biochem 13-15 protein dynamics
1/110
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
111 Terms
what is a catalyst
a compound that increases the rate of a chemical reaction
catalyst to what to activation free energy (ΔG)
lowers
what is activation energy
the difference in energy between the reactant in its ground state and in its transition state
what is transition state
"setting up the reaction" by breaking existing bonds and forming new ones
catalysts combine transiently with what, which promotes what
They combine transiently with the reactants promoting a reactive transition state condition
what does catalyst do to ΔG
does not alter
average energy of reaction does not change
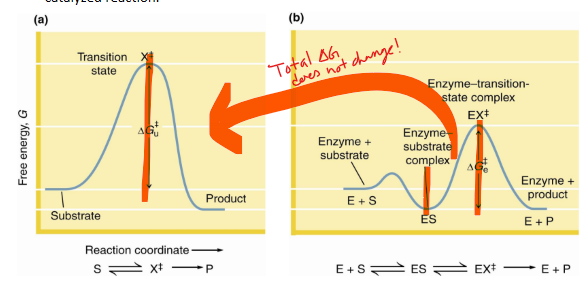
enzymatic catalysis offers what 3 things
Acceleration under mild conditions
High specificity
Possibility for regulation
no enzyme vs enzyme catalyzed reaction graph
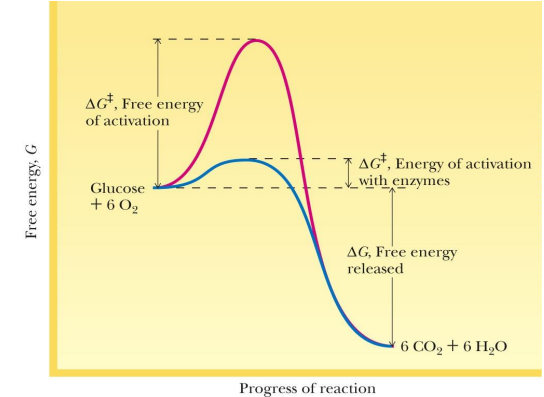
enzymes catalyze what types of reactions; causes them to do what
Thermodynamically favorable reactions
Causing them to proceed at extraordinarily rapid rates
Living systems use enzymes to accelerate and control the rates of vitally important biochemical reactions
enzymes and metabolic function
Enzymes are the agents of metabolic function
the reactants in an enzyme catalyzed reactions are referred to as
substrates
what is catalytic power
Catalytic power is defined as the ratio of the enzyme-catalyzed rate of a reaction to the uncatalyzed rate
Relationship between catalyzed reaction and the uncatalyzed, how much faster is reaction with enzyme compared to without enzyme
Enzymes can accelerate reactions as much as 10^26 over uncatalyzed rates
Urease is a goof example
Catalyzed rate: 3x10^4/sec
Uncatalyzed rate: 3x10^-10/sec
Ratio (catalytic power) is 1x10^14
specificity
is the term used to define the selectivity of enzymes for their substrates
regulation of enzyme activity ensures
that the rate of metabolic reactions is appropriate to cellular requirements
Not break down all we are making before we get to use it
what are coenzymes and cofactors
are nonprotein components essential of enzyme activity
Vits, minerals, ATP, NADH
Not in the amino acid sequence but needed for the protein function (enzyme)
enzyme specificity
Enzymes selectively recognize proper substrates over other molecules
Produce products in very high yields- often much greater than 95%
specificity is controlled by
structure
The unique fit of substrate with enzyme controls (2)
The unique fit of substrate with enzyme controls the selectively for substrate and the product that is formed
what are the major classes of enzymes (6)
Oxidoreductases - oxidation reduction reactions
Transferases - transfers a functional group between 2 compounds
Hydroxylases- hydrolyzes (adds water to split molecule)
Lyases- break it by removing something
Isomerases - making isomer
Ligases - combining things using ATP
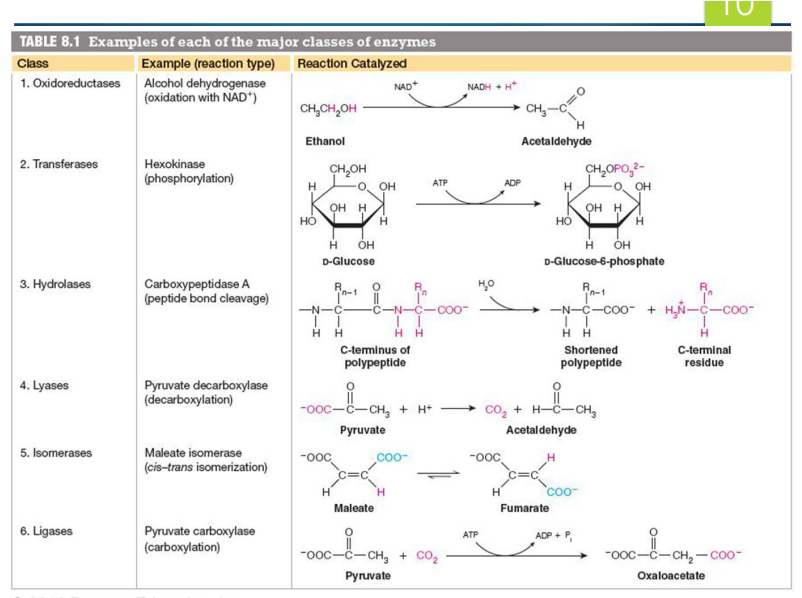
what is a cofactor
Inorganic ions (Fe+2, Mg+2, Zn+2)
what is a coenzyme
A complex organic or metalorganic molecule that act as a carrier of functional groups (biotin in carboxylation reaction)
Usually derived from vitamins
coenzymes and cofactors are __________ changed
reversibly
They need to be able to go back to original state to be able to redo reaction
So like F2+ became F3+ after reaction, so then needed to go back to F2+ to be reused
what is a prosthetic group
A coenzyme or metal ion tightly or covalently bound to the enzyme (heme)
what is a haloenzyme
enzyme together with its bound coenzyme/metal ion (glutathione reductase with Se-)
enzyme is whole so it has its cofactor or coenzyme
what is apoenzyme/apoprotein
the protein part of an enzyme (lipoproteins)
without its cofactor or coenzyme
covalent modification
Covalent attachment via phosphorylation, glycosylation, etc.
