Population Genetics
1/29
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
30 Terms
What do the pie charts represent in regards to black tip sharks collected in the gulf and other side of Florida?
each color in the pie chart = unique genetic form of mtDNA
Haplotype distribution tells us that there’s distinct populations which we may not expect
More diverse (genetic forms) in the Gulf than Atlantic
What are the most common genetic markers?
mitochondrial DNA
Microsatellite DNA (chromosomal) or nuclear DNA
Characteristics of mtDNA
2,000 base pairs in sharks - circular strand
12s & 16s regions are conserved → they are unique to each particular species
D-loop: mutates easily, even in same species can be different
Analysis of mtDNA
determine individual genetic IDs
Sequence DNA
Determine hapotypes by aligning & comparing sequences
Determine polymorphic nucelotide positions (looking at variable positions in the sequence) and assigning haplotypes
Haploid & mtDNA
A haplotype - mtDNA is haploid because it is only inherited maternally
an individual is an “a” rather than “a”
Represents a unique nucleotide sequence for specific region in mtDNA genome
PCR
Polymerase chain reaction
use a thermocycler
Heat up and pull apart strands→ creates 2 copies
Cool down and repeat the cycle→ amplifies the region (or strands) that are being copied which makes lots of copies
Use TAQ polymerase to stick the strands together → Thermus aquaticus (a high temp resistant bacteria)
What is mtDNA used for?
Determining stock/population structure or philopatry homing by comparing haplotype frequencies
important for management and fishery regulations
What are the three hypothesis for genetic diversity for black tip sharks
Separate population at each location
