8- Dna alteration, Regulation of transcription and translation, Gene expression and cancer
1/37
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
38 Terms
What is a gene mutation?
A change in the base sequence of DNA (on chromosomes)
Can arise spontaneously during DNA replication (interphase)
What is a mutagenic agent?
A factor that increases rate of mutation e.g. UV light or alpha particles
Explain how a gene mutation can lead to the production of a non-functional protein or enzyme (general)
Changes sequence of base triplets in DNA so changes sequence of codons on mRNA
So changes sequence of amino acids in the encoded polypeptide
So changes position of hydrogen/ ionic/ disulphide bonds (between amino acids)
So changes tertiary structure (shape) of protein
Enzymes- active site changes shape so substrate can’t bind, E-S complex can’t form
Describe the different types of gene mutations
Substitution= a base/ nucleotide is replaced by a different base/ nucleotide in DNA
Additon= 1 or more bases/ nucleotides are added to the DNA base sequence
Deletion= 1 or more bases/ nucleotides are lost from the DNA base sequence
Duplication= a sequence of DNA bases/ nucleotides is repeated/ copied
Inversion= a sequence of bases/ nucleotides detaches from the DNA sequence, then rejoins at the same position in the reverse order
Translocation= a sequence of DNA bases/ nucleotides detaches and is inserted at a different location within the same or a different chromosome
Explain why not all gene mutations affect the order of amino acids
Some substitutions change only 1 triplet code/ codon which could still code for the same amino acid
As the genetic code is degenerate (an amino acid can be coded for by more than one triplet)
Some occur in introns which do not code for amino acids as they are removed curing splicing
Explain why a change in amino acid sequence is not always harmful
May not change tertiary structure of protein (if position of ionic/ disulphide/ H bonds don’t change)
May positively change the properties of the protein, giving the organism a selective advantage
Explain what is meant by a frameshift
Occurs when mutations (addition, deletion, duplication or translocation) change the number of nucleotides/ bases by a number not divisible by 3
This shifts the way the genetic code is read, so all the DNA triplets/ mRNA codons downstream from the mutation change (so significant effects)
> Effects on the encoded polypeptide are significant
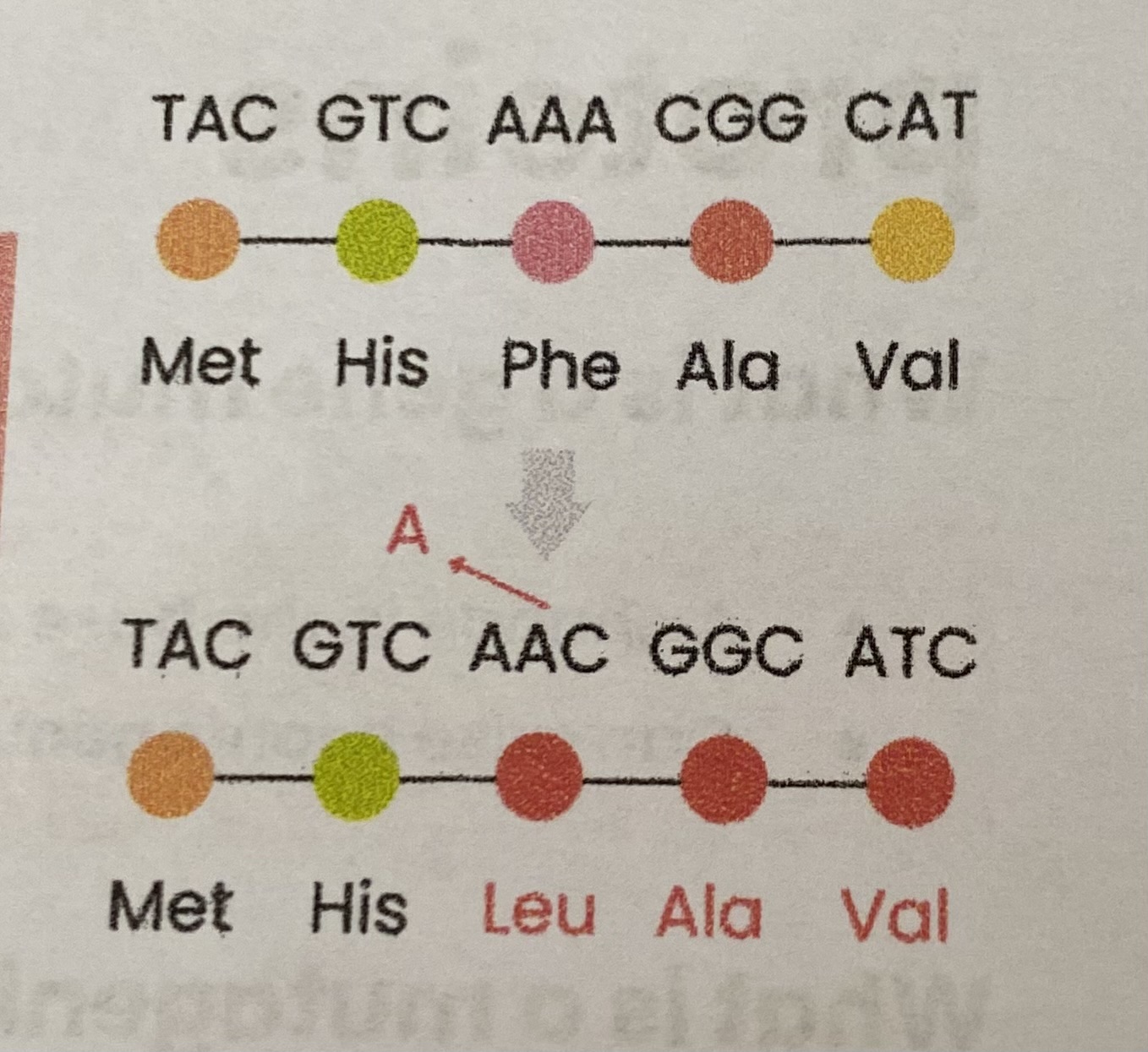
Explain how mutations can lead to production of shorter polypeptides
Deletion or translocation→ triplet(s)/ codon(s) missing so amino acid(s) missing
Substitution, addition, deletion, duplication, inversion or translocation→ premature stop triplet/ codon (doesn’t code for amino acids; terminates translation) so amino acids missing at end of polypeptide
What are stem cells?