alters enzyme activity or provides regulation
now onto kinetics …
what is kinetics (and what is its 2 goals)
is the study of the rate at which compounds react
Enzymes kinetics seeks to determine the max reaction velocity that enzymes can attain
and the binding affinities for substrates and inhibitors
what effects the enzymatic reaction rate (3)
Enzyme
Substrate
Temperature - slow down at cold and speed up at warm, yet too high denatures
why do we measure enzymatic kinetics (5)
Quantitative description of enzymes
Determine the order of binding of substrates (like in metabolic pathways )
Understand catalytic mechanism
Find effective inhibitors (or activators)
Understand regulation of activity
what is E, S, P, K
enzyme
substrate
product
rate of reaction
expression (equation of enzymes)
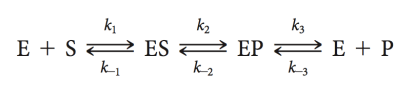
equation if done at initial rate
(before high [P])
Look at the initial rate because at the beginning you have more energy, if we do it later than most energy already spent (like class at 9:30am vs class at 2:30pm)
![<p><span><strong>(</strong></span>before high [P])</p><ul><li><p><span><em>Look at the initial rate because at the beginning you have more energy, if we do it later than most energy already spent (like class at 9:30am vs class at 2:30pm)</em></span></p></li></ul><p></p>](https://knowt-user-attachments.s3.amazonaws.com/755db5cb-9705-4596-8a6f-6243e60012ca.png)
how do we do analysis of initial rate
Mix the same [E] but varying [S]
Measure initial rates of [S] disappearance
Plot change in [S] vs time
Calculate initial rate based on slopes
![<ul><li><p><span><strong>Mix the same [E] but varying [S]</strong></span></p></li><li><p><span>Measure <strong>initial rates </strong>of [S] disappearance</span></p></li><li><p><span>Plot <strong>change in [S] vs time</strong></span></p></li><li><p><span>Calculate <strong>initial rate </strong>based on slopes</span></p></li></ul><p></p>](https://knowt-user-attachments.s3.amazonaws.com/ec2d0ffc-2920-440c-b23c-d2e8170c4126.png)
For E +S <--> ES→ E+P concentrations vary of time for (which parts of the equation)
Substrate [S]
Free energy [E]
Enzyme-substrate [ES]
Product [P]
