Bio 2 Test 3
1/84
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
85 Terms
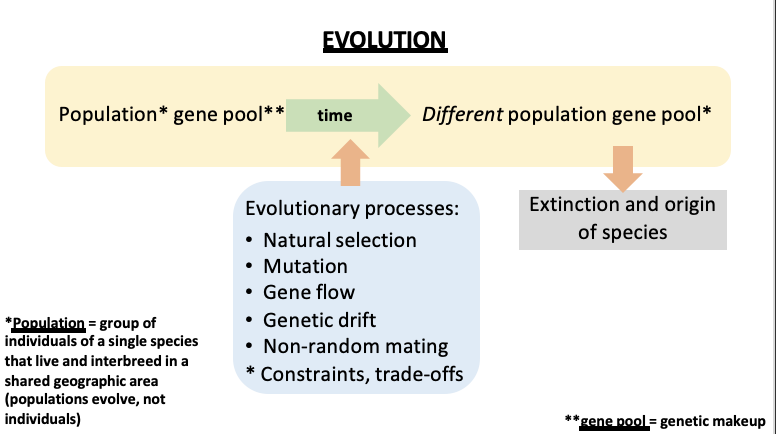
What does evolution start with and affect? What is it?
Evolution affects populations, not individuals
Evolution starts with a population (gene pool)
A population is a group of individuals of a single species that live and interbreed in a shared geographic area
Over time, in the bounds of constraints and trade-offs, evolutionary changes happen due to….. What is the result of this?
Evolutionary processes
Natural selection
Mutation
Gene flow
Genetic drift
Non-random mating
A different gene pool (genetic makeup) is created that can further drive the origination or extinction of a species.
How do we know evolution is happening?
Vast evidence from geological, morphological, behavioral, and molecular data that can be observed in the lab or natural populations.
Who proposed natural selection and how did they come upon it?
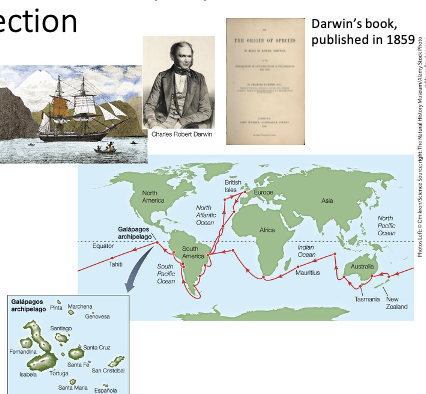
Charles Darwin and Alfred Wallace- Natural Selection
The goal of their mission: chart the oceans; collect oceanographic and biological information
Darwin’s Findings:
He observed differences in the animals in Europe versus those in South America
Animals in temperate and tropical regions in South America were similar
In the Galapagos Islands, some animals were only found in that area
Island-to-island differences
What were Darwin’s 3 primary evolutionary frameworks?
Species are not immutable
They change over time
Descent with modification
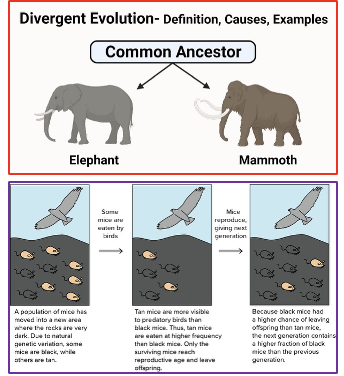
Divergent species share a common ancestor
Species diverged from one another gradually over time
Changes in species over time can be explained by natural selection
Evolution can be cause by ___ ___. Explain this
Natural Selection
Increased relative individual fitness (survival and reproduction success over a lifetime) of certain individual phenotypes within a population:
Genetic Variation in a population (due to mutations)
Heritable Traits
The traits are related to fitness (survival and reproduction)
The result/consequence
The favored phenotypes (and the corresponding underlying genetic differences) are spread throughout the population and to future generations: leading to evolution
A population’s genetic structure can be described by ___ and ___ frequencies. Explain what this structure is.
Allele, genotype
The Gene Pool’s Structure
2 Key descriptions:
Allele Frequency
The proportion of individual alleles in a population
Example: Frequency of allele X1 vs. allele X2 in a population
Genotype Frequency
The proportion of genotypes (which are composed of multiple alleles)
Example: Frequency of the Genotype X1X2 vs. X1X1 in a population
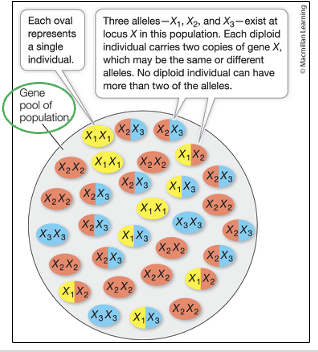
Evolution is defined by changes in ____ frequencies, NOT ____ frequencies
Allele, genotype
What are adaptations?
Favored traits that are spread through a population by natural selection
They tend to be favorable for a specific organism in a particular habitat
Describes a trait and a process
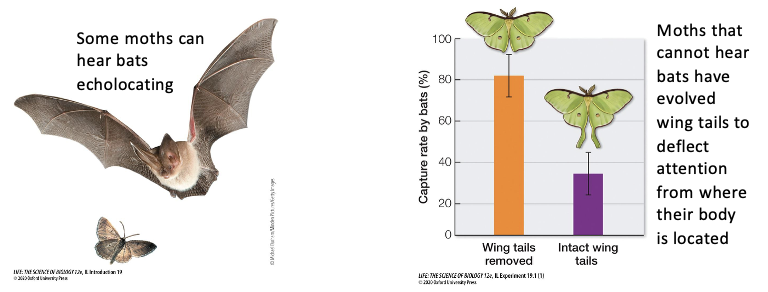
Some moths can hear bats echolocating
Moths that cannot hear bats have evolved wing tails to deflect attention from where their bodies are located
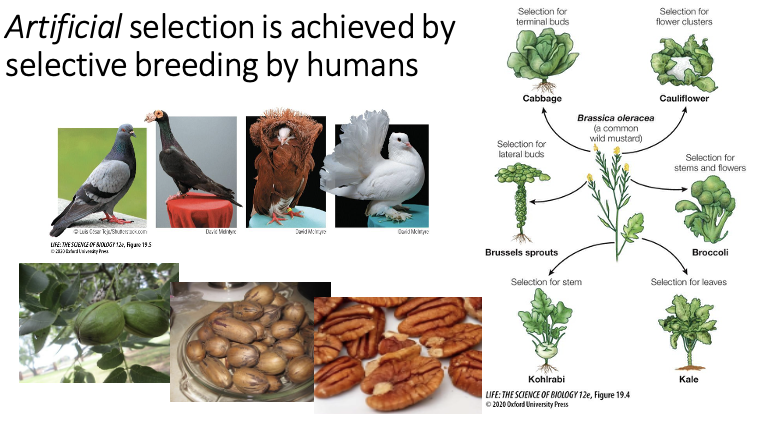
What is artificial selection?
When certain traits are selected for by humans
What do mutations generate? Where are they common? Are they maintained?
Genetic variation: raw material for evolution by selections
Basis for genetic variation
At the population level, mutations are common (in contrast to where mutations are a more rare event in individuals)
Mutations can stem from unfixed errors in DNA replication
Mutations are maintained in a population when the mutation is beneficial (via natural selection)
Can be a form of adaptation
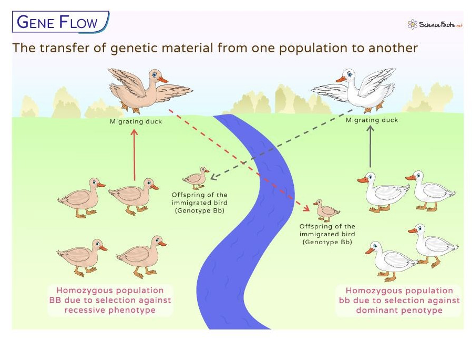
What is gene flow? What can it change?
Gene flow is the exchange of genetic material between populations through migration of individuals or movement of gametes (i.e., pollen dispersal)
This exchange of allele frequencies in a population
Populations are not isolated from each other: they can experience emigration and immigration.
Two different populations of ducks. Immigration/emigration. Dispersal of gametes is like in pollination. If they can breed, they can introduce new alleles into the respective population. A homozygous dominant cross with a homozygous recessive.
What is genetic drift? What does it impact the most and why?
Random evolutionary process that impacts small populations the most.
Why? Chance events have a proportionately larger impact on their smaller gene pools, leading to rapid, significant shifts in allele frequencies.
Is a process of random changes in allele frequencies over generations
This impacts the ratio of deleterious vs beneficial alleles.
In small populations, advantageous alleles might be lost, and harmful ones spread.
Blue little a allele is starting to be removed over time if the only male breeder is the red beetle.
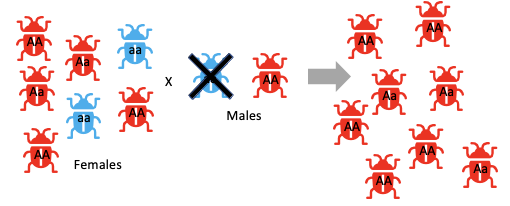
What is the effect of genetic drift on large populations? What are the two types?
In large populations, frequencies of neutral alleles (no effect on fitness) might change.
Founder effect and bottleneck effects
What is a population bottleneck?
Result of an environmental event (Volcano, Tornado, etc.)
A population starts with equal allele frequencies, but after an event in which a small group of individuals gets isolated from the original population, the allele frequency changes.
Loses genetic variation.
Happened with the greater prairie chicken in Illinois
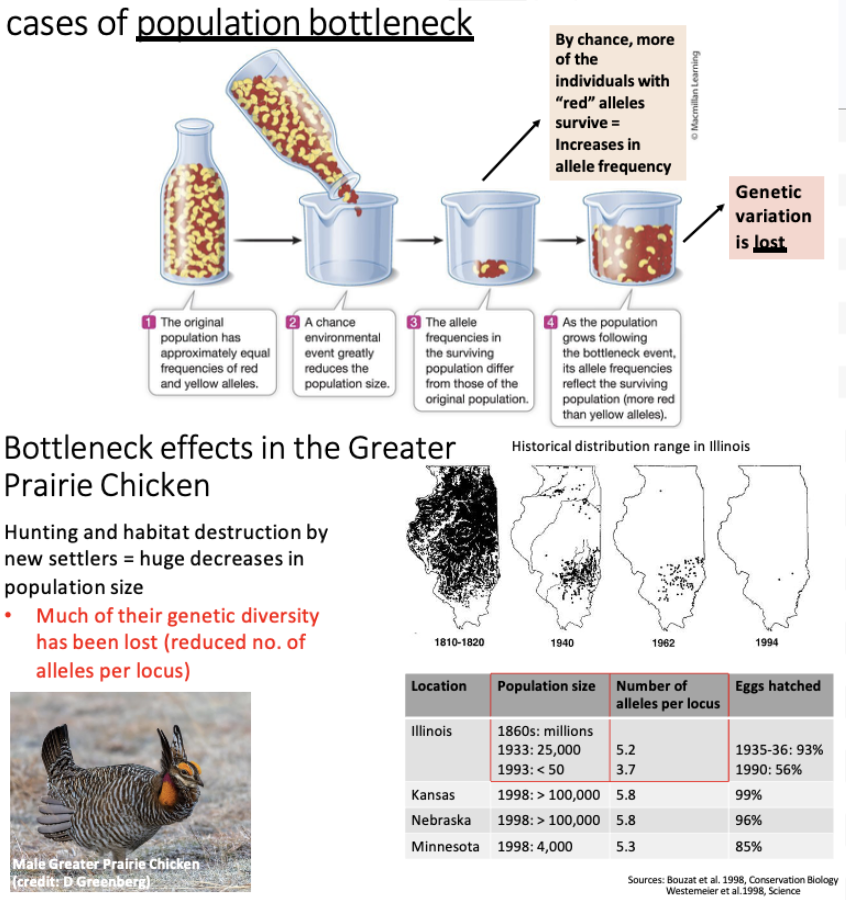
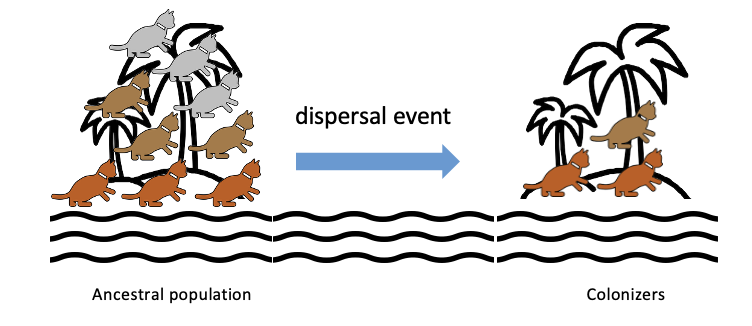
What is the founder effect?
Result of a Dispersal Event (migration, colonization)
A small portion of the population leaves and populates a different area, causing only certain alleles to be present in the new population
Genetic variation decreases
Because only a subset of the population is present on the new island = lower genetic variation compared to ancestral population
No grey individuals, hence lower than the ancestral that did include the grey.
What is non-random mating? What can it change?
Based on individuals’ preferentially choosing specific mating partners
Nonrandom mating changes genotype frequencies: no evolution
Nonrandom mating changes allele frequencies: evolution
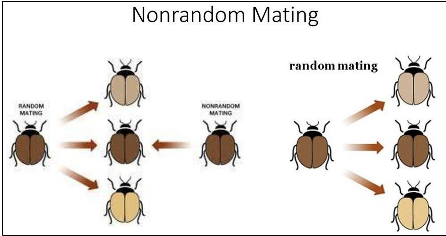

What is non-random mating’s impact on genotypic frequencies?