Undifferentiated/ unspecialised cells capable of:
Dividing (by mitosis) to replace themselves indefinitely
Differentiating into other types of (specialised) cells
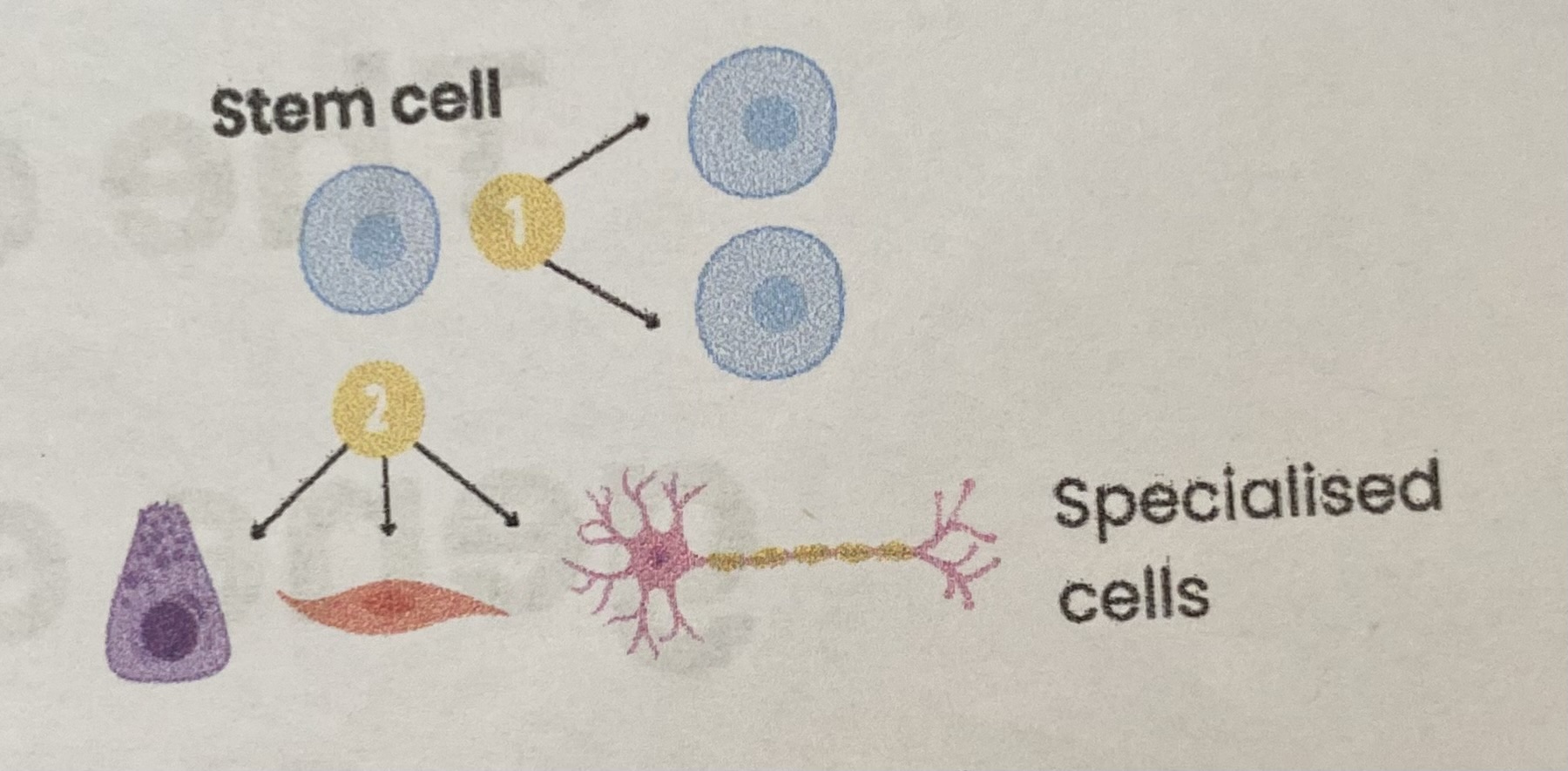
Describe how stem cells become specialised during development
Stimuli lead to activation of some genes (due to transcription factors)
So mRNA is transcribed only from these genes and then translated to form proteins
These proteins modify cells permanently and determine cell structure/ function
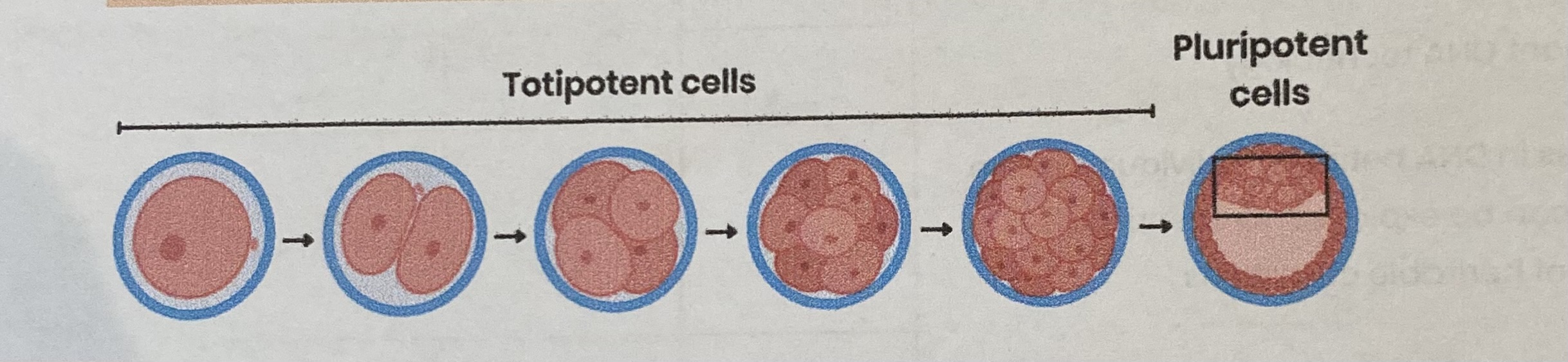
Describe totipotent cells
Occur for a limited time in early mammalian embryos
Can divide AND differentiate into any type of body cell (including extra- embryonic cells e.g. placenta)
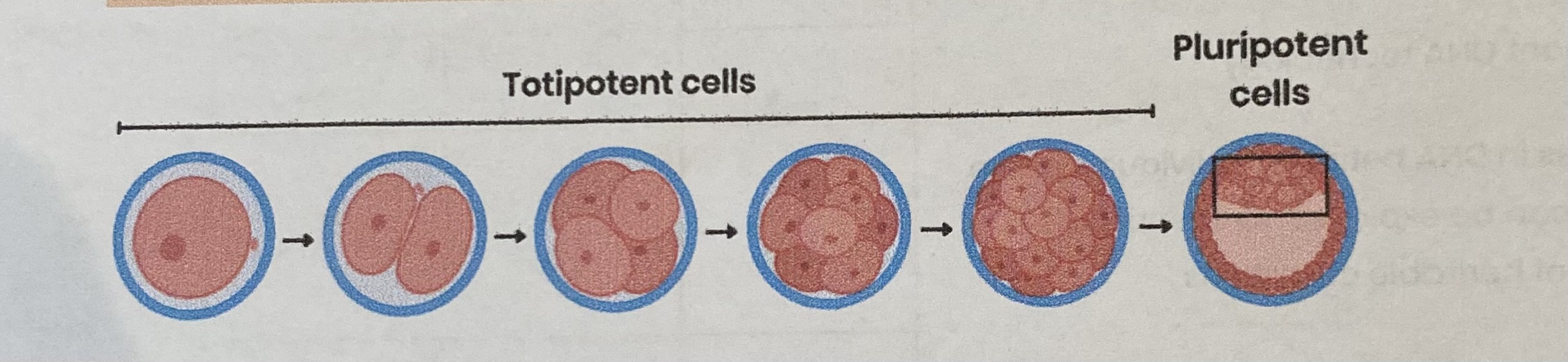
Describe pluripotent cells
Found in mammalian embryos (after first few cell divisions)
Can divide AND differentiate into most cell types (every cell type in the body but not placental cells)
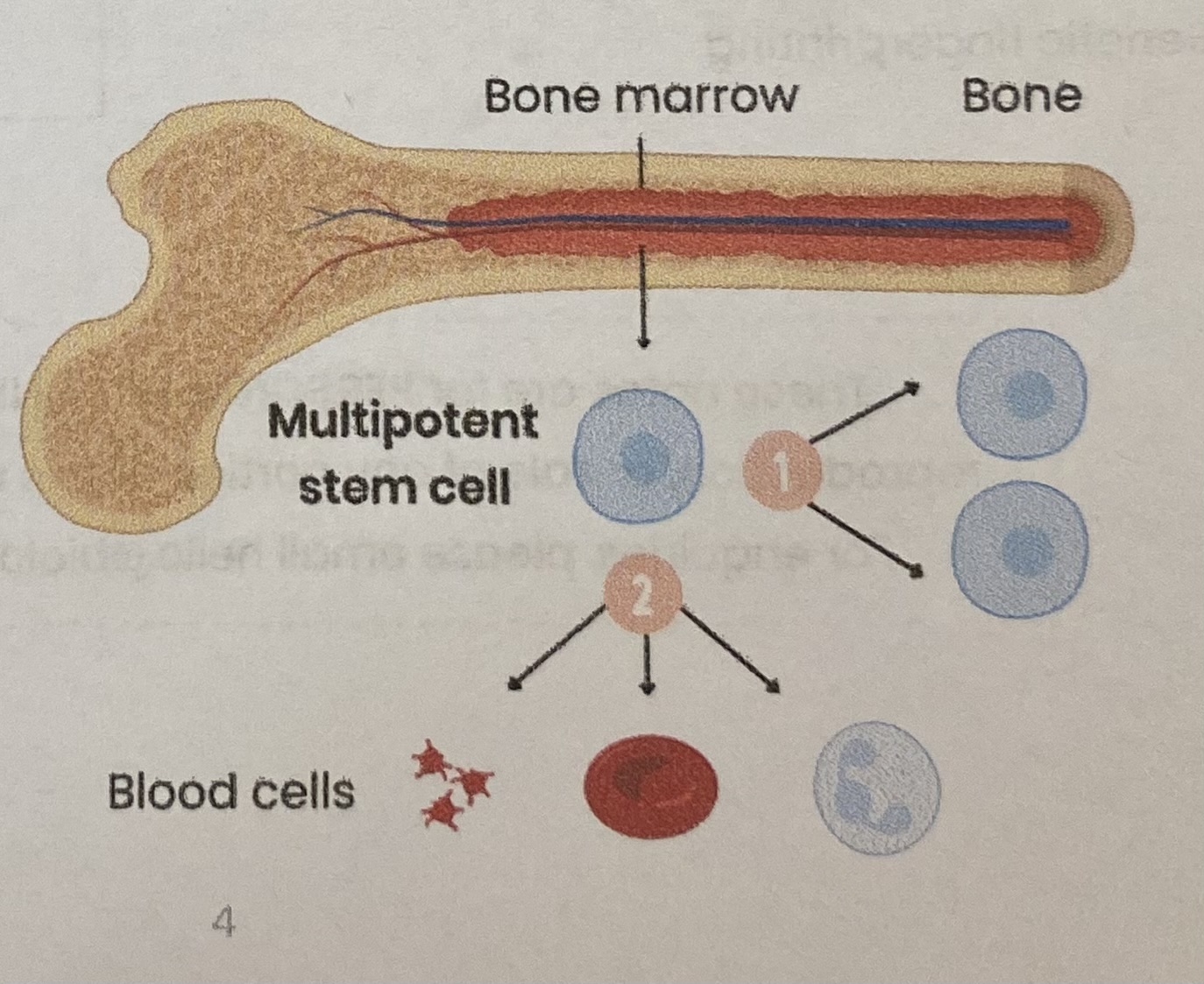
Describe multipotent cells
Found in mature mammals
Can divide AND differentiate into limited number of cell types
EXAMPLE: multipotent cells in bone marrow can divide and differentiate into different types of blood cell
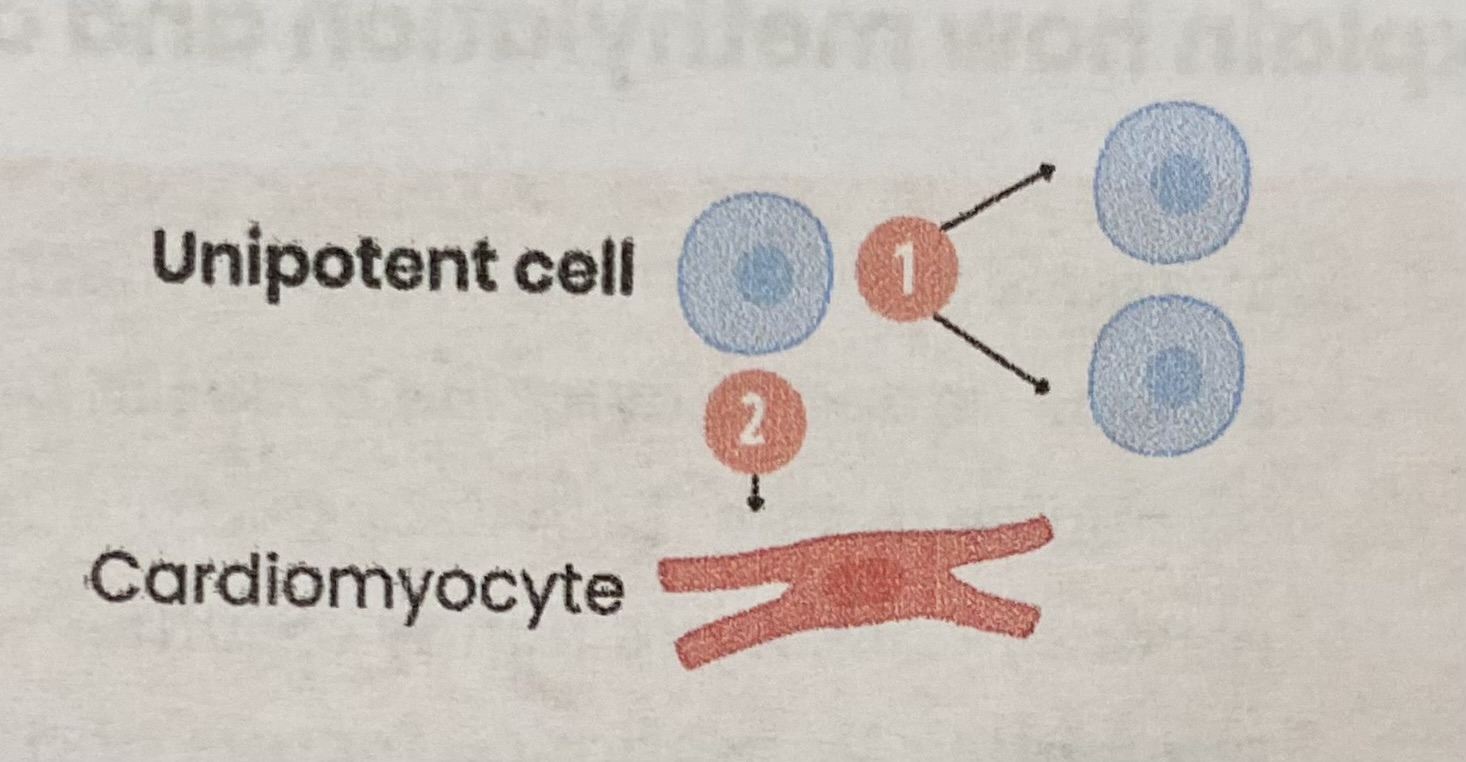
Describe unipotent cells
Found in mature mammals
Can divide AND differentiate into just one cell type
EXAMPLE: unipotent cells in the heart can divide and differentiate into cardiomyocytes (cardiac muscle cells)
Explain how stem cells can be used in the treatment of human disorders
Transplanted into patients to divide in unlimited numbers
