BIOC 3560
1/42
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
43 Terms
signals
specific - signal molecule fits binding site on it complementary receptor, other signals dont fit, some receptor are present only in certain cell types
amplification - when enzymes activate enzymes the number of affected molecules increase in an enzyme cascade
modularity - proteins with multivalent affinities form diverse signaling complexes from interchangeable parts, phphylation provides reversible points of interaction
desensitization/adaptation - receptor activation triggers a feedback circuit that shuts off the receptor or removes it from cell surface, when stimulus falls below certain threshold the system is active again
integration of signals - when 2 signals have opposite effects that outcome results from the input on both receptors, unified response
types of signal transducers
G protein-coupled receptor - external ligan binds to GTP-binding protein which produces second messenger
receptor enzymes - ligand binding activates activity by autophph
gated ion channel - opens or closes in response to concentration of signal ligand or membrane potential
nuclear receptor - hormone binding allows receptor to regulate expression of specific genes
neuron ion gradient
neuron has high [K+] and low [Na+]'
at rest inside cell is ΔΨ=-60mV
action potential carries electrical signal down axon and neurotransmitter carries signal to next cell
nicotinic acetylcholine receptor (AchR)
example of neurotransmitter
passage of electrical signal from motor neuron to muscle fiber at neuromuscular junction
acetylcholine released by motor neuron diffuses to pm of myocyte → binds AchR
conformational change in AchR → open and activated
inward movement of cations (ion channels go down concentration gradient)
trigger muscle contraction
has 5 subunits with 4 helices in each
neural transmission
acetylcholine (Ach) opens Ach receptors (ligand gated Na+/Ca2+ channel)
Na+ flows in (down gradient) → depolarization
adjacent voltage-gated Na+ channels open: Na+ rushes in → ΔΨ=+30mV
Na+ channels inactivated
K+ channels opens: K+ flows out (down gradient) → ΔΨ=-75mV
K+ channels inactivated → ΔΨ=-60mV
heterotrimeric G protein-coupled receptors
glucogon and epinephrine
has pm receptor with 7 transmembrane helices
heterotrimeric guanosine nucleotide-binding protein (G-protein)