Two types of mating preferences
Mating with their own genotypes:
Self-Fertilization in Hermaphroditic Species
Result: Increase in homozygous genotype in a population
Mating with different genotypes:
Result: Increase in heterozygous genotype
What is non-random mating’s impact on allele frequencies?
Only in species with sexual reproduction and 2 or more sexes.
Individuals of one sex mate preferentially with particular individuals of the other sex(es):
Intersexual selection- Based on desirable traits
Individuals of one sex prefer to mate with individuals of another sex that show certain phenotypes
Intrasexual selection- Based on competition
Individuals of one sex compete among each other to access mates
Interaction between sexes
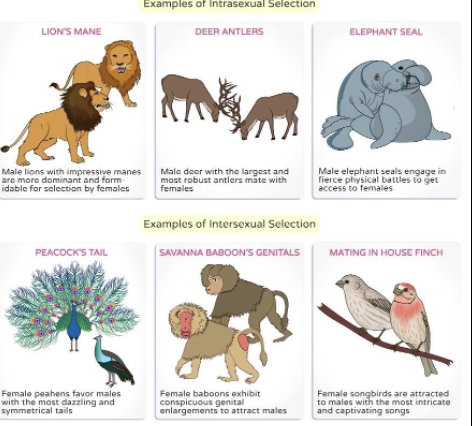
What are the two main reproductive strategies and their subsets?
Asexual reproduction
Vegetative reproduction
From non-sexual tissues: clones
Parthenogenesis
From germ cells: clones, genetically variable
Sexual reproduction
Isogamy
Similar gametes in both parents
Anisogamy
Different gametes, often variable in size: gametic sex
Possible sexual dimorphisms (2 or 2+ sexes) by sex determination systems: sexual chromosomes, haplodiploidy, physical environment, social environment
Mating systems, sexual selection, sex roles
Hermaphroditism
Sequential OR Simultaneous (sometimes self-fertilization)
Produce both large AND small gametes
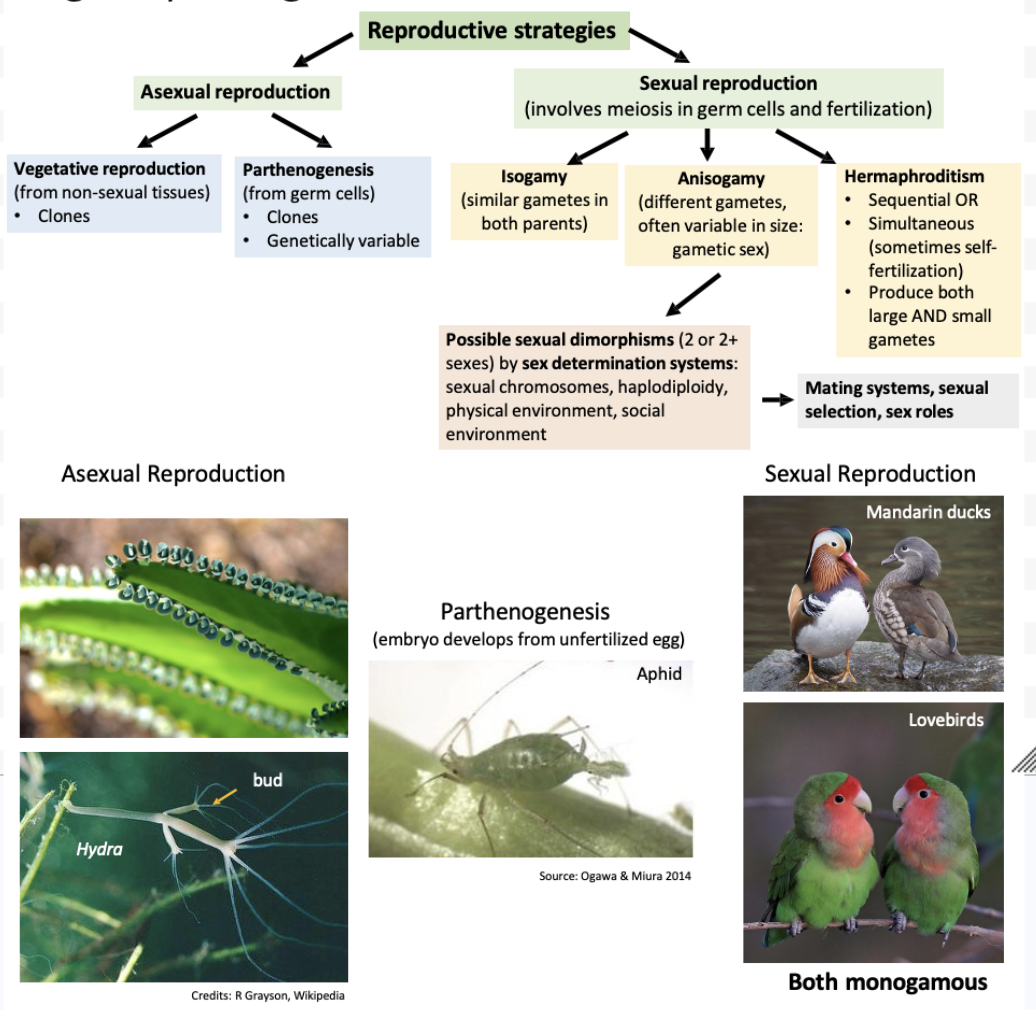
Explain hemaphroditeisms that exist.
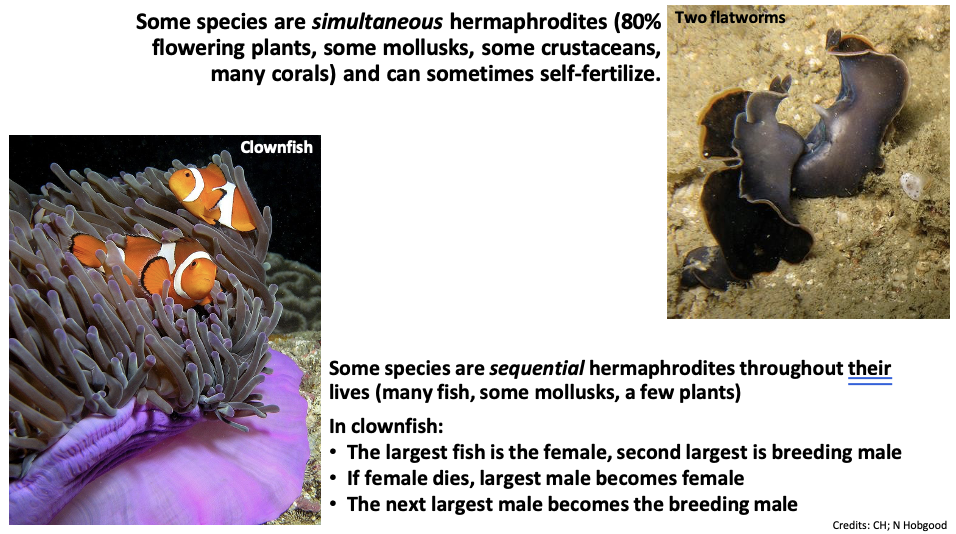
Some species are simultaneous hermaphrodites (80% flowering plants, some mollusks, some crustaceans, many corals) and can sometimes self-fertilize.
Some species are sequential hermaphrodites throughout their lives (many fish, some mollusks, a few plants)
In clownfish:
The largest fish is the female, the second largest is the breeding male
If a female dies, the largest male becomes female
The next largest male becomes the breeding male

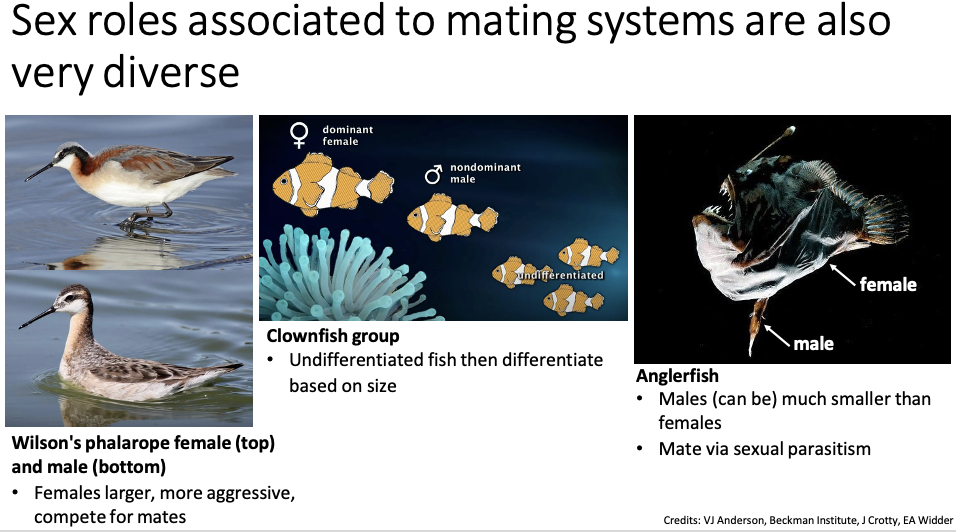
Sex roles associated to mating systems are also ___ ____
Very diverse!
Wilson's phalarope female (top) and male (bottom)
Females are larger and more aggressive, and compete for mates
Clownfish group
Undifferentiated fish then differentiate based on size
Anglerfish
Males can be much smaller than females
Mate via sexual parasitism

Constraints and Trade-Offs Shaping Evolution: Preexisting Traits
Preexisting Traits can limit the outcome of selection
Physical and chemical constraints limit evolution (e.g., thermodynamics, cell functioning and cell size, etc.)
All evolutionary innovations are modifications of previously existing structures
Can only evolve if its pre-exisisted in some fashion.
Lack of the necessary genetic variation prevents the evolution of certain traits
Example: Flounder versus Stingray

Constraints and Trade-Offs Shaping Evolution: Trade-offs
Trade-offs balance costs and benefits in the evolution of adaptations.
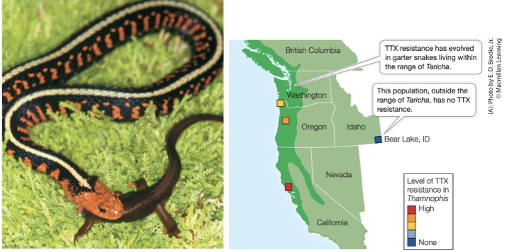
Case Study: Garter Snakes
When garter snakes consume rough-skinned newts, a toxin kills them
Certain snakes have evolved to not die from the toxin by means of a modified sodium channel that resists the toxin
The benefit: They are resistant to the toxin
The cost: They generally move more slowly than non-resistant snakes, especially after consuming a newt.
The result: Because the resistant snakes are slower, they are more susceptible to predation, which overrides the benefit of being resistant to the toxin. This resistance is therefore selected against.
Explain how natural selection ties into traits.
Natural selection acts directly on the phenotype, and thus indirectly on the genotype
Each phenotype within a population has a relative fitness
Fitness - relative rates of survival + reproduction of the phenotype in the population
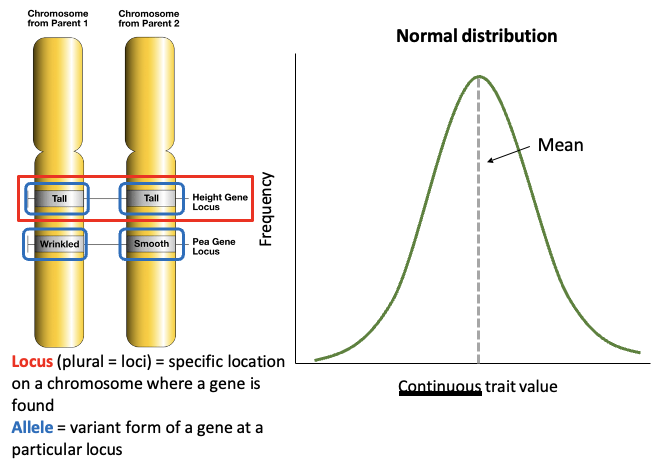
Traits can be qualitative or quantitative, explain. What are Allele vs Locus?
Traits affected by one locus (place on the chromosome) are qualitative
Discrete variation (smooth vs wrinkled peas)
Traits affected by more than one locus are quantitative
Continuous variation (height vs weight)
The different types of selection apply to quantitative!