Then differentiate into required healthy cells (to replace faulty/ damaged cells)
EXAMPLES:
Potential treatment of Type 1 diabetes by creating healthy islet cells that produce insulin
Bone marrow stem cell transplant for sickle cell disease/ blood cancers
Destroy patient’s bone marrow before treatment→ so no faulty cells are produced
Transplant stem cells from healthy person→ divide and differentiate into healthy cells
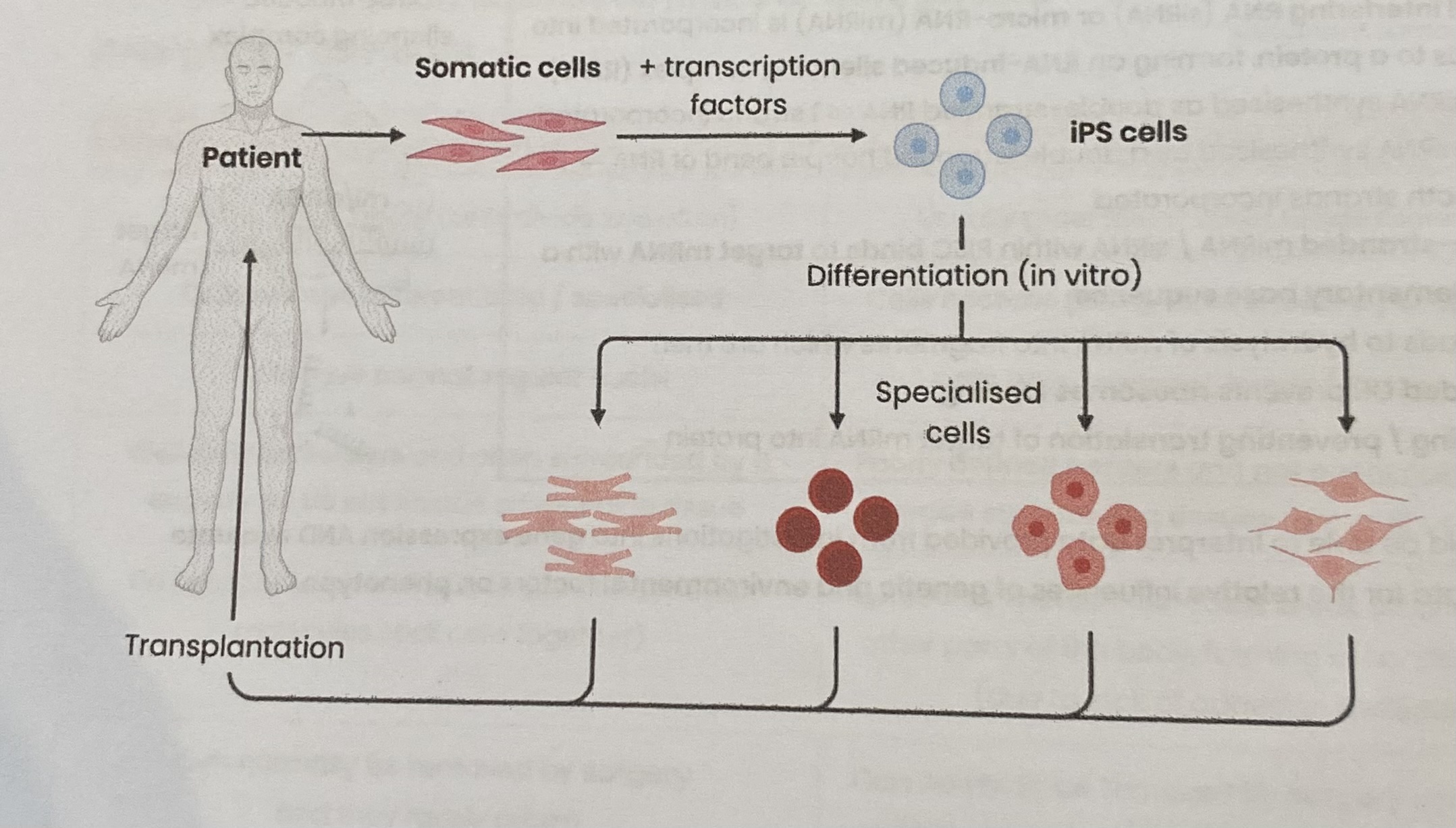
Explain how induced pluripotent stem (iPS) cells are produced
Obtain adult somatic (body) cells (non-pluripotent cells or fibroblasts) from patient
Add specific proteintranscription factors associated with pluripotency to cells so they express genes associated with pluripotency (reprogramming)
Transcription factors attach to promotor regions of DNA, stimulating or inhibiting transcription
Culture cells to allow them to divide by mitosis
> Once made, iPS cells can divide and differentiate into healthy cells to be transplanted into the same patient
Evaluate the use of stem cells in treating human disorders
FOR:
Can divide and differentiate into required healthy cells, so could relieve human suffering by saving lives and improving quality of life
Embryos are often left over from IVF and so would otherwise be destroyed
iPS cells unlikely to be rejected by patient’s immune system as made with patient’s own cells
iPS cells can be made without destruction of embryo and adult can give permission
AGAINST:
Ethical issues with embryonic stem cells as obtaining them requires destruction of an embryo and potential life (embryo cannot consent)