Female philopatry→ females never leave location even if they mate w/males from other locations, the populations aren’t separate then bc interbreeding but it wouldn’t show up in mtDNA bc that isn’t passed on from the male
Regional natal homing→ females could migrate but come back & pup where they were born
if females stay there will be region specific mutations & genetic drift
Haplotype Diversity
Chance that 2 randomly selected individuals have different haplotypes

Figuring out haplotype diversity?
Which pops have high intermediate or low diversity?
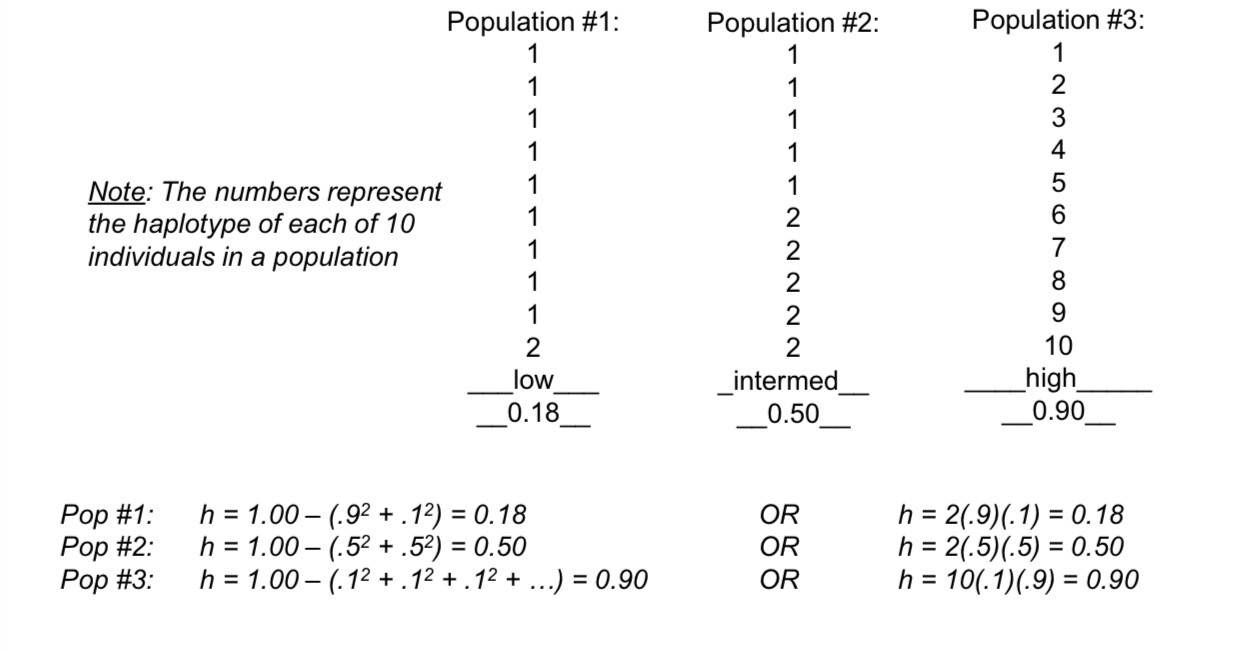
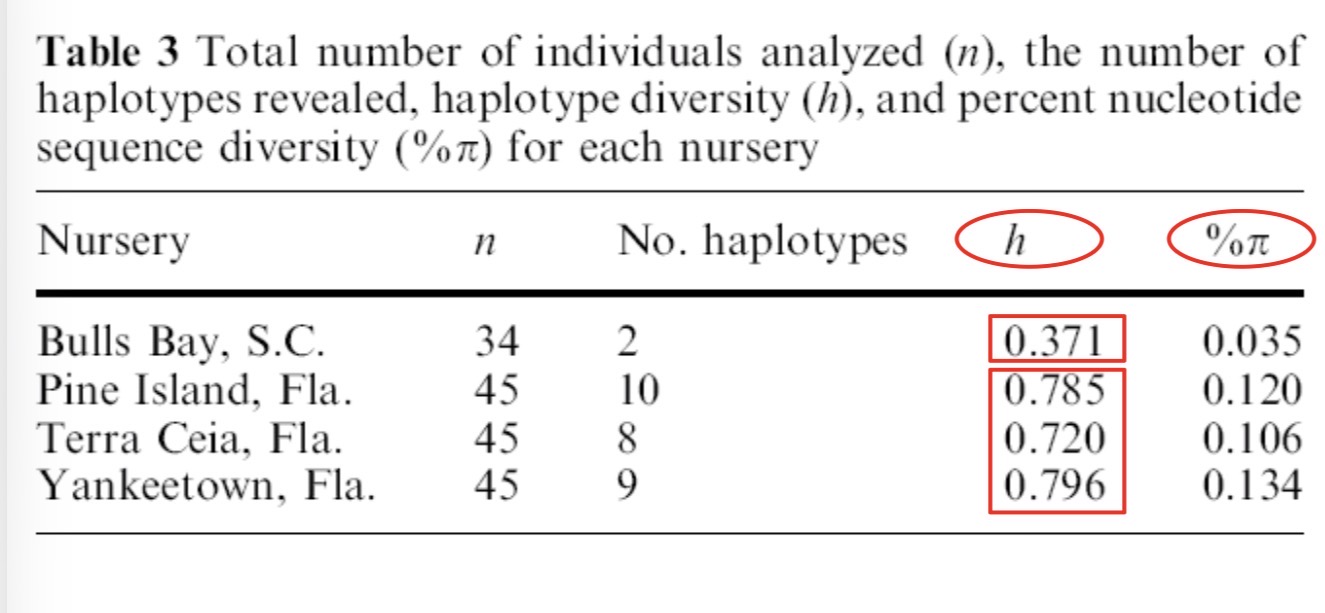


From the paper about Blacktip shark haplotypes
What is the probabity of 2 sharks from Bulls Bay, SC in 2000 having different haplotypes?
Unlikely
SC haplotype diversity= 0.371
Florida Gulf Side= 0.710 (way higher)
10 polymorphic sites out of 1000 base pairs, 990 of those positions were same in nucleotides
Conclusions from black tip paper
10 polymorphic sites out of 1000 base pairs, 990 of those positions were same in nucleotides
Did not detect significant structuring of haplotypes among 3 gulf nurseries→ did btw Gulf & Atlantic
They know that they seasonally migrate→ they think female philopatry for nursery areas which means nursery areas should be treated as separate management areas
Microsatellites
Short nucleotide sequences repeated over & over (ATATAT..) noncoding regions
repeated different number of times on homologous chromosomes
Microsatellite locus flanked by gene loci
There could be individual genotypes such s 8/12, 8/14, 16/16,… (Lots of diff combos)
What does it mean that microsatellites re hypervriable regions?
In Lemon sharks, looked at 4 loci→ range of 19-43 alleles at each locus→ high number of alleles t each locus (in western Atlantic lemon sharks)
Hyper variable!!!