Allele = variant form of a gene at a particular locus
Locus = specific location on a chromosome where a gene is found
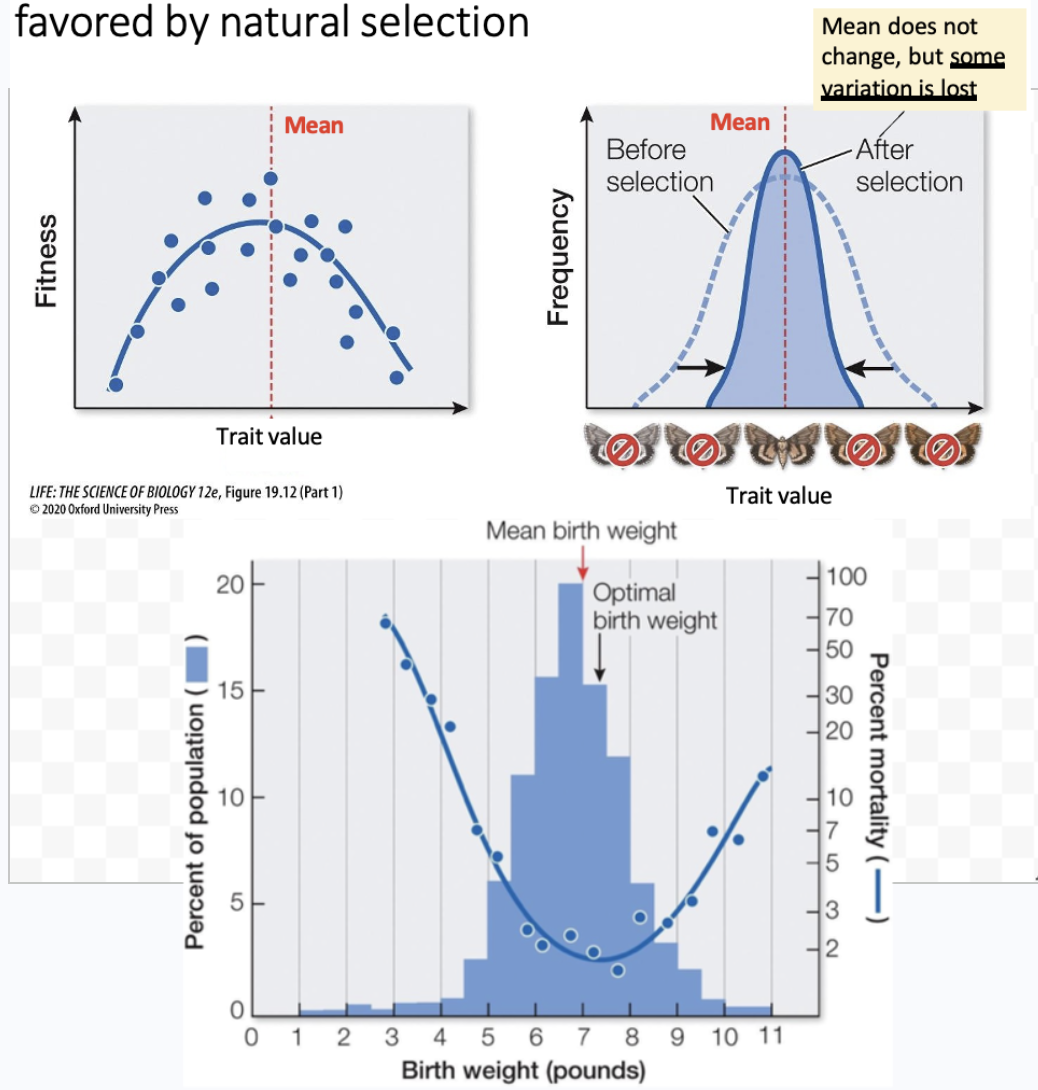
What is stabilizing selection? Example?
Quantitative selection where the mean phenotype is favored by natural selection, while variation is reduced
Sometimes referred to as purifying selection
Birth weight in humans.
Babies born with weights < or > than the mean have higher death rates than those closer to the mean
Selection against deleterious mutations or consequences
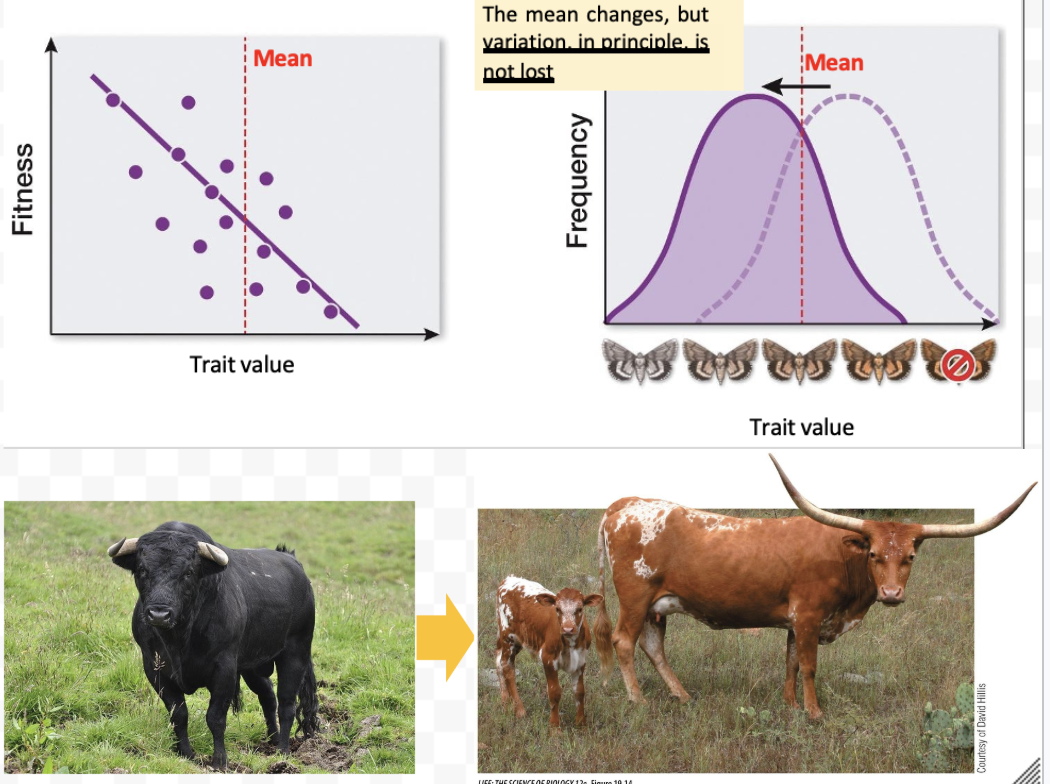
What is directional selection? Example?
Quantitative selection where one phenotype other than the mean is favored by natural selection
The mean changes, but the variation stays the same
Texas longhorn = result of directional selection on cattle introduced from Europe
Directional selection in horn length favored protection against local predators in Texas and the SW (mountain lions, bears and wolves)
A form of “positive selection”
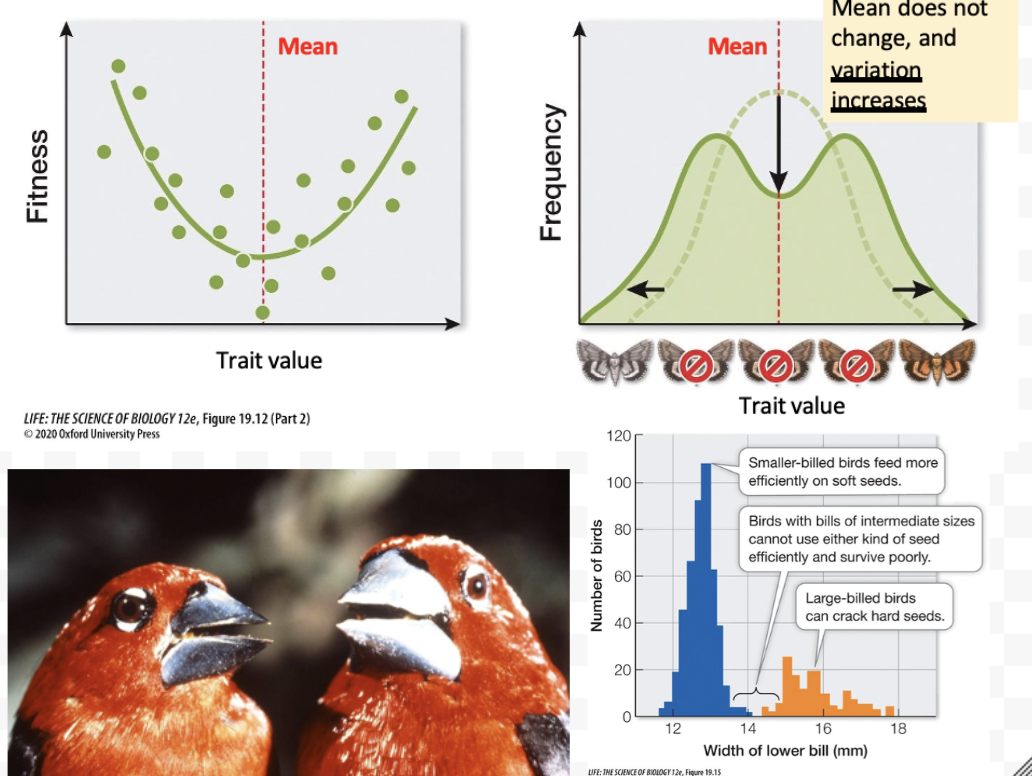
What is disruptive selection? Example?
Quantitative selection where both extremes are favored as phenotypes vary in both directions from the mean.
Mean stays the same, but variation increases
Black-bellied seedcrackers have two morphs as a result of disruptive selection (credit: T Smith, UCLA)
Feed on seeds with different morphologies from the same genus of tree
Those with an intermediate bill/beak size do not survive
Frequency-dependent selection maintains genetic variation within populations. Explain.
Genetic variation normally results in having two or more phenotypic variations within the same population.
The frequency of each phenotype determines the individual fitness
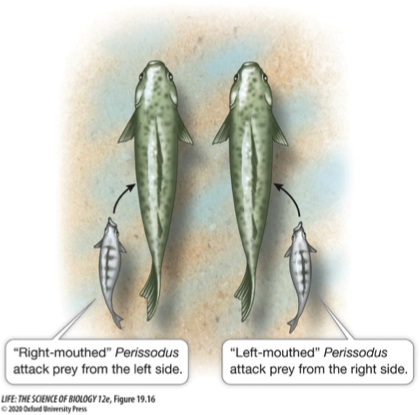
Think back to the example of right and left mouthed scale eaters
Prey fish are more likely to be attacked if they have to watch both sides for predators
This favors equal numbers of right- and left-mouthed scale-eaters in a population

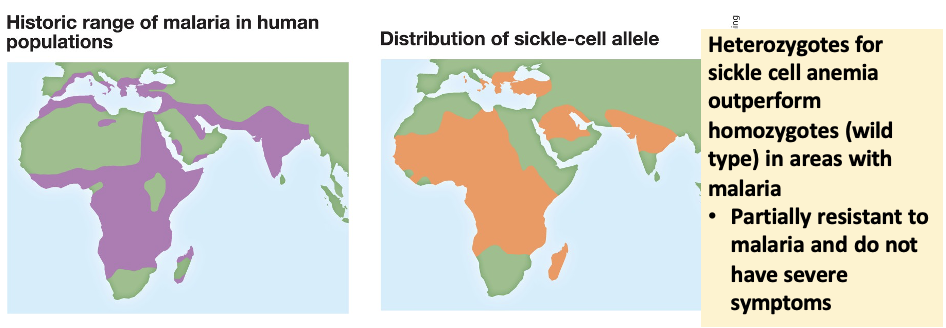
Heterozygote advantage maintains polymorphic loci. Explain.
Heterozygote individuals (with 2 alleles) could outperform homozygotes (with 1 allele) under variable environmental conditions.
Sickle-cell anemia provides heterozygotes with immunity to malaria
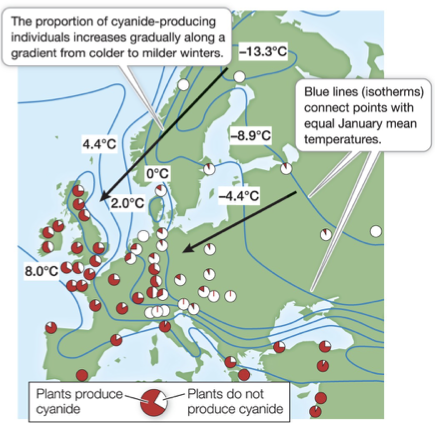
Genetic variation within species maintained in geographically distinct populations
Populations are under different local selection pressures (differing environments)
Temperature, sunlight, predation, etc.
More cyanide production = in warmer environments
Cyanide production in clover (red) protects from herbivory, but makes the plants susceptible to frost
Damages cell membranes, releases cyanide into their own cells
Clover plants. Eat the leaves; they contain cyanide. The downside is that when they live where it’s cold, it bursts their cell membranes and they essentially kill themselves. In warmer climates, it survives just fine.
END OF CHAPTER 19
END OF CHAPTER 19
All things share what?
A common ancestor.
What are evolutionary relationships called? How are they presented?
Phylogeny - the history of how organisms are related
Historical relationships represented among lineages using a phylogenetic tree
A lineage = line of ancestors and descendants
What can a phylogenetic tree be used for, and how is it determined? What are branches and what do it use?
Can be used for major evolutionary groups (i.e., insects), populations, species, genes, etc.
Usually determined via morphological (physical) and/or molecular data, but can also use behavioral, physiological, developmental, and ecological.
Uses DNA
Branches form when a new species evolves over a period of time
Mutations can cause these splits
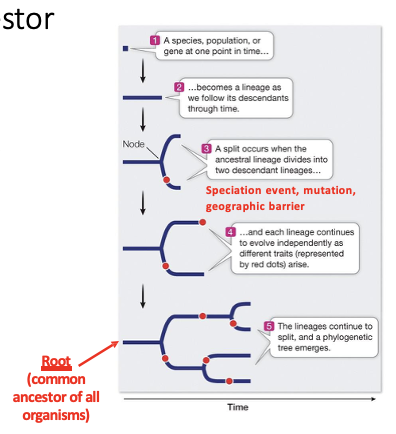
What does matter when reading a phylogenetic tree? What doesn’t?