Immune system could reject cells and immunosuppressant drugs are required
Cells could divide out of control, leading to formation of tumours/ cancer
What are transcriptional factors?
Proteins which regulate (stimulate or inhibit) transcription of specific target genes in eukaryotes
By binding to a specific DNA base sequence on a promotor region
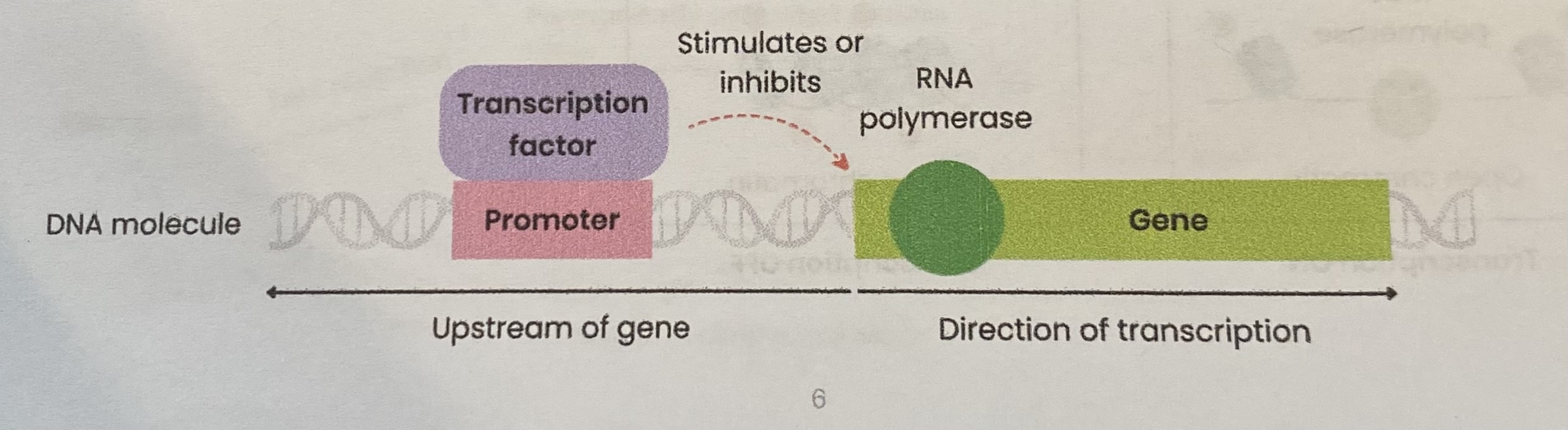
Describe how transcription can be regulated using transcriptional factors
Transcription factors move from cytoplasm to nucleus
Bind to DNA at a specific DNA base sequence on a promotor region (before/ upstream of target gene)
This stimulates or inhibits transcription (production of mRNA) of target gene(s) by helping or preventing RNA polymerase binding
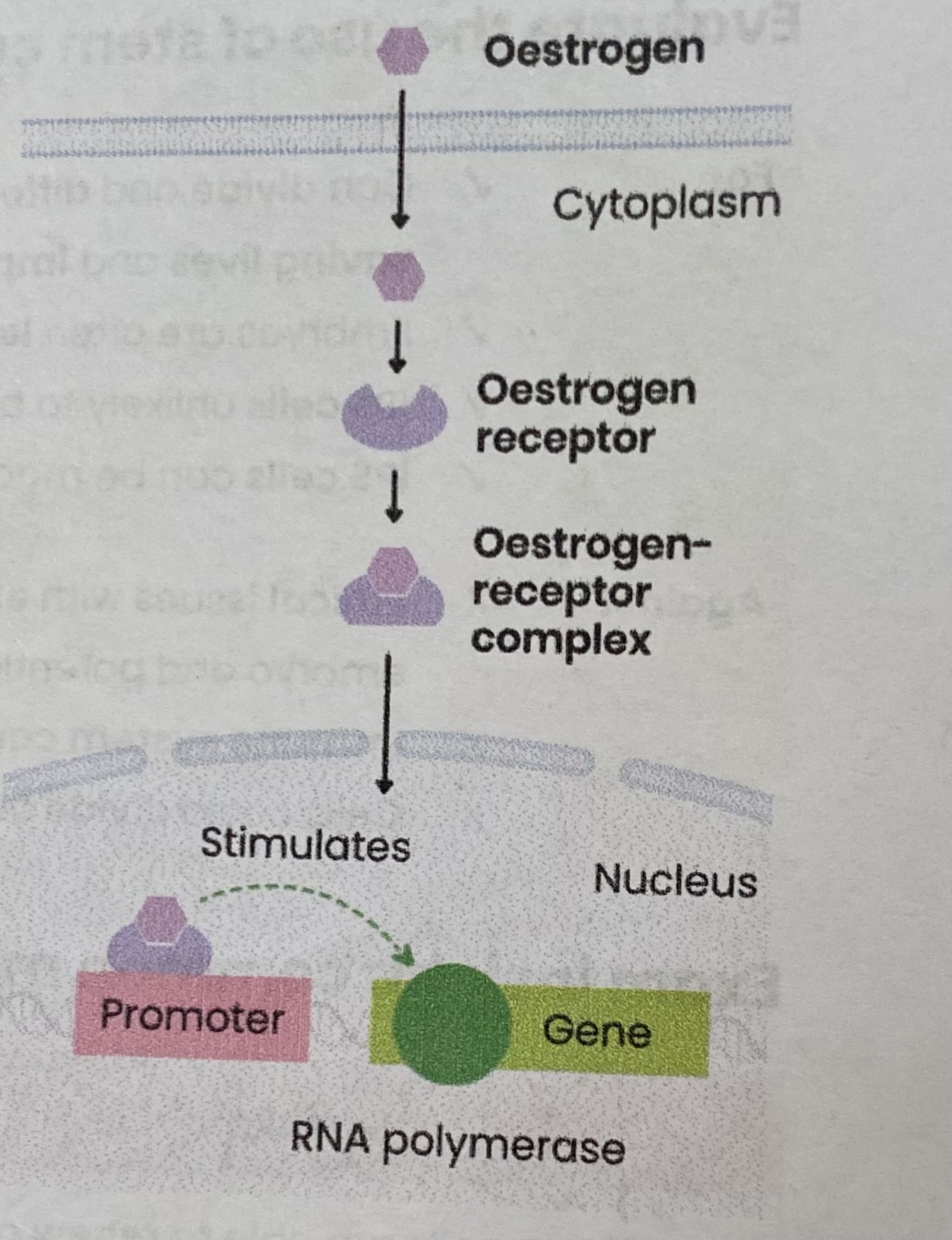
Explain how oestrogen affects transcription
Oestrogen is a lipid-soluble steroid hormone so diffuses into cell across the phospholipid bilayer
In cytoplasm, oestrogen binds to its receptor, an inactive transcriptional factor, forming an oestrogen- receptor complex
This changes the shape of the inactive transcriptional factor, forming an active transcriptional factor
The complex diffuses from cytoplasm into the nucleus
Then binds to a specific DNA base sequence on the promotor region of a target gene
Stimulating transcription of target genes forming mRNA by helping RNA polymerase to bind
Explain why oestrogen only affects target cells
Other cells do not have oestrogen receptors
Describe what is meant by epigenetics
Heritable changes in gene function/ expression without changes to the base sequence of DNA
