BIO1140 Midterm 2 Notes
1/89
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
90 Terms
Signal transduction
The conversion of one type of signal into another.
Typically, when an extracellular signal is converted into an intracellular signal that a cell can act on
Signal transduction pathway
A series of molecular events by which a cell converts an external signal into a specific internal response.
The process is:
A receptor binds the extracellular signal
Intracellular signalling molecules activate one another in sequence
The final molecule activates an effector protein (eg: enzyme, transcription factor, cytoskeletal proteins)
This leads to a cellular response (such as altered metabolism, gene expression or cell movement.
Intracellular signaling molecule
Proteins or small messenger molecules inside the cell.
Eg: cAMP, cGMP.
They relay, amplify, integrate, distribute and regulate the signal within the signalling pathway.
They pass the signal from the receptor to downstream targets
Some act as molecular switches (on/off via phosphorylation or GTP binding
Many amplify the sinal so a small external signal produces a large cellular response
Sources of signals a cell is exposed to
Cells are typically exposed to chemical signalling molecules (such as hormones, neurotransmitters, growth factors, gases or peptides)
These signals can originate from:
Nearby cells in the same tissue
The same cell that releases the signal
Neurons transmitting signals aross synapses
Endocrine glands releasing hormones into the bloodstream
The signals bind to specific receipts on or inside target cells, triggering cellular responses
Main types of signal-mediated cell-cell communication
Paracrine signaling (local signaling)
Synaptic (neuronal) signaling. (short but targetted)
Contact-dependent signaling (direct contact)
Endocrine signaling (long-distance signaling)
Paracrine signaling
Signaling molecules are released and act on nearby cells.
The molecules diffuse through the extracellular fluid.
Examples include growth factors, inflammatory factors, and gases.
Acts over short distances.
Synaptic (neuronal) signaling
Occurs between neurons and their target cells.
An electrical signal travels along the neuron, causing release of a neurotransmitter at the synapse.
The neurotransmitter binds receptors on the target cell.
Very fast and highly specific.
Contact-dependent signaling
The signaling molecule is attached to the membrane of the signaling cell.
The signal is transmitted only when cells directly touch.
Important in cell development and immune responses.
Endocrine signaling
Hormones are released by endocrine cells into the bloodstream.
The hormones travel long distances to reach target cells with the appropriate receptors.
Slower but affects many cells throughout the body.
Intracellular vs. cell-surface receptors in terms of ligand type
Intracellular receptors:
Bind small, hydrophobic (lipid-souble) molecules that can cross the plasma membrane.
Examples: Steroid hormones (eg cortisol), some gases.
Cell-surface receptors:
Bind large or hydrophilic molecules that cannot cross the plasma membrane.
Examples: proteins, peptides, neurotransmitters, growth factors.
Intracellular vs. cell-surface receptors in terms of location and function of the receptor
Intracellular receptors:
Located inside the cell (cytoplasm or nucleus).
After ligand binding, the receptor-ligand complex often acts as a transcription factor, binding DNA and regulating gene transcription.
Cell-surface receptors:
Located on the plasma membrane.
When a ligand binds, the receptor activates intracellular signaling pathways (signal transduction) through proteins such as G-proteins, enzymes, or ion channels.
Intracellular vs. cell-surface receptors in terms of type of cellular response
Intracellular receptors:
Usually slower but longer-lasting responses because they involve changes in gene expression and protein synthesis.
Cell-surface receptors:
Usually faster responses because they trigger intracellular signaling cascades that modify existing proteins or activate enzymes.
Intracellular vs. cell-surface receptors in terms of receptor classes
Intracellular receptors:
Steroid hormone receptors
Cell-surface receptors:
G-protein-coupled receptors, enzyme-linked receptors, ion-channel-coupled receptors
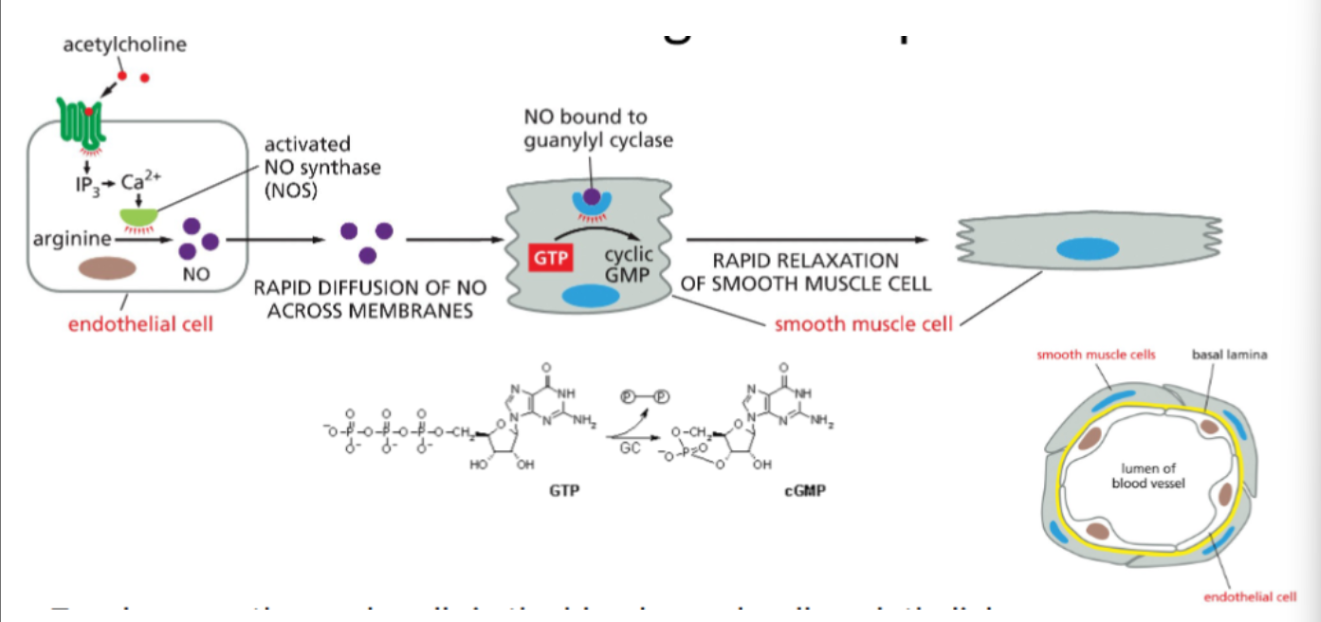
Nitric oxide signaling and smooth muscle relaxation
1. Signal reception in endothelial cells
A signaling molecule such as acetylcholine binds to receptors on endothelial cells lining the blood vessel.
2. Production of nitric oxide (NO)
The signal activates the enzyme nitric oxide synthase (NOS), which converts arginine into nitric oxide (NO).
3. Diffusion of NO
NO is a small gas molecule that diffuses rapidly across cell membranes from the endothelial cell into nearby smooth muscle cells in the vessel wall.
4. Activation of intracellular receptor
In the smooth muscle cell, NO binds to the intracellular enzyme guanylyl cyclase.
5. Production of second messenger
Activated guanylyl cyclase converts GTP into cyclic GMP (cGMP).
6. Smooth muscle relaxation
Increased cGMP triggers signaling processes that cause relaxation of smooth muscle cells, leading to dilation of the blood vessel (vasodilation).
Outline how cortisol acts to elicit a cellular response.
1. Cortisol enters the cell
Cortisol is a steroid hormone and is hydrophobic, so it can diffuse directly through the plasma membrane of the target cell.