DOES MATTER
Order of nodes
Branch Length represents time (longer branches=longer time)
DOES NOT MATTER
The vertical distance between branches is irrelevant
No connection to the relationship among groups
The order of neighboring branches (i.e., chimp and human) d
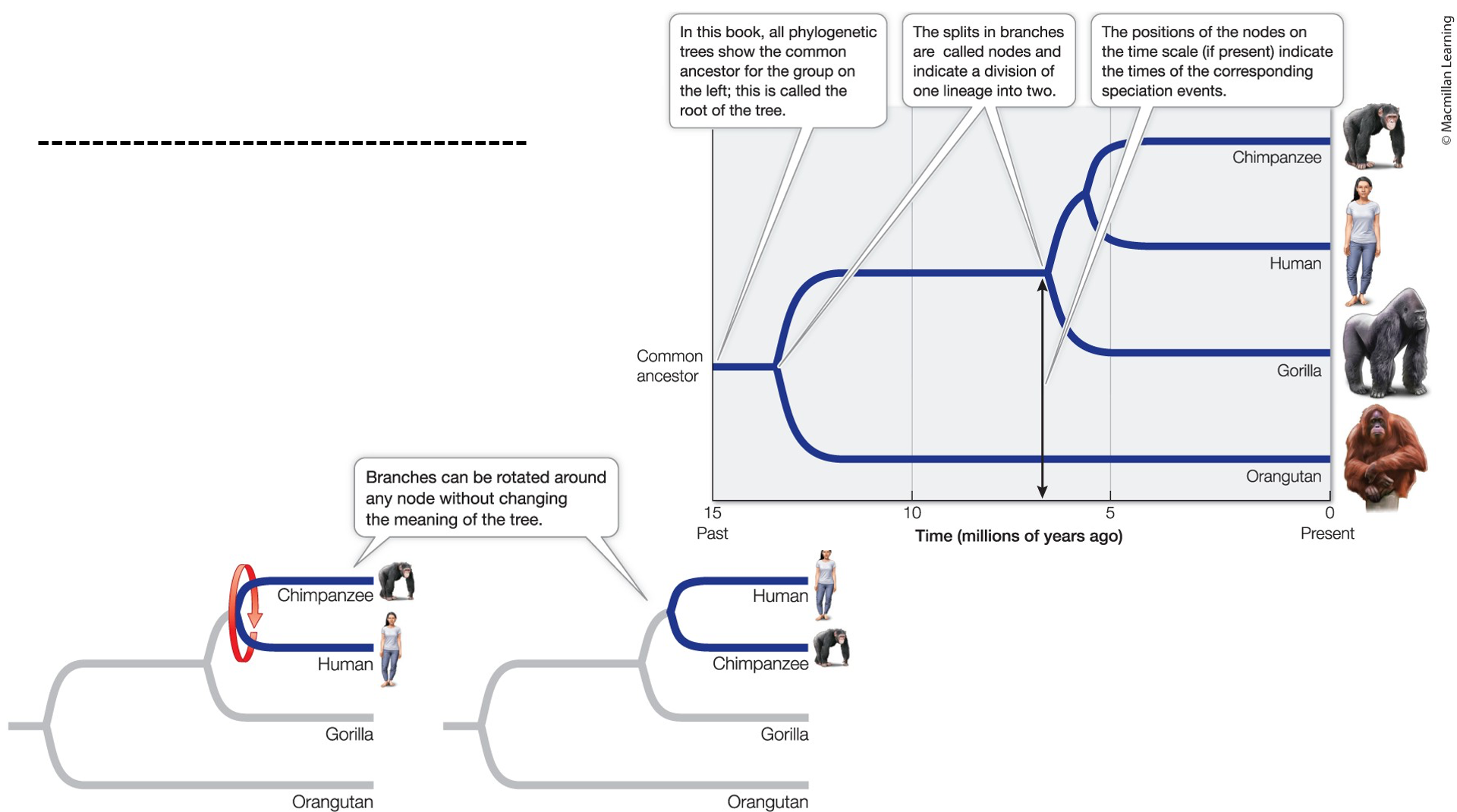
How are organisms named?
Binomial nomenclature: a system made by Carolus Linnaeus
Each one has a genus+species
Both italicized; genus is capitalized, species is not
Homo sapiens Linneaus: Genus is Homo, sapiens is the species, whole is italicized, last part is who proposed it
Written in Latin

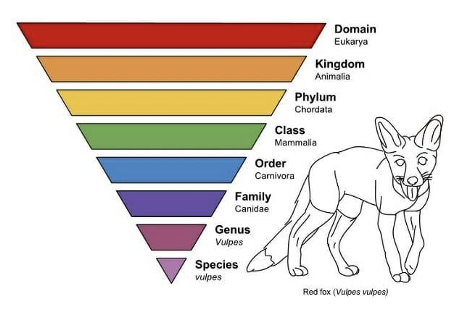
What is taxonomy?
The primary system used to classify and organize living things
Organisms are grouped into categories based on similarities (traits, DNA, behavior, etc.)
A group at the same classification/taxonomic level = taxon
A clade is a group that includes a common ancestor and all of its evolutionary descendants
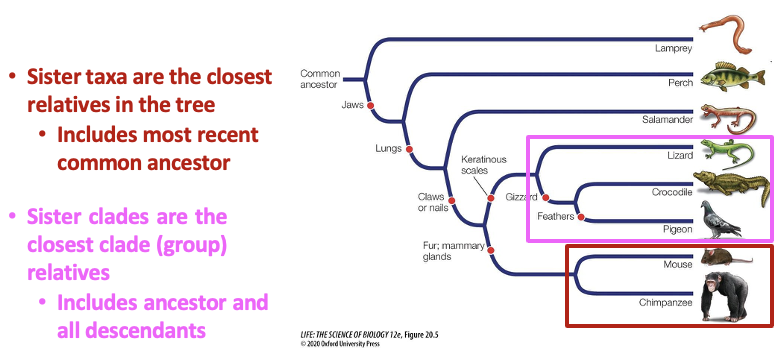
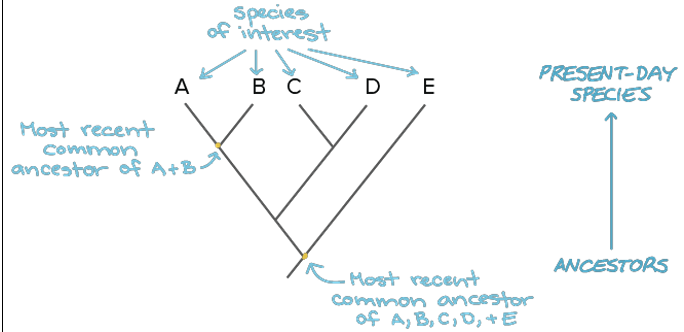
Phylogenetic trees and sister grouping. Explain.
Sister taxa are the closest relatives in the tree
Includes the most recent common ancestor
Sister clades are the closest clade (group) relatives
Includes ancestor and all descendants
Know that the sister is the closest because they have the most recent common ancestor. Clades share a common ancestor, but it’s not the most recent one. Shared ancestor is big.
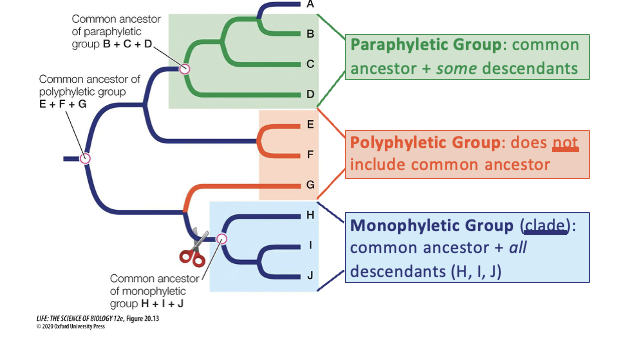
Explain the three types of grouping on a phylogenetic tree.
Polyphyletic Group: does not include a common ancestor
Paraphyletic Group: common ancestor + some descendants
Monophyletic Group (clade): common ancestor + all descendants (H, I, J)
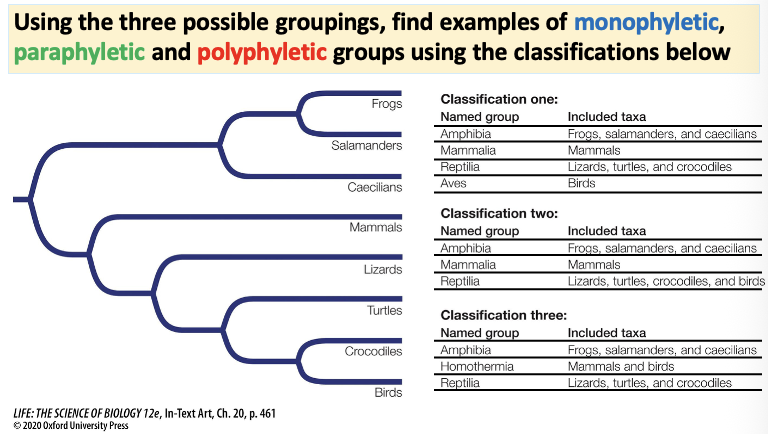
Answer this question diva!
Answer to thy phylogenetic tree that you seek yass queen.
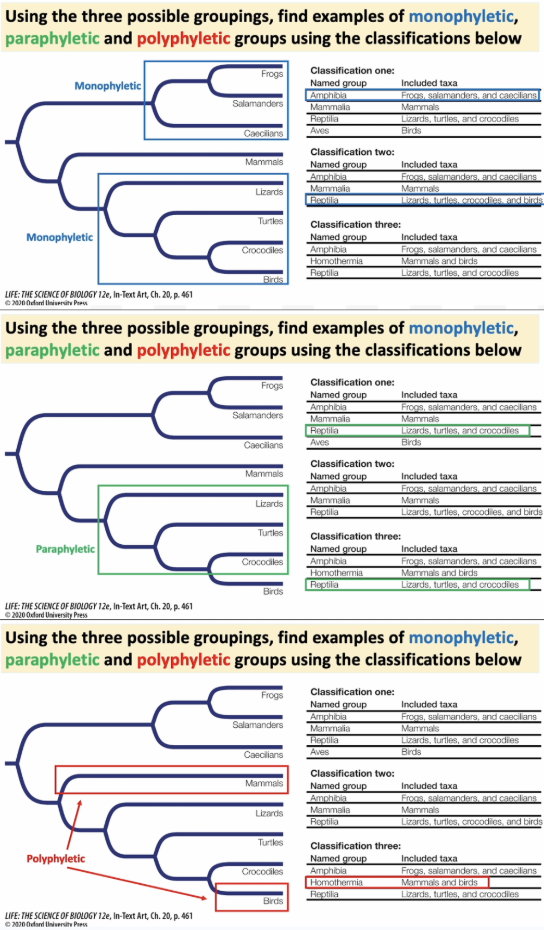
Phylogenies help scientists study and compare how traits evolved over time. Explain in terms of how they are derived and examined.
They compare shared traits across different groups (taxa), and thus study the evolution of traits
Current traits are derived, meaning they evolved from an ancestral trait
Scientists can estimate and reconstruct the ancestral state of traits by looking at the trait values of the extant (currently living) species.
Synapomorphies = shared derived traits that evolved from a common ancestor and are passed down to its descendants.
These are the red dots on the phylogenetic tree!
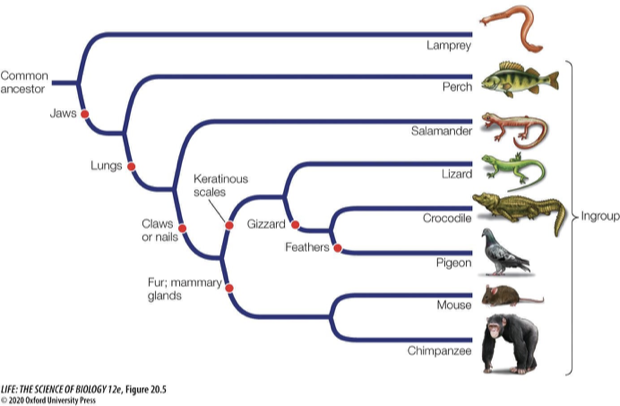
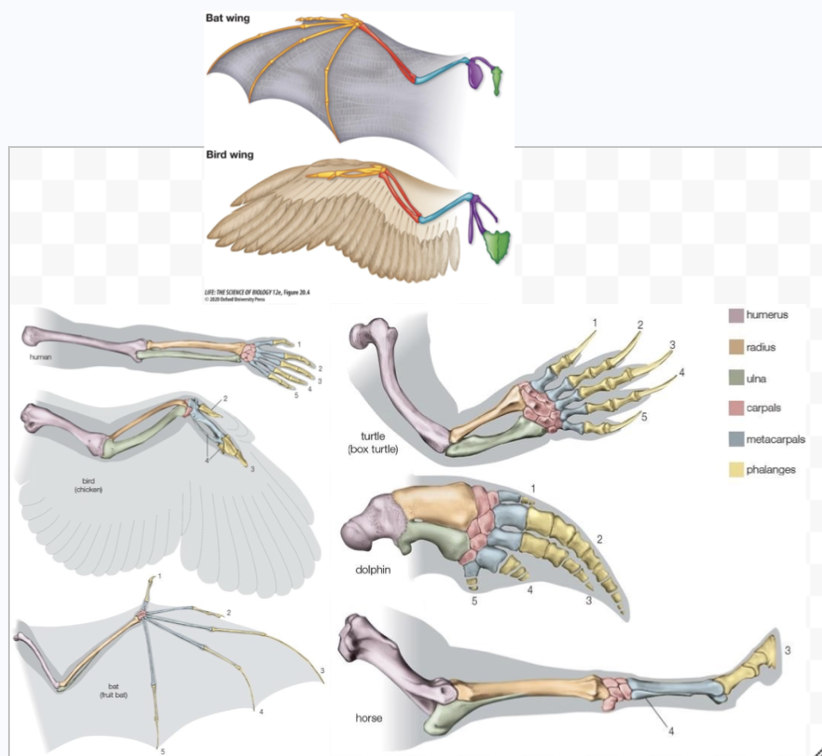
Explain homologous and analogous traits. What can analogous traits be an example of?
A similar trait function does not always involve shared ancestry
Homologous traits = features that come from the same ancestor.