Caused by changes in the environment (e.g. diet, stress, toxins)
Describe what is meant by epigenome
All chemical modification of DNA and histone proteins- methyl groups on DNA and acetyl groups on histones
Summarise the epigenetic control of gene expression in eukaryotes
To INHIBIT transcription:
increased methylation of DNA
decreased acetylation of histones
To ALLOW transcription:
decreased methylation of DNA
increased acetylation of histones
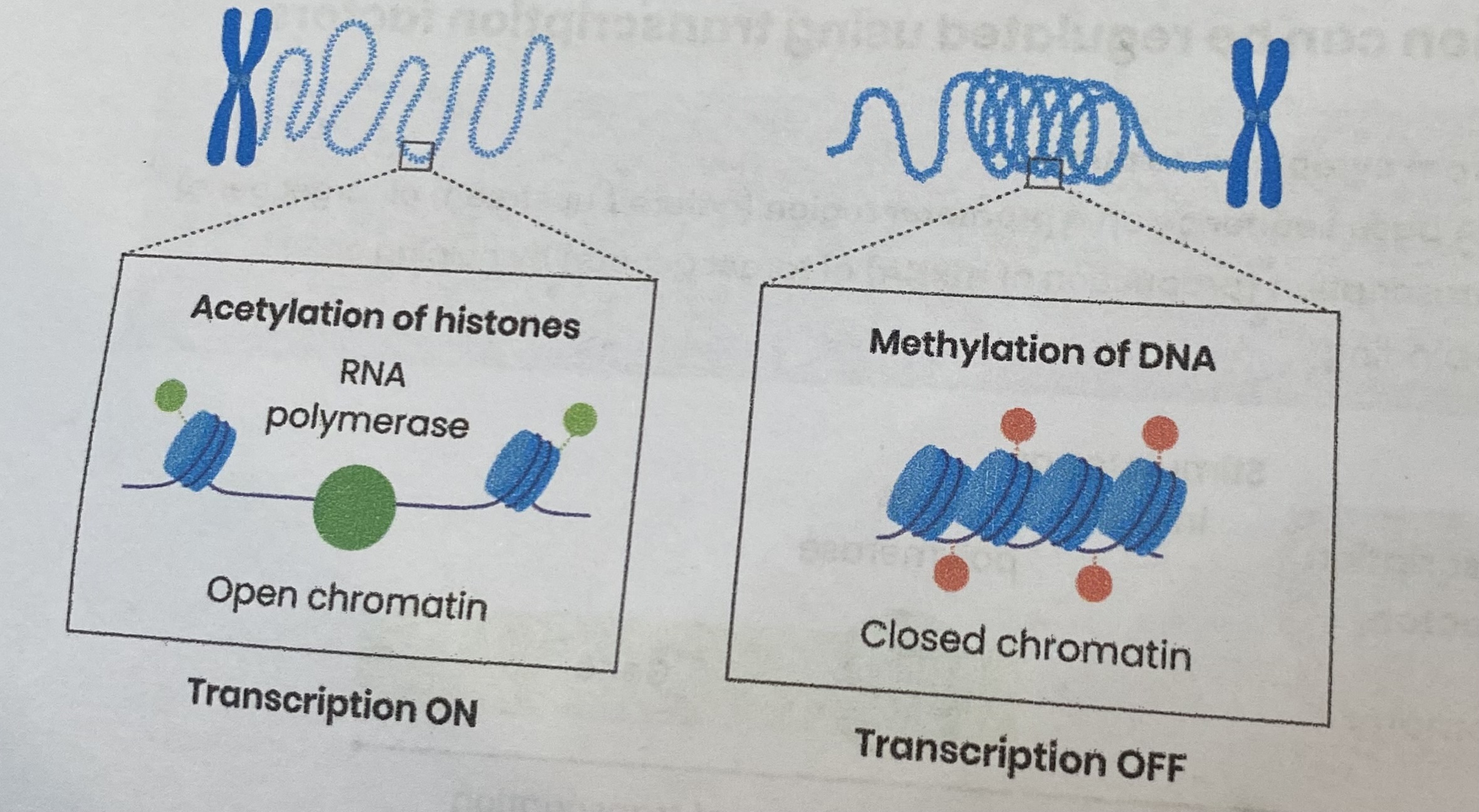
Explain how methylation can inhibit transcription
Increased methylation of DNA- methyl groups added to cytosine bases in DNA
So nucleosomes (DNA wrapped around histone) pack more tightly together
Preventing transcriptional factors and RNA polymerase binding to promotor
Explain how acetylation can inhibit transcription
Decreased acetylation of histones increases positive charge of histones
So histones bind DNA (negatively charged) more tightly
Preventing transcriptional factors and RNA polymerase binding to promotor
Explain the relevance of epigenetics on disease development and treatment
Environmental factors (e.g. diet, stress, toxins) can lead to epigenetic changes
These can stimulate/ inhibit expression of certain genes that can lead to disease development
Increased methylation of DNA OR decreased acetylation of histones inhibits transcription
Decreased methylation of DNA OR increased acetylation of histones stimulates transcription
Diagnostic tests can be developed that detect these epigenetic changes before symptoms present
Drugs can be developed to reverse these epigenetic changes
What is RNA interference (RNAi)?
