Topic 7: Structure and function of the cell and membrane
1/132
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
133 Terms
How are all organisms are classified into three major domains?
based on molecular and evolutionary evidence
3 major domains
1. Bacteria (Eubacteria)
2. Archaea (Archaebacteria)
3. Eukaryota (Eukarya)
3 factors of classification for domains of life
1. Ribosomal RNA sequence analysis
2. Molecular phylogeny
3. Biochemical characteristics
Bacteria and Archaea
both prokaryotic
What domain is Archaea evolutionarily closer to?
eukaryotes
domains in phylogenetic trees
- Archaea and Eukarya cluster together
- Bacteria form a separate branch
evolutionary insight major biochemical implications for:
1. Gene expression machinery
2. Membrane structure
3. Evolution of cellular complexity
Prokaryotes - General Characteristics
- Bacteria and Archaea
- unicellular organisms
- Lack a membrane-bound nucleu
Cellular Organization of prokaryotes
- Genetic material located in nucleoid region
- Cell surrounded by cell envelope
- Cytoplasm contains Ribosomes, Soluble proteins & small metabolites
Surface Structures of prokaryotes
- Flagella (tail-like structures) → Allow movement in aqueous environments
- Pili (fimbriae) → Enable adhesion to solid surfaces
Gram Staining
- Developed by Hans Christian Gram
- Classifies bacteria based on differences in cell envelope structure into: 1. Gram-positive 2. Gram-negative
Gram-positive bacteria
- thick PG layer
- retain crystal violet stain
-lack outer membrane
- staphylococcus aureus
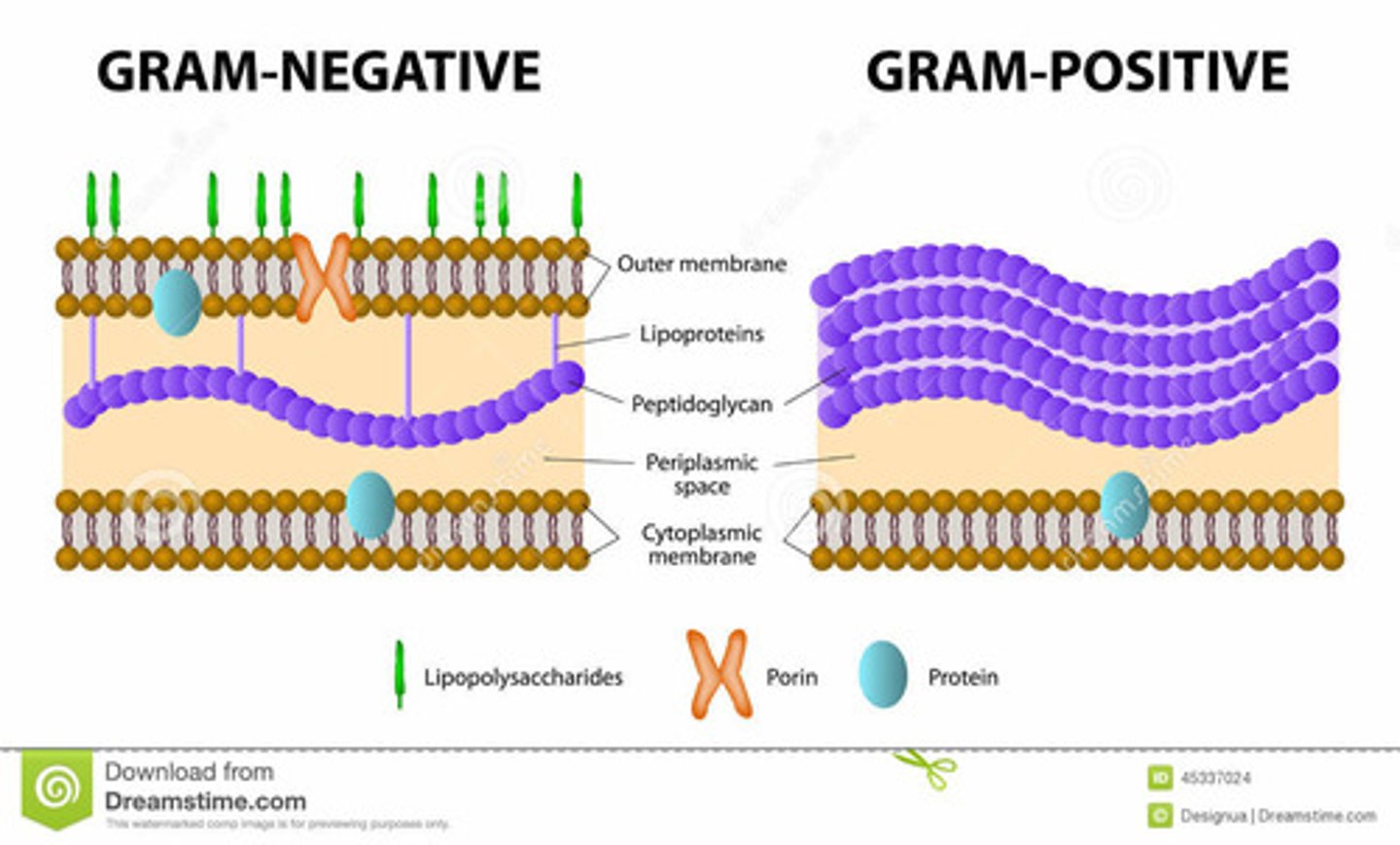
gram-negative bacteria
- thing PG layer
- have an outer membrane
- retain safrinin stain
- Porins in outer membrane → Allow diffusion of small molecules
- Example: Escherichia coli (E. coli)
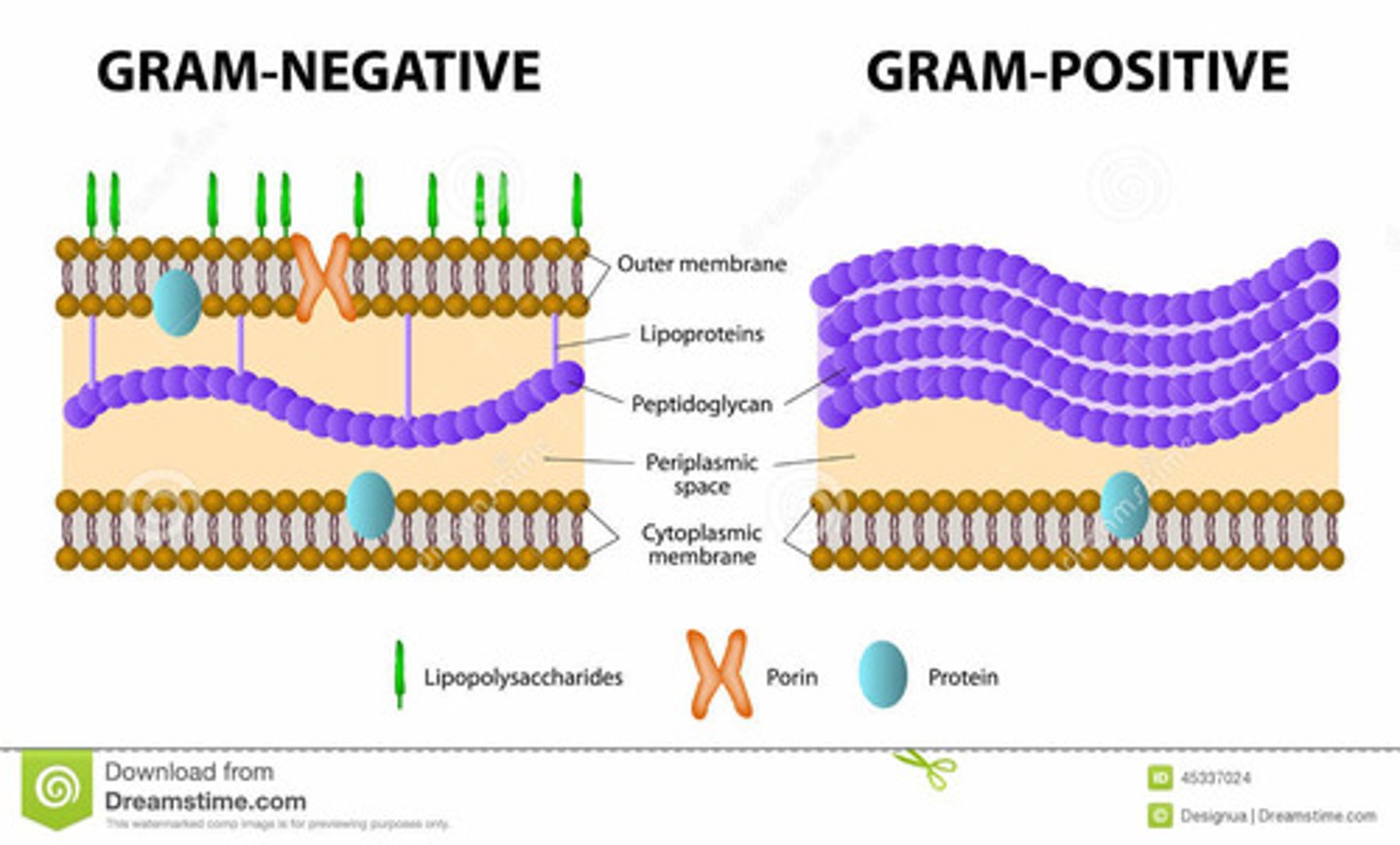
Why do Gram-Negative Not Retain Stain?
