micro exam 3
1/65
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
66 Terms
what are the two main divisions of metabolism? compare.
catabolism: breaks down molecules
anabolism: assembles macromolecules
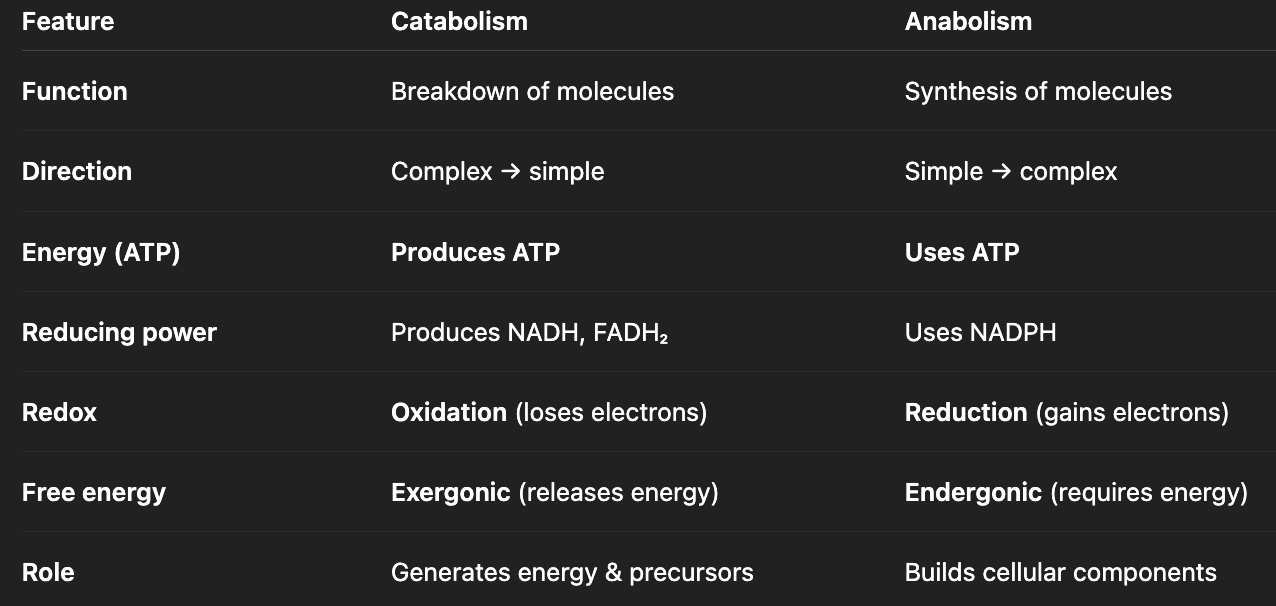
how are catabolism and anabolism coupled processes?
catabolism: provides ATP, reducing power, and building blocks
anabolism: uses them to build cellular components
what are the 12 key precursor metabolites?
glycolysis (6): G6P, F6P, G3P, 3PG, PEP, pyruvate
TCA (4): acetyl-CoA, a-ketoglutarate, succinyl-CoA, oxaloacetate
PPP (2): E4P, R5P
which precursors are involved in peptidoglycan synthesis?
NAM-NAG
F6P, PEP, acetyl-CoA
what does a negative vs. positive reduction potential signify?
negative (-) reduction potential: donates electrons
positive (+) reduction potential: accepts electrons
what are the sources of energy for catabolism?
phototroph: uses light to generate ATP via. photophosphorylation (e.g., photosynthesis) (light → excited electrons → PMF → ATP)
chemotroph: uses chemicals to generate ATP via. oxidation reactions (electron carriers → ETC → PMF → ATP)
what are sources of electrons for catabolism?
lithotroph: inorganic electron donors; low energy yield; may require reverse electron flow to make NADH (e.g., H2, NH3, H2S, Fe2+)
organotrophs: organic electron donors; high energy yield; (e.g., glucose)
what are sources of carbon for catabolism?
autotroph: CO2, carbon fully oxidized; requires ATP and NADH for carbon fixation
heterotroph: organic carbon; carbon already reduced and easily used in biosynthesis
what type of metabolism type are humans?