![<ul><li><p><span>Substrate [S]</span></p></li><li><p><span>Free energy [E]</span></p></li><li><p><span>Enzyme-substrate [ES]</span></p></li><li><p><span>Product [P]</span></p></li></ul><p></p>](https://knowt-user-attachments.s3.amazonaws.com/ab721cb5-5ce0-4ed2-b227-9dd25ad7fed7.png)
steady state in enzyme kinetics
After a very brief initial period, [ES] reaches a steady rate
ES is consumed approximately as fast as it is formed
Allows us to calculate the velocity by assuming steady-state conditions
WE STUDY THE INITIAL RATE BECAUSE OTHERWISE THE ES IS STEADY, AND WE CANT REALLY OBSERVE ANYMORE WHAT IS HAPPENING
![<ul><li><p>After a very brief initial period, <strong>[ES] reaches a steady rate</strong></p><ul><li><p><strong>ES is consumed approximately as fast as it is formed</strong></p></li><li><p>Allows us to <strong>calculate the velocity</strong> by assuming <strong>steady-state conditions</strong></p></li></ul></li></ul><ul><li><p><span><em>WE STUDY THE INITIAL RATE BECAUSE OTHERWISE THE ES IS STEADY, AND WE CANT REALLY OBSERVE ANYMORE WHAT IS HAPPENING</em></span></p></li></ul><p></p>](https://knowt-user-attachments.s3.amazonaws.com/9737a89e-e4f5-42e1-91e4-ddabd1b9ab71.png)
michealis menton equation has what parts
Km and Vmax
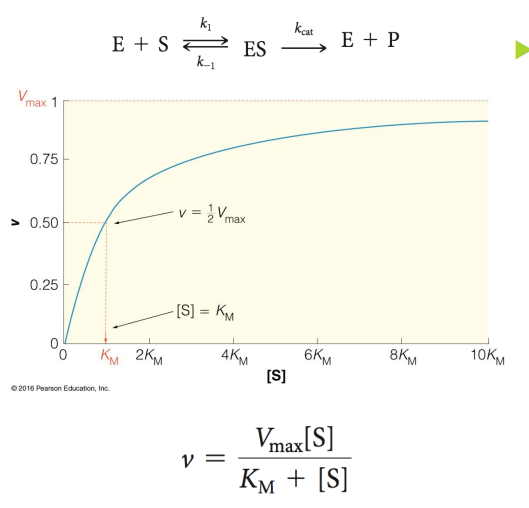
what is Km
Michaelis constant
Specific for each E acting on a given S
Equal to substrate concentration is 1/2 the Vmax
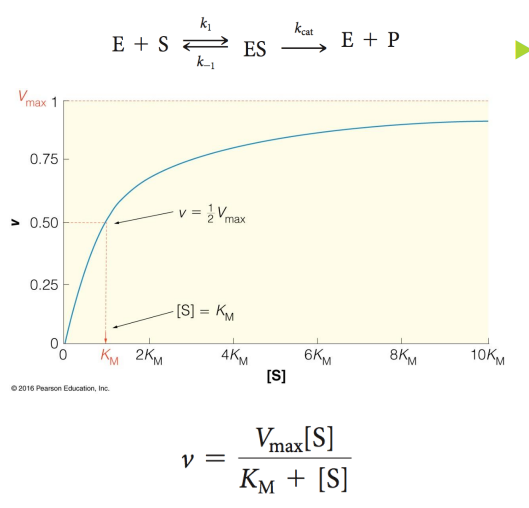
Vmax
maximum velocity
Point at which E is saturated
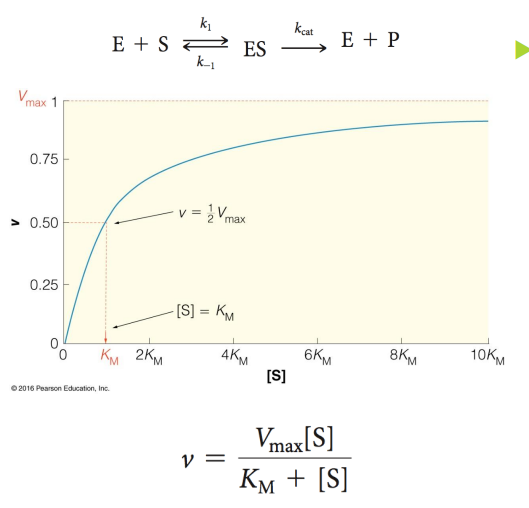
Km is the [S] when V is what
1/2Vmax
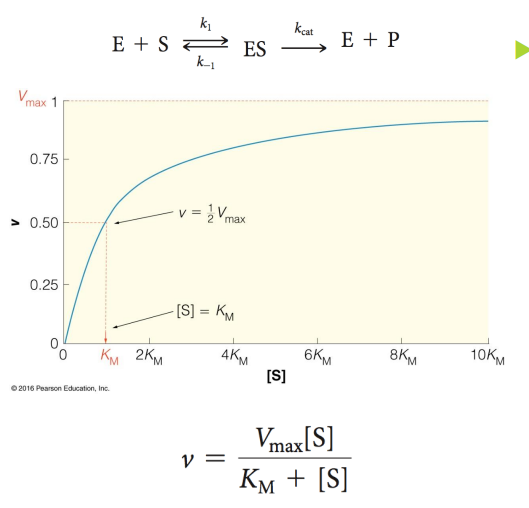
when [S] is above Km, enzyme activity is
high
Km is also knows as
kinetic activator constant
small Km means ______ affinity
Small Km means high affinity of the enzyme for its substrate
Very sensitive to low amounts of that substrate
Needs less substrate to reach Vmax than if Km is high
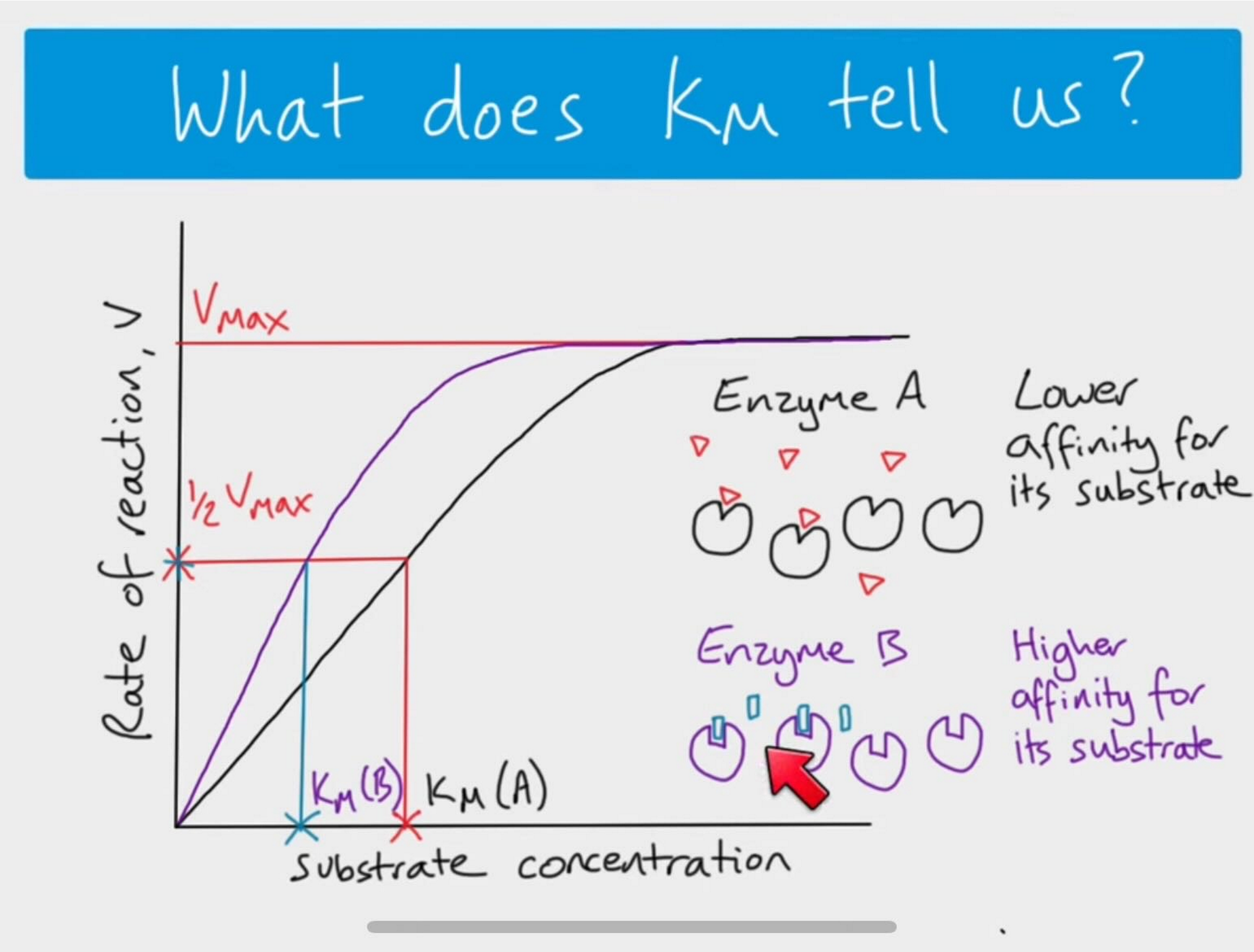
high Km means ____ affinity
High Km means low affinity of the enzyme for its substrate
Needs a higher amount of substrate present to reach Vmax
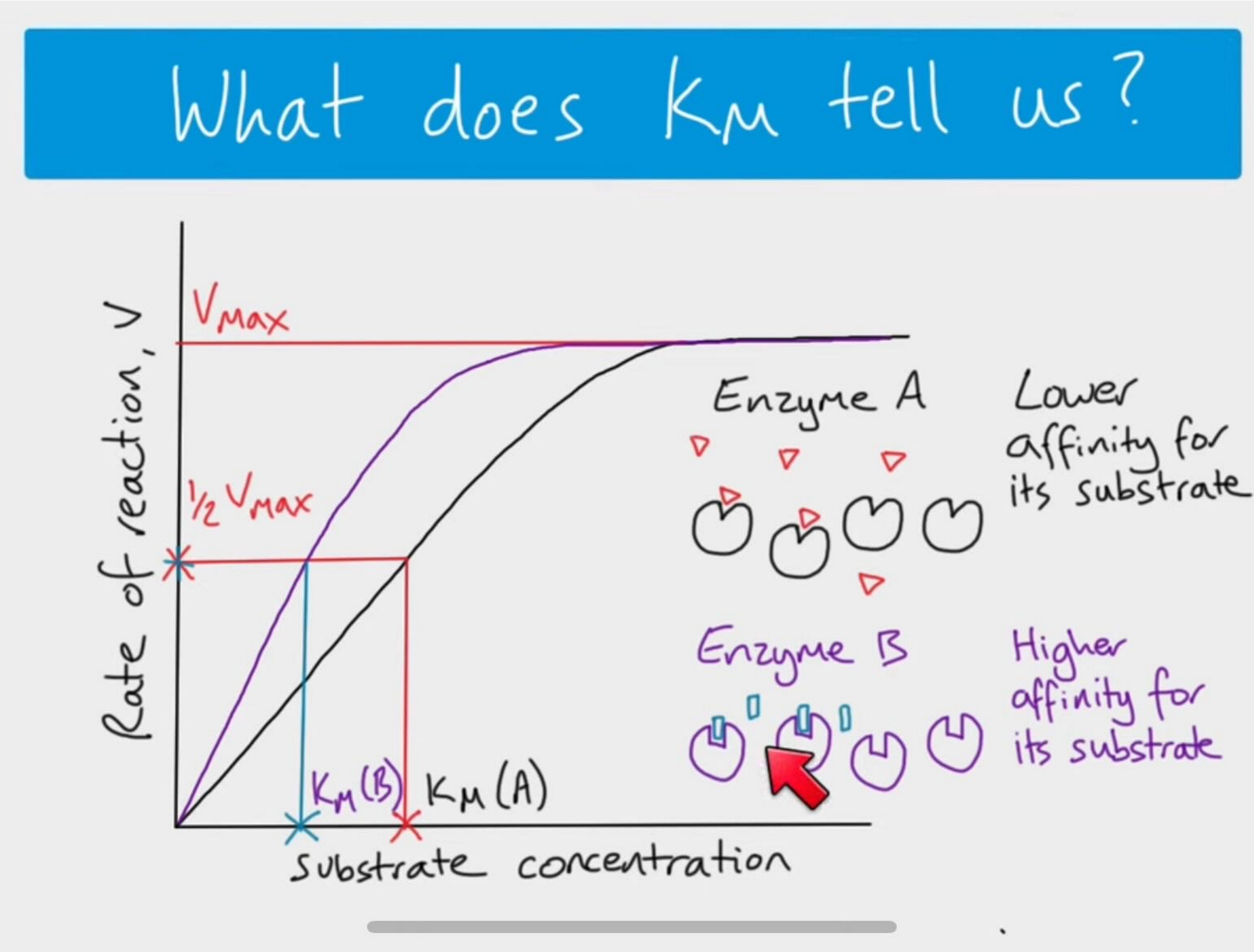
can Vmax be reached
Vmax is the theoretical maximal rate of the reaction- but it is NEVER achieved in reality
To reach Vmax would require that ALL enzyme molecules have substrate bound
