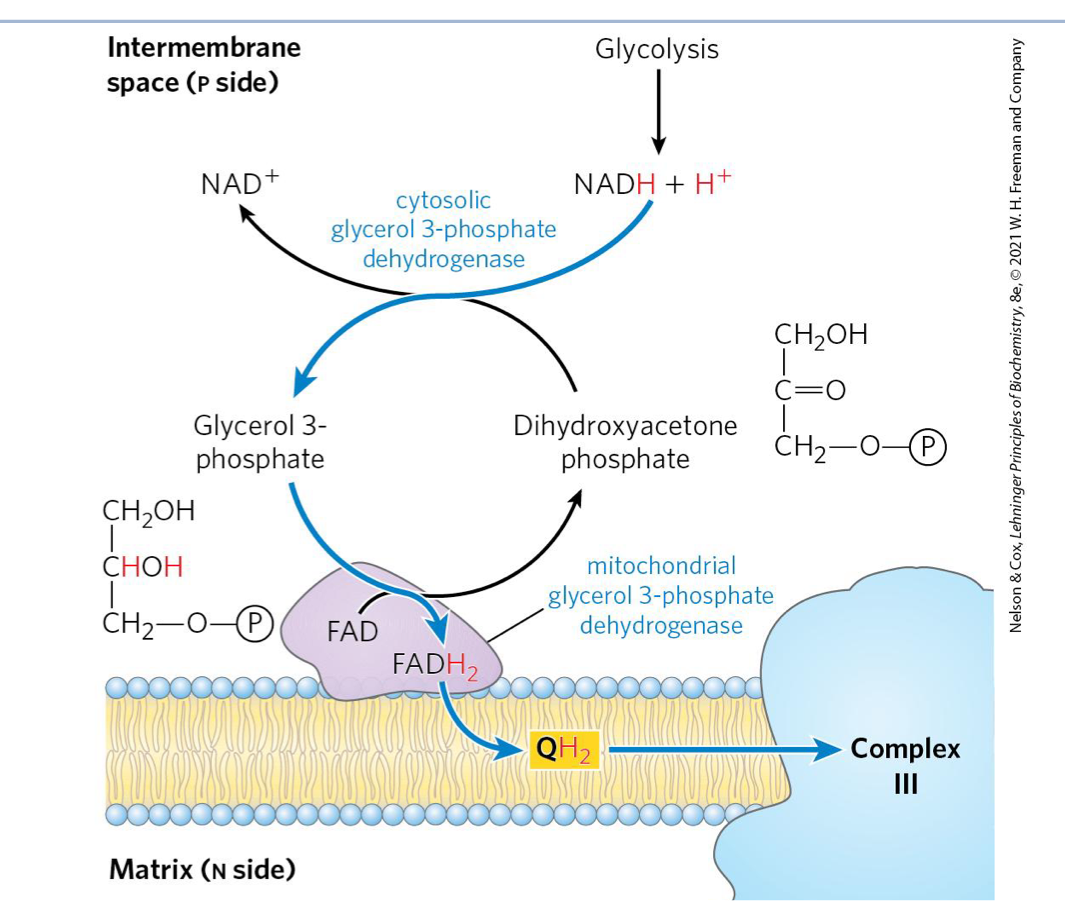
Describe steps for glycerol-3-phosphate shuttle
occurs in skeletal muscle and brain
cytosolic glycerol-3-phosphate dehydrogenase reduces dihydroxyactone phosphate (DHAP) to glyercol-3-phosphate oxidzing NADH → NAD +
glyercol-3-phosphate is carrying the e-
mitochondiral glycerol-3-phosphate dehydrogenase (bound to IMM) oxidizes glycerol-3-phosphate back to DHAP in the process transferring 2e- to FAD → FADH2
FADH2 passes 2e- to Q reducing it to QH2
QH2 → CII
this is similar to CII
e- by pass CI entering as FADH2
therefore, use FADH2 p: o value)
describe steps for malate-asparatate shuttle
occurs in liver, kidney, heart
oxaloacetate is reduced by NADH to form malate through the activity of cytosolic malate dehydrogenase (MDH)
Malate (carrying 2e- is transported to matrix by the malate) alpha-KG tranlocase
Malate is then oxdised back to oxaloactate by mitochondrial MDH, generating NADH
NADH → CI BUT oxaloacetate cannotdirectly go back to the cytosol
oxaloacetate is transaminated by glutamate to form aspartate and alpha-KG in the matrix
the glutamate/aspatase translocase moves asp into the cystol alphaKG is also moved into the cytosol by mal/alphaKG translocase
in the reverse of 4, asp transminates alphaKG, reforming oxaloactate and Glu
goes back to step 1
describe ATP, ADP, Pi as transport across the IMM
page 41.