2. Binding to an intracellular receptor
Inside the cytoplasm, cortisol binds to a cortisol receptor (an intracellular receptor protein).
The receptor is initially associated with a chaperone protein.
3. Receptor activation
When cortisol binds, the receptor changes shape and releases the chaperone protein, activating the receptor.
4. Movement into the nucleus
The cortisol–receptor complex then enters the nucleus of the cell.
5. Regulation of gene expression
The complex binds to specific DNA sequences, acting as a transcription factor that regulates gene transcription.
6. Cellular response
This leads to the production of specific mRNA and proteins, resulting in changes in cell activity such as metabolism, stress response, and immune regulation.
Extracellular signal relay
When a signaling molecule binds to a receptor on the cell surface, the signal is relayed through a series of intracellular signaling proteins.
Each activated molecule activates the next molecule in the pathway, passing the signal from the receptor deeper into the cell until it reaches target proteins.
Extracellular signal amplification
Signals are amplified during the signaling cascade.
One activated receptor can activate many intracellular signaling molecules, and each of those can activate many more downstream molecules.
This means a small external signal can produce a large cellular response.
Extracellular signal distribution
The signal can be distributed to multiple pathways inside the cell.
A single signaling molecule may activate several different proteins, allowing the signal to trigger multiple cellular responses simultaneously, such as changes in metabolism, gene expression, or cytoskeleton structure.
Extracellular signal modulation by feedback
Signaling pathways are regulated by feedback mechanisms:
Positive feedback enhances or strengthens the signal.
Negative feedback reduces or shuts down the signaling pathway after the response has occurred.
These feedback mechanisms help ensure the signal is controlled, balanced, and appropriate for the cell’s needs.
How multiple signaling pathways integrate information to produce coordinated cell response
Cells receive many extracellular signals at once, so the signals are processed through different signal transduction pathways which can interact with one another
Multiple signals received
A cell can detect several extracellular signals because it has different types of receptors on its surface or inside the cell.
Activation of different signaling pathways
Each signal activates its own intracellular signaling pathway, involving different signaling molecules and molecular events.
Integration of signals
Inside the cell, some intracellular signaling proteins can integrate signals from two or more pathways.
This can occur through a shared signaling protein or a signaling complex that receives inputs from multiple pathways.
Coordinated cellular response
The integrated signals determine which effector proteins are activated, producing a coordinated outcome such as:
survival
growth and division
differentiation
secretion
apoptosis (cell death)
Dependence on signal combination
The final response depends on the combination of signals received, meaning the same signal may cause different responses in different cells depending on other signals present.
Intracellular signaling proteins as molecular switches
They act as a molecular switch by alternating between an active (ON) and inactive (OFF) state.
When ON (active):
the protein stimulates or inhibits downstream proteins in the signaling pathway.
When OFF (inactive)
the protein does not transmit the signal.
How intracellular signaling proteins switch states
Phosphorylation / Dephosphorylation
Protein kinase adds a phosphate group (P) from ATP → often switches protein ON.
Protein phosphatase removes the phosphate group → switches protein OFF.
GTP binding / Hydrolysis
GTP binding → protein becomes active (ON).
GTP hydrolysis to GDP → protein becomes inactive (OFF).
Same signal molecules causing different responses
Same signal molecules (ligand) can cause different responses in different target cells mainly because the cells differ in receptors, intracellular signaling molecules and effector proteins.
Eg: Acetylcholine.
In skeleton muscle cells, it causes muscle contraction
In heart pacemaker cells, it decreases heart rate
In Salivary glands, it simulates secretion
Eg: Cortisol
In some cells, changes gene expression related to metabolism
In immune cells, suppresses inflammation
Why signal responses differ
Different receptors
Different cell types may have different receptors for the same signal molecule.
Each receptor activates a different signaling pathway, producing different outcomes.
Different intracellular signaling pathways
Even if cells have the same receptor, they may contain different intracellular signaling proteins, leading to different cellular responses.
Different effector proteins
The final response depends on the effector proteins present in the cell, such as:
metabolic enzymes
transcription factors
cytoskeletal proteins
GCPR’s
A large family of cell-surface receptors.
They are composed of a single polypeptide chain that spans the plasma membrane 7 times. (7 transmembrane helices).
These receptors detect extracellular signals and transmit them into the cell by interacting with G proteins on the cytosolic side.
Their function is to transmit signals by activating G proteins, which then regulate enzymes or ion channels inside the cell.
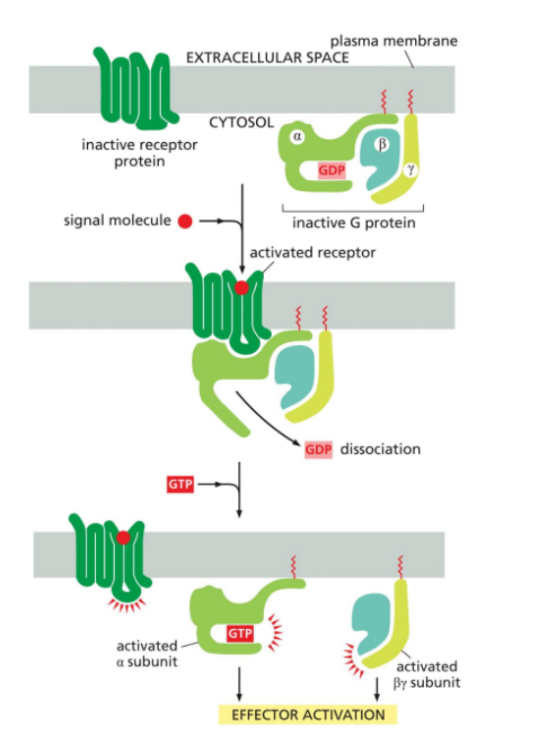
Trimeric G Proteins
Membrane-associated proteins made up of three subunits. Alpha, beta and gamma.
The alpha subunit has GTPase activity, meaning it can bind and hydrolyze GTP
The beta and gamma complex remains associated as a unit.
In their inactive state, the three subunits are bound together with GDP attached to the alpha subunit.
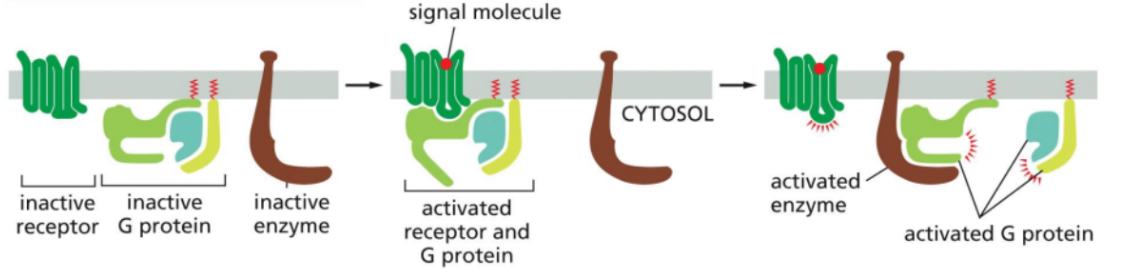
Activation of G proteins by GPCR
This occurs when a ligand (signal molecule binds to GPCR.
The GPCR undergoes a conformational change.
The activated GPCR interacts with the inactive trimeric G protein.
GDP on the alpha subunit is exchanged for GTP.
GTP binding activates the G protein.
The alpha subunit dissociated from the beta-gamma complex.
Both alpha (GTP-bound) and the beta-gamma complex can then interact and regulate target proteins such as enzymes or ion channels in the membrane.