Vmax is asymptotically approached as substrate is increased
how does pH affect enzyme activity
Enzyme- substrate recognition and catalysis are greatly dependent on pH
Enzymes are usually active only over a limited range of pH
The effects of pH may be due to effects on Km or Vmax or both
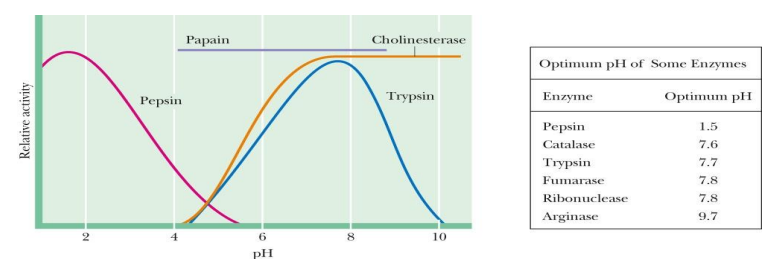
how does pH effect structure and active site
Enzymes have a variety of ionizable side chains that determine their secondary and tertiary structure and also affect events in the active site—> so pH can mess with the side chains and therefore mess with structure and the active site
how are substrates affected by pH
they also may have ionizable groups
and remember pH messes with this (tritation of amino acids)
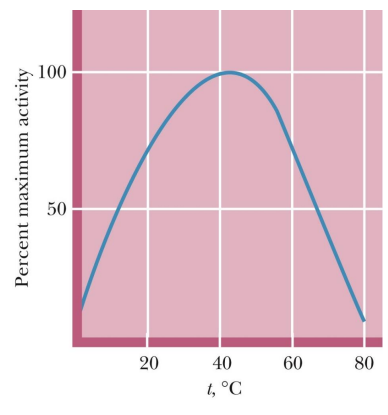
how does temp affect enzyme
Rates of enzyme-catalyzed reactions generally increase with increasing temperature
However, at temperatures above 50° to 60°C, enzymes typically show a decline in activity --> because of denaturing
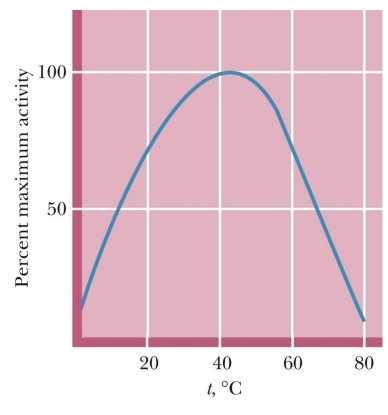
what 2 effects does temp have
Enzyme rate typically doubles in rate for every 10ºC rise in temperature as long as the enzyme is stable and active (as long as hasn’t denatured)
At higher temperatures, the protein becomes unstable and denaturation occurs
what are the 2 types of inhibitor classes
inhibitors (noncovalently bound)
Can be removed or come and then leave
Irreversible inhibitors (covalently bound)
Cannot be removed
what are the 3 types of reversible inhibition
competetive, uncompetivite, mixed
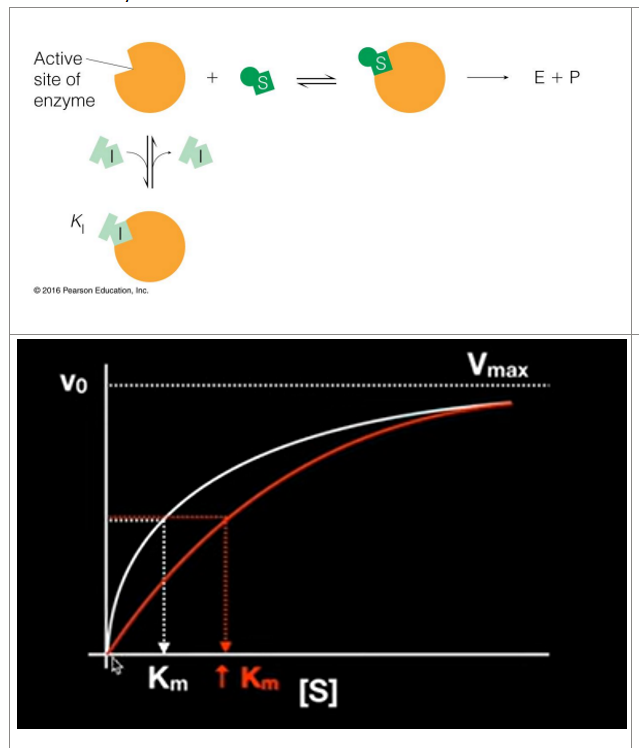
competitive inhibitors
Competes with substrate for the active site
Binds to active site (competes for the active site)
Does not effect catalysis
(does not change the Vmax, but increases km)
What happens is that in order to get to same rate as with no inhibitor, you will need to add a lot more substrate
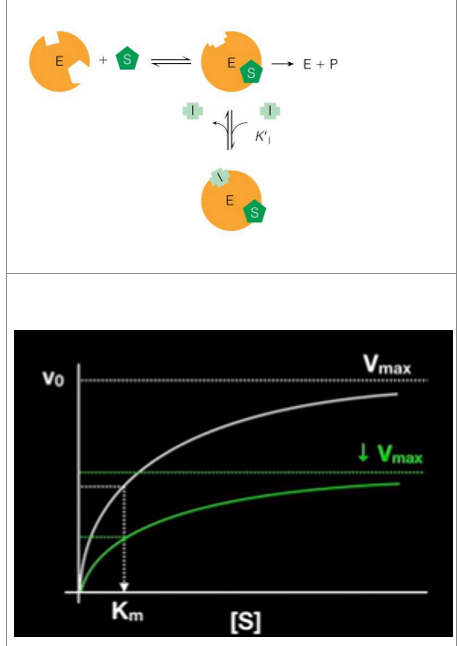
uncompetitive inhibition
Only binds to ES complex
Does not affect substrate binding(binds somewhere else on the enzyme)
Only binds once the substrate binds to the enzyme
Inhibits catalytic function
Decreases Vmax (overall catalytic activity) but Km of enzyme stays the same