chemoorganoheterotrophs
chemo-
organo-breaks down nutrients for energy
heterotrophs-
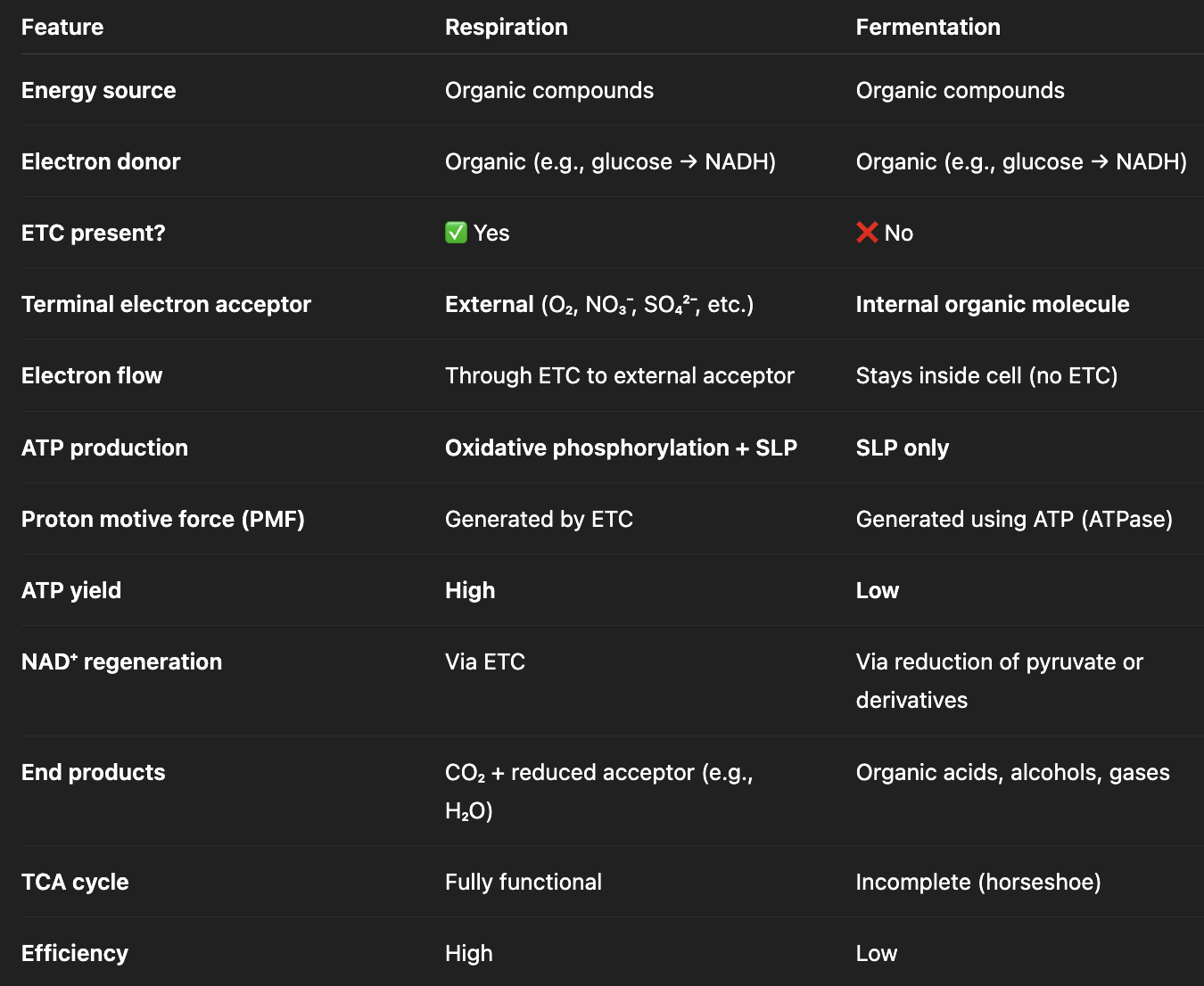
Compare and contrast respiration and fermentation as chemoorganotrophic fueling processes.
key similarities:
use organic compounds for energy + electrons
both start with glycolysis → pyruvate
both must regenerate NAD+
key differences:
uses external electron acceptor vs. uses internal electron acceptor (e.g., pyruvate)
oxidative phosphorylation & substrate-level phosphorylation (presence of ETC) vs. ONLY substrate-level phosphorylation (absence of ETC)
PMF generated from ETC vs. ATPase (reverse)
high vs. low ATP yield
fully functional vs. horseshoe TCA cycle
fully oxidized carbon vs. partially oxidized
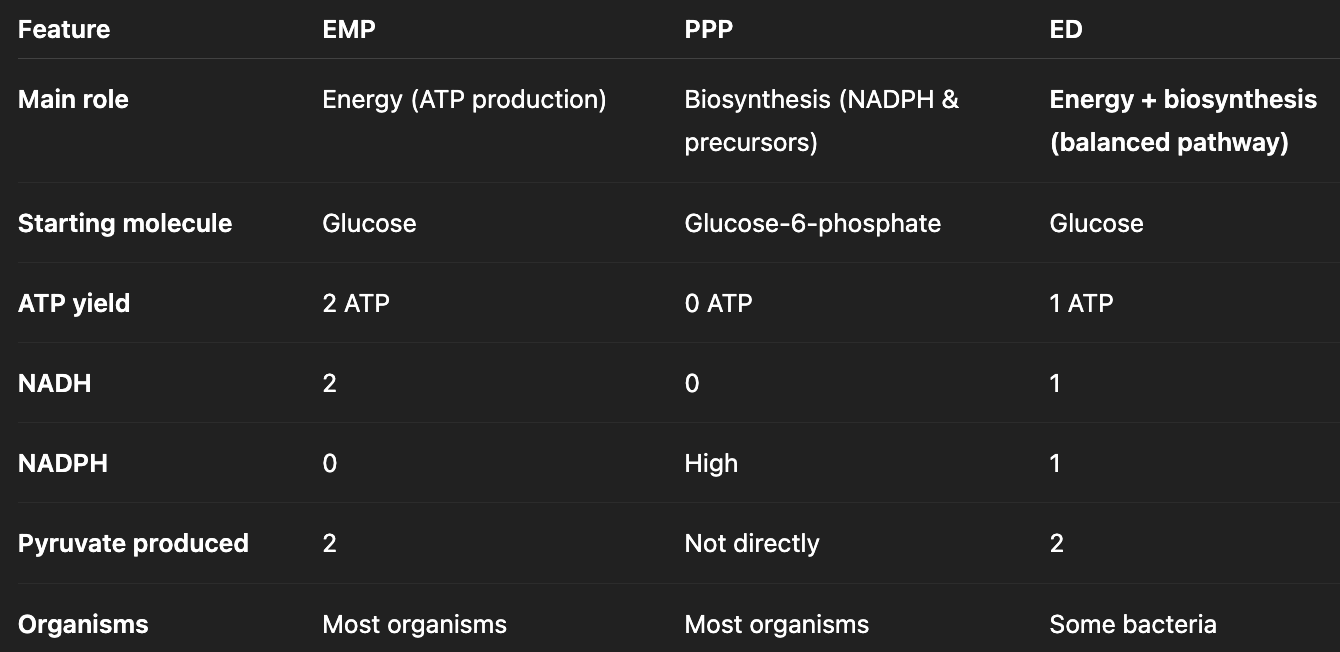
what are the three glycolytic pathways?