Inhibition of translation of mRNA produced from target genes, by RNA molecules e.g. siRNA, miRNA
This inhibits expression of (silencing) a target gene
> This happens in eukaryotes and some prokaryotes
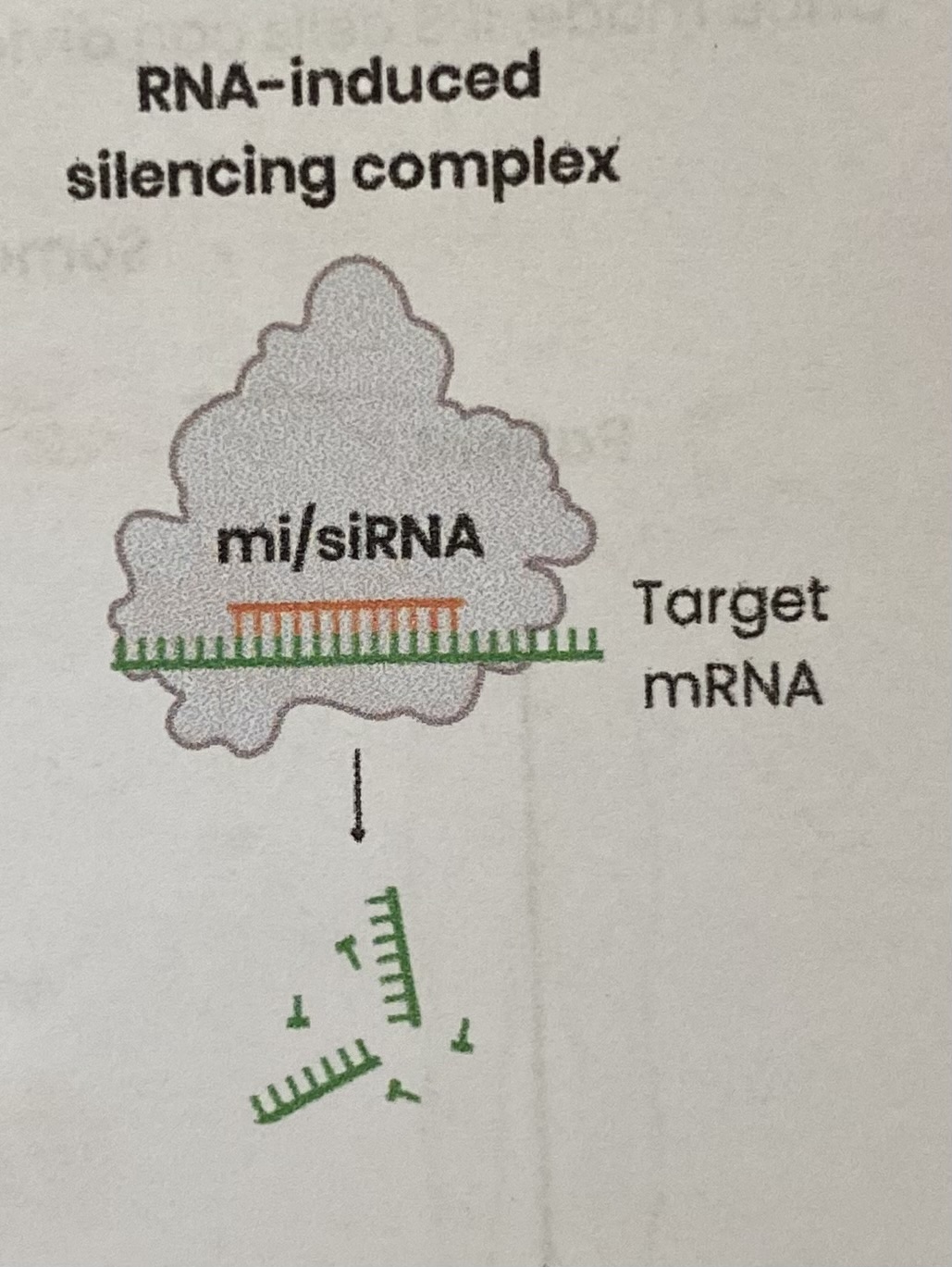
Describe the regulation of translation by RNA interference
Small interfering RNA (siRNA) or micro-RNA (miRNA) is incorporated into/ binds to a protein, forming an RNA-induced silencing complex (RISC)
siRNA synthesised as double-stranded RNA→ 1 strand incorporated
miRNA synthesised as a double-stranded haripin bend of RNA→ both strands incorporated
Single-stranded miRNA/ siRNA within RISC binds to target mRNA with a complementary base sequence
This leads to hydrolysis of mRNA into fragments which are then degraded OR prevents ribosomes binding
Reducing/ preventing translation of target mRNA into protein
Describe how tumours and cancers form
Mutations in DNA/ genes controlling mitosis can lead to uncontrolled cell division
Tumour formed if this results in mass of abnormal cells
Malignant tumour= cancerous, can spread by metastatsis
Benign tumour= non-cancerous
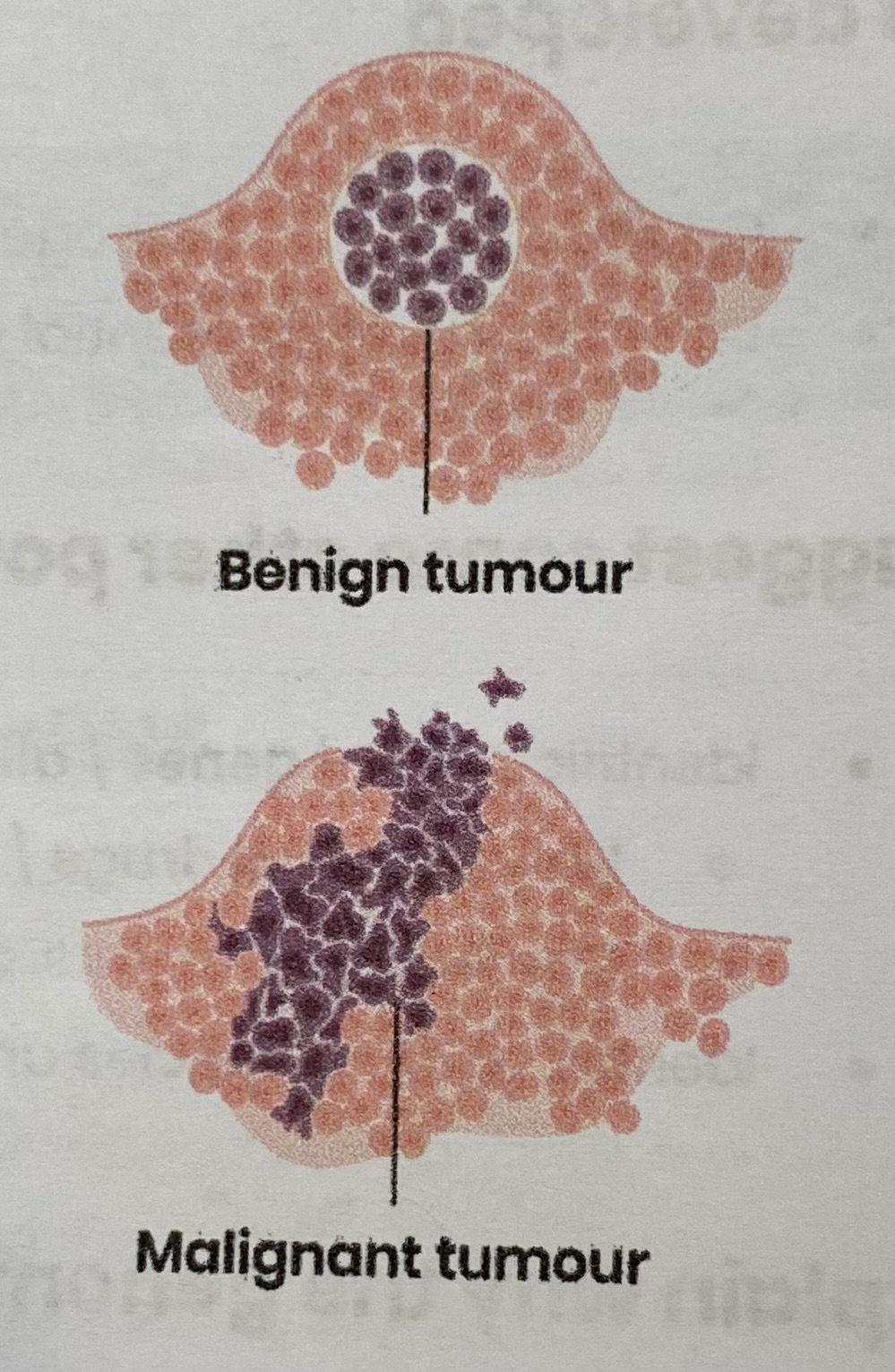
Compare the main characteristics of benign and malignant tumours
BENIGN TUMOURS:
Usually grow slowly (cells divide less often)
Cells are well differentiated/ specialised
Cells have normal, regular nuclei
Well defined borders and often surrounded by a capsule so do not invade surrounding tissue