- Peptidoglycan layer is thin
- Covered by outer MB
- Dye cannot bind effectively
Archaebacteria (Archea)
- Plasma MB
- Pseudopeptidoglycan layer: chemically different from bacterial peptidoglycan, but structurally similar
- Unique MB lipids (ether-linked)
Cyanobacteria
- Photosynthetic Gram-negative bacteria
- Additional internal membrane system
- Photosynthetic machinery embedded in internal membranes
General Characteristics of Eukaryotic cells
- Include animals, plants, fungi, protists
- Larger than prokaryotes: 10-100 μm (vs 1-5 μm in prokaryotes)
- Contain MB-bound organelles
- Surrounded by a plasma MB
Cellular Components of Eukaryotic cells
- Cytoplasm = cytosol + organelles + ribosomes
- Genetic material (DNA) stored in nucleus
nucleus structure
- Surrounded by double membrane (nuclear envelope)
- Contains nuclear pores
- 2 regions: Nucleolus → rRNA synthesis & Nucleoplasm → rest of nuclear interior
nucleus function
- DNA replication
- RNA synthesis (transcription)
- Regulated transport via nuclear pore complexes: small molecules diffuse freely & large molecules require transport proteins
Endoplasmic Reticulum (ER)
- Network of MB sacs
- lumen continuous with nuclear envelope
Rough ER (RER)
- Ribosome-bound
- Synthesizes: Secreted proteins, MB proteins
- Protein processing & glycosylation
Smooth ER (SER)
- Lipid & cholesterol synthesis
- Detoxification (liver cells)
Golgi Apparatus structure