**all glycolytic pathways lead to producing pyruvate
**pyruvate can feed into TCA cycle, fermentation products, amino acid synthesis, fatty acid synthesis, etc
Embden-Meyerhof-Parnas (EMP) Pathway
Pentose Phosphate Pathway (PPP)
Entner-Doudoroff (ED) Pathway
What is Embden-Meyerhof-Parnas (EMP) Pathway?
purpose: energy production (main pathway); produce precursor metabolites that contribute to anabolic/biosynthetic pathways (e.g., amino acids, phospholipids, NAM-NAG)
input: glucose, 2 ADP + 2 Pi, 2 NAD+
output: 2 pyruvate, 2 ATP (net), 2 NADH
6C Phase (investment): glucose (C6) → 2 phosphoglyceraldehydes (aka PGALD, G3P, C3)
3C Phase (payoff): oxidation of G3P → pyruvate
What is Pentose Phosphate Pathway (PPP)?
purpose: biosynthesis of precursors for nucleotides + amino acids & reducing power (NADPH)
does not directly generate ATP but can feed intermediates (F6P, G3P) into glycolysis
input: glucose-6-phosphate (G6P), NADP+
output: ribose-5-phosphate (R5P), NADPH (major product-reducing power), CO2, intermediates (F6P, G3P) **NO ATP production
Stage 1. oxidation-decarboxylation of G6P → Ru5P
Stage 2: isomerization of ribulose Ru5P → Xu5P + R5P
Stage 3: sugar rearrangement reactions → F6P + G3P (glycolysis), R5P (nucleotides, histidine), E4P (aromatic amino acids)
What is Entner-Doudoroff (ED) pathway
purpose: alternative glycolysis pathway (similar to EMP) specific to prokaryotes; mix of energy synthesis + biosynthesis
input: glucose, ADP + Pi, NAD+ + NADP+
output: 2 pyruvate, 1 ATP (less efficient), 1 NADH, 1 NADPH
used when growing on aldonic acids (gluconate, galactonate)
glucose → G6P → 6-phosphogluconate
6-phosphogluconate → KDPG
KDPG splits into 1 pyruvate (amino acid synthesis) + 1 G3P (lipid synthesis)
G3P enters bottom half of EMP → pyruvate
what is pyruvate oxidation?
purpose: produce precursor (Acetyl-CoA) for TCA cycle
input: pyruvate, NAD+, CoA
output: acetyl-CoA (precursor metabolite for TCA cycle), NADH, CO2
enzyme complex: pyruvate dehydrogenase
what is TCA cycle?
purpose: regenerate reducing power (NADH, FADH2) for ETC; provides intermediates for biosynthesis/anabolic pathways (e.g., amino acids)
input: 1 acetyl-CoA (2C), 3 NAD+, FAD, GDP/ADP+Pi, oxaloacetate (regenerated, not consumed)
output: 2 CO2, 3 NADH, 1 FADH2, 1 GTP/ATP, CoA (recycled)
what are anaplerotic pathways?
Anaplerotic pathways: replenish TCA cycle intermediates that have been used in biosynthesis, ensuring the cycle can continue functioning
**pyruvate → oxaloacetate
amino acid → TCA intermediates (e.g., glutamate → a-ketoglutarate)
what are the two types of respiration?