noncompetitive inhibition reduces the overall catalytic activity of the enzyme by affecting its function, but it does not prevent substrate binding. This leads to a lower Vmax while leaving the Km unaffected.
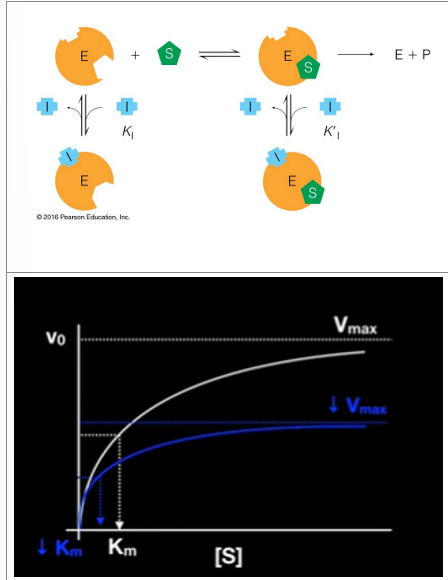
mixed inhibition
Binds to enzyme with or without substrate
Binds to regulatory site (not the active site)
Inhibits both regulatory binding and catalysis
Decreases Vmax and Km
If binds before substrate binds to enzyme, then changes active site
If binds to the ES at secondary site, then affects catalysis because not binding active site
irreversible inhibition
Substances combine covalently with enzymes→ toxic
irreversible inhibition
React with functional group in active site
Many are transition state analogs
what are the enzyme substrate models
lock and key
induced fit
lock and key hypothesis
the first explanation for specificity (explains substrate specificity)
does not explain catalysis or transition state
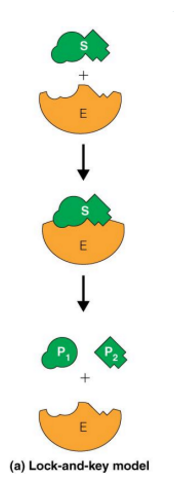
induced fit hypothesis
provides a more accurate description of specificity
Induced fit favors formation of the transition state
allows catalysis to occur (speed up reaction and lower activation energy)
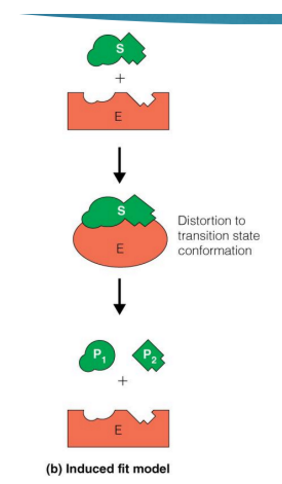
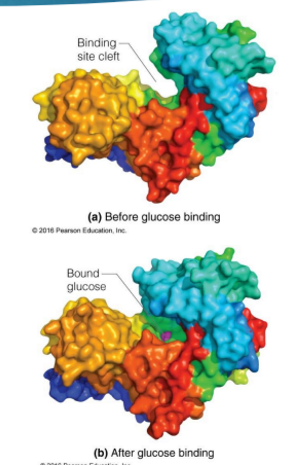
hexokinase and specificity
Specificity and reactivity are often linked
In the hexokinase reaction, binding of glucose in the active site induces a conformational change in the enzyme that causes the 2 domains of hexokinase to close around the substrate, creating the catalytic site
are all enzymes proteins
no
robozymes
RNA molecules that display enzyme activity in the absence of protein
Examples: RNase P and peptidyl transferase
abzymes
antibodies raised to bind the transition state of a reaction of interest
The large rate accelerations of enzymes correspond to (what does enzyme do that increases rate fo reaction)
large decreases in the free energy of activation for the reaction
all reactions pass through a _____________________________ on the reaction pathways
transition state
what is the transition state
state of most free energy required
not the product yet, but changes occurring to drive to the product
active sites of enzymes bind more tightly to what than what; why?