Main enzymes targeted by G proteins
Adenylyl cyclase
Second messenger produced is cAMP (cyclic AMP)
Phospholipase C (PLC)
Second messenger produced is IP3 (inositol 1,4,5-triphosphate) and DAG (diacylglycerol)
Production of cAMP following G protein activation
A ligand (signalling molecule) binds to a GPCR, resulting in the receptor activating
GPCR activates a trimeric G protein, resulting in GDP being swapped for GTP on the alpha subunit.
The activated alpha subunit (GT-bound) dissociated from the beta-gamma complex.
The alpha subunit binds to and activates adenylyl cyclase.
Adenylyl cyclase converts ATP to cAMP
cAMP acts as a second messenger and activates downstream signalling.
Caffeine’s effect on the cAMP signalling pathway
In normal cells, cAMP is continuously being produced (by adenylyl cyclase) and broken down (by the enzyme cAMP phosphodiesterase) into AMP, turning off the signal.
Caffeine inhibits cAMP phosphodiesterase.
As a result, cAMP is not broken down as quickly and begins to accumulate in the cell.
This results in a stronger and longer-lasting signalling effect because the second messenger (cAMP) remains active and continues activating downstream targets.
Basically, caffeine amplifies and prolongs cAMP signalling.
Function of IP3 (Inositol 1,4,5-triphosphate)
It’s a secondary messenger generated by activated phospholipase C.
It’s soluble, so it diffuses through the cytosol and binds to IP3 receptors on the Endoplasmic Reticulum (ER).
This causes Ca2+ to be released into the cytoplasm, which increases the intracellular calcium levels.
This increase in calcium levels triggers various cellular responses, such as secretion or contraction.
Function of DAG (diacylglycerol)
It’s a secondary messenger generated by activated phospholipase C.
It’s hydrophobic; as a result, it remains embedded in the plasma membrane.
Its main function is to help activate protein kinase C (PKC), especially in combination with the increased Ca2+ levels produced by IP3 signalling.
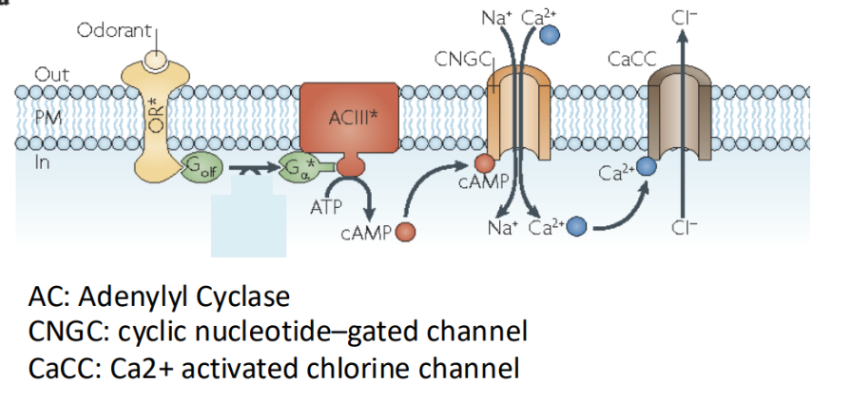
GPCR’s role in odour perception
In this, odorant molecules bind to GPCRs located in olfactory receptor cells.
This activates a G protein, which then stimulates adenylylcyclase to produce cAMP.
The increase in cAMP opens cyclic nucleotide-gated ion channels, resulting in depolarization of the cell and initiation of a nerve signal to the brain.
GPCR’s role in taste perception
In this (specifically for sweet, bitter and umami tastes), molecules bind to GPCRs on taste receptor cells.
This activates the phospholipase C pathway.
This results in the roduction of IP3, which increases intracellular Ca2+ levels.
The rise in calcium triggers the release of neurotransmitters that send signals to sensory neurons.
GPCRs role in light perception
In this, GPCRs are involved in vision through a receptor called rhodopsin (found in rod cells of the retina).
When light activates rhodopsin, it activates a G protein (transducin), which activates an enzyme that breaks down cGMP.
The decrease in cGMP results in ion channels closing, resulting in hypepolarization of the cell, and a change in neurotransmitter release.
This change in NT release is interpreted as a light signal.
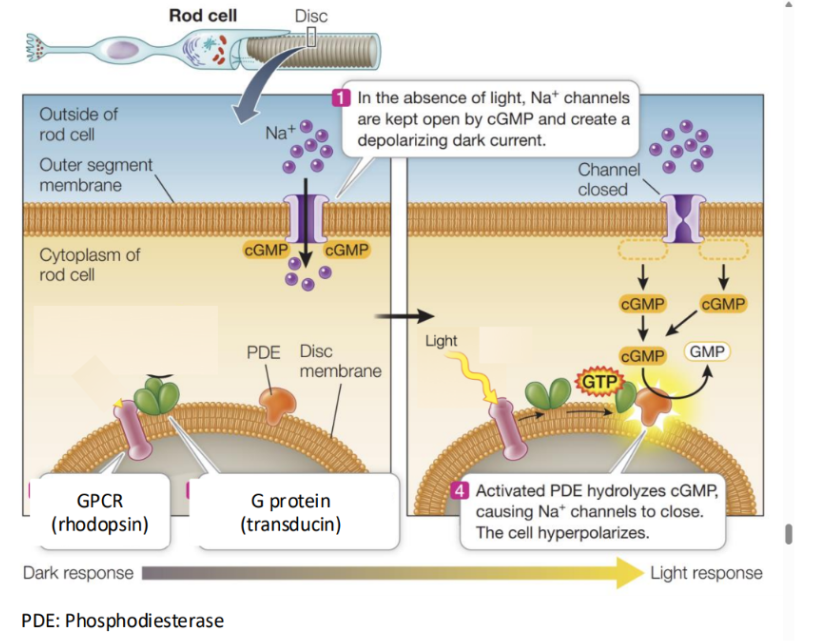
How light signals are amplified in rod photoreceptor cells.
This amplification is done through a GPCR-based signaling cascade.
When light activates the GPCR rhodopsin, it activates many molecules of the G protein transducin.
Each activates transducin then actgivates a molecule of cGMP phosphodiesterase (PDE).
This creates a cascade effect.
Basically each step multiplies the signal:
One activated rhodopsin activates hundreds of g proteins.
Each G protein activates PDE.
Each PDE rapidly breaks down many molecules of cGMP.
As cGMP levels drop, a large number of cGMP-gated Na+ channels close, resulting in a strong hyperpolarization.
Cell-surface receptors
Fall into three main classes.
G protein-coupled receptors (GPCRs)
Enzyme-coupled receptors
Ion-channel-coupled receptors.
They differ in both structure and how they transmit signals across the membrane.
Structure & Function of Ion-channel-coupled receptors
Structure:
They form transmembrane channels that can open or close in response to ligand binding
Function:
They function more directly by opening or closing ion channels. This allows ions to flow across the membrane and rapidly change the cell’s membrane potential.
Structure & Function of enzyme-coupled receptors
Structure:
They typically have a signal transmembrane domain and a cytoplasmic portion that either has enzymatic activity itself or is directly associated with an enzyme.
Function:
They function by activating intracellular enzymes or signalling pathways (such as phosphorylation cascades), resulting in longer-term cellular responses like growth, differentiation or survival.
RTK activation
Occurs when a signalling molecule, such as a growth factor or hormone, binds to the extracellular domain of the receptor.
This binding causes two RTK molecules to come together, forming a dimer.