**terminal electron acceptor differs but both use ETC
aerobic respiration: final electron acceptor = O2, high energy yield
anaerobic respiration: final electron acceptors = NO₃⁻, SO₄²⁻, Fe³⁺, CO₂, etc; lower energy yield
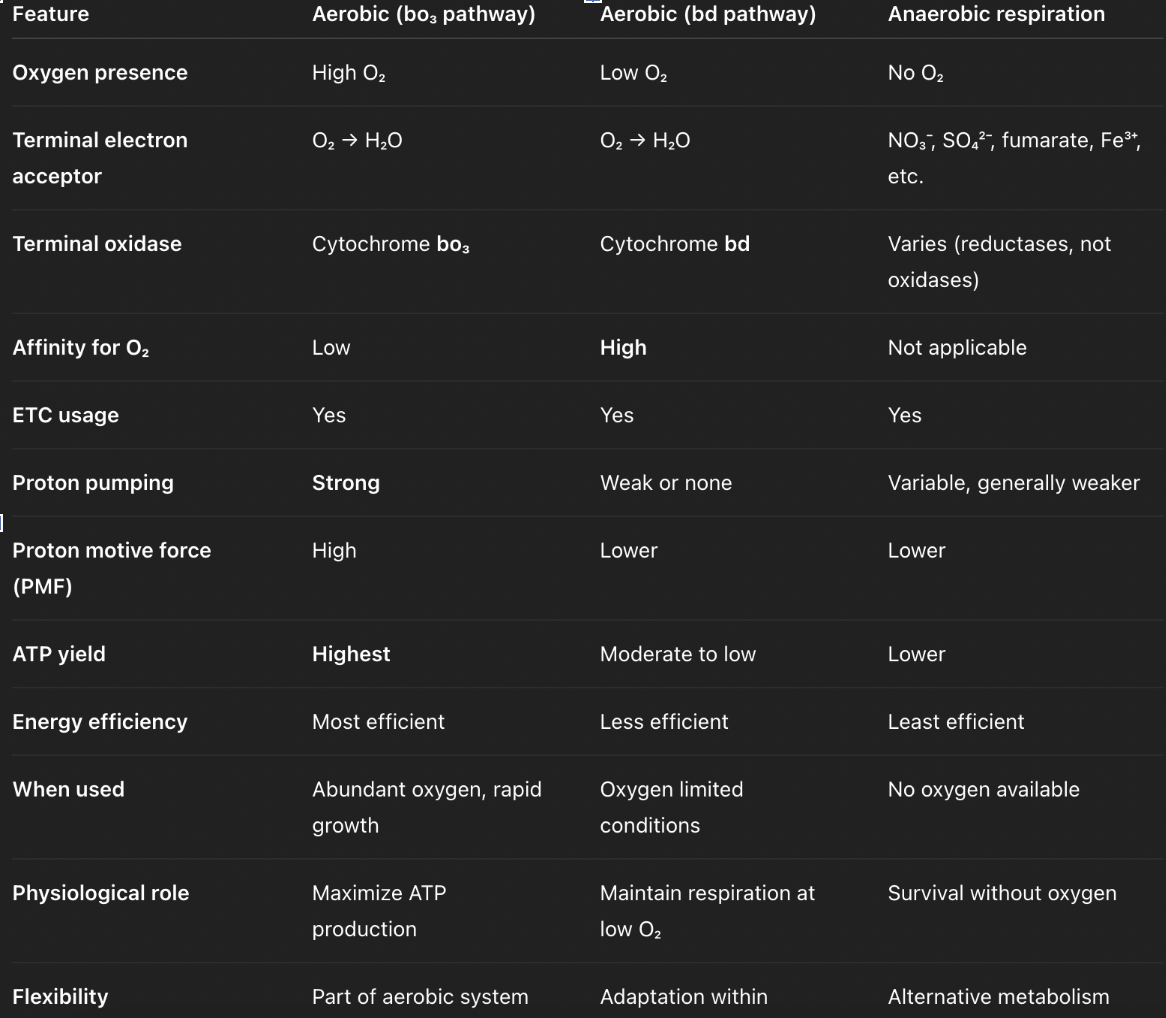
compare aerobic respiration in eukaryotes vs. e.coli
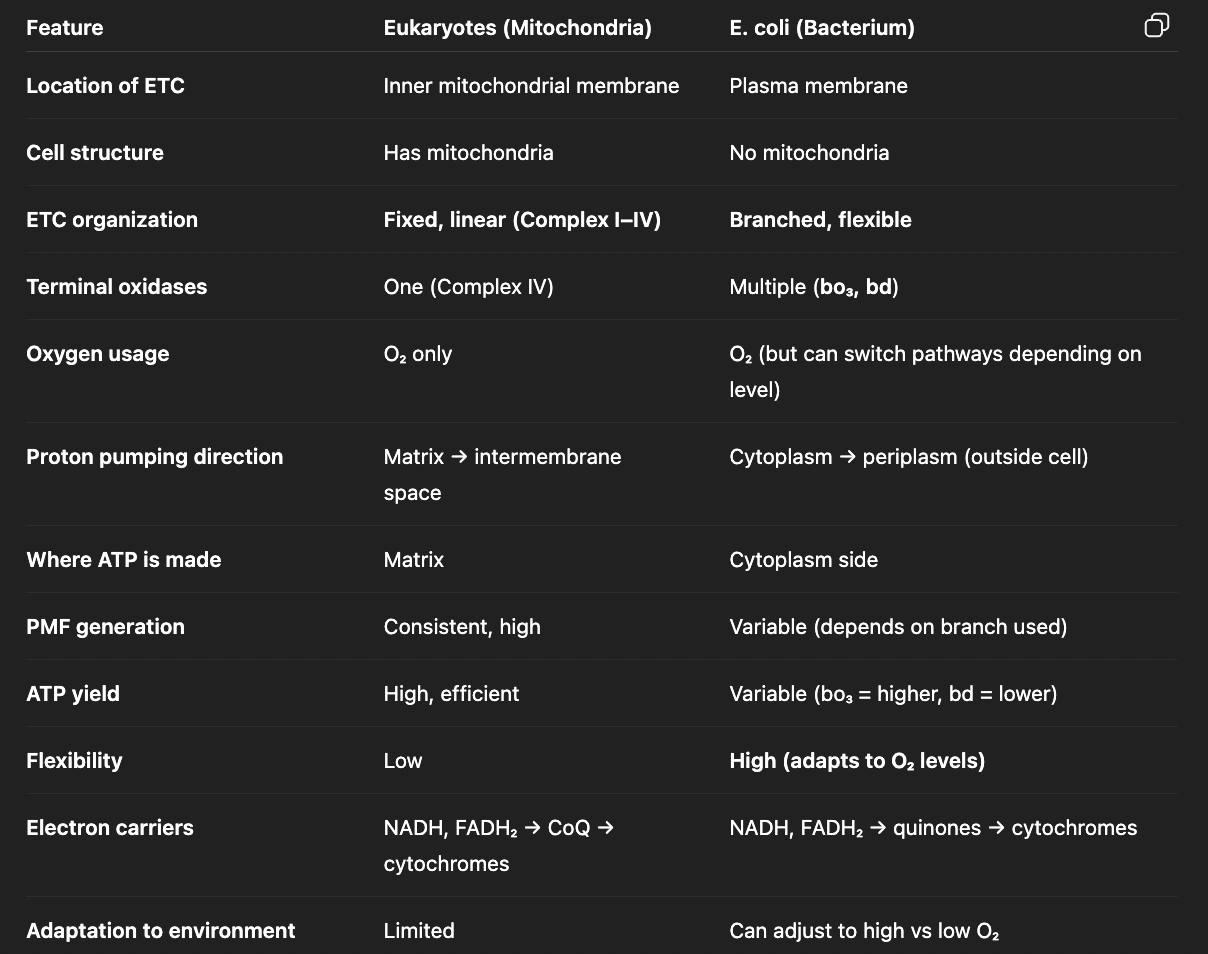
how does e.coli switch pathways depending on O2 levels?
high O2 levels (log phase) → bo3 branch → high ATP yield, promotes growth
low O2 levels (stationary phase) → bd branch (minimal proton pumping) → low ATP yield, promotes survival
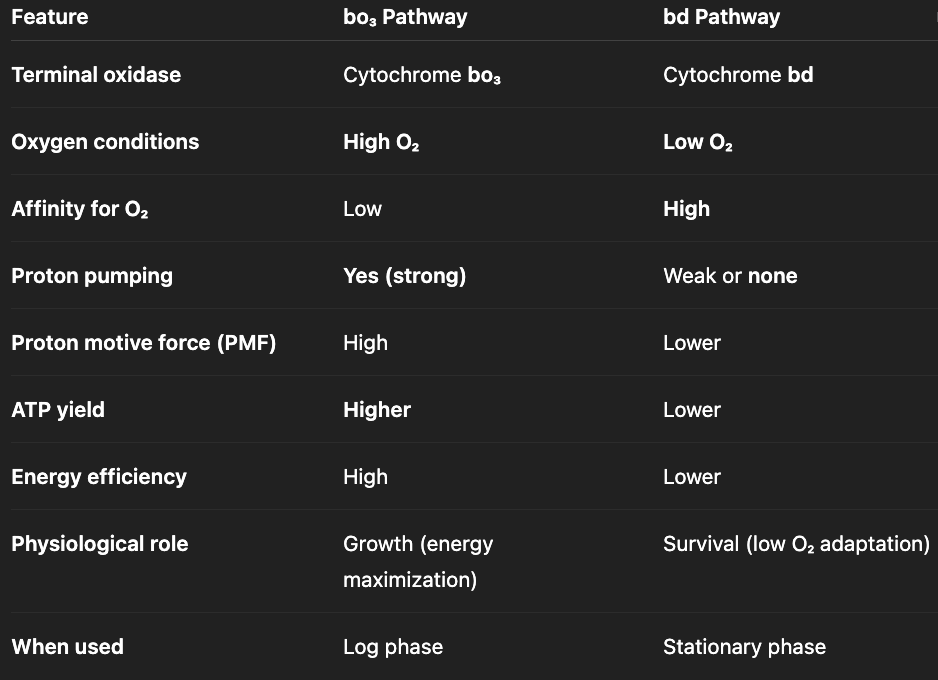
what is electron transport chain?
series of membrane-bound carriers that transfer electrons to terminal electron acceptor
what is proton pumping?
pumping H+ across the membrane (cytoplasm → outside) as electrons are being transferred
what is PMF?
protein motive force: gradient of protons across the membrane that stores potential energy
what is chemiosmosis?
movement of protons back across the membrane (outside → inside) down their gradient
what is oxidative phosphorylation?
protons flowing through the ATP synthase that drives ATP synthesis
what is the purpose of fermentation?
to regenerate NAD+ to feed back into glycolysis
reduces pyruvate and reoxidizes NADH
what type of species undergo fermentation?