Same-colored bones
Analogous traits = features that evolved separately/independently but do the same job and are intended for the same function
Convergent evolution
Like wings in bats and birds
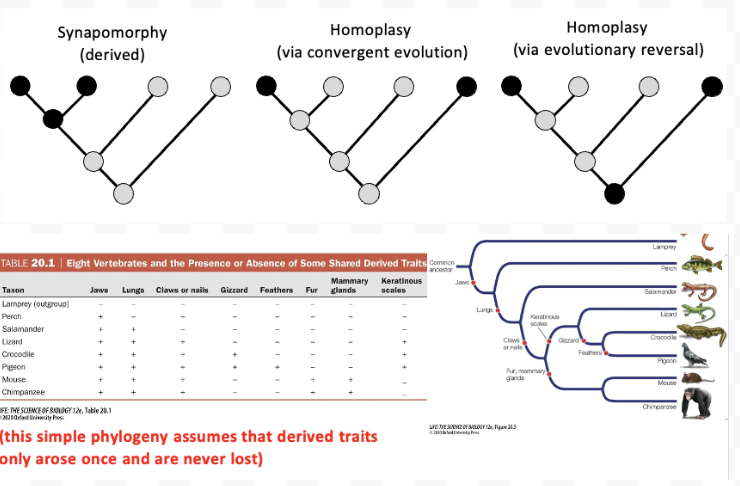
What are homoplasies? What is snapomorphy?
They are similar traits that arise from convergent evolution or evolutionary reversals, not ancestry
Shared derived trait from a common ancestor. normal evolution within a lineage.
“syn” + “apo” + “morphies” = shared + derived + form
Phylogenetic trees help scientists figure how relationships have _____. Thanks to these trees, ___ ____ can be reconstructed. This in turn shows how species have ____ over time, can help trace when ____ first appeared, and can show when groups ____ form a ___ ancestor
Evolved
Past events
Evolved, traits, split, common
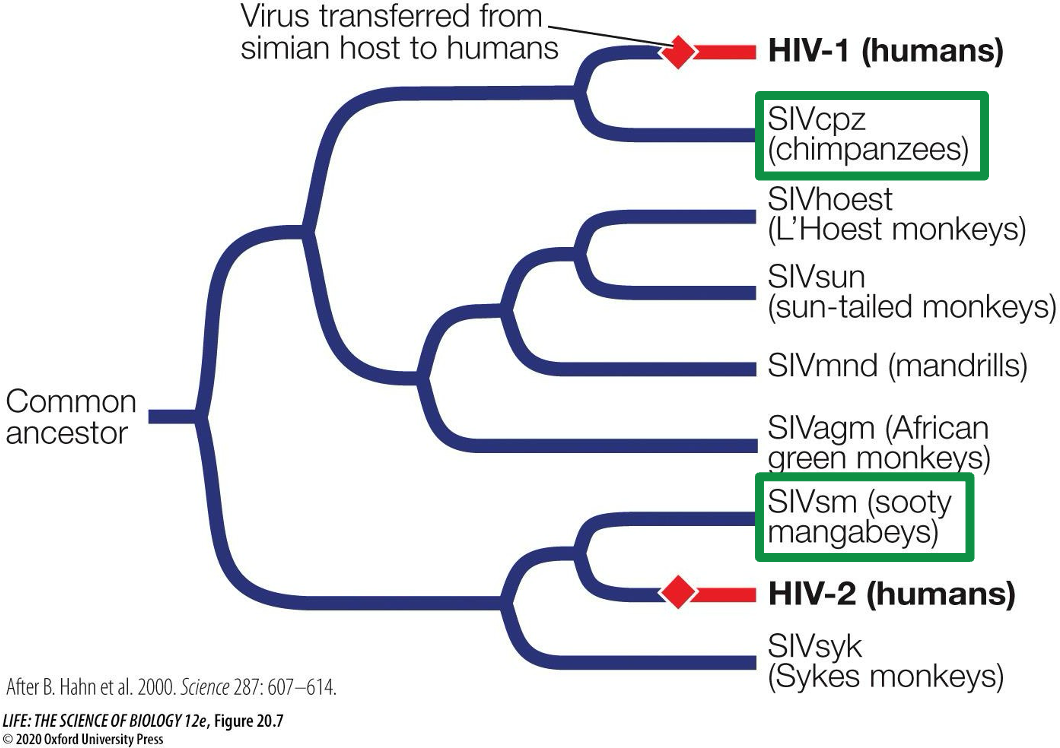
What was used to investigate HIV and why?
Phylogenetic analysis can help figure out where a virus came from
HIV likely came from a monkey host, but scientists do not know exactly which one it came from. Likely came from the sooty mangabeys and chimpanzees through blood transmission.
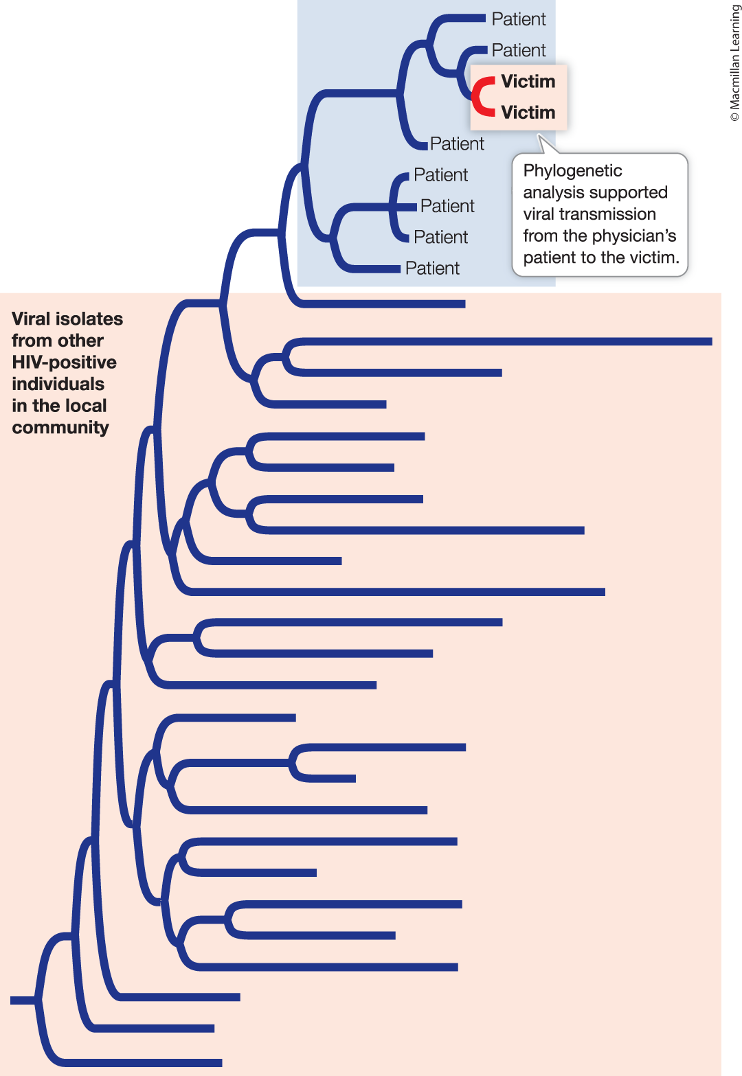
Phylogenetic trees can also be used in forensic investigations, explain with physician HIV.
The physician is accused of injecting blood from his HIV-positive patient into his girlfriend
Phylogenetic analysis can show if the HIV strain found in the girlfriend is present in a subset of his patients
Why are molecular clocks useful?
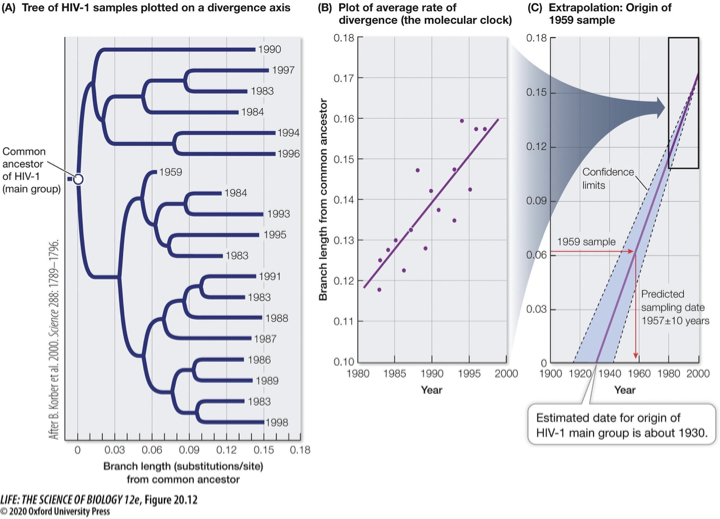
Molecular clocks use how fast DNA changes (mutation rate) to estimate when events happened in evolution.
Example: When did HIV jump from chimpanzees to humans?
Most samples of HIV-1 were collected from human specimens in the 1980s
Some samples from as early as the 1950s
Scientists can use the observed changes in HIV-1 over the past several decades to predict the common ancestor of all HIV-1 isolates
We can estimate when HIV-1 first entered human populations from chimps
Molecular clocks and closely related species
In closely related species, a given gene evolves at a reasonably constant rate
Not ALL genes evolve at the same rate
The corresponding protein accumulates amino acid replacements at a relatively constant rate: the rate of change is the molecular clock (slope of the line)
A molecular clock needs to be calibrated in the evolutionary timeline (e.g., fossil record, biogeographic dates)
Collect many HIV samples from different years, compare the number of substitutions and mutations (length exemplifies this), translate the tree into a line graph to show a molecular clock, and track this slope back in time to track when the virus jumped from monkeys to humans.
Visual chart for building a phylogenetic tree
Visual chart for building a phylogenetic tree
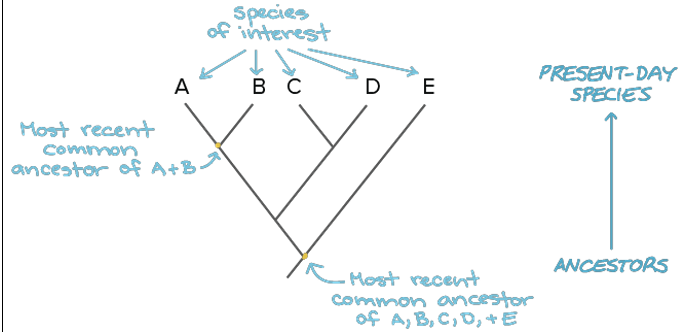
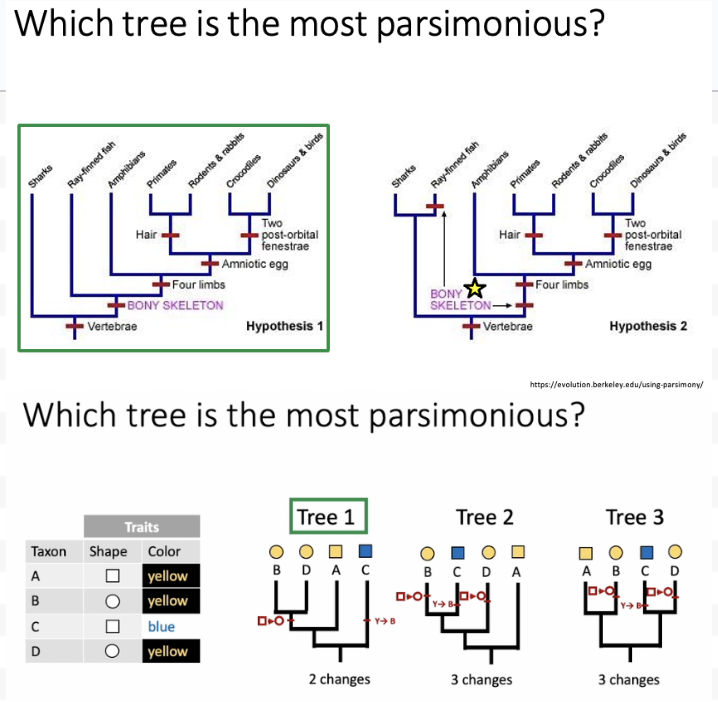
The principle of parsimony, explain.
The best phylogenetic tree is the one that assumes the fewest evolutionary changes.
Scientists prefer the simplest explanation
The best tree has the lowest number of independent trait changes
This avoids too many homoplasies (traits that evolved separately in different groups)
Phylogenetic trees can be built based on many traits
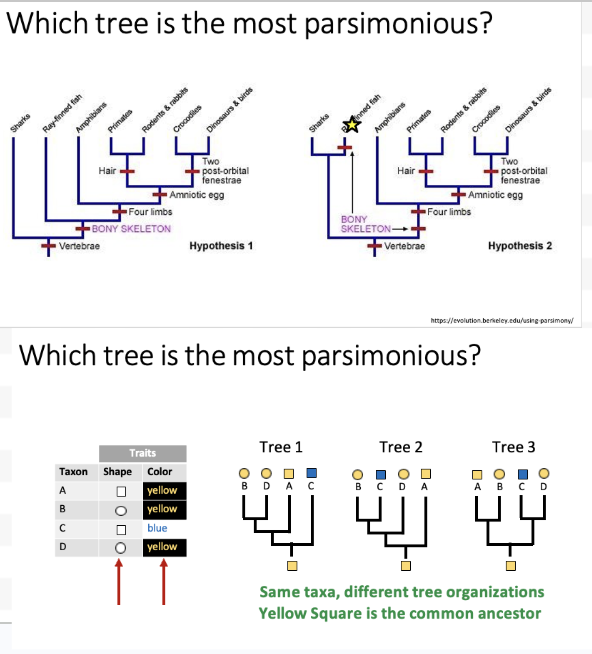
Parsimony practice question
The left tree because it doesn’t have as many red lines on it. The bony skeleton appears twice on the right vs only once on the left.