Dimerization activates their cytoplasmic kinase domains, allowing each receptor to phosphorylate tyrosine residues on the other receptor’s tail. (known as autophosphorylation)
These phosphorylated tyrosines then serve as docking sites for intracellular proteins, which bind to the receptor and become activated.
This initiates downstream signalling pathways.
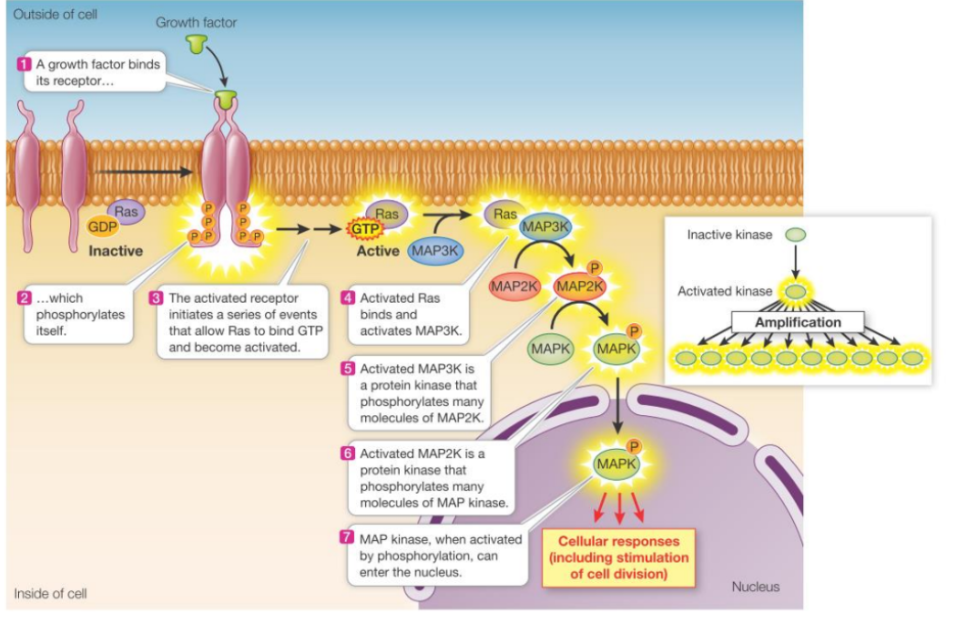
Ras-MAPK pathway
A major pathway activated by RTKs
After RTK activation, adaptor proteins and a guanine nucleotide exchange factor (Ras-GEF) are recruited to the receptor.
Ras-GEF activates the small (monomeric) G protein Ras by promoting the exchange of GDP for GTP.
The activated Ras (GTP-bound) then triggers a phosphorylation cascade involving a series of protein kinases. (MAP kinase cascade).
This cascade ultimately leads to changes in protein activity and gene expression, such as promoting cell growth & division.
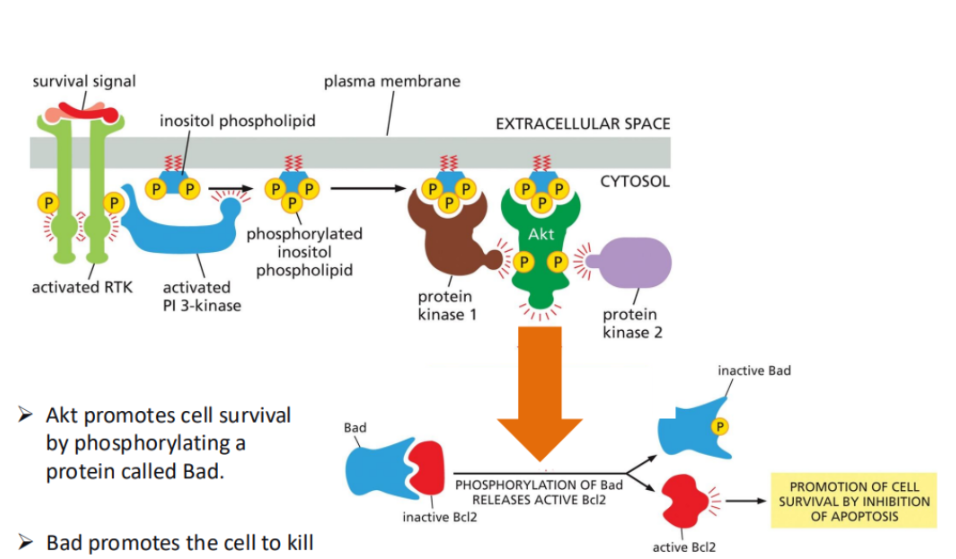
PI3K-Akt pathway
A major pathway activated by RTKs.
In this, activated RTKs recruit and activate PI3K-kinase (PI3K).
Pi3K phosphorylates a membrane lipid (PIP2) to produce PIP3, which acts as a docking site for signalling proteins like Akt (protein kinase B).
Once activated, Akt promotes cell survival by phosphorylating target proteins, such as inhibiting pro-apoptotic proteins (e.g., Bad), thereby preventing cell death.
Mechanisms for terminating RTK signalling
These exist because signalling through RTK’s must be tightly controlled to prevent excessive or uncontrolled cellular responses.
RTK dephosphorylation
RTK removal from the cell membrane.
Ras inactivation (turn off)
RTK dephosphorylation
A mechanism for terminating RTK signalling.
In this, the RTK receptors are dephosphorylated (removing phosphate groups that are added during activation).
This inactivates the receptor’s signalling ability, since the phosphorylated sites are required for recruiting and activating downstream signalling proteins.
RTK removal from the cell membrane
A mechanism for terminating RTK signalling.
In this, RTK’s are removed from the cell membrane, often through internalization (endocytosis).
Once removed, they are either degraded or recycled, which prevents them from continuing to interact with signalling molecules.
Ras inactivation (turn off)
A mechanism for terminating RTK signalling.
In this, signalling is shut off at the level of downstream proteins. Particularly Ras.
Ras is a protein that becomes inactive when it hydrolyzes its bound GTP back to GDP. (When its GTP turns into GDP).
This hydrolyzation process is accelerated by Ras-GAP (GTPase-activating protein), which helps turn Ras “off”.
This stops the signalling cascade.
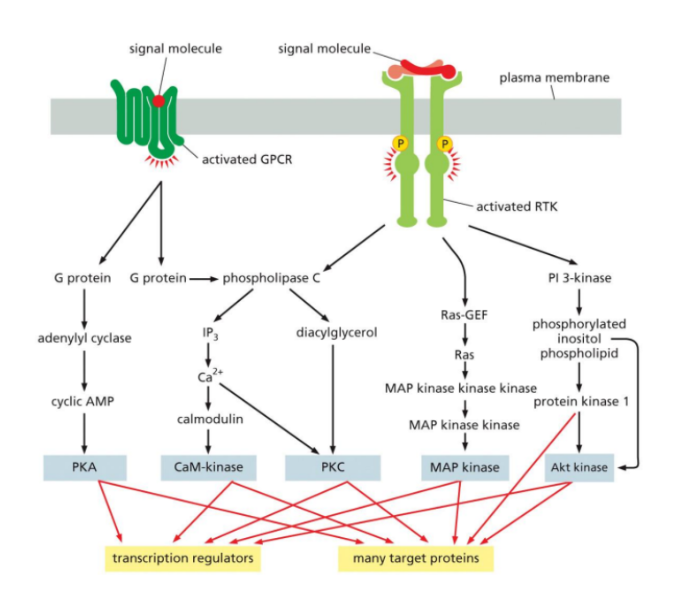
Interaction between the components of a signal transduction pathway
The signal transduction pathway begins when a ligand binds to a specific cell-surface receptor. (Such as GPCR or RTK).