How to use microsatellites to determine paternity?
using two siblings and potential dad
For each individual look at a chart of genotypes found at multiple loci

Can use microstellites to determine mating system
if you compre the genotypes of the embryos to the moms genotype and then identify which allele the embryos have from the moms you will be left with all the alleles that came from the dad
Depending on how many different alleles there are can help determine number of mates the mom had
Newborn bonnetheads in public aquarium found from a tank that only had 3 juvenile female bonnetheads raised from birth and 1 male sandbar shark
What type of genotype did the pup had and what are the implications for paternity?
they found that the pup that was born was homozygous at each loci
If one allele came from a dad it would have had to be the exact same s the mom and the chances of that are really low→ also found one of the alleles to be super rare so even more unlikely the dad would have it
Why is the pup homozygous for all loci when it comes from a mom that is heterozygous in only some loci?
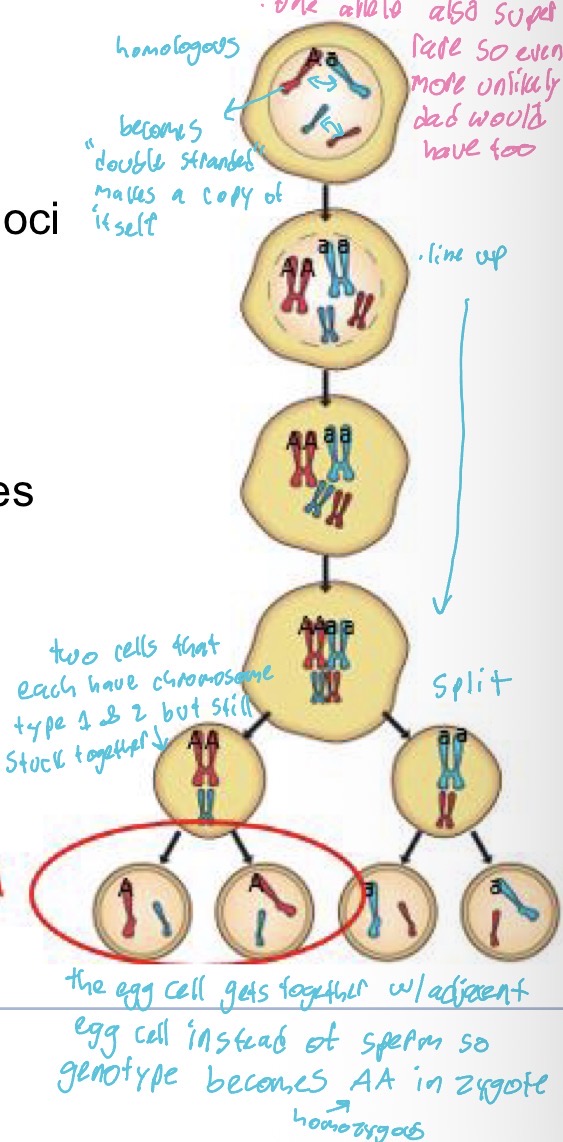
(Explain steps of cell division)
automictic parthenogenesis produces high levels of homozygosity
Have homologous chromosomes that become “double stranded” and makes a copy of themselves so A & a
Then they line up with eachother so is AA & aa and go through division process (split along the middle)
Forms two cells that each have a chromosome type 1 & 2 but theyre each still stuck to together
Then the egg cell gets together w/adjacent egg cell instead of sperm so genotype becomes AA in zygote (homozygous)
Inbreeding depression
Greater chance both offspring will express recessive harmful allele if both their parents are siblings or related that carry it but is being covered up by a dominant allele
What are the 3 reasons populations will different genetically?
Random mutations- mutations on both sides of geographic area that is split will be random so lead to different genetics
Genetic drift- random changes in allele frequencies
Natural selection- 2 different environments so different alleles & genes may be found to be more well suited and prevalent
What do we expect to see when comparing allele and genotype frequencies geographically?
with no barrier to gene flow in a well-mixed population, gene and genotype frequencies are the same on both sides (proportions of genes are the same)
With a barriere, genetic differences develop between populations due to mutations, genetic drift, and natural selection
What are the characteristics and conditions of a Hardy Weinberg Population?