Tree 1 – 2 changes, a color change, and a shape change. Tree 2 – 3 changes: shape change, shape change, and a color change. Tree 3 – 3 changes: shape change, color change, and a shape change.
So tree #1 is most parsimonious!
Phylo trees can be based on many traits, explain.
Hundreds or thousands of traits from morphology, paleontology, development, physiology, ecology, behavior, and molecular data
Phenotype can be influenced by the environment
Using more data helps account for these possible changes
Results are usually not the same depending on the traits used, so a consensus tree (showing the most common characteristics) is computed
DNA and molecular data can be leveraged along with ecology, behavior, etc. Trees with multiple kinds of data produce different results, and all look different. Computer software can create a consensus tree to make this simpler.
Scientists can study evolution by comparing DNA and protein sequences. Explain how and why.
They compare large amounts of molecular data between ancestors and descendants to build the trees
Homologous DNA sequences (like nuclear or mitochondrial DNA) are aligned and compared
Helps with the study of gene evolution: mutations accumulate over time, at different paces across organisms
Mathematical models (like maximum likelihood) are used to find the tree that best fits the data
There are different models of evolutionary change, so this method is more flexible than parsimony
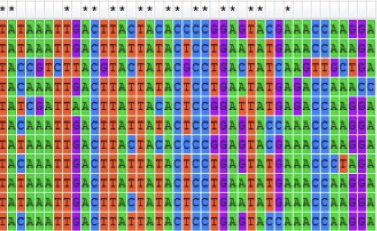
When comparing DNA sequences using a mathematical model, some changes can be missed or underestimated. Explain why this is.
Multiple substitution events might have happened, but they are now undetectable.
We do not know the ancestral sequence
BUT we can compare observed sequences and predict how these sequences have evolved from the ancestral sequence
In a mathematical model, we consider possible rates of change (e.g., transitions vs. transversions, introns vs. exons).
How would the mathematical model be carried out?
Count the number of differences at homologous nucleotide positions
Reflects the minimum number of nucleotide changes that have occurred since the two sequences diverged from a common ancestral sequence
BUT this count likely underestimates the actual number of changes that have occurred since the sequences diverged
Any given change counted by comparing DNA sequences may result from multiple substitution events that occurred at a given nucleotide position over time
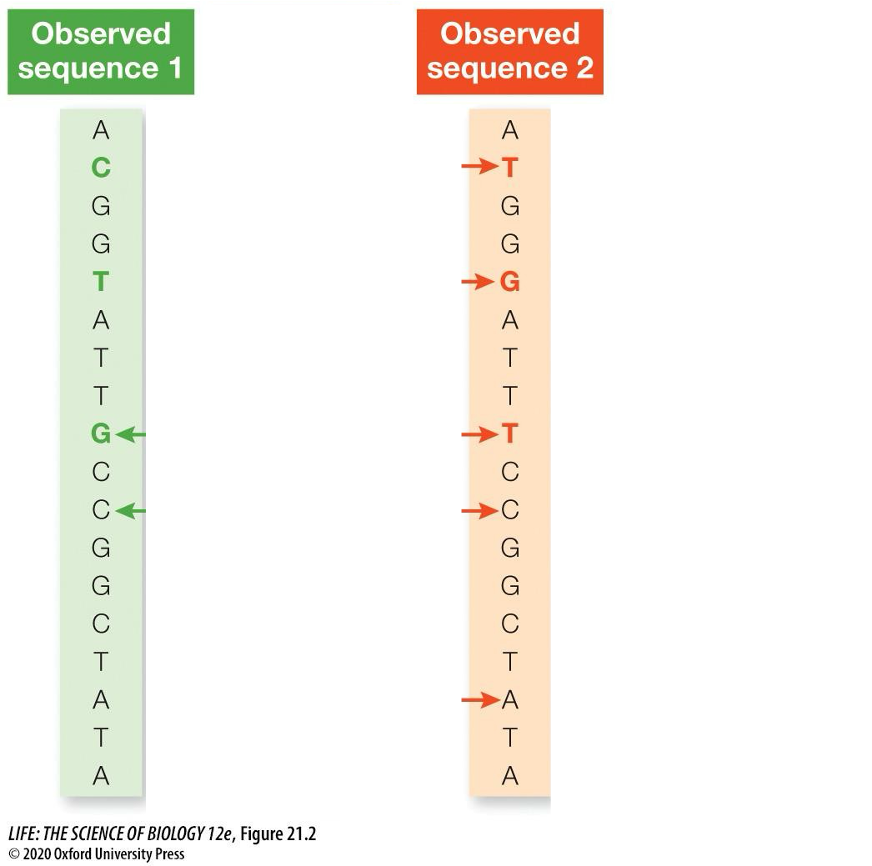
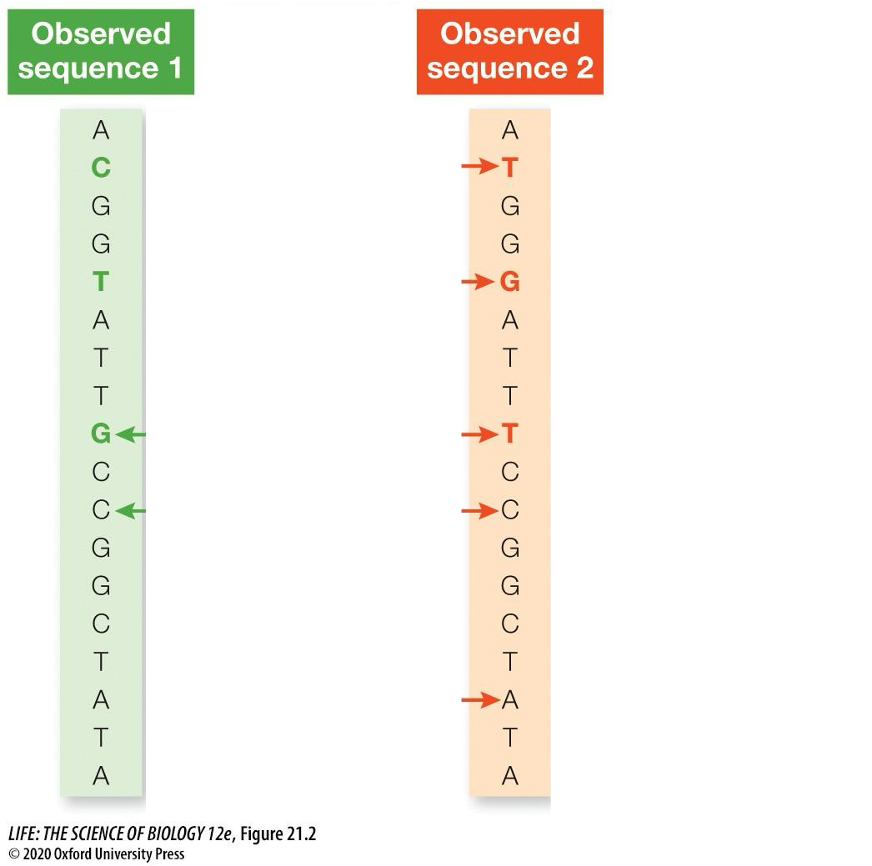
Continually using this approach, how is the ancestral sequence determined?
Two observed descendant sequences, which share a common ancestor, have undergone a series of substitutions
The two observed sequences (green, red) differ by only three nucleotides (colored letters)
BUT these three differences result from a total of nine substitutions (arrows)
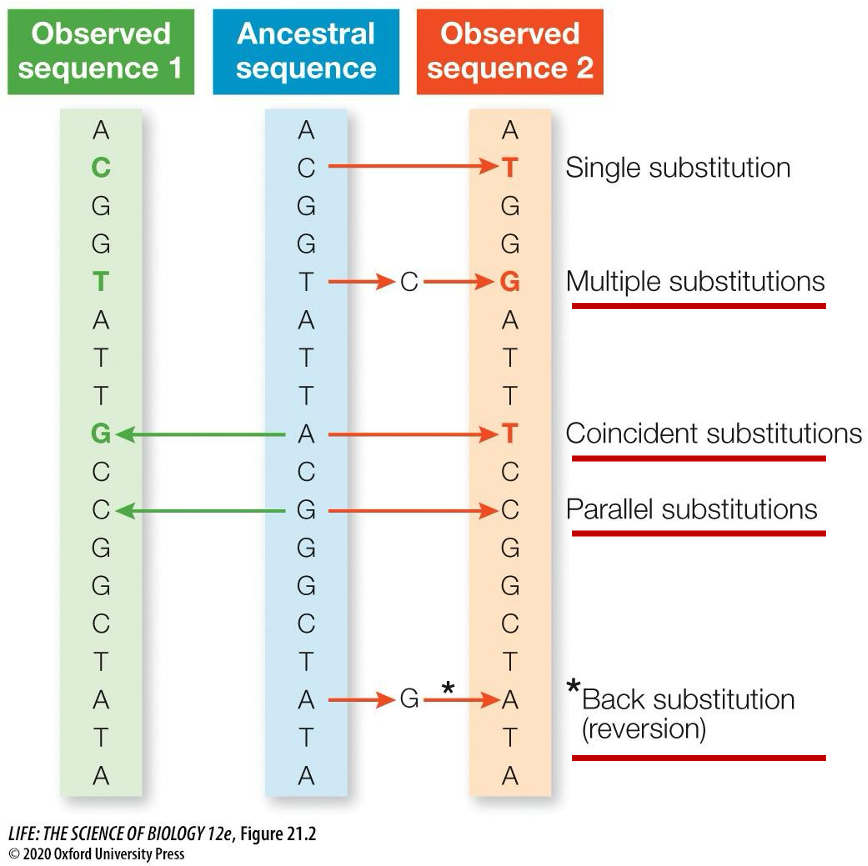
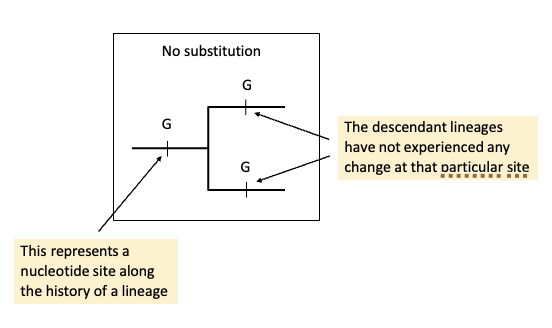
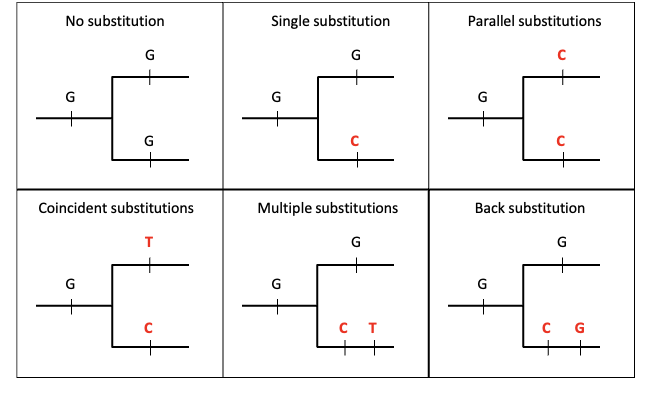
How can these substitutions be visually represented? No, single, parallel, coincident, multiple, and back.
Substitution Type | What Happens | Example in Diagram | Key Idea |
|---|---|---|---|
No substitution | The nucleotide stays the same throughout evolution. | G → G → G | No mutation occurs. |
Single substitution | One mutation happens on one branch of the tree. | G → C in one lineage | Only one evolutionary change. |
Parallel substitutions | The same mutation occurs independently in two separate lineages. | G → C and G → C | Example of convergent homoplasy. |
Coincident substitutions | Different mutations occur at the same site in different branches. | G → T and G → C | Multiple independent mutations. |
Multiple substitutions | Several mutations occur sequentially along the same lineage. | G → C → T | A site mutates more than once over time. |
Back substitution | A mutation occurs and later changes back to the original base. | G → C → G | Example of evolutionary reversal (homoplasy). |
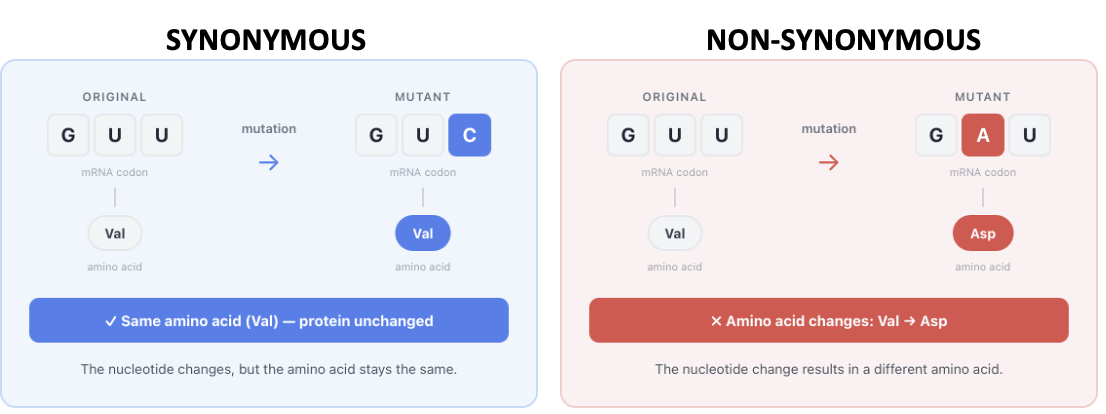
Genetic changes can affect organisms in different ways, synonymously explain.