This activates the receptor and initiates a series of intracellular events.
The activated receptor then interacts with intracellular signaling proteins, such as G proteins or adaptor proteins, which relay the signal further into the cell.
From here, the signal is transmitted through a network of pathways.
For example, in GPCR signaling, activated G proteins can stimulate enzymes like adenylyl cyclase or phospholipase C, producing second messengers (such as cAMP, IP₃, or DAG).
In RTK signaling, adaptor proteins activate Ras, which triggers a protein kinase cascade (Ras-MAPK pathway), or activate PI3K, leading to Akt activation.
These pathways often interconnect and branch, meaning one signal can activate multiple pathways at once.
The signal is amplified as it moves through cascades (especially phosphorylation cascades), allowing a small initial signal to produce a large cellular response.
Finally, the signal reaches target proteins, such as enzymes or transcription regulators. These targets produce the final cellular response, such as changes in gene expression, protein activity, cell growth, or survival.
Cytoskeleton
A highly dynamic network of protein filaments that extends throughout the cytoplasm, and plays a central role in maintaining the overall structure & function of the cell.
It’s essential for cell shape (due to providing mechanical support), and supports the shape through structures like microfilaments and intermediate filaments.
Important for internal organization, cell function and cell reproduction.
It’s composed of microtubules, intermediate filaments, and microfilaments (actin filaments)
Importance of the cytoskeleton for internal organization
It helps position organelles within the cell, and provides tracks for their movement.
Microtubules, for example, orgsnize the placement of organelles such as the golgi apparatus, and endoplasmic reticulum.
Motor proteins transport vesicles and other materials along these filaments.
This ensures that cellular components are properly distributed, and can function efficiently.
Importance of the cytoskeleton for cell function
It enables processes such as intracellular transport, cell movement and muscle contraction.
Microtubules allow the movement of vesicles and organelles, while microfilaments are involved in cell motility and contraction.
Structures like cilia and flagella, which are built from microtubules, allow cells to move or move fluids across their surface.
Importance of the cytoskeleton for cell reproduction.
During cell division, microtubules form the mitotic spindle, which separates chromosomes into daughter cells.
Microfilaments also form a contractile ring during cytokinesis, allowing the cell to physically divide into two.
Without these cytoskeletal components, proper cell division wouldn’t occur.
Microtubules
The largest filaments (approx 25nm in diameter).
Are made of tubulin dimers (alpha & beta) that assemble into hollow, rigid cylinders.
They are polar structures, with a positive and negative end that can grow and shrink dynamically.
Location: Found throughout the cytoplasm and often extend outward from centrosomes.
Function: Involved in intracellular transport, organelle positioning, cel division (mitotic spindle) and movement of cilia and flagella.
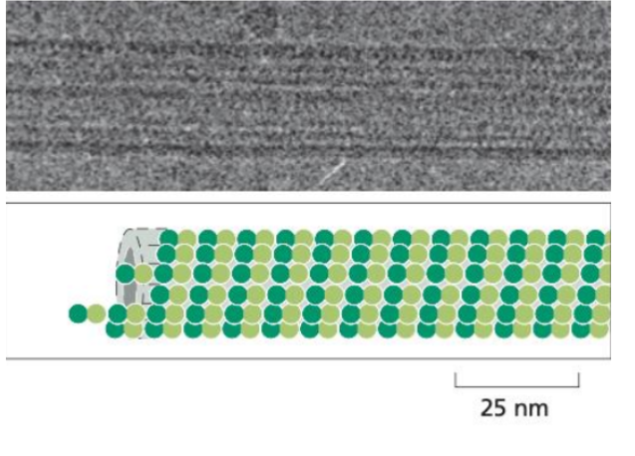
Microfilaments (Actin filaments)
The smallest filaments (7nm)
Composed of actin proteins arranged into helical polymers.
They are polar, with a positive and negative end that allows rapid assembly and disassembly.
Location: Mainly located in the cell cortex (just beneath the plasma membrane)
Function: Responsible for cell shape, cell movement, muscle contraction and cytokinesis.
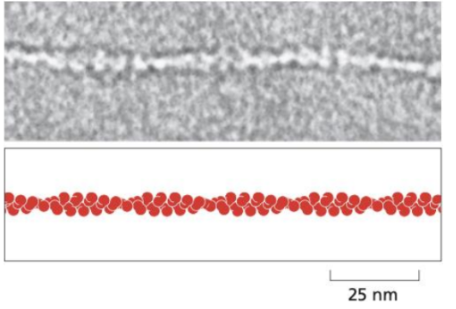
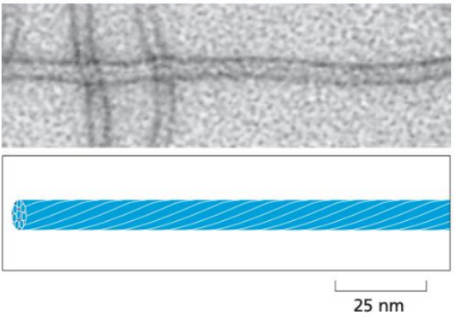
Intermediate filaments
About 10nm in diameter
Made of fibrous proteins that form rope-like structures.
Non-polar, and more stable as they don’t rapidly assemble and disassemble like the other two.
Location: They form a network that spans the cytoplasm, often connecting cell-cell junctions and providing structural integrity across the cell.
Function: Primarily provide mechanical strength and support, helping cells withstand stress.

How the structure of intermediate filaments give it strength and durability
They are made of fibrous proteins that assemble into rope-like fibres, which are both flexible and tough.
This rope-like arrangement allows them to withstand stretching and mechanical strain without breaking.
These filaments are non-polar and less dynamic than the other filaments. This stability allows them to form long-lasting networks that span the cytoplasm and connect cell-cell junctions.
At the mechanical level:
Intermediate filament proteins first form coiled-coil dimers, which then assemble into tetramers.
Multiple tetramers (about 8) associate laterally to form a thick, stable filament.
This layered, staggered assembly distributes force, resulting in the filament being highly resistant to breakage.
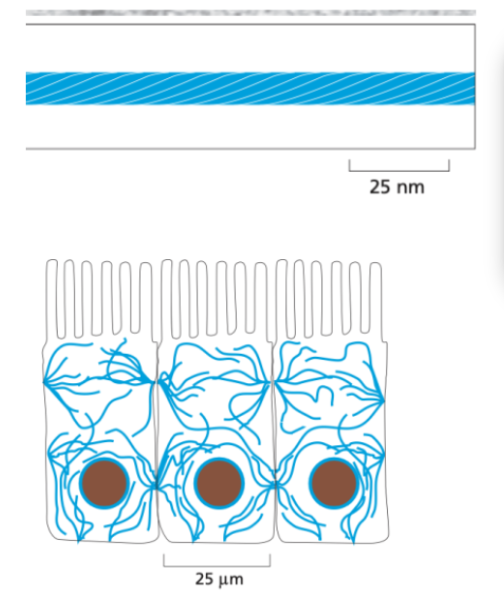
Microtubule assembly & disassembly
They assemble from tubulin dimers (alpha & beta) that polymerize to form protofilaments, which then combine into a hollow tube.
This assembly mainly occurs at the plus end, and they exhibit dynamic instability.
Dynamic instability means they rapidly switch between growth (polymerization) and shrinkage (depolymerization).