- Flattened membrane sacs
- Regions: Cis, Medial & Trans
Golgi Apparatus Function (3)
1. Protein processing
2. Sorting
3. Vesicular trafficking
mitochondria structure
- Double MB
- Inner MB folds → cristae has electron transport chain
- Matrix contains citric acid cycle enzymes
Major function of mitochondria
ATP production
chloroplast structure
- Double MB
- Internal thylakoid MBs
- Thylakoids stack → grana
- Stroma contains CO₂-fixing enzymes
Where does CO₂ fixation occur in the chloroplasts?
stroma (dark reactions)
Where do the light reactions occur in the chloroplasts?
thylakoid membrane
Lysosomes structure (Animal cells)
- Acidic pH (~5)
- Contain hydrolytic enzymes
lysosome functions
- Degrade macromolecules & damaged cell components
What protects the cytosol of lysosmes?
Acidic environment
Vacuoles Structure (Plant cells)
- Large central organelle
- occupy up to 80% of cell V
Vacuoles Function (Plant cells)
Maintain turgor pressure (5-20 atm)
Store pigments (anthocyanins)
Peroxisome functions (4)
- Oxidative rxns
- Detoxification
- Generate H₂O₂
- Catalase converts 2H₂O₂ → 2H₂O + O₂
Cytoskeleton Functions (4)
1. Structural support
2. Cell shape
3. Intracellular transport
4. Cell movement
3 cytoskeleton components
1. microtubules
2. microfilaments (Actin)
3. intermediate filaments
Microtubules
- Tubulin polymers
- 24 nm - 25 nm diameter
- Originate from centrosome (MTOC)