Do not spread by metastasis (as cell adhesion molecules stick cells together)
Can normally be removed by surgery and they rarely return
MALIGNANT TUMOURS:
Usually grow faster (cells divide more often)
Cells become poorly differentiated/ unspecialised
Cells have irregular, larger/ darker nuclei
Poorly defined bordersand not encapsulated so can invade surrounding tissues (growing projections)
Spread by metastasis- cells break off and spread to other parts of the body, forming secondary tumours (due to lack of adhesion molecules)
Can normally be removed by surgery combined with radiotherapy/ chemotherapy but they often return
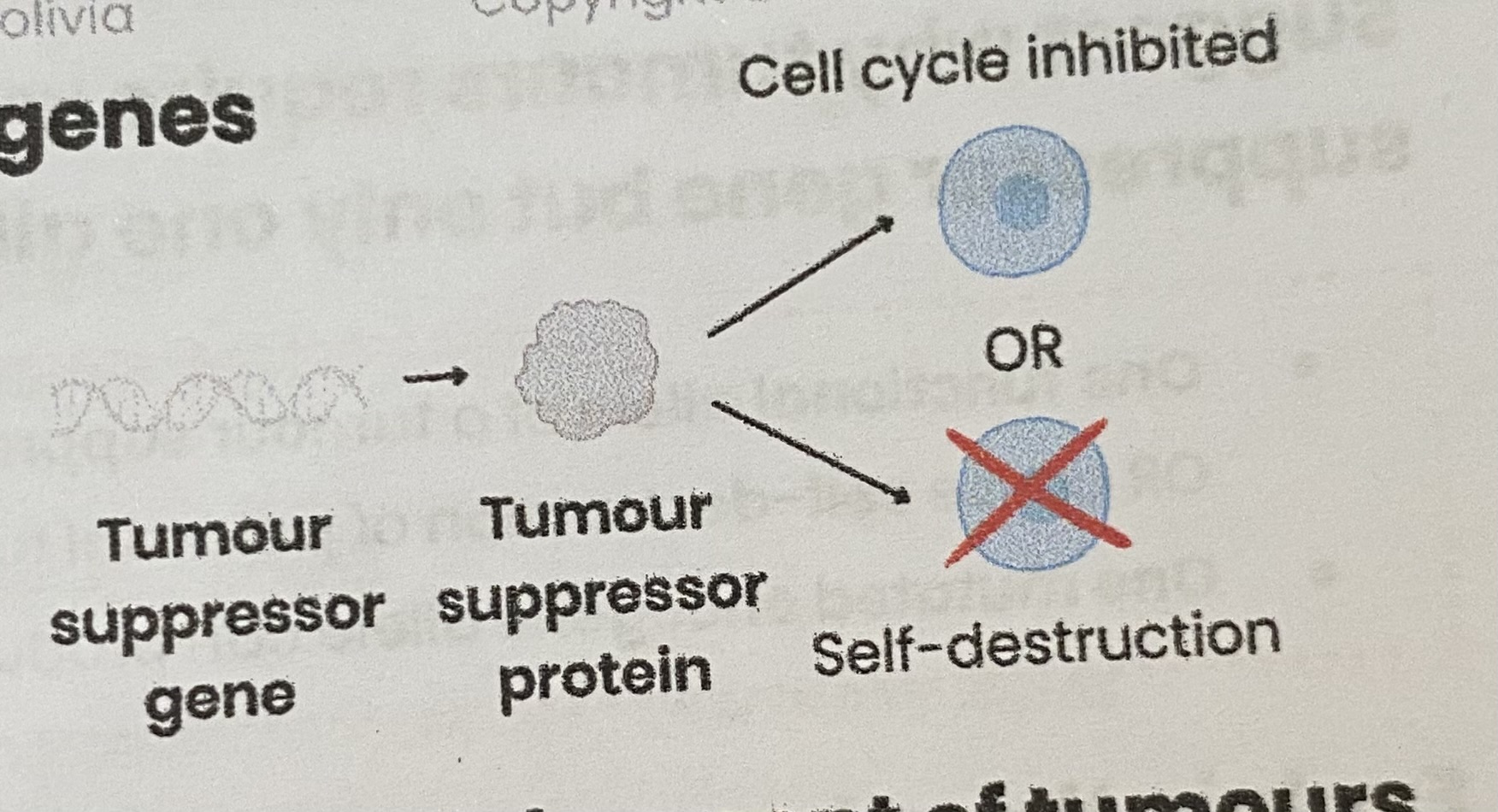
Describe the function of tumour suppressor genes
Code for proteins that:
Inhibit/ slow cell cycle (e.g. if DNA damage detected)
OR cause self-destruction (apoptosis) of potential tumour cells (e.g. if damaged DNA can’t be repaired)
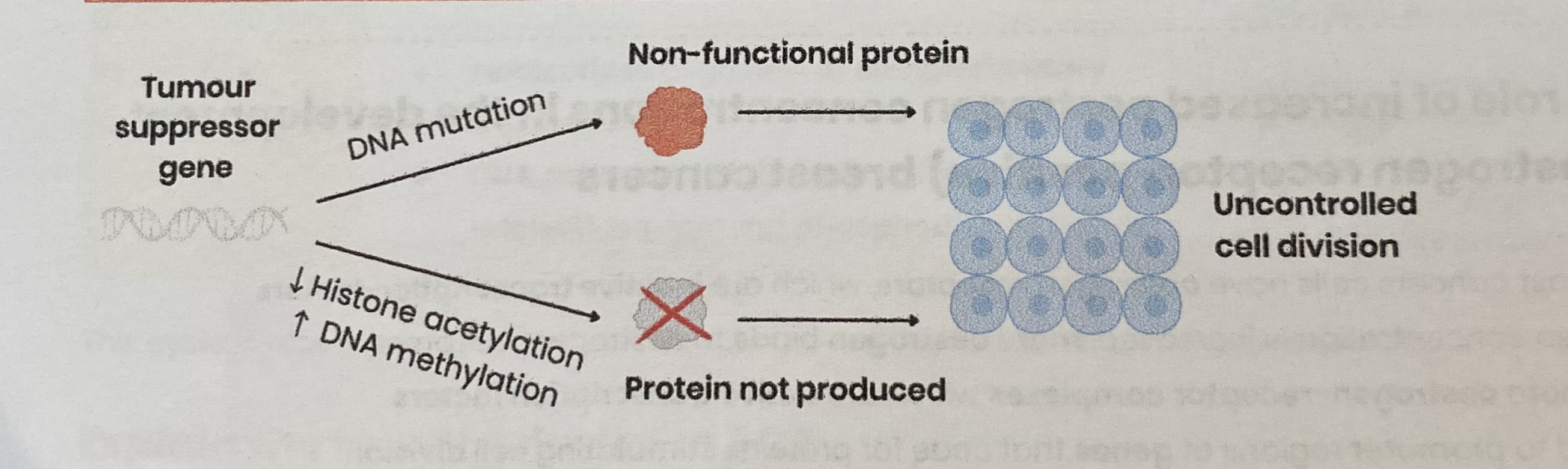
Explain the role of tumour suppressor genes in the development of tumours
Mutation in DNA base sequence→ production of non-functional protein
By leading to change in amino acid sequence which changes protein tertiary structure
Decreased histone acetylation OR increased DNA methylation→ prevents production of protein
By preventing binding of RNA polymerase to promotor region, inhibiting transcription
Both lead to uncontrolled cell division (cell division cannot be slowed)
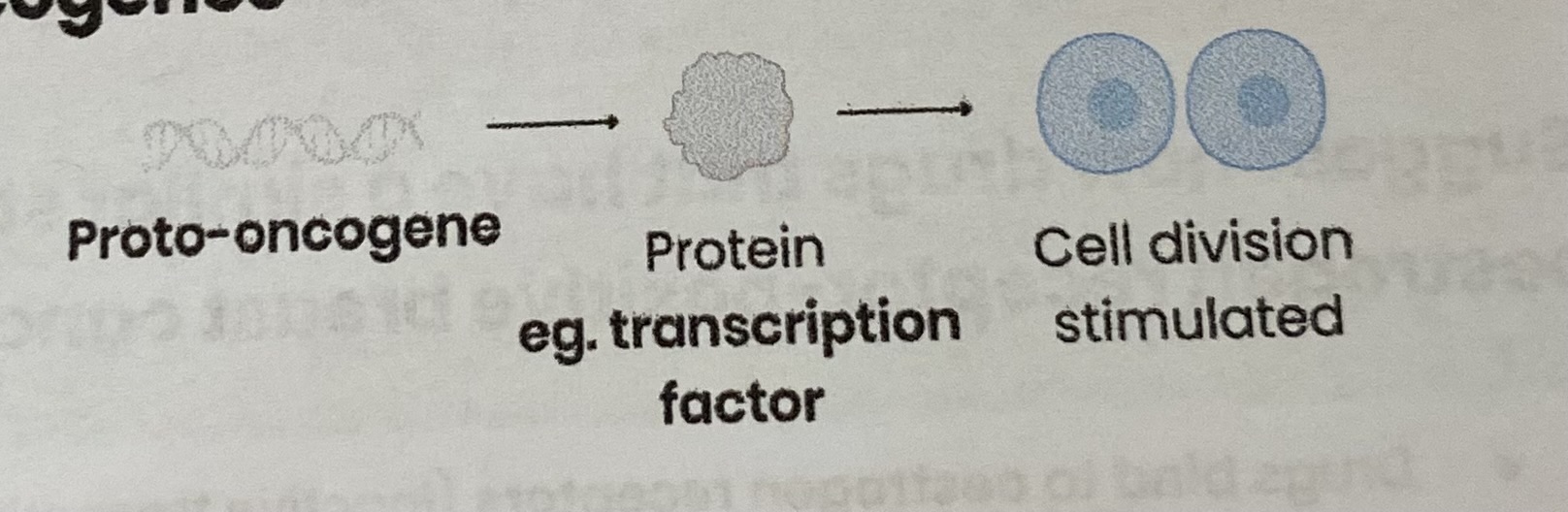
Describe the function of (proto-)oncogenes