This behaviour is regulated by GTP binding and hydrolysis on tubulin, allowing microtubules to quickly reorganize when needed. (eg, cell division)
Microfilament assembly & disassembly
These assemble from actin monomers that form helical polymers.
Due to their polarity, they continuously undergo polymerization & depolymerization.
Their dynamics are regulated by nucleotide binding (ATP on actin), allowing rapid remodelling for processes like cell movement and shape changes.
Intermediate filament assembly & dissassembly
These assemble from fibrous proteins that form dimers, tetramers, and then thicker rope-like structures.
Due to being non-polar and not relying on nucleotide hydrolysis (no ATP or GT), their assembly is more stable and less dynamic, and they don’t undergo rapid cycles of growth and shrinkage.
This makes them more resistant to disassembly and better suited for providing long-term structural support.
Motor proteins
Specialized proteins that move along cytoskeletal filaments to transport materials within the cell.
Main ones are kinesins (move towards + end) and dyneins (move towards - end), which are associated with microtubules. They act as tracks for intracellular transport.
The result is that they transport cargo in opposite directions.
Both proteins are responsible for transporting vesicles, organelles and macromolecules throughout the cell.
Motor proteins associated with microfilamins (actin), such as myosin, function mainly in muscle contraction and cell movement, rather than long-distance intracellular transport.
These proteins interact with actin filaments to generate force and movement, such as sliding filaments during contraction.
Kinesins
A motor protein.
Move towards the + end of microtubules
Moves cargo outward from the center of the cell toward the periphery.
Dyneins
A motor protein
Moves towards the - end
They move cargo inward toward the centrosome.
Also has an additional role in cilia and flagella movement, where its activity causes bending of microtubules to generate motion.
Dynamic instability
Refers to the ability of microtubules to rapidly switch between periods of growth (polymerization) and shrinkage (depolarization).
Instead of being stable, each microtubule constantly switches between the two states, allowing the cytoskeleton to be highly adaptable and reorganize quickly when needed.
This is essential because it allows them to quickly reorganize the internal structure of the cell, which is especially important for cell division, intracellular transport, and changes in cell shape.
Control of dynamic instability
It’s controlled by GTP hydrolysis on tublin dimers, which add to the growing + end of a microtubule in a GTP-bound ofrm, which stabilizes the structure.
After incorporation, the GTP is hydrolyzed to DGP.
GDP-bo0und tublin is less stable, so if the GTP cap at the growing end is lost, the microtubuline becomes unstable and rapidly depolymerizes (a process called catastrophe).
If new GTP-bound tubulin is added again, growth can resume.
The four phases of the eukaryotic cell cycle
G1, S, G2, (these three make up interphase) and M (mitotic) phase.
Each phase has specific roles in preparing the cell for division.
G1 Phase (Gap 1)
The first stage of the eukaryotic cell cycle
In this, the cell grows in size, produces new proteins and organelles, and begins duplicating its centrosome.
This phase prepares the cell for DNA replication in the next phase.
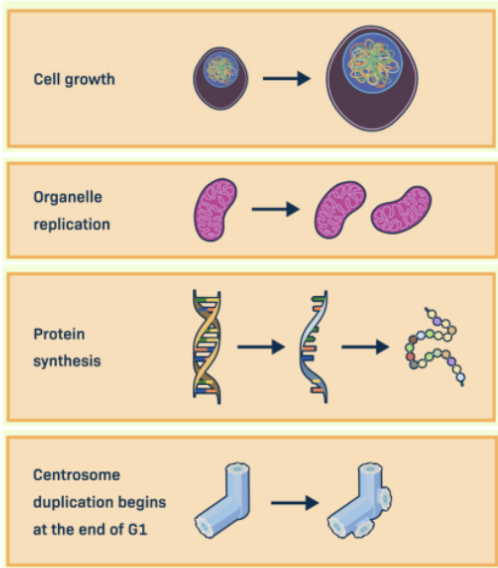
S Phase (synthesis)
The second stage of the eukaryotic cell cycle.
In this, the cell replicates its DNA, producing a complete copy of each chromosome (forming sister chromatids).
The cell also continues to grow and duplicate organelles and centrosomes.
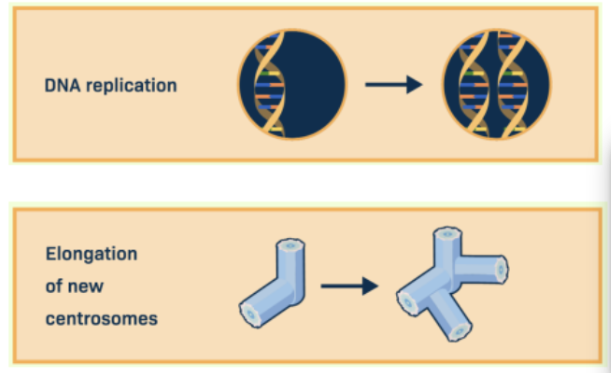
G2 Phase (Gap 2)
The third stage of the eukaryotic cell cycle
In this, the cell continues to grow and prepares for division by producing ATP, synthesizing proteins required for mitosis, and maturing duplicated centrosomes.
This phase ensures the cell is ready to enter mitosis.
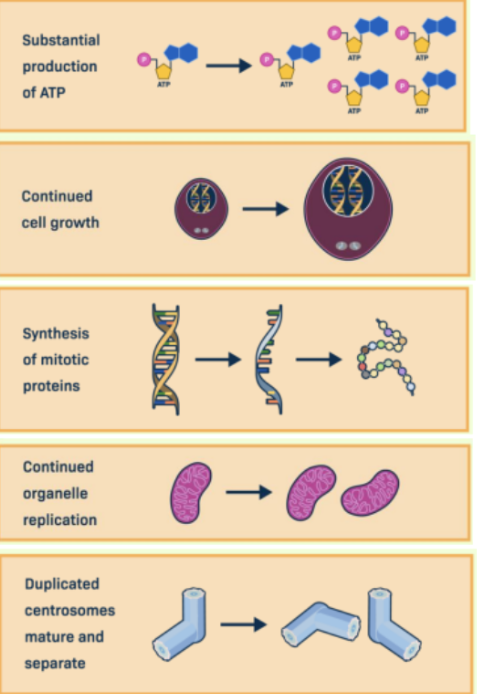
M Phase (Mitotic Phase)
The fourth stage of the eukaryotic cell cycle.
This phase includes mitosis and cytokinesis.
In mitosis, the duplicated chromosomes are separated into two identical sets.
During cytokinesis, the cytoplasm divides to form two daughter cells.
Five stages of mitosis
Prophase, prometaphase, metaphase, anaphase and telophase.
Each stage involves specific changes to ensure accurate chromosome separation.
Prophase
The first stage of mitosis
In this, chromosomes begin to condense and become visible.
Cohesin proteins hold sister chromatids together, while condensin proteins help compact the chromosomes.
At the same time, the mitotic spindle starts to form between the two centrosomes.