- Long-range transport
Microfilaments (Actin)
- Cell periphery
- Shape & local transport
Intermediate Filaments
- Mechanical strength
Differential Centrifugation Purpose
In vitro studies often require isolation of specific organelles
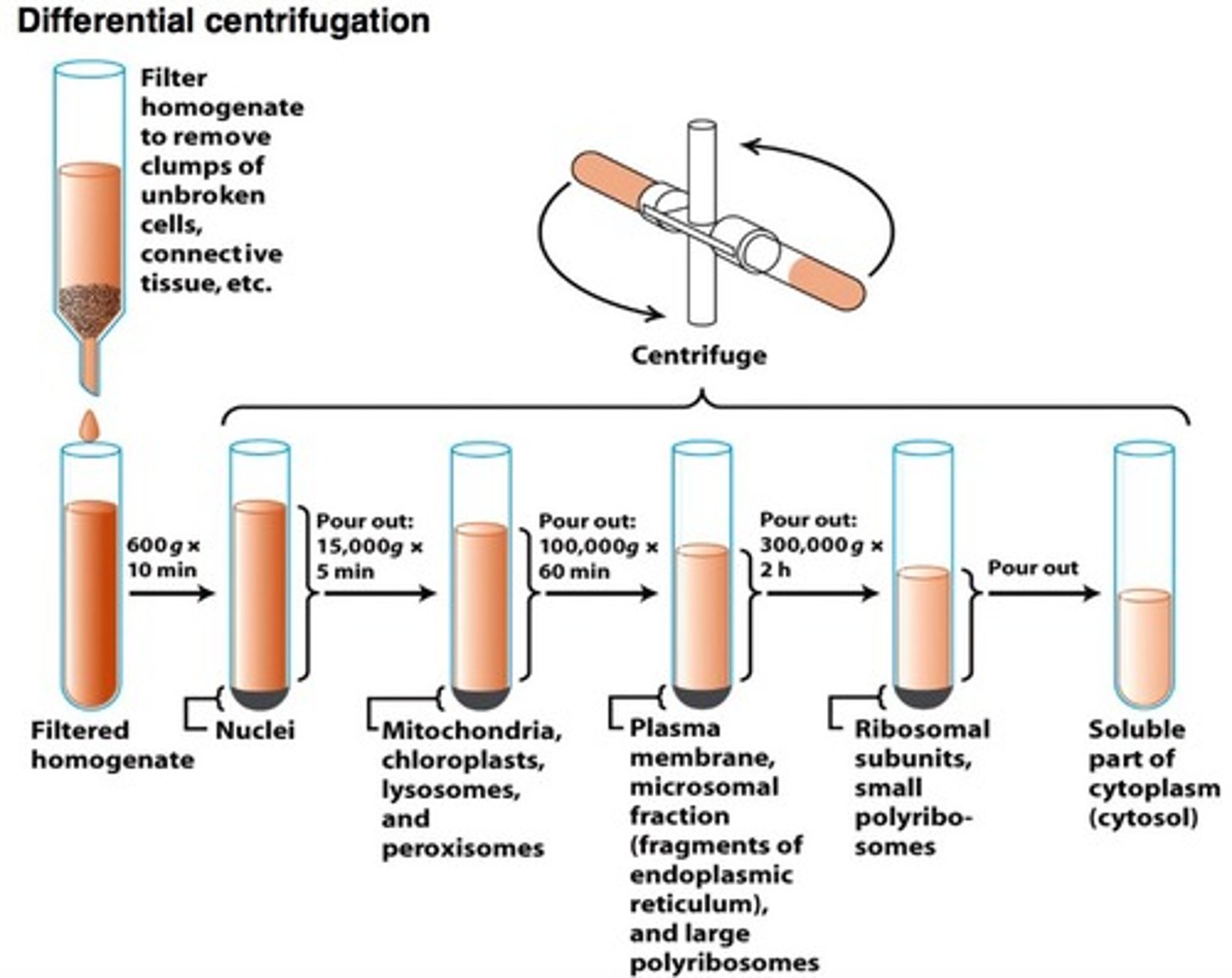
2 steps of Differential Centrifugation
1. Homogenization
2. Differential Centrifugation
Step 1: Homogenization
- Cells/tissues suspended in isotonic solution. Example: 0.25 M sucrose
- Maintains osmotic balance
- Cells disrupted using high-speed blender OR sonication (ultrahigh-frequency sound
- Resulting mixture = homogenate
Step 2: Differential Centrifugation
- sequential centrifugation at increasing speeds
- Supernatant from each step is centrifuged again at higher speed
- Separation based on size and mass
speeds in step 2: differential centrifugation
- 1000g (10 min) → Pellets nuclei, intact cells, large debris
- Higher speeds → mitochondria, lysosomes, peroxisomes
- Even higher speeds → microsomes, ribosomes
Density Gradient (Isopycnic) Centrifugation Purpose
Further purification of organelles based on density
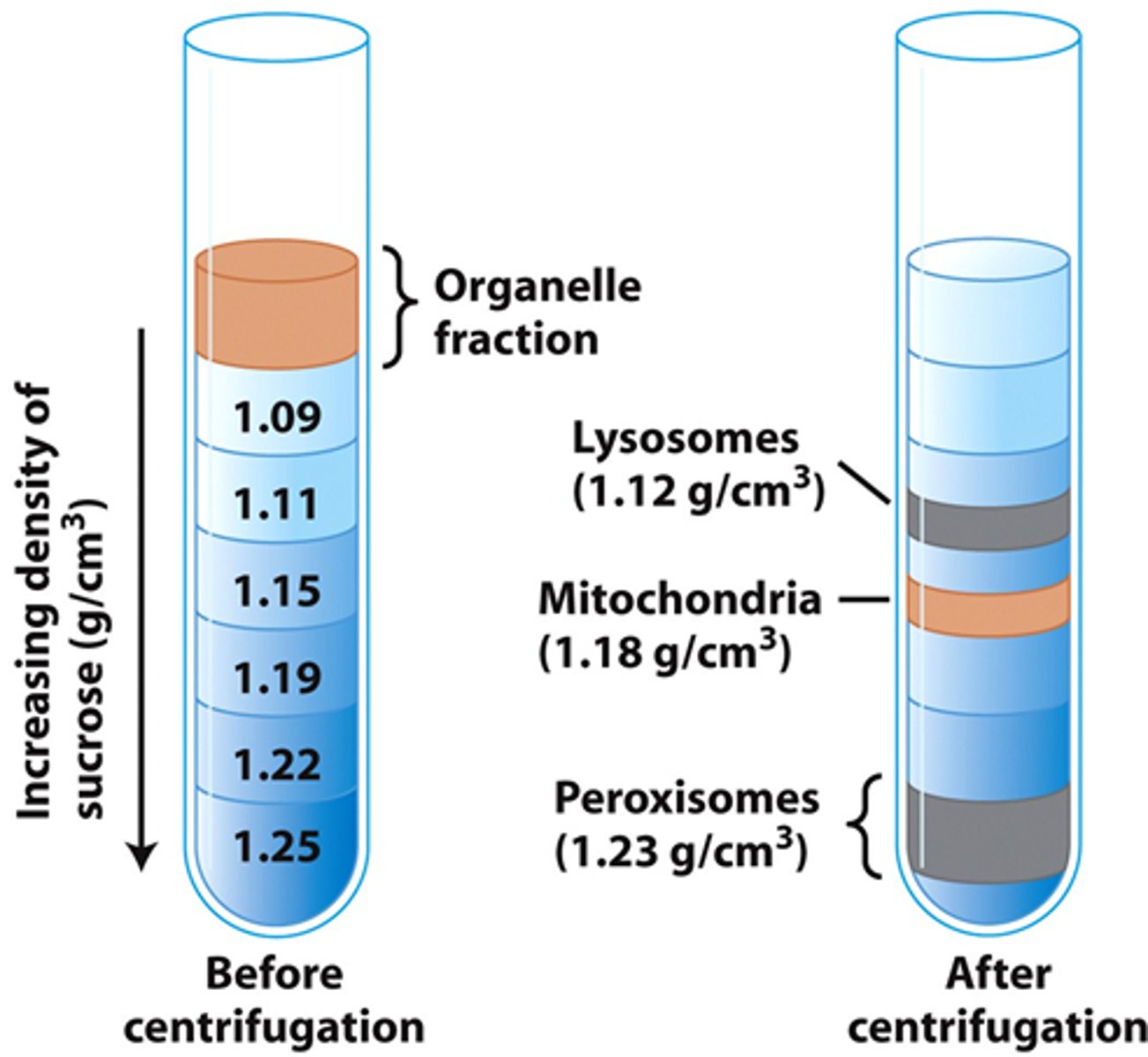
Density Gradient Centrifugation Method
- Tube filled w layers of inc [sucrose]→ Density inc from top to bottom
- Organelle mixture layered on top
- Centrifuged at high speed
Density Gradient Centrifugation Principle
- Each organelle migrates to a position where its density = surrounding sucrose density