intracellular enzyme that generates second messenger
Gs is stimulatory
Gi is inhibitory
epinephrine signal transduction pathway
Gα has GDP bound at rest
epinephrine binding receptor promotes GTP-binding at Gα, activating it
Gα dissociates from receptor moving to adenylyl cyclase and activates it
catalyzes formation of cAMP
cAMP activates PKA which phph cellular protein
what drives epinephrine receptor (internal)
Internalization of the epinephrine receptor is induced by phosphorylation of the receptor by β-adrenergic receptor kinase (βARK) and subsequent binding of β-arrestin.
insulin receptor
insulin bind externally and activates tyrosine kinase activity in intracellular domain
β-chains are autophph which opens up active site
insulin receptor phph Tyr
SH2 domain of Grb2 binds to phphTyr of IRS1
Sos binds to Grp2 than Ras releasing GTP
activated Ras binds and activated Raf-1
Raf-1 phph MEK which phph ERK
what blocks insulin receptor
tyrosine kinase catalytic site is blocked by its activation loop when its inactive
loop is Tyr which H bonds with Asp to keep in position
when insulin bind Tyr kinase phph all Tyr to stabilize loop so it wont block catalytic site
insulin activates what in muscle cells
increase glucose transport by recruiting GLUT4 to membrane
induce synthesis of hexokinase
activate glycogen synthase by phph of GSK3
effects of insulin
reduced phph of glycogen synthase → increased activity and glycogen synthesis by glycogen synthase
movement of glucose transporter to pm → brings more glucose into cell
modulation of insulin-responsive transcription factors
nuclear hormone receptor
hydrophobic hormone diffuses across membrane and binds to receptor protein in nucleus
binding causes conformational change of receptor to form dimer - receptor has zinc finger to allow it to bind to specific DNA sequences
receptor attracts transcription regulating proteins to either increase or decrease mRNA formation
cellular response
phosphoglycerides
glycerol backbone
2 FA in ester link
alcohol head group
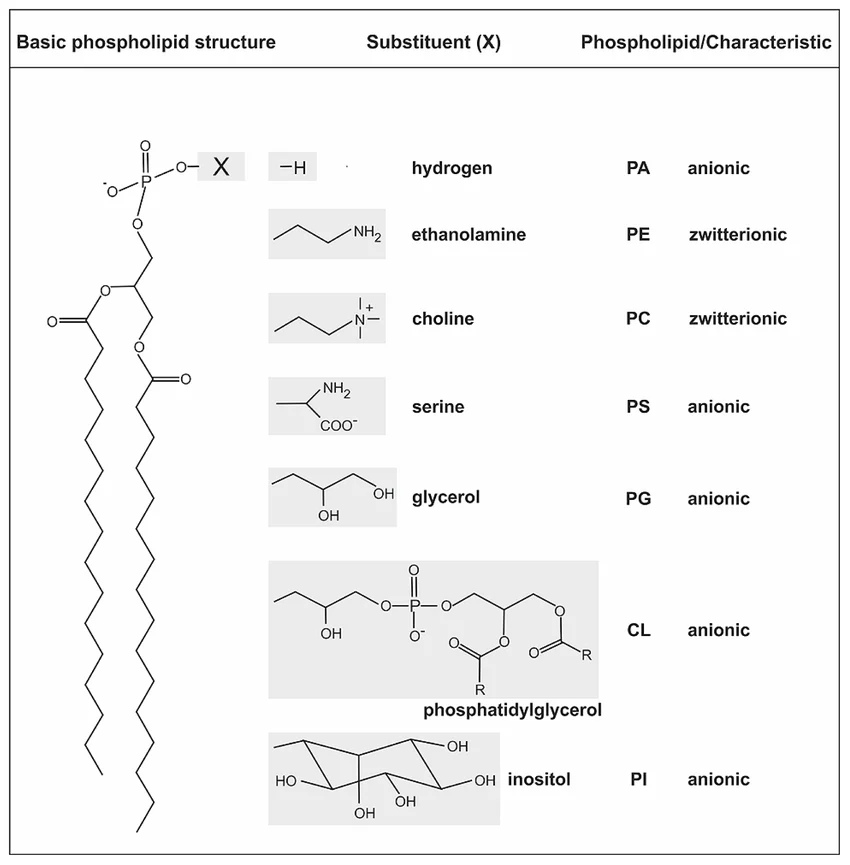
sphingolipids
sphingosine backbone
1 FA in amide link
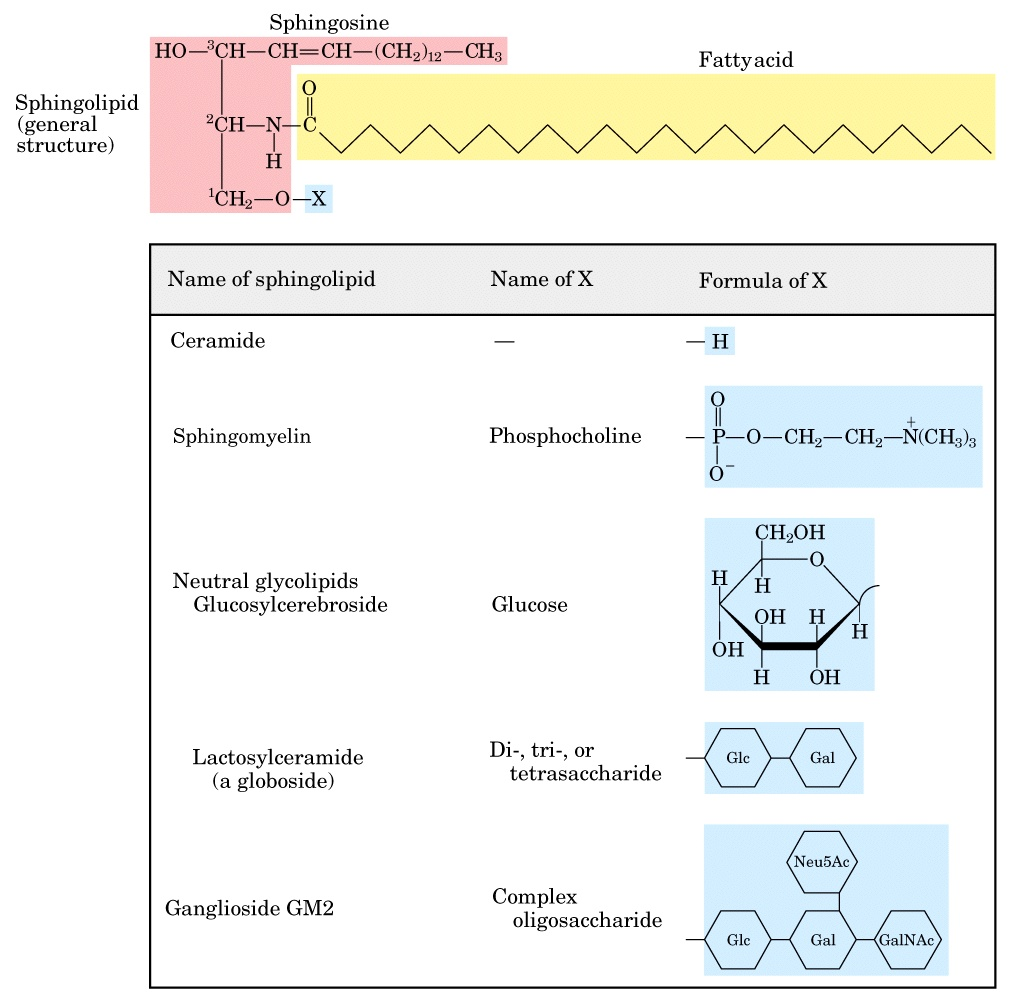
glycolipids
sphingolipids with carbohydrate headgroups
what is the major lipid in all membranes? what is major component of pm?
phosphatidyl choline
cholesterol
how can membrane proteins be removed?