aerotolerant anaerobes — ALWAYS ferments bc no ETC; can tolerate O2 but does not use O2
facultative anaerobes — flexible; switches between aerobic respiration, anaerobic respiration, and fermentation
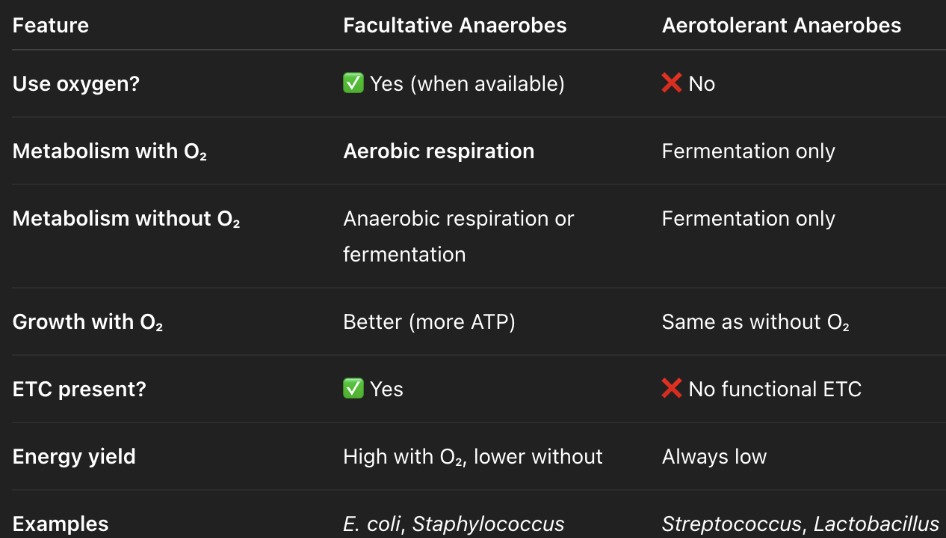
what are the possibilities of metabolism that a facultative anaerobe undergoes when in an anoxic environment?
fermentation
anaerobic respiration
uses anaerobic respiration first, then switches to fermentation when terminal electron acceptor runs out
what are the different types of fermentation?
alcohol fermentation
lactic acid fermentation
mixed acid fermentation
ABE (acetone-butanol-ethanol)
what are the steps of alcohol fermentation?
pyruvate → acetaldehyde + CO2
acetaldehyde + NADH + H+ → ethanol + NAD+
what are the steps of lactic acid fermentation?
pyruvate + NADH + H+ → lactate + NAD+
what is homofermentative vs. heterofermentative?
homofermentative: produces only lactate
heterofermentative: can do both alcohol and lactic acid fermentation; yields lactate + CO2 + ethano
what is mixed acid fermentation?
occurs when multiple pathways are active at once, yielding diverse products
lactic acid fermentation (LDH) → forms lactate
alcohol fermentation (ADH) → forms alcohol
pyruvate formate lyase (PFL) → forms acetyl-CoA + formate
formate hydrogen lyase (FHL) breaks down formate → CO2 + H2 gas
what is ABE?
mainly carried out by a certain species of clostridia (strict anaerobe)
pathway triggered by acid stress, nutrient depletion, waste accumulation, etc → stationary phase → sporulation
mechanism:
acid-producing phase - produces acetate, butyrate, ATP; pH drops
solvent-producing phase - acids converted into acetone, butanol (main product), ethanol; regenerates NAD+
products: acetone, butanol (main product), ethanol, NAD+
what are two problems that arise from fermentation? what is the cause and solution?
internal acidification
cause: fermentation produces organic acids (lactate, acetate, etc)
solution: pump H+ out of cytoplasm via. reverse ATPase; hydrolyzes ATP to ADP
struggle to produce enough biosynthetic precursors
cause: no functional ETC bc no terminal electron acceptor → NADH not efficiently reoxidized → TCA cycle can’t run fully → lack of TCA intermediates for biosynthesis
solutions: horseshoe TCA cycle (run partial pathways to generate needed intermediates), anaplerotic pathways
what is horseshoe TCA cycle?
horseshoe TCA cycle: split TCA cycle into two partial pathways to generate needed intermediates
oxidative branch: acetyl CoA + oxaloacetate → a-ketoglutarate → biosynthetic precursors for amino acids
reductive branch: oxaloacetate → malate → fumarate → succinate; regenerates NAD+ and prevents NADH buildup
how does the goal of anaplerotic pathways differ between respiration and fermentation?
respiration: refill intermediates to keep TCA cycle running
fermentation: refill intermediates to be used for anabolic/biosynthetic pathways, not to run full TCA cycle?