Synonymous substitutions (silent mutations) are neutral to natural selection and don’t change the protein and usually don’t matter, so they build up over time
Non-synonymous substitutions do change the protein
Most non-synonymous substitutions are harmful and get removed (deleterious)
Some are almost neutral (small effect with similar AA or outside coding region)
A few are beneficial and can create a new or improved function (gain of function)
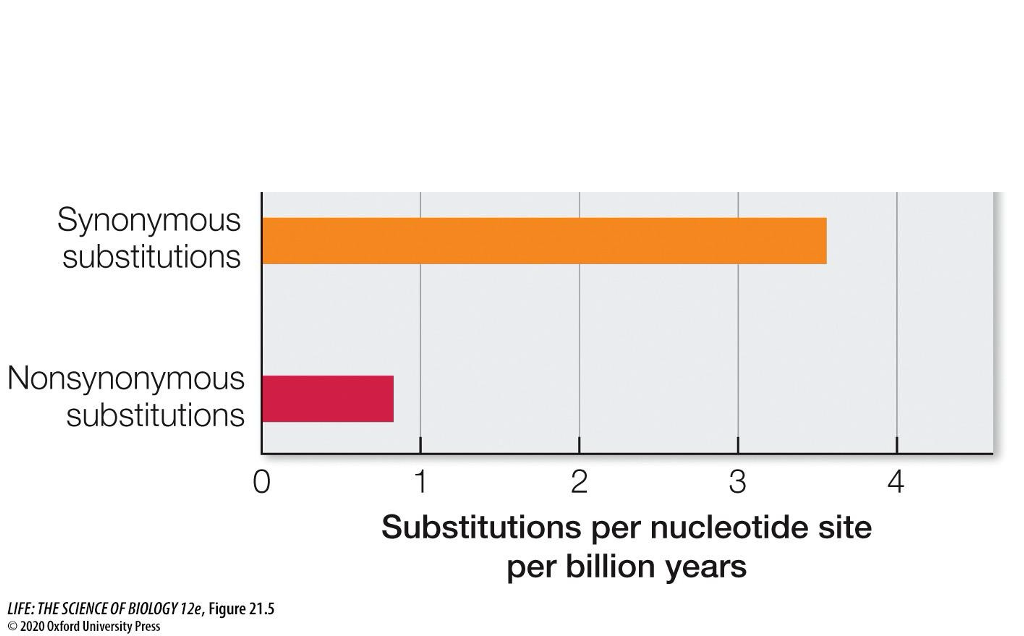
Selection in ____ among _____ populations or species can be detected by comparing the rate of _____ and ___ _____ substitutions.
Genes, different, synonymous and non-synonymous
What is advantageous in comparing the rates of these substitutions? What do they represent?
Can help detect natural selection
Rate of synonymous/*non-synonymous substitutions:
Compare the rate of non-synonymous changes to synonymous changes (dN/dS ratio)
If the rate is close to 1, the mutations have no effect on fitness (neutral to selection)
* = There are about 3 times more possible non-synonymous substitutions than synonymous ones
A few are beneficial and can create a new or improved function (gain of function)
What does it mean when one synonymous is more present than the other?
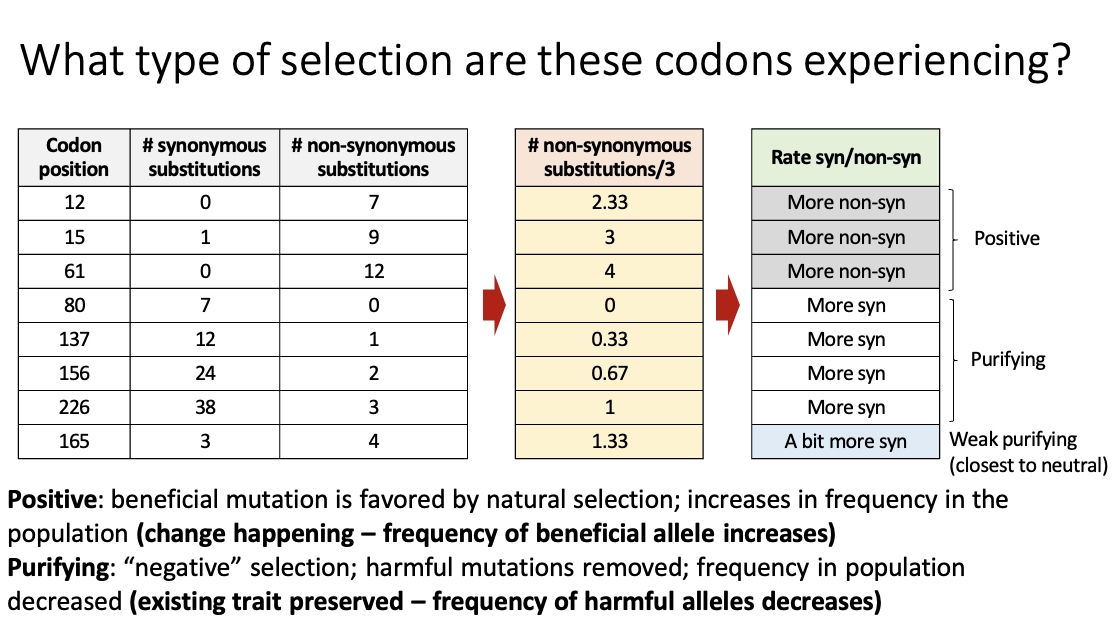
Synonymous < non-synonymous (more non-synonymous): positive selection
Beneficial mutations that change an amino acid are favored, changing the protein sequence, driving evolution
Synonymous > non-synonymous (more synonymous): purifying selection
Deleterious non-synonymous mutations removed; conserves protein sequence

What type of selection are these codons experiencing?
Codons experience answers
END OF CHAPTER 20
END OF CHAPTER 20
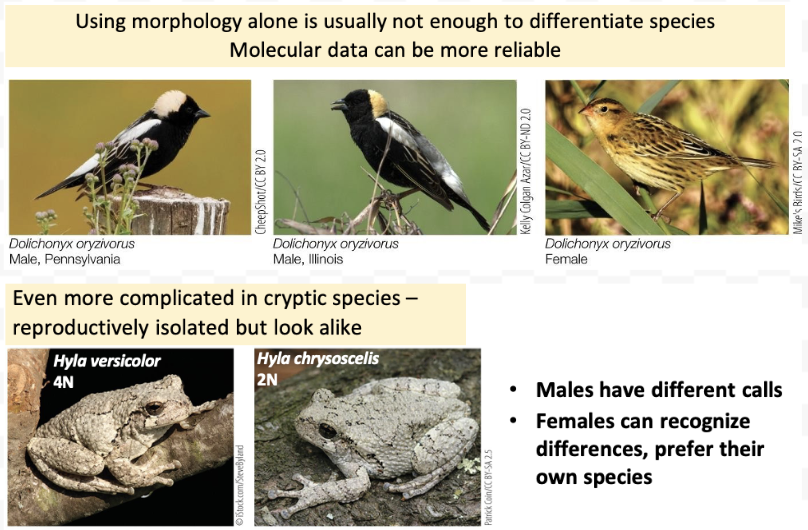
How is a species defined? - Physically/appearance. Drawbacks?
Morphological species concept
A species is a group of individuals that look alike
Linnaeus named thousands based on this
Drawback: members of the same species do not always look alike, an issue for cryptic species
How to define a sexually reproducing species on a biological level? What is the criteria and how does speciation play into this? Drawbacks?
A group of actually or potentially interbreeding natural populations that is reproductively isolated from other groups
A species is a group of interbreeding individuals that are reproductively isolated
Known as the biological species concept (via reproductive isolation).
Species result from speciation: divergence of lineages and emergence of reproductive isolation between lineages.
There’s no gene flow to make them their own species

How is a species defined? - Lineage-wise. What is it a series of?,
A species is a group of organisms that share a pattern of ancestry and descent
Forms a unique branch on the tree of life, emphasizing evolutionary history and distinct lineages
Ancestor-descendant series
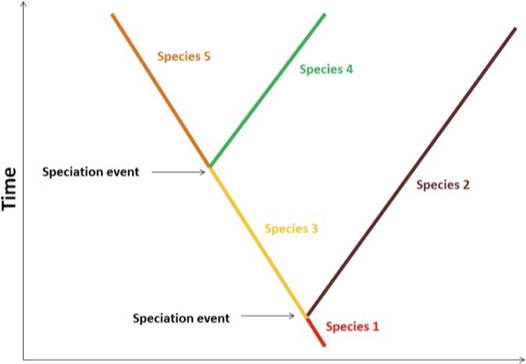
Start with the speciation event
End with an extinction event OR another speciation
Is gradual - could take thousands of years
How can reproductive isolation be reached? What is required for speciation to occur?
Speciation requires the interruption of gene flow (i.e., the sharing of genetic material) within a species that formerly exchanged genes
A result of a genetic change
Gene incompatibility can result in reproductive isolation
What is the Dobzhansky-Muller model and what does it explain?
Explains the evolution of reproductive isolation
Let’s start with an ancestral lineage and focus on two loci (1 and 2) with one allele each (a and b).

When is the ancestral lineage separated? What do they form?
Ancestral lineage is separated if there is a barrier to gene flow
In this example, the isolation is caused by a mountain range (geographic isolation)
They form independently evolving descendant lineages
•New mutations and alleles can independently arise in separate loci.
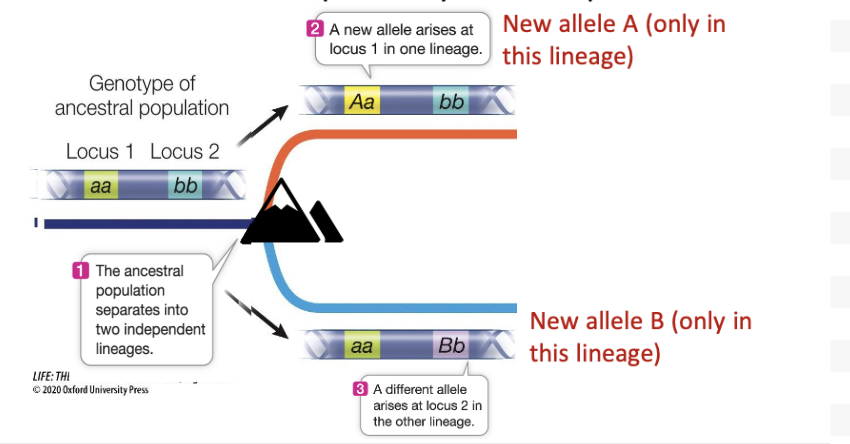
What happens over time as a result of this separation?
The loci can experience allele fixation over time
Loss of polymorphism (ex. AA, not Aa), resulting in a single allele in a locus (just a single A or B)
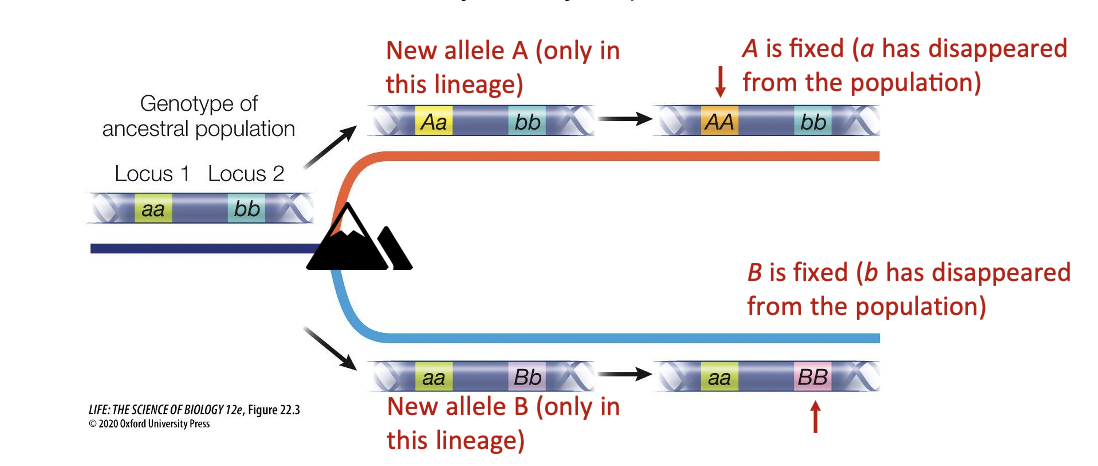
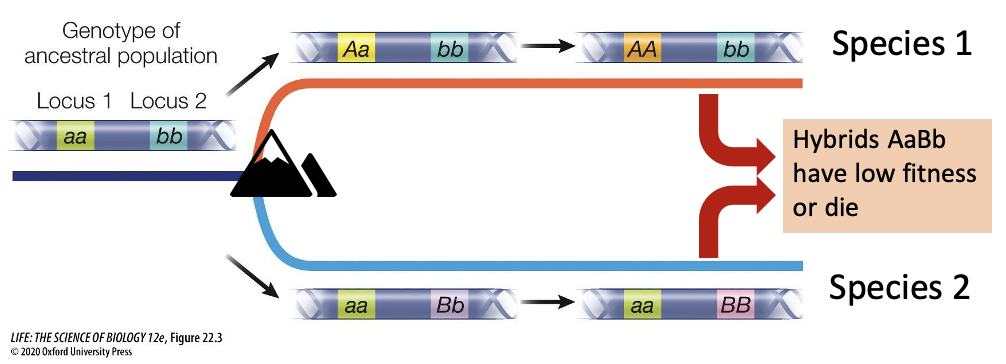
What happens now if the two lineages try to interbreed?