- Organelles form distinct bands
- Separation based on Density of organelles
Other names for Density Gradient Centrifugation Method
- Equilibrium density-gradient centrifugation
- Isopycnic centrifugation
Membrane Composition
1. Lipid bilayers (amphipathic lipids)- 2 lipid layers (leaflets)
2. MB proteins
3. Carbohydrates (attached to lipids or proteins) located on the outer surface of the plasma MB
Amphipathic molecules composition
- Hydrophilic (polar) head
- Hydrophobic (nonpolar) tails
Amphipathic Lipids/Molecules in Aq environments
- Hydrophobic regions cluster together
- Hydrophilic regions interact with water
What are Glycerophospholipids derived from?
phosphatidic acid
Glycerophospholipids structure
- Glycerol backbone
- 2 fatty acids attached to C1 and C2 via ester bonds
- Phosphate group attached to C3 & linked to a polar or ionic head group
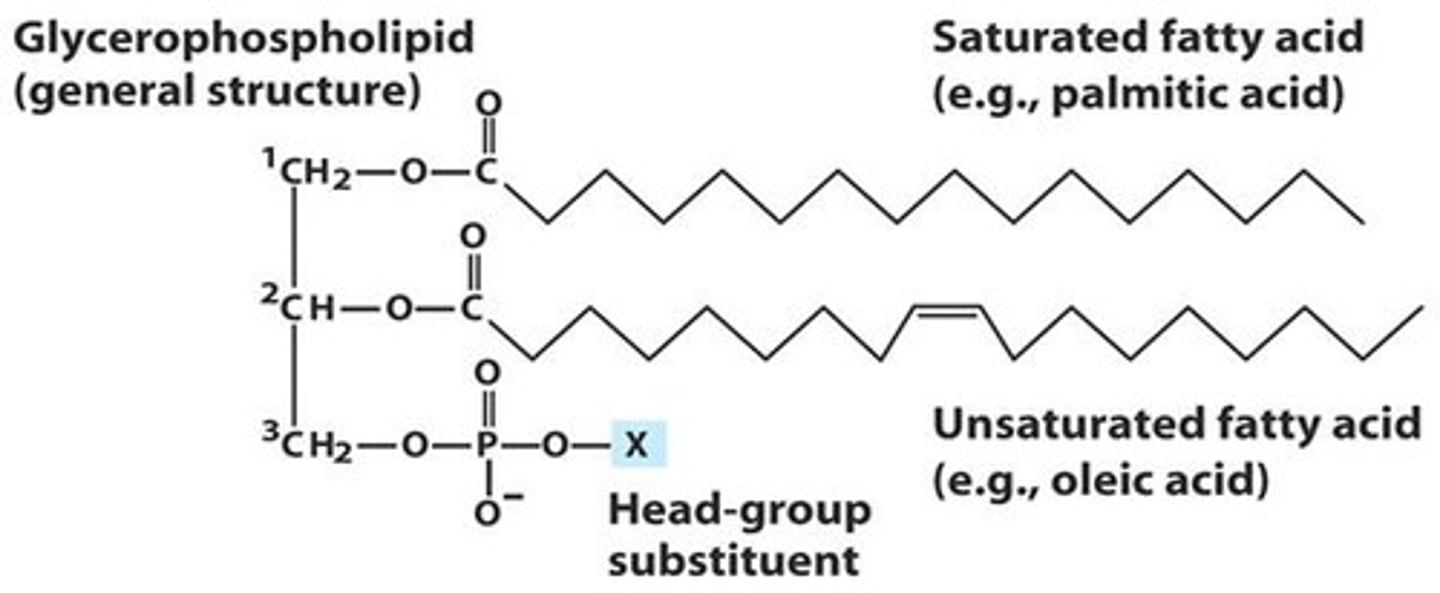
Fatty Acids structure
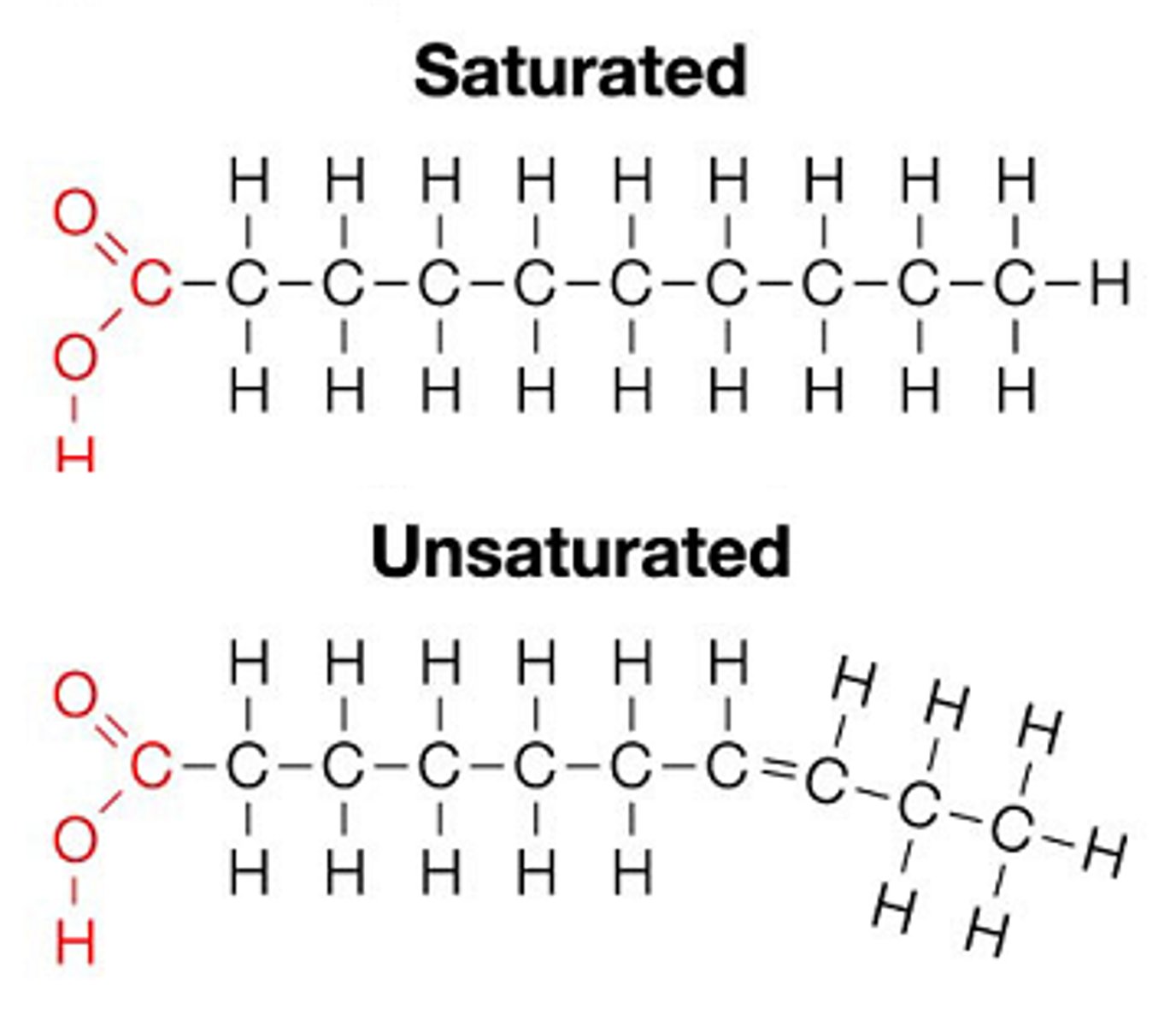
- Usually contain even # of Cs
- Saturated (no double bonds) OR unsaturated (cis double bonds common)
- C1 & C2 fatty acids can be the same or different types
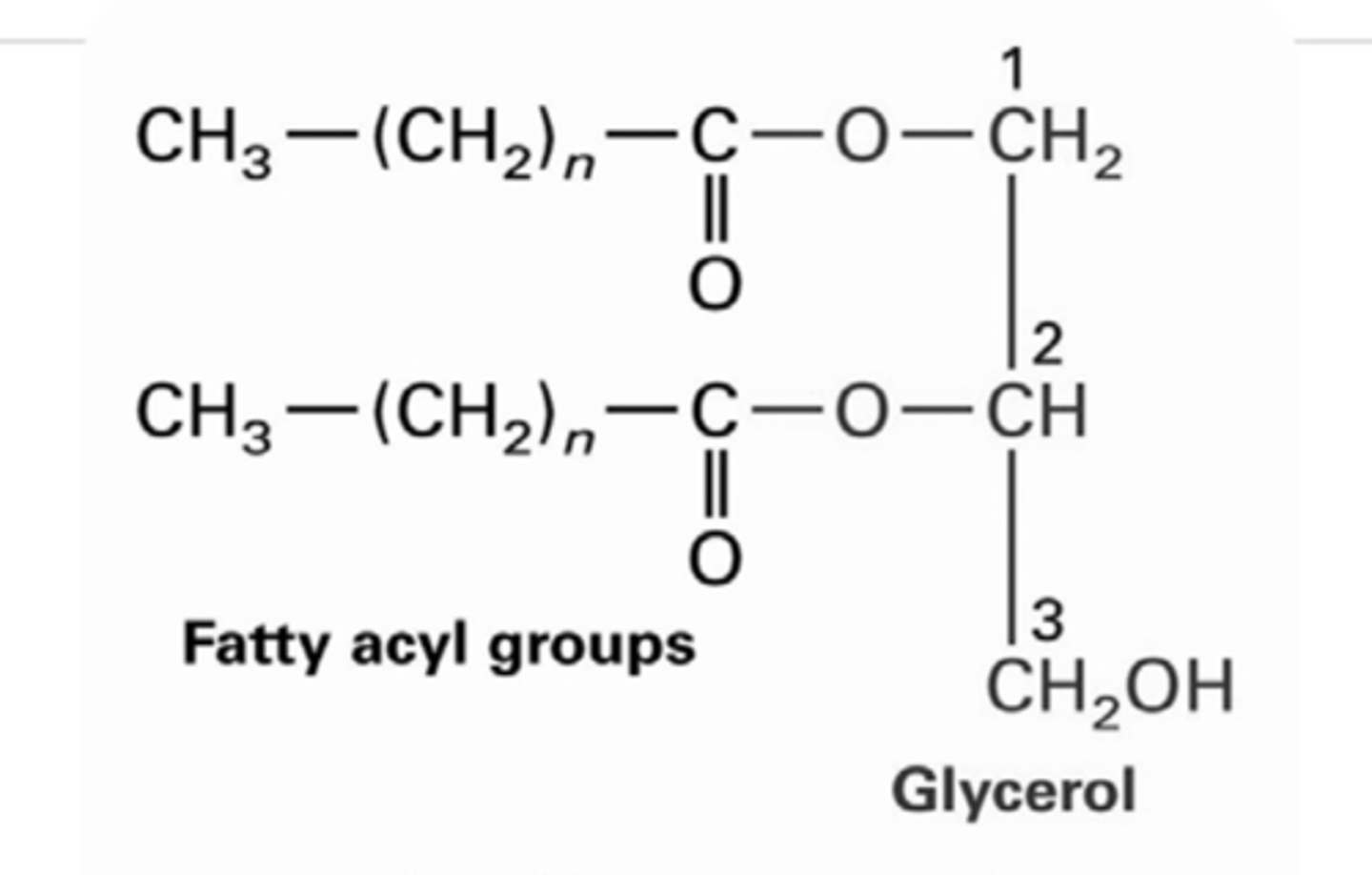
Diacylglycerol (DAG)
Glycerol + 2 fatty acids
Phosphatidic acid
DAG + phosphate group at C3
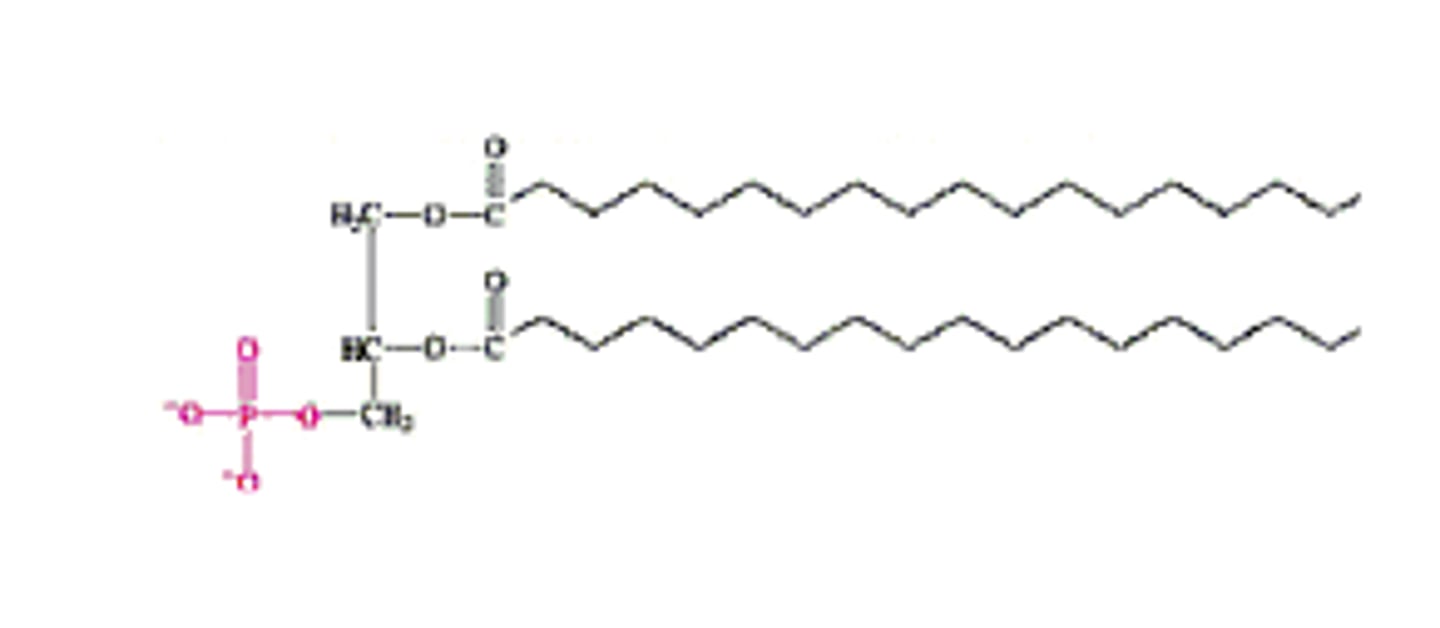
Common Glycerophospholipids
- Named based on head group attached to phosphate