compare and contrast between chemoorganotrophs and chemolithotrophs
similarities:
use chemical compounds for energy instead of light
use redox reactions and ETC
generate ATP, reducing power, and proton motive force
support biosynthesis
differences
organic molecule provides energy, electrons, and carbon
inorganic molecules provides energy + electrons but need to fix CO2 (require ATP + NADPH)
energy yield: higher vs. lower
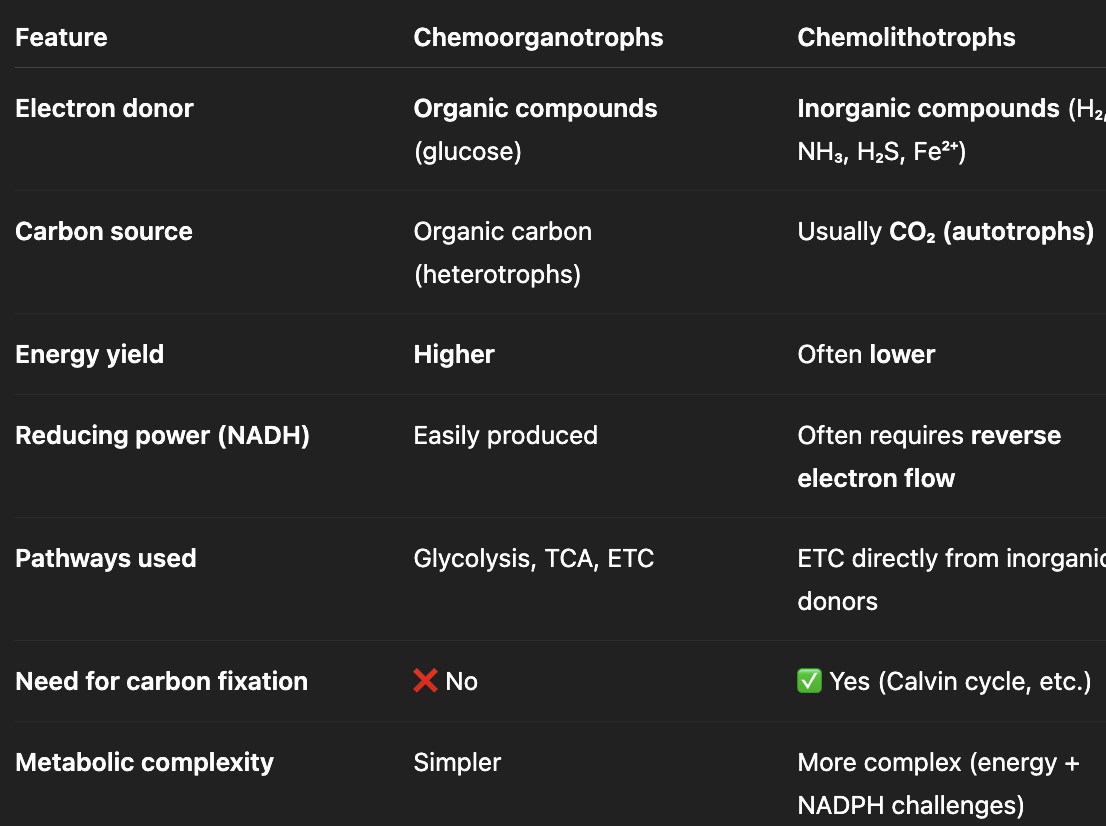
what are chemolithotrophs?
uses inorganic material to provide energy and electrons
CO2 (inorganic carbon) needs to be fixed to be used in anabolic pathways
weak electron donors requires reverse electron flow to make NADH
what are examples of chemolithotrophy?
hydrogen oxidation
nitrification (ammonia oxidation, nitrite oxidation)
sulfur oxidation
iron oxidation
what are the relative energy yields of chemolithotrophy?
energy yield depends on difference in reduction potential
bigger difference = more energy release + more ATP produced
H2 = strong electron donor → high energy yield
NO2-/Fe2+ = weak electron donor → low energy yield

what is hydrogen bacteria?
chemolithotroph
hydrogen oxidation: H2 + ½ O2 → H2O
high energy yield (strong donor + O2 acceptor)
need to find environment with presence of hydrogen and oxygen
problem: H2 is scare in oxygen-rich environment
what is sulfur bacteria?
chemolithotroph
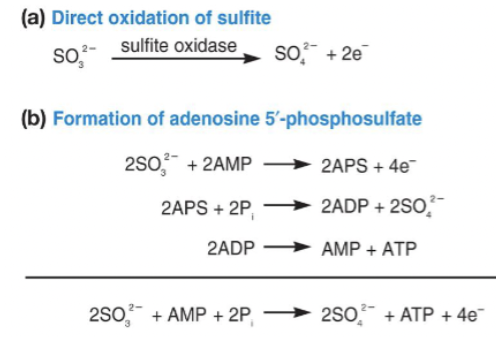
electron donors: H2S (hydrogen sulfide), S0 (elemental sulfur), S2O32- (thiosulfate), SO32- (sulfite)
final oxidation product: SO42- (sulfate)
pathways
Sox system (Sulfite oxidase): direct oxidation → electrons → ETC
APS pathway (adenosine 5’ triphosphate): produce ATP via. substrate-level phosphorylation
species found in very acidic environments
what is nitrifying bacteria?
chemolithotroph
low energy yield → slow growth
requires reverse electron flow to generate NADH
ammonia oxidizers: NH4+ + O2 → NH2OH → NO2- ; bacteria & archaea
nitrite oxidizers: NO2- (nitrite) → NO3- (nitrate); bacteria
anammox (anaerobic ammonium oxidation): NH4+ + NO2- → N2 + 2 H2O ; bacteria
what is iron bacteria
electron donor: Fe2+
very low energy yield
common in iron-rich environments
requires reverse electron flow to generate NADH
when is reverse electron flow vs. reverse atpase used?