Predictable genotype frequencies
Unchanging genotype frequencies (across many generations)
Conditions:
Large population- no genetic drift
No migration (no immigration)
No mutation
No natural selection
Random mating- a well-mixed population, therefore if genotype frequencies are predictable it implies a singe wel-mixed population
How to calculate hardy-Weinberg?
predictable genotype frequencies
Calculate p² (AA), 2pq (Aa), q² (aa)
p= proportion/ frequency of A allele 0.40 = (0.40)²
Q= proportion of a allele, 0.60= (0.60)²
2pq= 2(0.6)(0.4)
Haplotype diversity
Measure of mtDNA genetic diversity
Genetic Variability
Proportion of individuals in the population/sample that are heterozygous
heterozygosity is the most common measure of genetic variability for nuclear (chromosomal) DNA like microsatellites
Pops with lots of heterozygotes have lots of genetic diversity
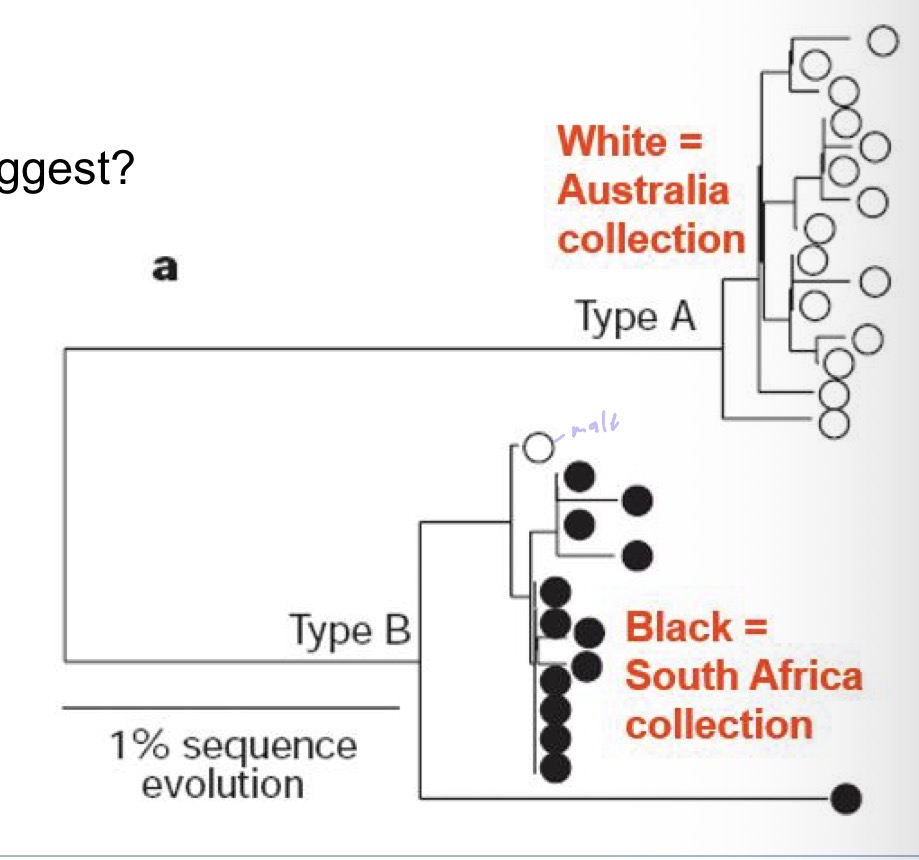
Paper about white sharks in Australia & South Africa mtDNA
What does the data suggest
either 2 different pops
Or females migrate
Or male swam to africa
What did they actually find?
Used locus, alleles, and observed heterozygosity and found that there was no significant difference btw the loci so none between the populations
What happens is that females swim back and forth between Australia & Africa but the individuals have the same mtDNA because of natal homing, so the nuclear DNA is different but not mtDNA so the females swim across the ocean and back to pup
Why study “low genetic differentiation across Three major ocean populations of the whale shark”?
Because they are swimming and mating with eachother across the three major oceans so there is a need to globally protect them not just have individual regional protections
used # of alleles, observed heterozygosity, only one locus different from hardy wiengburg equilibrium implying one pop in three oceans
Lemon Sharks in Bimini- natal homing
every year in Bimini researchers caught every newborn pup and figured out microsatellites DNA - 6 females pupping in Bimini were born there,
Were able to figure out genotypes of moms & dads, know every year how many moms there were
Also found that the moms were coming back & pupping where they were born with a reproductive cycle every two years and geographic distinction btw north & south Bimini → 2 were directly capture when pupping and their genotype matched genotypes of pups caught decades earlier, about half pupping females were born in Bimini (choosing to come back)
The homing mechanism is unknown but lemon sharks have been shown to be capable of homing behavior
First demonstration of natal homing in an elasmobranch!!!