1. Phosphatidylcholine (PC)
2. Phosphatidylethanolamine (PE)
3. Phosphatidylserine (PS)
4. Phosphatidylglycerol (PG)
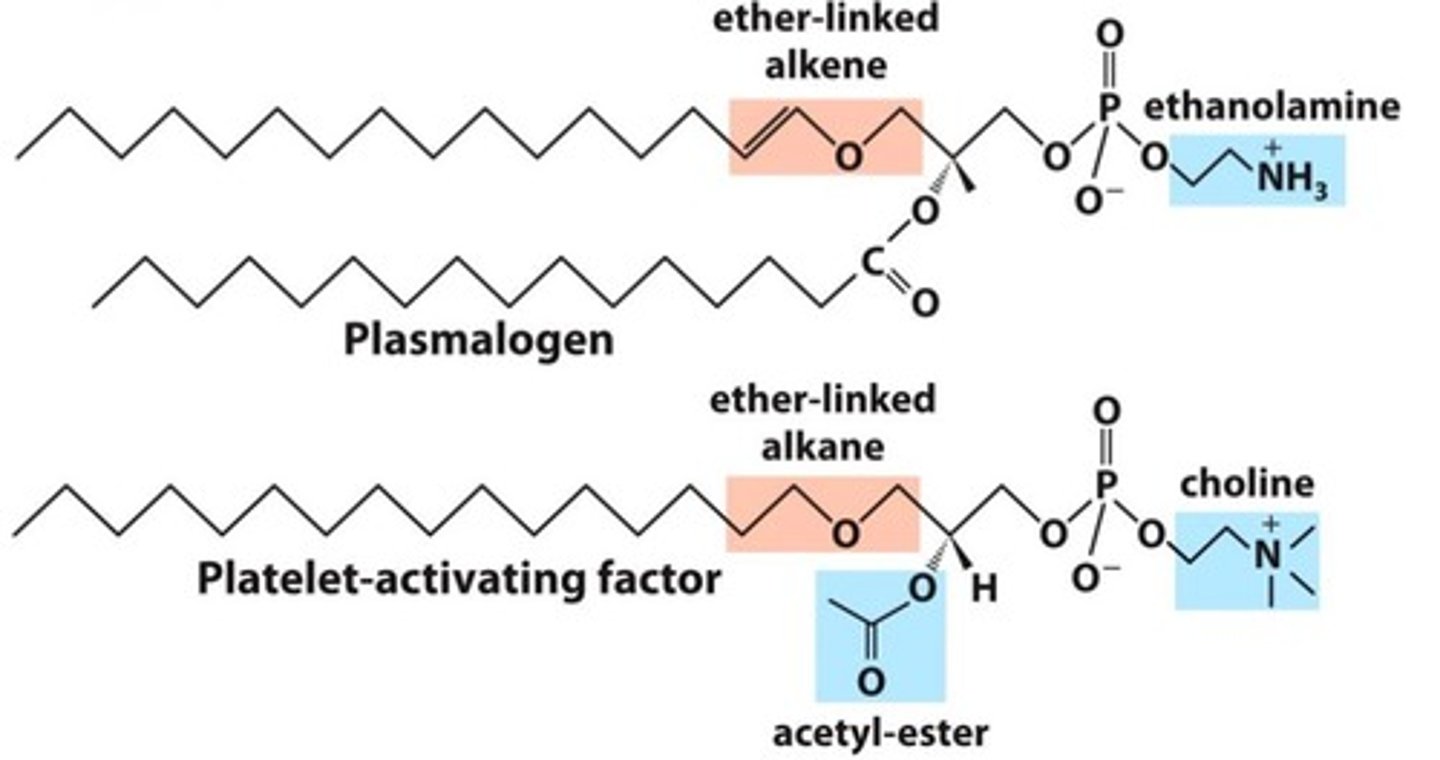
Ether Lipids (Variation)
- At C1, fatty acid replaced by alcohol & connected via ether linkage
- Examples: Plasmalogens (in archea), Platelet-activating factor (PAF)
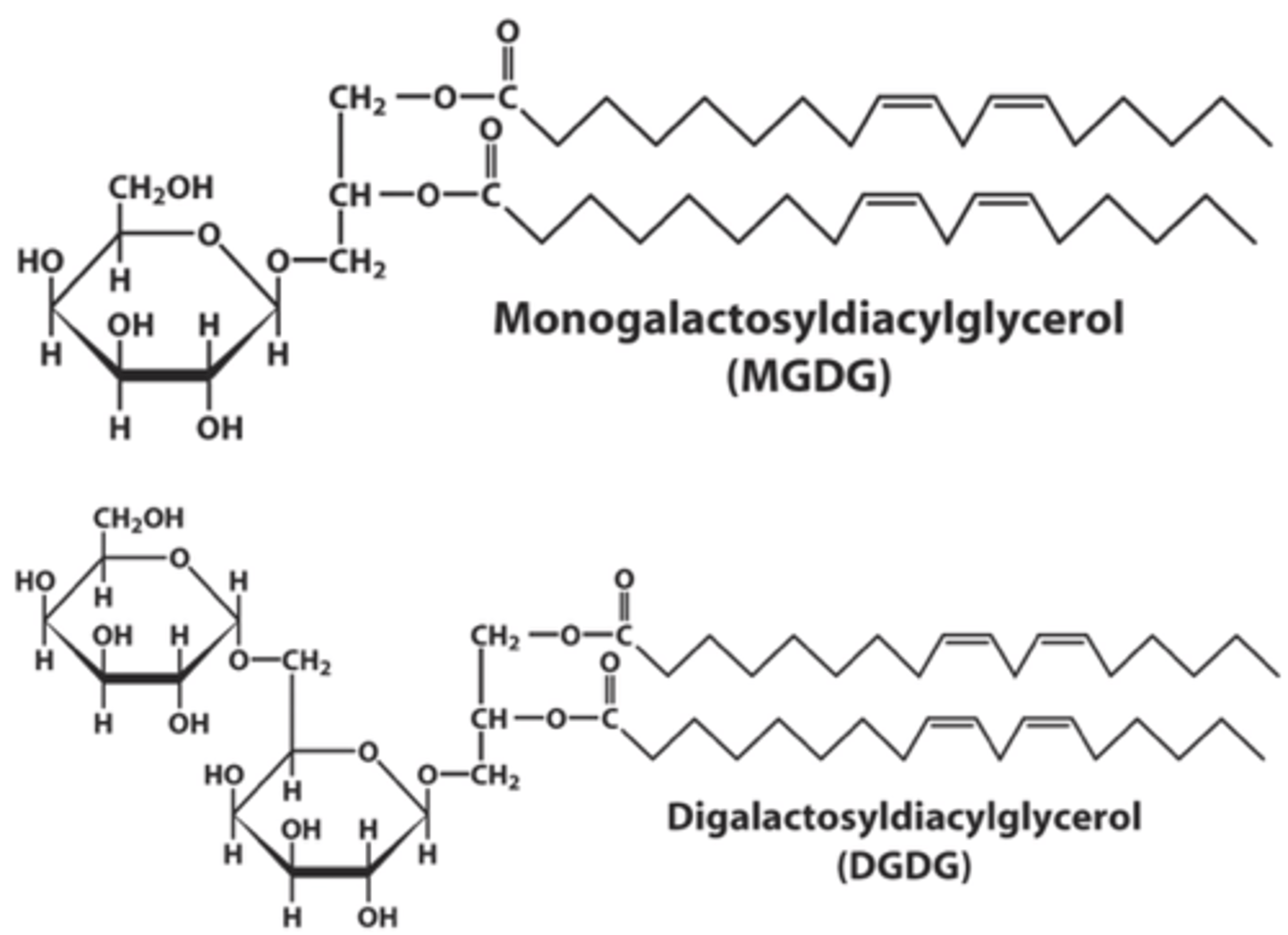
Glycolipids
MB lipids in with sugar groups attached to diacylglycerol via glycosidic bonds
Galactolipids
- attached to 1 (monogalactosyldiacylglycerol) or 2 (digalactosyldiacylglycerol) galactose residues
- Attached to the C3 hydroxyl group of diacylglycerol
- Linkage type: glycosidic bond
- Predominantly found in thylakoid MBs of chloroplasts
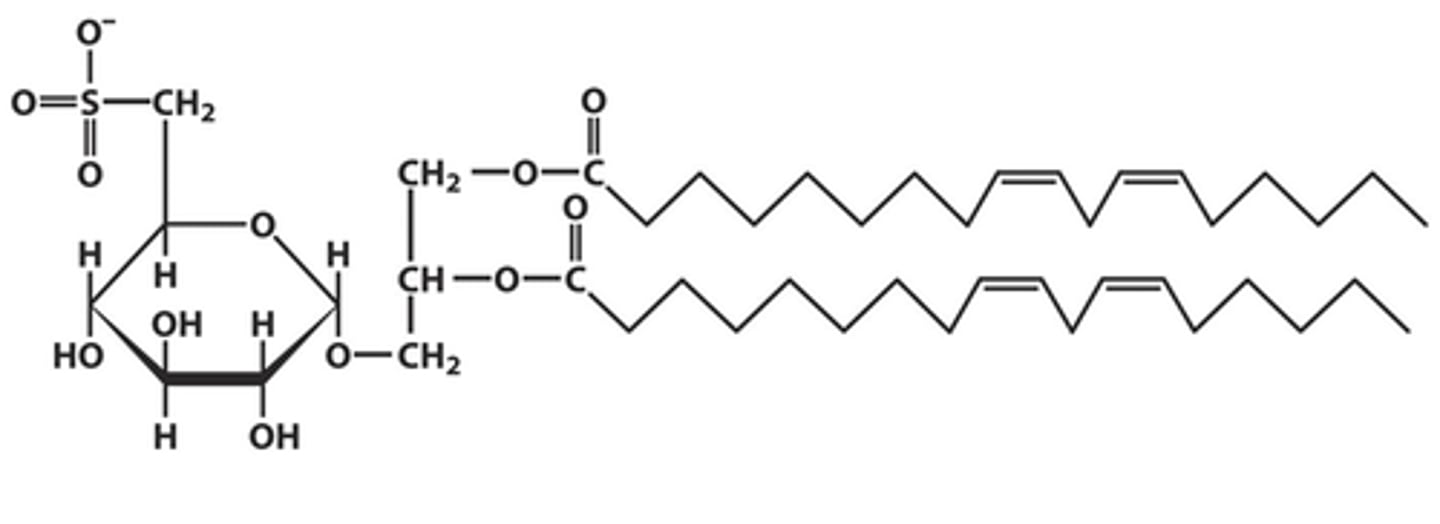
Sulfolipids
- sulfonated glucose attached to the C3 hydroxyl group of diacylglycerol linked via glycosidic bond
- Present in plant MBs
Sphingolipids
derived from sphingosine
sphingosine
- An 18-C linear amino alcohol
- OH at C1 and C3, Amino at C2 & Double bond at C4
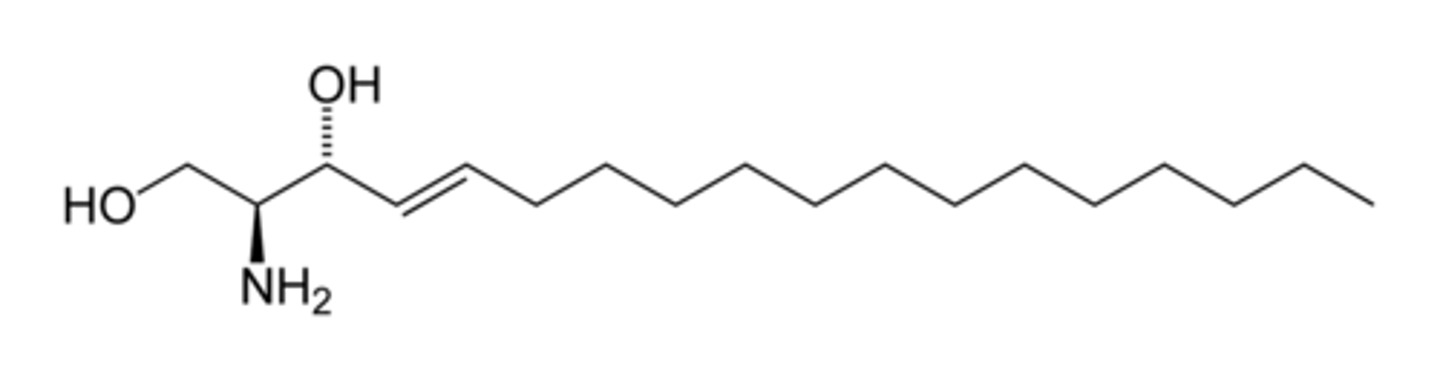
Ceramide Formation
-When a fatty acid attaches to the amino group at C2 of sphingosine
Ceramide
- 2 long hydrophobic chains
- Structurally similar to diacylglycerol (DAG) in hydrophobic character
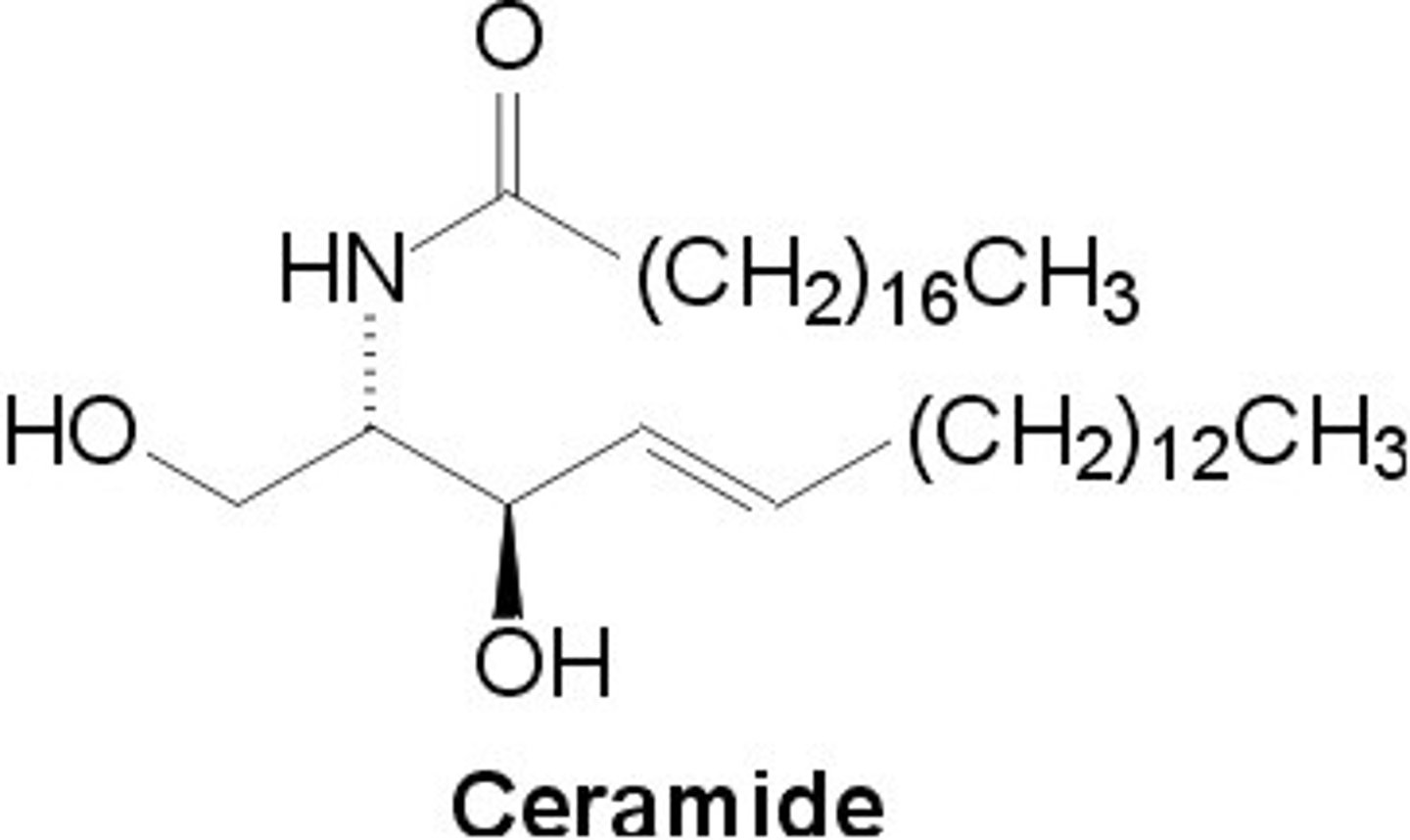
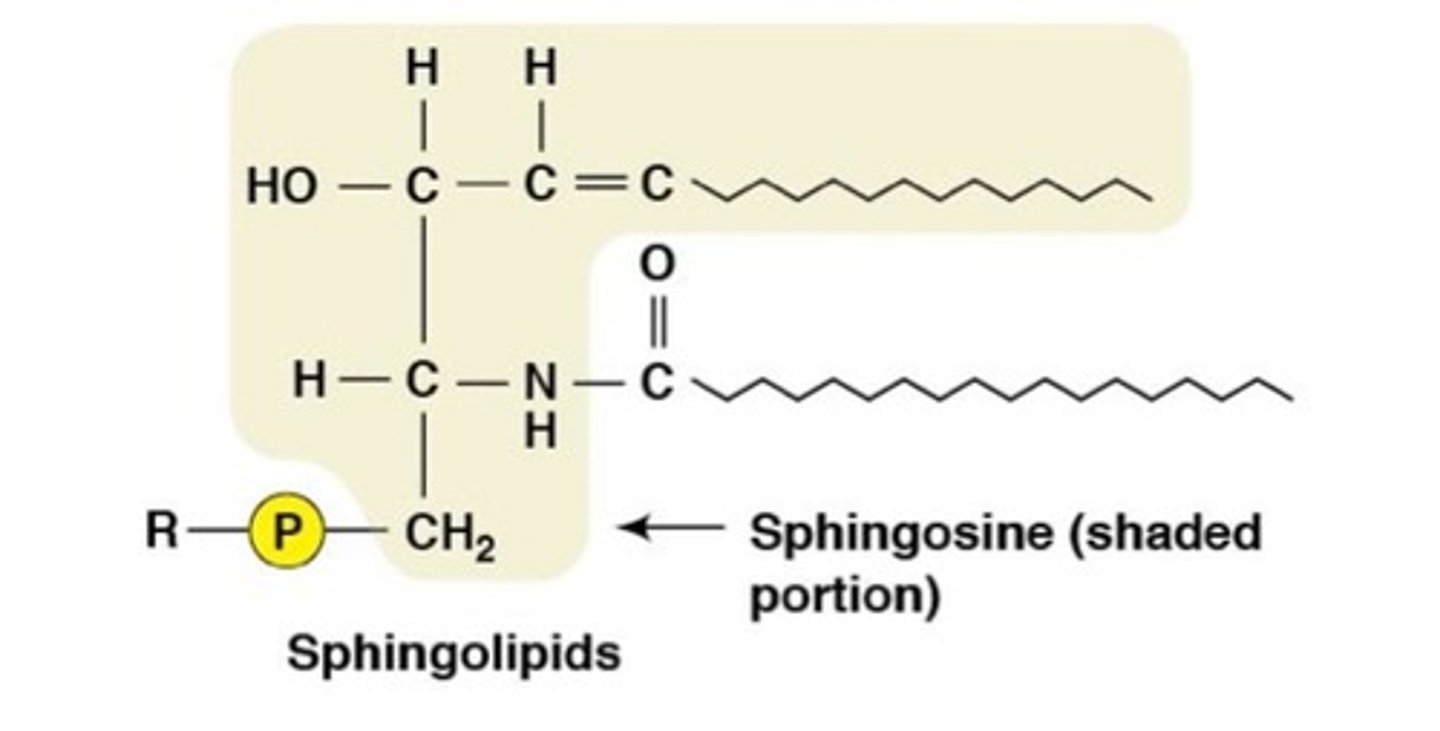
Sphingolipid Structure
A variable head group attaches to the C1 hydroxyl of ceramide.