The active sites of enzymes bind the transition state of the reaction more tightly than the substrate
It prefers the transition state over the substrate because this will help drive the reaction, if it binds more to the substrate then no reaction would occur.
the active sites of enzymes binding more tightly to the transition state than the substrate does what
the enzymes stabilize the transition state and lower the activation energy of the reaction
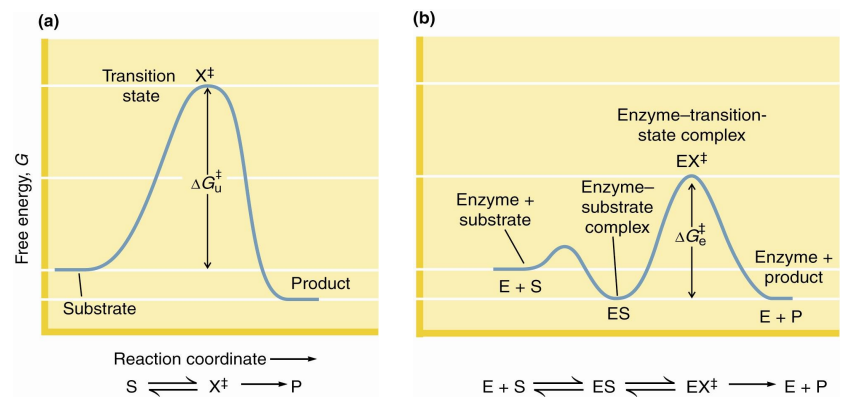
The catalytic role of an enzyme is
to reduce the energy barrier between substrate S and transition state X‡
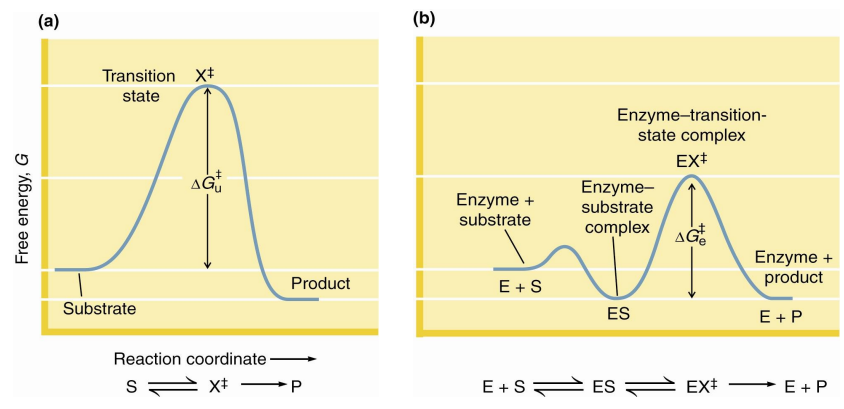
what does X‡ represent
transition state
what does rate acceleration of an enzyme mean
rate acceleration by an enzyme means that the energy barrier between ES and EX‡ must be smaller than the barrier between S and X‡
This means that the enzyme must stabilize the EX‡ transition state more than it stabilizes ES
If we were stabilizing ES, then nothing would happen, by moving it to transition state, this helps drive it forward by starting to change it which them moves it to continue changing until reaching the EP
how does formation of ES help with catalysis (in terms of transition state)
For a given energy of X‡, raising the energy of ES will increase the catalyzed rate
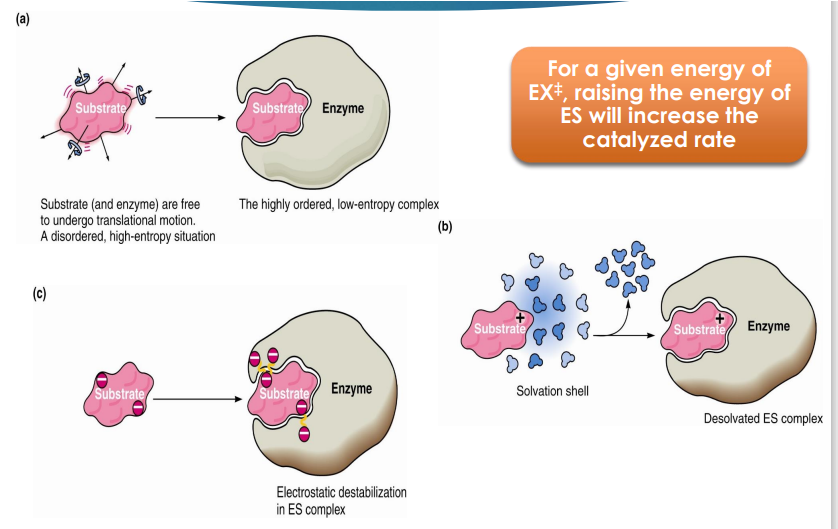
raising the energy of ES will _____the catalyzed rate
increase
HOW TIGHTLY DO TRANSITION-STATE ANALOGS BIND TO THE ACTIVE SITE?