reverse electron flow: in chemolithotrophs to generates NADH bc electron donors are weak; pushes electrons “uphill” to make NADH
reverse ATPase : used in fermentation when no ETC and no external electron acceptor to pump out acids
compare and contrast photolithoautotrophy and photoorganoheterotrophy
similiarities:
phototrophs - use light as energy source → ETC → PMF → ATP
differences:
Photolithoautotrophs: inorganic electron & carbon source; undergoes carbon fixation to make carbon source usable for biosynthesis
chlorophyll-based phototrophy
Photoorganoheterotrophs: uses organic carbon from environment that is ready for biosynthesis
rhodopsin-based phototrophy → does not form NADPH, requires reverse electron flow

compare chlorophyll vs. rhodopsin-based phototrophy
chlorophyll system: uses ETC, makes ATP + NADPH, supports autotrophy (CO2 fixation)
rhodopsin systems: uses light-driven proton pump directly, makes only ATP, only supports heterotrophs
what is oxygenic vs. anoxygenic photosynthesis?
oxygenic:
uses H2O → O2
2 photosystems (PSII + PSI)
produces ATP + NADPH
anoxygenic:
uses H2S, S0, organic compounds
1 photosystem
produces ATP only; must use reverse electron flow to make NADPH
compare the pigments used in photosynthesis?
chlorophyll (680nm):
higher energy light
oxygenic (H2O splitting forms O2)
2 photosystems
direct NADPH generated
bacteriochlorophyll (860 nm):
lower energy light
anoxygenic (no H2O splitting, does not form O2)
1 photosystem
reverse electron flow
how does oxygenic photolithotrophy work?
photosynthesis: light hits chlorophyll → electron excites and leaves PSII → H2O splits to O2 → ETC (generates PMF → ATP) → light excites PSI→ NADPH formation
PSII → splits water to form O2
PSI → makes NADPH (direct process)
how does anoxygenic photolithtrophy work?
anoxygenic photolithotrophy: couples photosynthesis with sulfur oxidation (getting electrons from sulfur);
anoxygenic photosynthesis: light excites bacteriochlorophyll → electrons flow through ETC → PMF is generated → protons flow back through ATP synthase → ATP is produced
electrons from sulfur oxidation reduces NADP+ to NADPH (separate process)
How do autotrophic microorganisms fix CO2 (in general)?
obtain energy (ATP) and reducing power (NADPH or NADH → NADPH) from light (phototrophs) or chemical reactions (chemolithotrophs)
CO2 is reduced and incorporated into an organic molecule
How does Calvin Cycle work?
carbon fixation: CO2 attached to RuBP (5C) → two 3-PG ; enzyme-Rubisco
reduction: 3PG → 1,3BPG → G3P
regeneration of RuBP
what is the difference between assimilatory vs. dissimilatory processes?
Assimilatory = for building biomass (anabolism/biosynthesis), requires ATP, incorporates end product into cell
Dissimilatory = for generating energy (catabolism), produces ATP, releases end product as waste
what is the assimilatory mechanism of inorganic sulfate?
reduce sulfate SO42- (inorganic, unusable) → sulfite SO32-, → H2S (organic, usable) → cysteine
what is the assimilatory mechanism of inorganic sulfate?
sulfur acts as a terminal electron acceptor in anaerobic respiration
SO42- → H2S (released)
what is the assimilatory process of inorganic nitrogen?
use nitrate reductase NAS enzyme (located in cytoplasm)
nitrate NO3- → nitrite NO2- → nitroxyl (NOH) → hydroxylamine (NH2OH) → NH3
after assimilatory process, can use ammonia (NH3) for amino acid synthesis
what is the dissimilatory mechanism of inorganic nitrogen?
use nitrate reductase NAR enzyme
NO3-→ N2 (denitrification)
what is nitrogen fixation?
converting nitrogen gas N2 to ammonia NH3 for biosynthesis
enzyme: nitrogenase—O2 sensitive, requires lots of ATP and electrons
what are mechanisms to shield nitrogenase enzymes from O2?
*nitrogenase are O2 sensitive; enzyme complex deactivates in the presence if O2
physical separation—heterocyst (cyanobacteria)
spatial separation (symbiosis)—leghemoglobin (produced in nodulation of plant roots)
high respiration rates—ex: Azotobacter species
how do heterocysts work?
separates photosynthesis from nitrogen fixation (either via. compartment or cell differentiating into specialized cell) → limits the effects of O2
how do leghemoglobin work?
leghemoglobin: O2 binding protein
produced in Rhizobium bacteria (found in root nodules of plant)
keeps O2 levels low enough to protect nitrogenase but high enough for respiration
how does Azotobacter protect nitrogenase?
rapidly consumes O2 via. respiration → keeps intracellular O2 low
accumulate PHB (polymer) reserve when low O2
mobilizes PHB polymers and secrete alginate (thick slime layer) → prevents/slows diffusion of oxygen into the cell
what is carboxysome?
compartmentalizes carbon fixation process