Examples of sphingolipids
- Phosphocholine → Sphingomyelin
- Phosphoethanolamine → Sphingomyelin
- These resemble: Phosphatidylcholine (PC) and Phosphatidylethanolamine (PE) because they contain two hydrophobic chains + phosphate head group
Biological Importance of Sphingomyelins
- major components of the myelin sheath
- Provide electrical insulation around neuronal axons
Glycosphingolipids structure
- ceramides w sugar groups attached to C1 hydroxyl
- located on the outer surface of plasma MB
- a type of glycolipid
ABO Blood Group System
- Determined by specific sugar residues
- Present on glycolipids & some MB protein
Key Concept of Glycosphingolipids & ABO Blood Groups
- The A, B, and O blood groups differ by a single terminal sugar residue - Type O → base structure
- Type A → additional N-acetylgalactosamine
- Type B → additional galactose
Biological Importance of ABO blood groups (3)
1. Cell recognition
2. Immune response
3. Transfusion compatibility
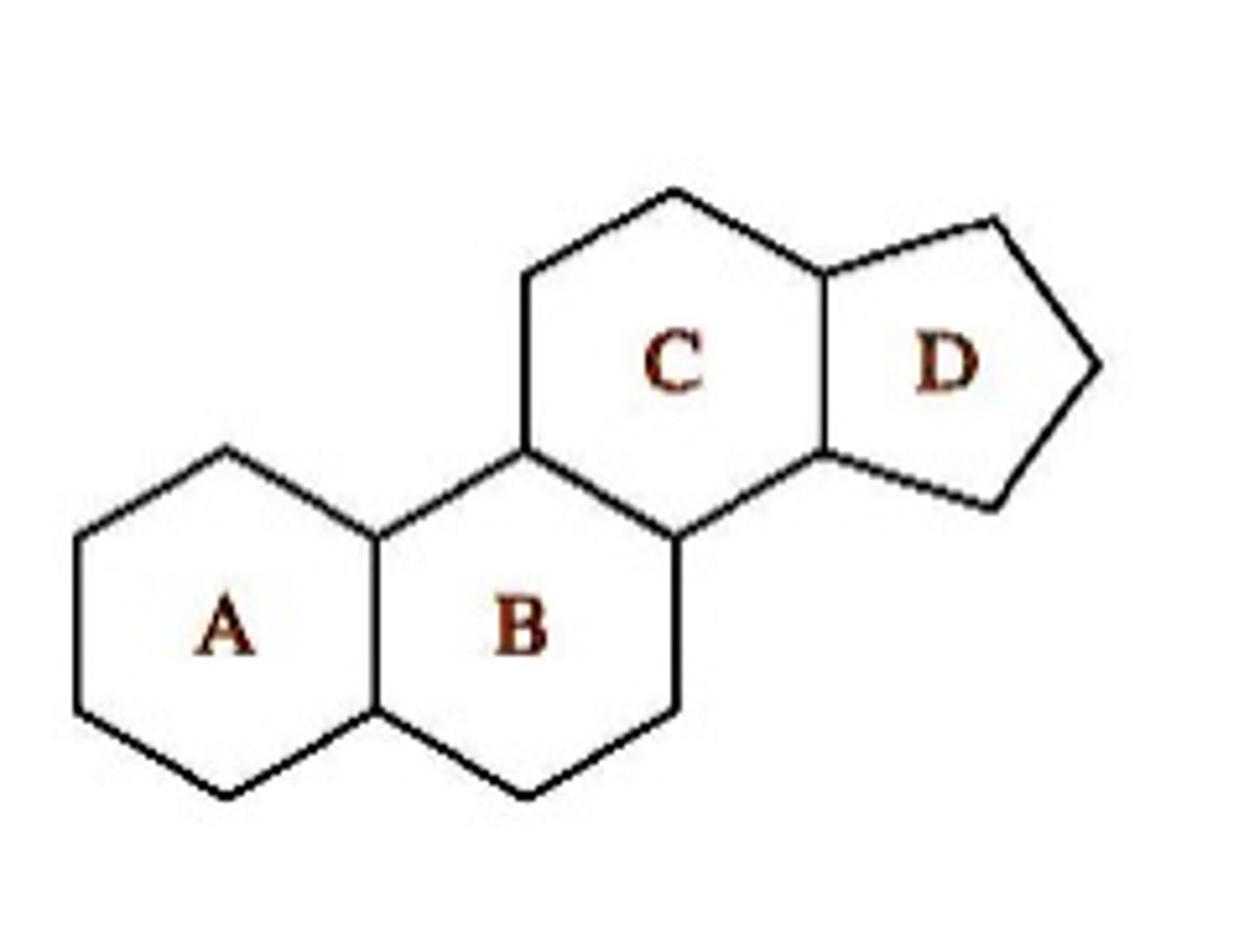
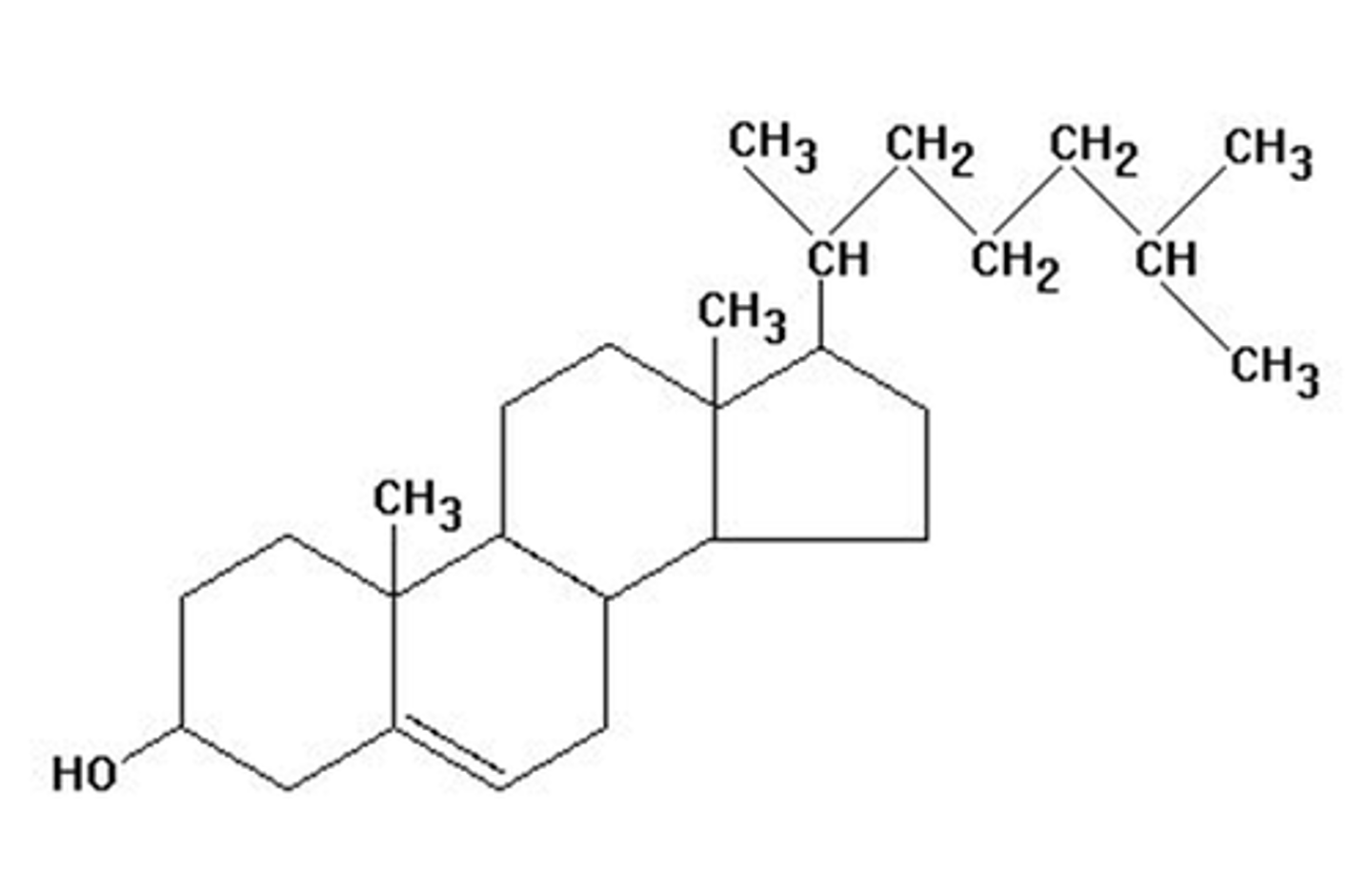
Steroid Structure
- derived from the steroid nucleus
- fused ring system consisting of three 6-carbon rings, one 5-carbon ring
- Rigid structure Hydrophobic core
Cholesterol
- Major steroid in animal cells
- Amphipathic: Hydroxyl (-OH) group → polar, hydrophilic
- Ring system + hydrocarbon tail → nonpolar, hydrophobic
= Located in animal cell membranes
Functional Role of Cholesterol (3)
1. Modulates MB fluidity
2. Stabilizes MB structure
3. Reduces permeability to small molecules
2 Other Steroids in Eukaryotes
1. Stigmasterol → Plants
2. Ergosterol → Fungi
(different types of chloesterol)
Role of Membrane Proteins (5)
1. transport
2. signaling
3. cell recognition
4. enzymes
5. structural support
What does protein composition vary by in cell membranes?