pH changes or Ca2+ is removed → peripheral membrane protein removed
detergent → integral membrane protein removed
phospholipase → lipid anchored membrane protein removed
amphitropic can attach and leave independently
types of integral membrane proteins
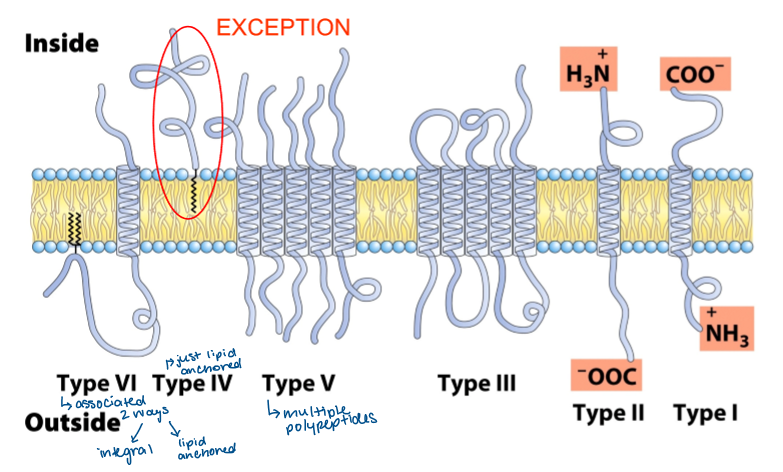
hydropathy plots
predict the number of transmembrane helices a protein might have
why dont beta barrel integral membrane proteins show up on hydropathy plots?
alternating hydrophobic, hydrophilic sheets that arent long enough
what aa are found where transmembrane protein meets headgroup of membrane
Tyr and Typ (hydrophobic and polar)
charged aa are unfavourable in hydrophobic environments → found mostly in aqueous phase
N-linked carbohydrate chain is attached to what
Asn side chain
O-linked carbohydrate chain is attached to what
Ser and Thr
what do the sugar groups of glycoproteins and glycolipids do?
contribute to cell surface recognition
function as receptors
lipid bilayer state changes
gel phase (cold) → all motion is constrained, lipids ordered in paracrystalline state
liquid-ordered state (physiological) → intermediate thermal motion, lateral movement in the plane of bilayer
liquid-disordered state (fluid) → hydrocarbons chains in constant motion with no regular organization
what state do different FA favour?
long chain, saturated FA pack into liquid ordered
short chain, unsaturated favour liquid disorders
behaviour of cholesterol on membrane fluidity
long saturated → cholesterol INCREASES fluidity -
unsaturated cis → cholesterol DECREASES fluidity
high temps → cholesterol DECREASES fluidity
low temps → cholesterol INCREASES fluidity
cholesterol is a fluidity buffer → does opposite of what composition/temp does to maintain good fluidity levels of membrane
enzymes that help move lipids in pm
flippase - P-type ATPase that moves PE and PS from outer to cytosolic leaflet
floppase - ABC transporter that moves phospholipid from cytosolic to outer leaflet
scramblase - moves lipids in either direction toward equilibrium (no ATP)
what restricts lipid/protein motion in pm
spectrin is part of cytoskeleton and links to membrane proteins and keeps lipids from diffusing freely
lipid rafts
sections of pm that are enriched in sphingolipids and cholesterol
simple diffusion
spontaneously from high to low concentration
what does a membrane transporter help with?
decreases the amount of free energy needed to transport hydrophilic solutes across membrane
membrane channels
pore that spans bilayer
solutes flow through rapidly compared to transporters
they are gated - open/close in response to stimuli
highly selective
membrane transporter classification
uniport → single molecule down concentration gradient
symport → 2 different molecules across in same direction (one down concentration gradient other up concentration gradient)
antiport → 2 different molecules across in different directions (one down concentration gradient other up concentration gradient)
passive transporters
transport one set of molecules at a time down concentration gradient
highly selective
not continuous pore through membrane
rate of transport is dependent on number of binding site for substrate
GLUT1 transporter
substrate binds on one side of membrane
conformational change takes place
site opens on other side of membrane and substate is released
conformational change takes place
active transporters
against concentration gradient
many powered by ATP hydrolysis
generate ion gradients across membrane
Na+ K+ ATPase
generates gradient
3 Na+ out and 2 K+ in
net negative charge in cell
both ions move up concentration gradient
functions:
control cell volume
drive active transport of other species
render nerve cells electrically excitable
example of secondary active transporter
Na+-glucose symporter
one ion down concentration gradient drives other ion up concentration gradient
differences between channels and transporters
rate of flux → high in channels limited only by diffusion
saturability → binding sites on transporters
channels are gated
ion channels present in pm of all cells
voltage gated K+ channel
tetramer
outer helices interact with bilayer
inner helices contributes to inner pore
K+ doesnt have to pass through pm
partially neg
K+ are spaced apart in the channel since pos charge repel each other
voltage gated Na+ channel
helix 6 forms pore
helices 1-4 help with gating mechanism
4 pore-forming helices arranged around pore
helix 4 has high net pos charge and is pulled into cell
depolarization moves helix 4 outside and Na+ enter
inactivation loop stopping ions from passing