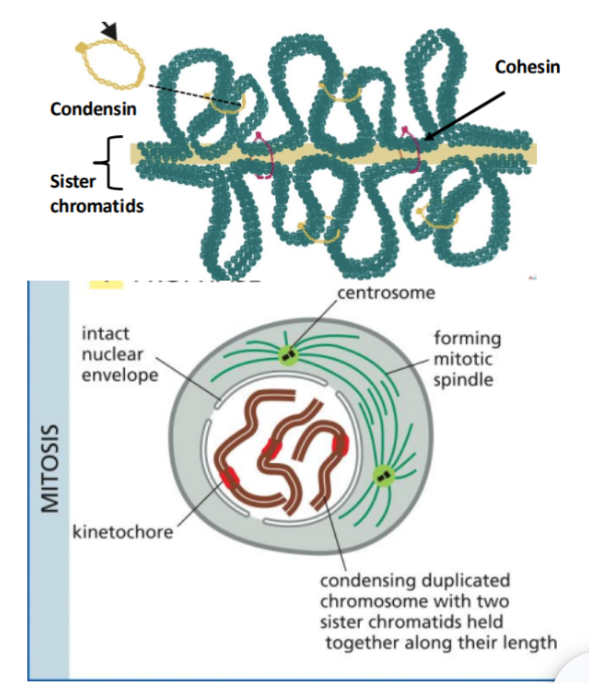
Prometaphase
The second stage of mitosis.
It begins with the breakdown of the nuclear envelope.
This allows spindle microtubules to access the chromosomes and attach to them at structures called kinetochores, which are located at the centromeres.
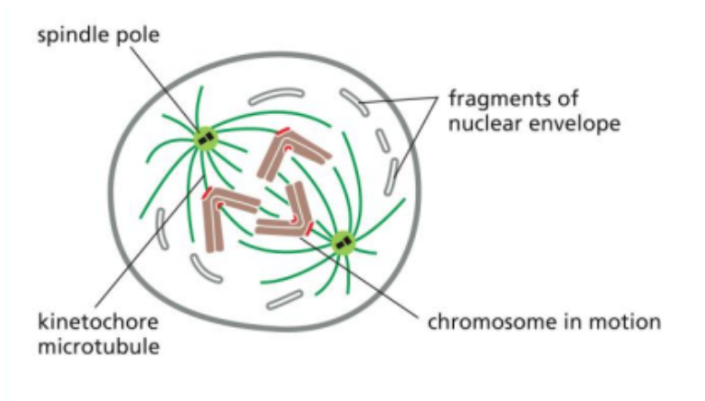
Metaphase
The third phase of mitosis
In this, all chromosomes are fully attached to spindle microtubules and aligned at the center of the cell (metaphase plate).
There are three types of spindle microtubules involved: kinetochore, interpolar, and astral microtubules.
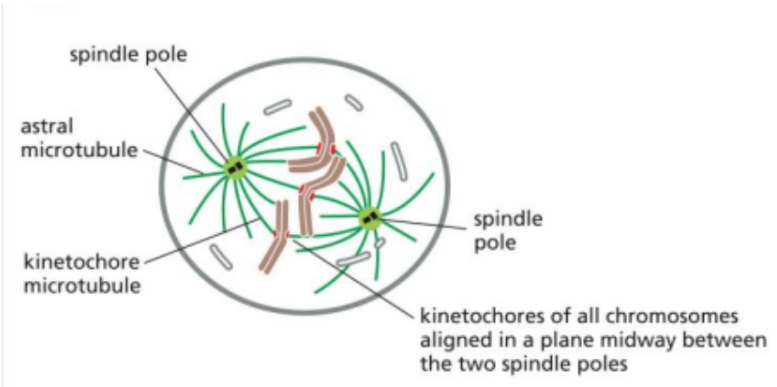
Anaphase
The fourth phase of mitosis
During this, the cohesin proteins are cleaved, allowing sister chromatids to separate.
The chromatids are then pulled toward opposite poles of the cell by shortening kinetochore microtubules and other forces within the spindle.
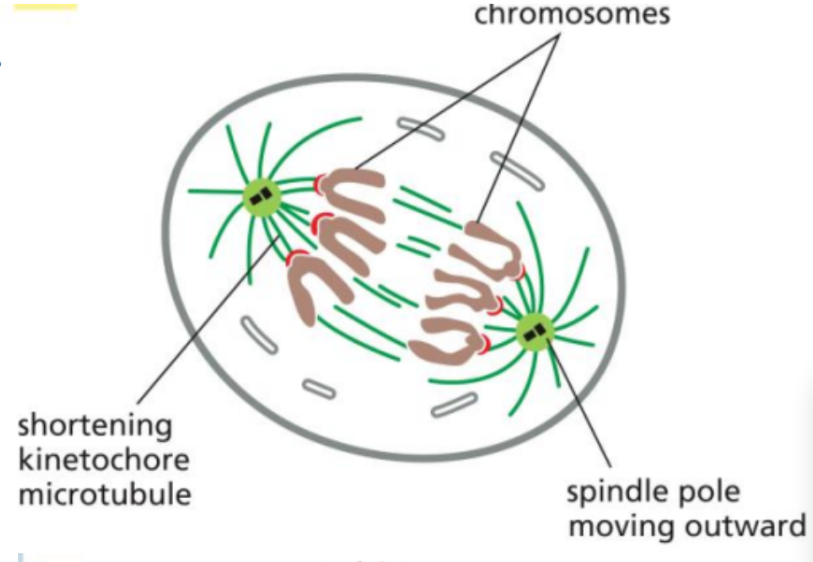
Telophase
The fifth stage of mitosis
In this, a new nuclear envelope reforms around each set of chromosomes at opposite poles.
The chromosomes begin to decondense, and the cell prepares to complete division as cytokinesis begins.
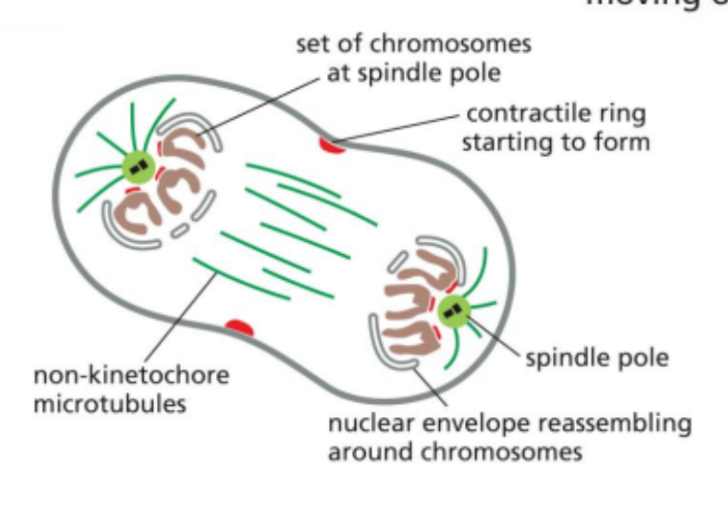
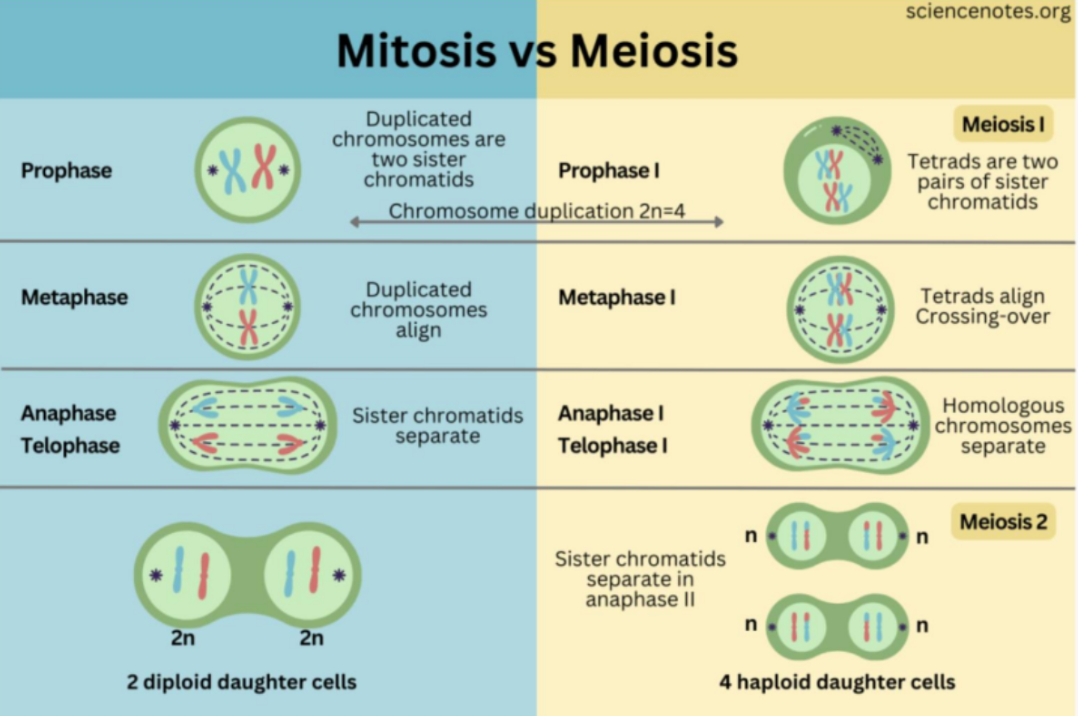
Chromosome segregation in meiosis vs mitosis
In mitosis:
Duplicated chromosomes (sister chromatids) align individually at the metaphase plate, and during anaphase, they are separated and pulled to opposite poles.