1. Cell type
2. Organelle
3. MB function
Integral/Transmembrane Membrane Proteins
- Span the lipid bilayer
- Contain 1 or more transmembrane domains
- Transmembrane regions are hydrophobic & often α-helices
3 Domains of Plasma Membrane Proteins
1. Extracellular domain → outside the cell (hydrophilic)
2. Transmembrane domain → within lipid bilayer (hydrophobic)
3. Cytosolic domain → inside the cell (hydrophilic)
How can transmembrane proteins be removed?
1. Detergents
2. Organic solvents (disrupt lipid bilayer)
Peripheral Membrane Proteins
- Located on MB surface & bound via Electrostatic interactions or Hydrogen bonding
- Interact with polar head groups & integral proteins
- hydrophillic
How can Peripheral Membrane Proteins be removed?
easily by High salt concentration (disrupt ionic interactions)
Examples of Peripheral Membrane Proteins
Enzymes, Anchorage, Transporters (carriers)
Lipid-Anchored Proteins
associate with MBs via covalent attachment to hydrophobic lipid groups, which insert into the bilayer
3 Common Lipid Modifications
Myristoylation, Palmitoylation, Prenylation
Myristoylation
- C14 fatty acid (myristoyl group) attached to N-terminal glycine via amide bond
Palmitoylation
- C16 fatty acid (palmitoyl group) attached to internal cysteine (thioester bond) & Serine (ester bond)
Prenylation
- Farnesyl (C15) or Geranyl (C20) attached to C-terminal cysteine via thioether bond
- these are isoprenoids (polymers of isoprene, C5)
Function of lipid anchors
allow proteins to associate with membranes without spanning them
How does Membrane composition vary?
- Organelle
- Cell type
- Organism
Membrane Asymmetry
- 2 leaflets of a MB are asymmetric
- Different lipid composition
- Different protein distribution
Fluid-Mosaic Model
- Developed to describe MB structure and dynamics
- MB is a lipid bilayer with proteins embedded within → "mosaic"
- the Lipids & many proteins can move laterally
- MBs behaves like a 2D fluid
Basis of Membrane Fluidity
- Due largely to unsaturated fatty acids
- Cis double bonds introduce kinks, disrupt tight packing & increase fluidity
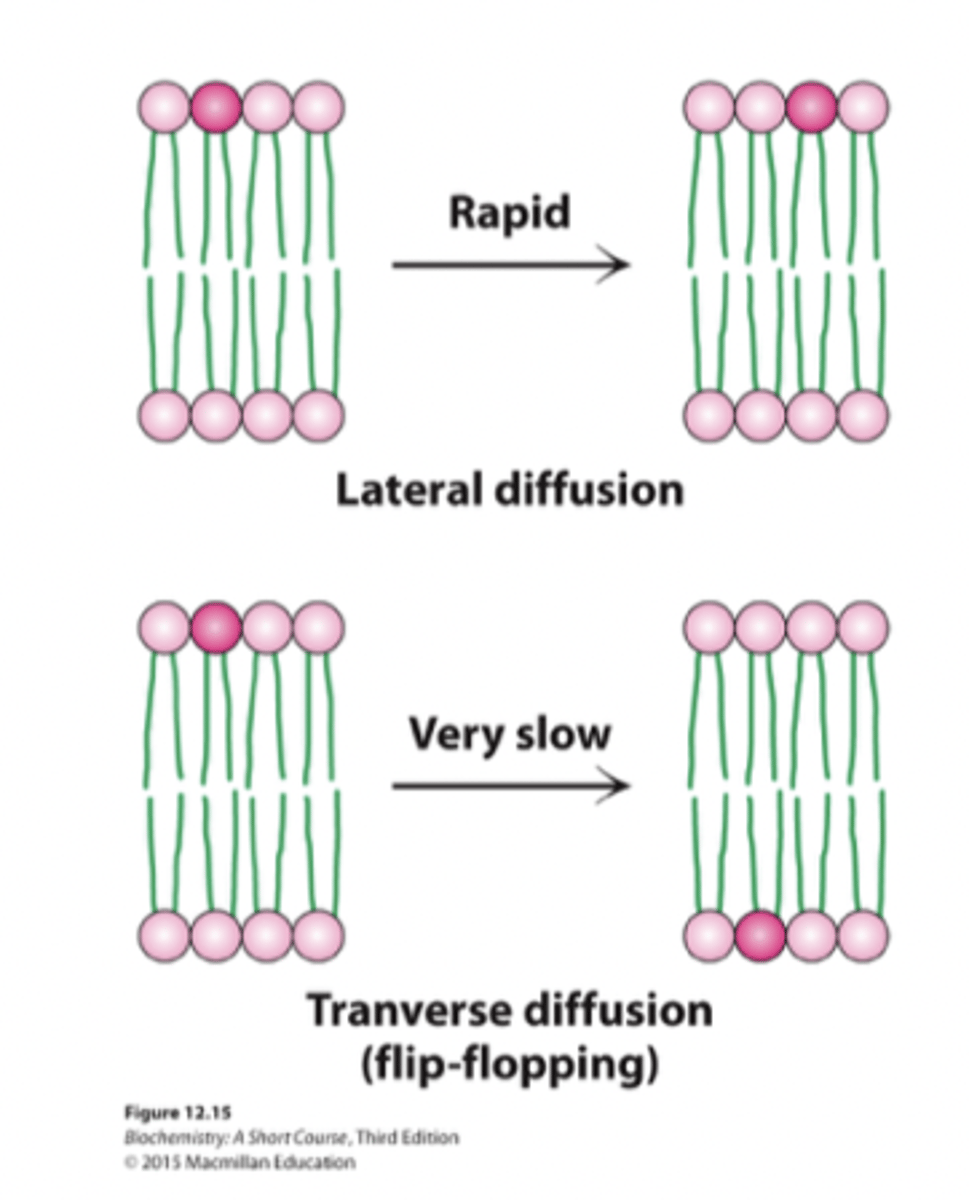
Lateral Diffusion
Membrane components move within the same leaflet
How is lateral diffusion demonstrated?
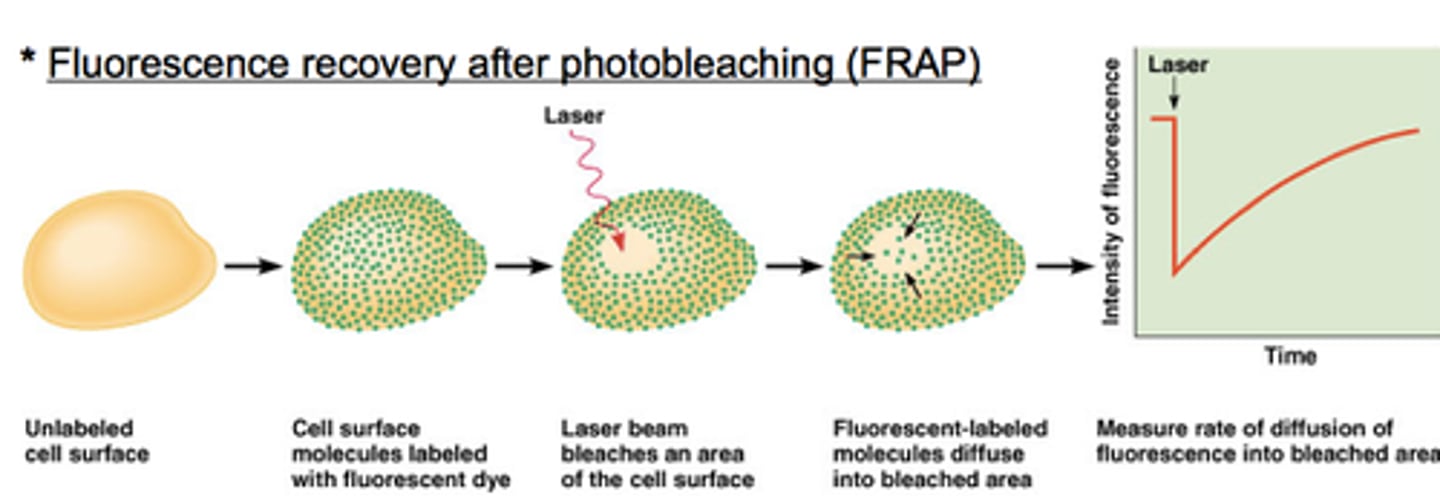
by the FRAP (Fluorescence Recovery After Photobleaching)
FRAP (Fluorescence Recovery After Photobleaching)
- Lipids labeled w fluorescent dye
- Small region bleached using laser
- Fluorescence returns as lipids diffuse laterally
- Confirms MB fluidity
Lateral movement
- within the same leaflet → easy