They have now become incompatible, as new alleles have never interacted (hybrids have low fitness or die
A combination of AB within the same individual is lethal or causes low fitness
Two separate species have formed, as now the A or B alleles will not spread throughout the other respective lineages
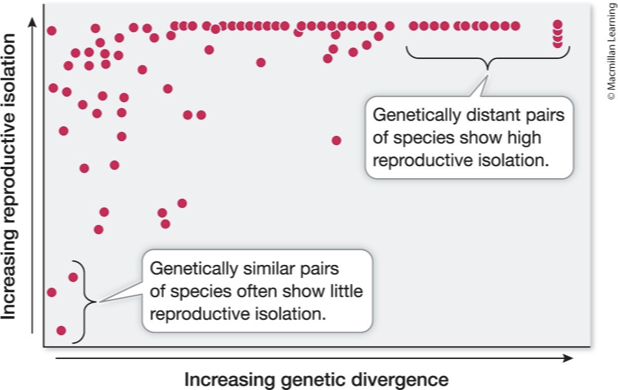
What does reproductive isolation increase with over time? What is it not?
Reproductive isolation increases with genetic divergence
Some species experience isolation with FEW genetic changes (left on the x-axis)
BUT others need more genetic divergence to achieve reproductive isolation (right on the x-axis)
Gene incapability drives this process, but it’s not one size fits all.
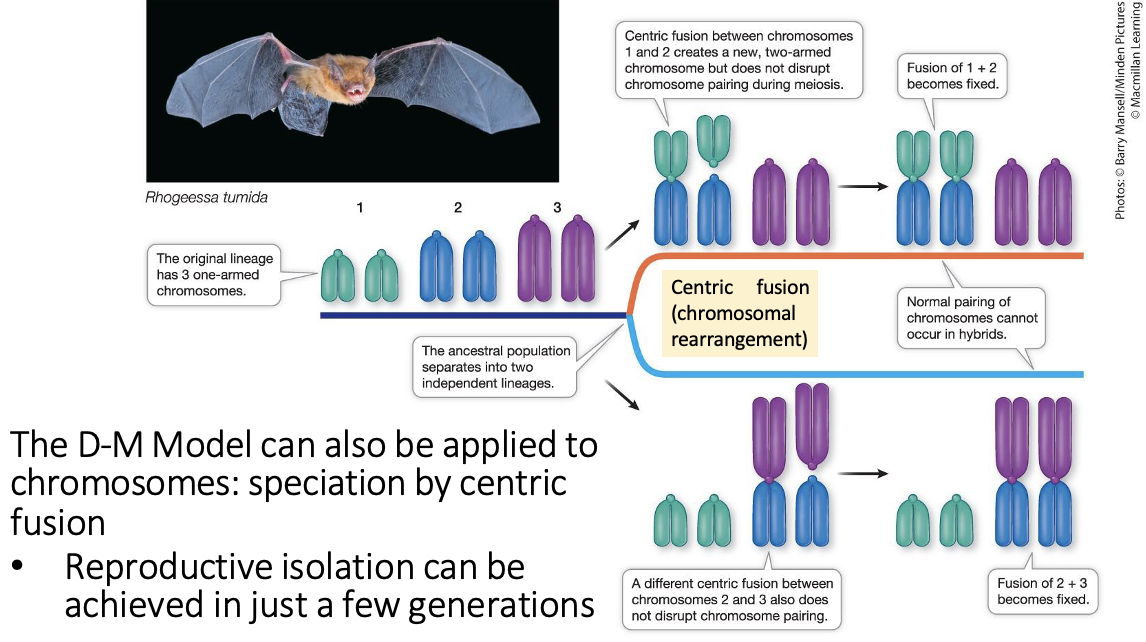
What else can the model apply to? How is it caused, and what is achieved? What are these unique as well?
Can apply to chromosomes
Speciation by centric fusion
Chromosomal rearrangement
Fusion doesn’t stop viable sex cells, but could stop breeding. Could not produce a functional zygote.
Isolated achieved in only a few generations (fatser than the DM)
Chromosomes are acrocentric: centromere very close to one end, not in the middle
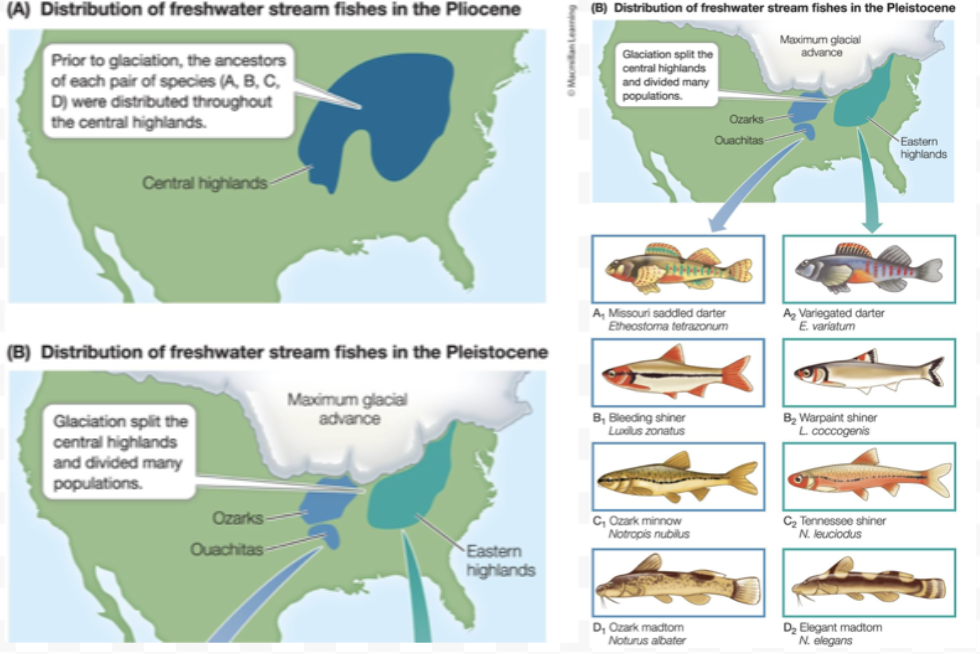
Speciation processes - Allopatric. How? When? What happens?
Occurs due to physical barriers (of inhospitable habitat) that separate a population
The most common mode of speciation
Usually over geological time (continental drift, sea level shifts)
Separated populations experience a mutation, genetic drift, and local adaptation
Lineages split, and reproductive isolation may arise
As a result, = Observe pairs of closely related sister species
Disruption of a population's habitation with something that is inhospitable. Glaciation split the central highlands into 2. Habitat in between became inhospitable, and gene flow became limited between the two populations. Will also adapt to their new local habitat, forming distinct species. Sister species very closely related.
Those (fish) are the sister species. They are different but also the same. Closely related but can’t interbreed.
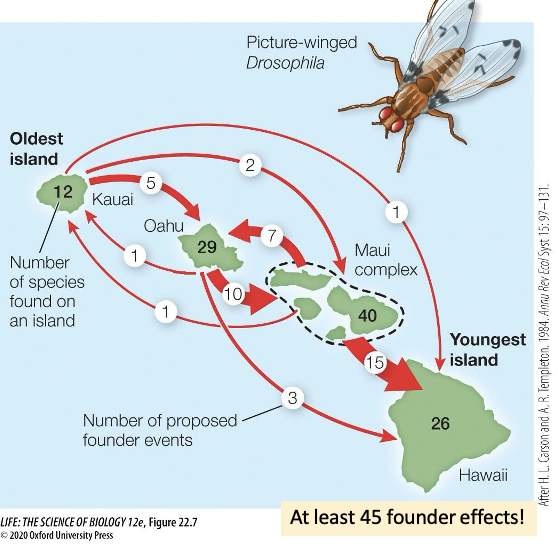
How else can allopatric speciation occur? Explain this.
Founder effects
A group of individuals crosses an existing barrier (like islands) and establishes a new population
Each colonization generated a new, isolated population
Genetic drift, mutation, and local adaptation occur in the founder population
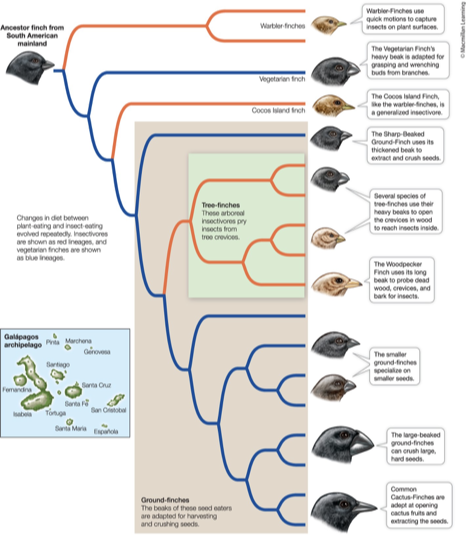
What is one of the most famous examples of the allopatric founder effect? What happened?
Darwin’s Finches!
Species across the Galapagos Islands differ greatly
Both from their ancestor and from one another
The islands are far apart, have unique and distinct geography and climate
Feeding specializations (i.e., beak size and shape) arose on these different islands to meet the environments of each island
Speciation processes - Sympatric. How? What are the two main mechanisms?
Occurs without physical barriers
Disruptive selection (resulting in ecological isolation)
Polyploidy
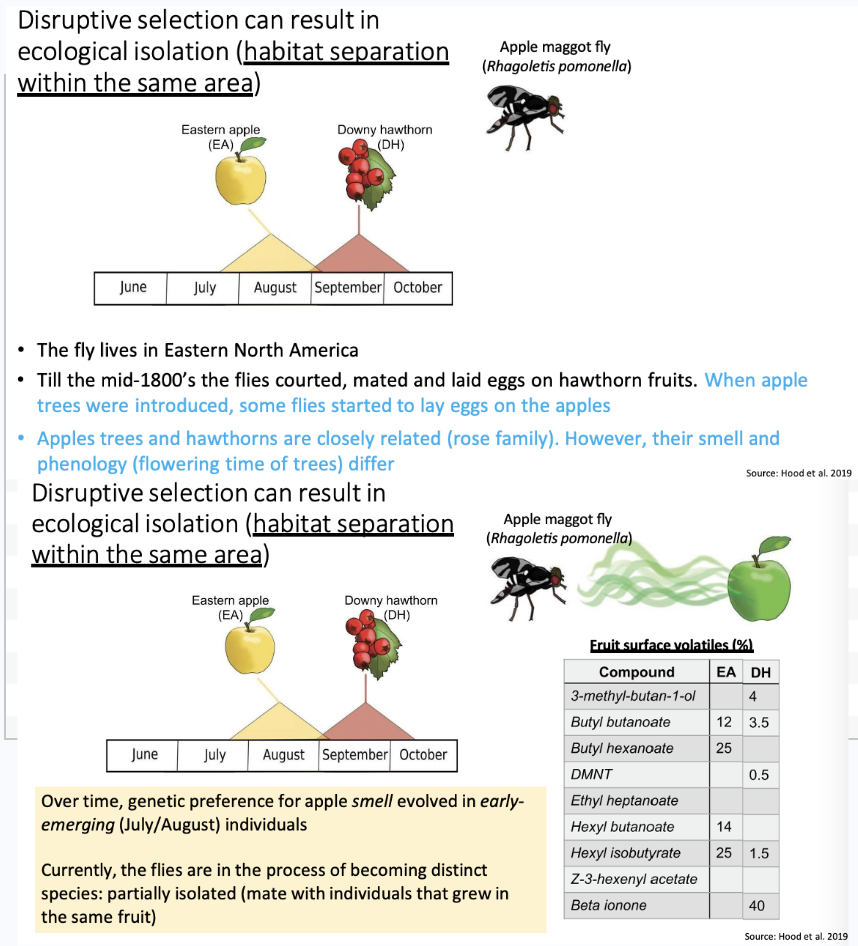
What can disruptive selection result in? What is the fly example?
Results in ecological isolation
AKA habitat separation within the same area
The chart reflects when the tree flowers and goes on to produce fruit.
The apple has earlier flowering, and so over time the flies will develop a smell preference for one or the other, and hence drive to mate in breed on the apple earlier or later on the downy hawthorn.
A change in when and where they are breeding causes reproductive isolation over time.
Nearing becoming a distinct species today.

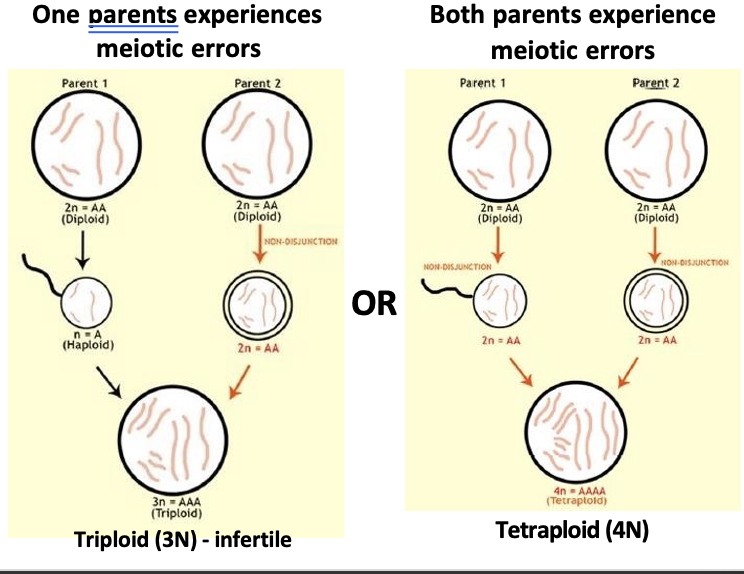
What does polyploidy cause? What is autoployploidy?
The phenomenon of when homologous chromosomes fail to separate from one another
This can cause speciation in just one generation
Autoployploidy
Two of the same species breed
One or both parents experience a meiosis error → A gamete has twice as many chromosomes as normal
Results in an offspring with chromosomal duplication. (3N or 4N vs 2N).