This results in two genetically identical daughter cells, with the same chromosome number as the parent. (diploid cell to diploid cell)
In meiosis, there’s two rounds.
In meiosis I, homologous chromosomes (each consisting of 2 sister chromatids) pair up and align together, and then the homologous chromosomes (not sister chromatids) are separated.
This reduces the chromosome number by half (diploid cell to hapolid cell).
In meiosis II, the process resembles mitosis, where sister chromatids are serrated.
The final result is four genetically distinct haploid cells.
The main difference is that meiosis introduces genetic variation, as crossing over occurs between homologous chromosomes (genetic info exchange)
The cell cycle control system
A network of regulatory proteins that drives the cell through different phases of the cell cycle and ensures events occur in the correct order.
It also has the ability to pause the cycle if a problem is detected, such as DNA damage or incomplete processes, ensuring proper cell division.
The main components of this system are cyclins, cyclin-dependent kinases (Cdks) and Cdk inhibitors (CKIs), each with a specific role.
Cyclins
Regulatory proteins whose levels rise and fall cyclically during the cell cycle.
They control the timing of cell-cycle events by activating Cdks only at the appropriate stages.
Cyclin-dependent kinases (Cdks)
Enzymes (kinases) that become active only when bound to a cyclin.
Once activated, they phosphorylate target proteins, triggering key processes such as DNA replication, mitosis and progression to the next phase of the cycle.
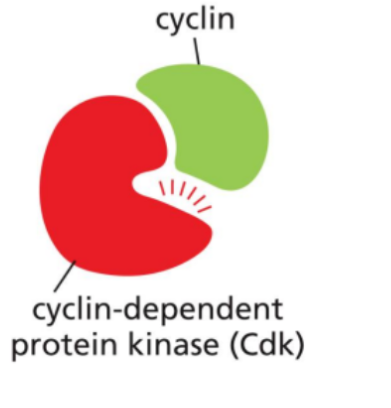
Cdk inhibitors (CKIs)
Proteins that bind to and inhibit cyclin-Cdk complexes, preventing cell-cycle progression when necessary.
They act as tumour suppressors by stopping the cycle if conditions are not suitable (e.g., DNA damage)
Mechanisms that regulate Cdk activity
3 main mechanisms
Cyclin degradation
Cdk phosphorylation and dephosphorylation
Binding to a Cdk inhibitor (ex:p27, p21)
Cyclin degradation
A mechanism that regulates Cdk activity
Since Cdk activity depends on cyclin binding, Cdks are inactive on their own and only become active when they bind to specific cyclins.
Since cyclin levels rise and fall during the cell cycle (through synthesis and degradation), this ensures that Cdks are only active at the appropriate stages.
Cdk phosphorylation and dephosphorylation
A mechanism that regulates Cdk activity
The addition or removal of phosphate groups on Cdks can either activate or inhibit their activity, providing another level of control over when they are turned on or off.
Binding to a Cdk inhibitor (eg: p27, p21)
A mechanism that regulates Cdk activity
Cdks are controlled by Cdk inhibitors (CKIs).
These proteins bind directly to cyclin-Cdk complexes and block their activity, preventing progression through the cell cycle when conditions are not suitable (such as DNA damage)
Cell-cycle checkpoints
These act as surveillance mechanisms to ensure that each stage of the cell cycle is completed correctly before the cell progresses to the next one.
They are the G1, G2, and M checkpoints.
G1 Checkpoint
A cell cycle checkpoint
It occurs before the cell enters the S phase.
At this point, the cell assesses whether conditions are favorable for DNA replication, such as sufficient cell size, nutrients, and the presence of growth signals (mitogens).
It also checks for DNA damage.
If conditions are not suitable, the cell cycle is paused or the cell may enter a resting state.
The G2 checkpoint
A cell cycle checkpoint
Occurs after DNA replication and before mitosis.
Here, the cell verifies that DNA has been fully and accurately replicated and checks again for DNA damage.
If problems are detected, the cycle is halted to allow for repair before entering mitosis.
The M checkpoint (spindle checkpoint)
A cell cycle checkpoint.
Occurs during mitosis, specifically before the separation of sister chromatids.
It ensures that all chromosomes are properly attached to spindle microtubules via their kinetochores.
If any chromosomes are not correctly attached, the checkpoint prevents progression to anaphase by blocking the activity of the anaphase-promoting complex (APC), ensuring accurate chromosome segregation.
Mitogens
Extracellular signalling molecules that stimulate cells to enter the cell cycle.
They do this by activating signalling pathways that lead to G1/S-Cdk activation.
This promotes expression of genes required for DNA replication (via factors like E2F), allowing the cell to enter the S phase.
Entry into the cell cycle is regulated by the cell-cycle control syste, which controls cyclin-Cdk activity.
If mitogens are present and conditions are favourable, Cdks are activated, and the cell progresses.
If not, Cdk inhibits block progression, preventing cell division.
The role of DNA damage in arresting cells in G1
If DNA damage is detected during the G1 phase, the cell-cycle control system activates mechanisms that pause the cell to prevent entry into the S phase.
This is mainly achieved through activation of CKIs (Cdk inhibitors), which bind to and inhibit G1/S-Cdk complexes, blocking their activity.
Since these Cdks are required to trigger DNA replication, their inhibition prevents the cell from progressing further in the cycle.
This arrest gives the cell time to repair the damaged DNA before replication occurs.
The effect of chromosomes failing to attach
During metaphase, all chromosomes must be properly attached to spindle microtubules at their kinetochores before the cell can proceed to anaphase.
If even one chromosome is not correctly attached, the M checkpoint (spindle checkpoint) is activated.
Unattached kinetochores generate a signal that inhibits the anaphase-promoting complex (APC), a key protein required to trigger the transition to anaphase.
Because APC remains inactive, cohesin proteins are not cleaved, so sister chromatids can’t separate.
As a result, the cell cycle is halted at metaphase until all chromosomes are properly attached.