Very tight binding to the active site
The binding constant of the enzyme for the transition state may be 10^ -20 to 10^-26 M!
Transition state analogs (TSAs)
stable molecules that are chemically and structurally similar to the transition state
That mimic the transition state
We use these because the enzyme prefers the transition state over the substrate—> enzymes will allow them to bind over substrate because they are already stable
Would be very good competitive inhibitors
Proline racemase was the first case observed
Transition-state analogs work well as what
enzyme inhibitors
-competitive
-irreversible
what 5 things that will cause enzymatic catalysis (one or more of these is needed) (5)
General acid/base catalysis (GABC)
Covalent catalysis
Electrostatic stabilization (Positive charge on enzyme helps stabilize the neg charge on substrate)
Proximity effects
Preferential stabilization of the transition state
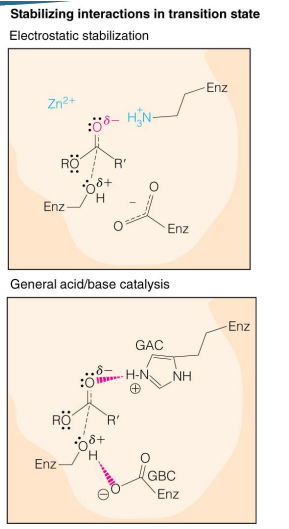
what else may play a role in catalysis
protein conformational changes
what is essential to enzyme catalysis
protein motions
protein movement (and importance for catalytic events)
Proteins are constantly moving –
Bonds vibrate, side chains bend and rotate, backbone loops wiggle and sway, and whole domains move as a unit
Enzymes depend on such motions to provoke and direct catalytic events
Active site conformation changes can - 4
Active site conformation changes can
Assist substrate binding
Bring catalytic groups into position
Assist in bond making and bond breaking
Facilitate conversion of substrate to product
HOW DO ACTIVE SITE RESIDUES INTERACT TO SUPPORT CATALYSIS
(main effect, secondary roles (4)
About half of the amino acids engage directly in catalytic effects in enzyme active sites
Other residues may function in secondary roles in the active site:
Raising or lowering catalytic residue pKa values
Orientation of catalytic residues
Charge stabilization
Proton transfers via hydrogen tunneling
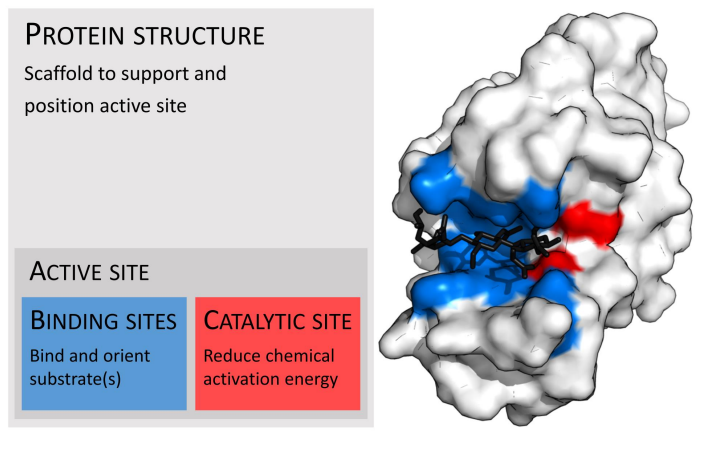
active site has what 2 parts
binding sites and catalytic sites
bind vs catalytic site
Blue is secondary roles
Red is actually catalytic activity place
what factors influence enzymatic activity (4)
The availability of substrates and cofactors
Genetic regulation of enzyme synthesis and decay
allosteric or covalent modification
Zymogens, isozymes, and modulator proteins
how does availability of substrates and cofactors effect enzyme activity
usually determines how fast the reaction goes
As product accumulates, the apparent rate of the enzymatic reaction will decrease