Code for proteins tat stimulate cell division (e.g. through involvement in signalling pathways that control cell responses to growth factors)
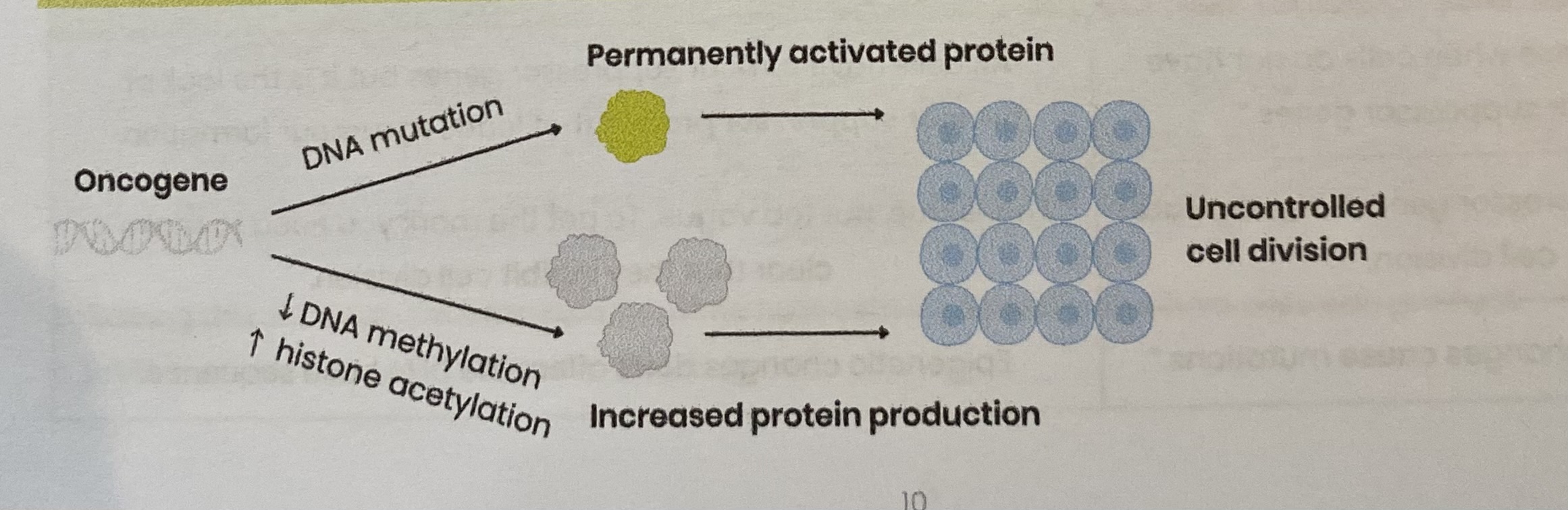
Explain the role of oncogenes in the development of tumours
An oncogene is a mutated/ abnormally expressed form of the corresponding proto-oncogene
Mutation in DNA base sequence→ overproduction of protein OR permanently activated protein
By leading to change in amino acid sequence which changes protein tertiary structure
Decreased DNA methylation OR increased histone acetylation→ increases production of protein
By stimulating binding of RNA polymerase to promotor region, stimulating transcription
Both lead to uncontrolled cell division (cell division is permanently stimulated)
Suggest why tumours require mutations in both alleles of a tumour suppressor gene but only one allele of an oncogene
One functional allele of a tumour suppressor gene can produce enough protein to slow the cell cycle OR cause self-destruction of potential tumour cells→ cell division is controlled
One mutated oncogene allele can produce enough protein to lead to rapid/ uncontrolled cell division
Explain the relevance of epigenetics in cancer treatment
Drugs could reverse epigenetic changes that caused cancer, preventing uncontrolled cell division. For example:
Increasing DNA methylation OR decreasing histone acetylation of oncogene
to inhibit transcription/ expression
Decreasing DNA methylation OR