how does Genetic regulation of enzyme synthesis and decay effect enzymatic activity
determines the amount of enzyme present at any moment
what are the 2 ways enzyme activity can be regulated
allosterically or covalent modification
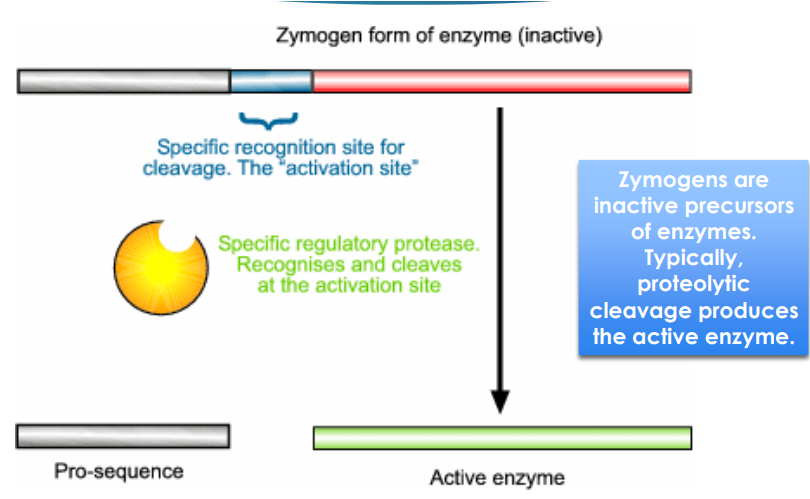
zymogen
inactive precursors of enzymes. Typically, proteolytic cleavage produces the active enzyme
So pretty much inactive enzyme that is activated by some sort of modification
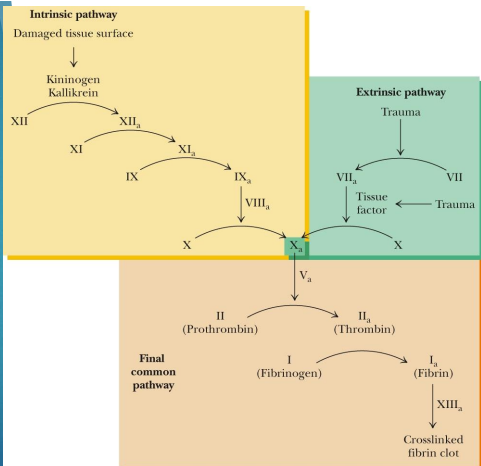
REGULATORY CASCADE OF BLOOD CLOTTING
Soluble fibrinogen is a zymogen that must be cleaved to form insoluble fibrin by thrombin
Some factors and proteins have Ca2+ binding sites which were formed via PTM of a Glu residue by Vitamin K dependent gamma -glutamy carboxylase
MAIN THING: We need PTMs and zymogens for our blood to clot - main point
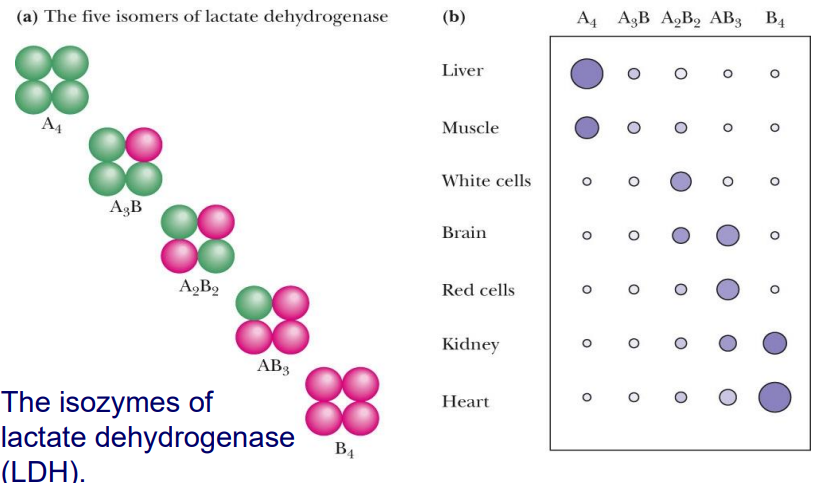
ISOENZYMES
ENZYMES WITH SLIGHTLY DIFFERENT SUBUNITS
Play same role but differ slightly depending on where they're made in the body
they often respond to different inhibitors and activators (was on slido)
Because of changes in subunits they may respond to different things
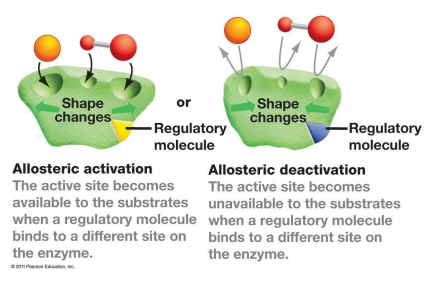
what is allosteric regulation
Action at "another site"
does not bind to active site to regulate
enzymes in metabolic pathways are modulated by
allosteric effectors
the effectors usually made where, and what roles do they play (2)
These effectors are usually produced elsewhere in the pathway
Effectors may be feed-forward activators or feedback inhibitors
the effectors are usually products in some part of the pathway or result of lack of product
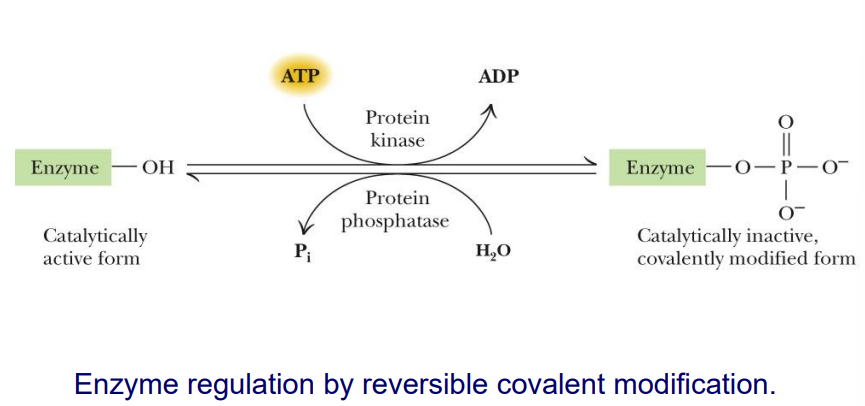
what is the most common covalent modification used to regulate activty of enzyme
reversible phosphorylation
This is the MOST prominent form of covalent modification in cellular regulation
phosphorylation (in terms of regualtion)
Phosphorylation is accomplished by protein kinases
Each protein kinase targets specific proteins for phosphorylation
Phosphoprotein phosphatases catalyze the reverse reaction – removing phosphoryl groups from proteins
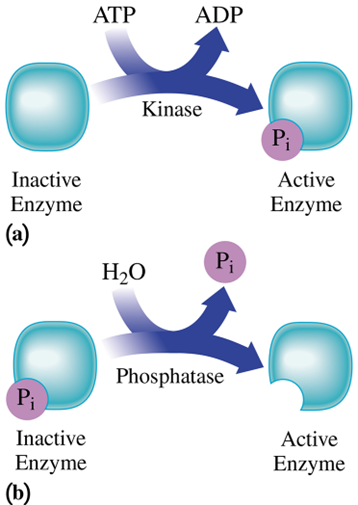
protein kinases
targets specific proteins for phosphorylation
Phosphoprotein phosphatases
catalyze the reverse reaction – removing phosphoryl groups from proteins
Protein kinases phosphorylate what residues in target protein
Ser, Thr, and Tyr
kinases recognize what in their target
amino acid sequence
Despite this specificity, all kinases share a common catalytic mechanism based on a conserved core kinase domain of about 260 residues