11- Translation(●'◡'●)
1/39
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
40 Terms
Key ingredients for translation and what the purpose is
purpose: Production of protein from mRNA template
ribosomes
tRNAs
aas (aa and tRNA= aminoacyl tRNAs)
transcript mRNA
What’s a polysome (polyribosome)
multiple ribosomes translating single mRNA at same time
Occurs when gene transcriptionally active
Ribosome purpose, location and general structure
Site of translation
Found in multiple locations in cell
cytoplasm, ER membrane, outer nuclear envelope, MT matrix, chloroplast stroma
Composed of 2 subunits (made of rRNA and proteins)
large (LSU) and small (SSU)
Ribosome composed of which rRNA (and proteins) in proks vs euks
rRNA folding pattern highly conserved in all organisms
Euks: 28S, 5.8S, 5S, 18S (expansions at surface)
Proks: 23S, 5S, 16S
THESE ARE rRNA inside the ribosome that build each subunit
Famous rRNA features-LSU
(23S/28S)
Interacts w/ tRNA
contains Peptidyl transferase center (PTC)=ribozyme= catalytic core
Catalyzes peptide bond formation (ribozymal activity)
a-sarcin-ricin loop (a seq)= highly conserved, target for toxins (23/28S)
Catalytic core is a highly conserved ribozyme = LUCA
Famous rRNA fts-SSU
SSU rRNAs= decoding center (interact w/ mRNA)
3’ end of 16S bp(anti-SD) with Shine Dalgarno seq
16S barcode= microbiome studies
18S interacts with mRNA (IRES, Kozak maybe)
LSU proteins and SSU proteins
LSU→ RPLs
SSU→ RPSs
both RP but then L for large, S for small
Compare prokaryotic vs eukaryotic ribosomes.
Prokaryotes | Eukaryotes |
|---|---|
30S + 50S = 70S | 40S + 60S = 80S |
Shine-Dalgarno | Kozak sequence |
fMet initiator | Met initiator |
Fewer protein factors | MANY eIFs |
Organism | Small subunit | Large subunit | Whole ribosome |
|---|---|---|---|
Prokaryote | 30S (16S) | 50S (23S + 5S) | 70S |
Eukaryote | 40S (18S) | 60S (28S + 5.8S + 5S) | 80S |
Ribosome 4 key sites
mRNA binding site (prokaryotes only)
E (exit site)
P (peptidyl site)
A (aminoacyl site)
Roles
Scaffold
Proofreading
Catalytic activity
tRNA will
bring amino acids to ribosome
each t RNA binds 1 specific amino acid
ester bonded to 3’ adenine
Recognizes and binds to 1 or more codon sites, has anticodon
Anticodon
3 base seq on tRNA that bp with mRNA codons via H bonding
anticodon forms a complementary RNA duplex with mRNA codon
Why can the anticodon in tRNA bind with more than 1 mRNA codon
Flexibility in 3rd position of codon
The wobble hypothesis
3rd position wobble effect where.. in the anticodon
G can pair w/ C or U
U can pair w/ A or G
I can pair w/ U, C or A
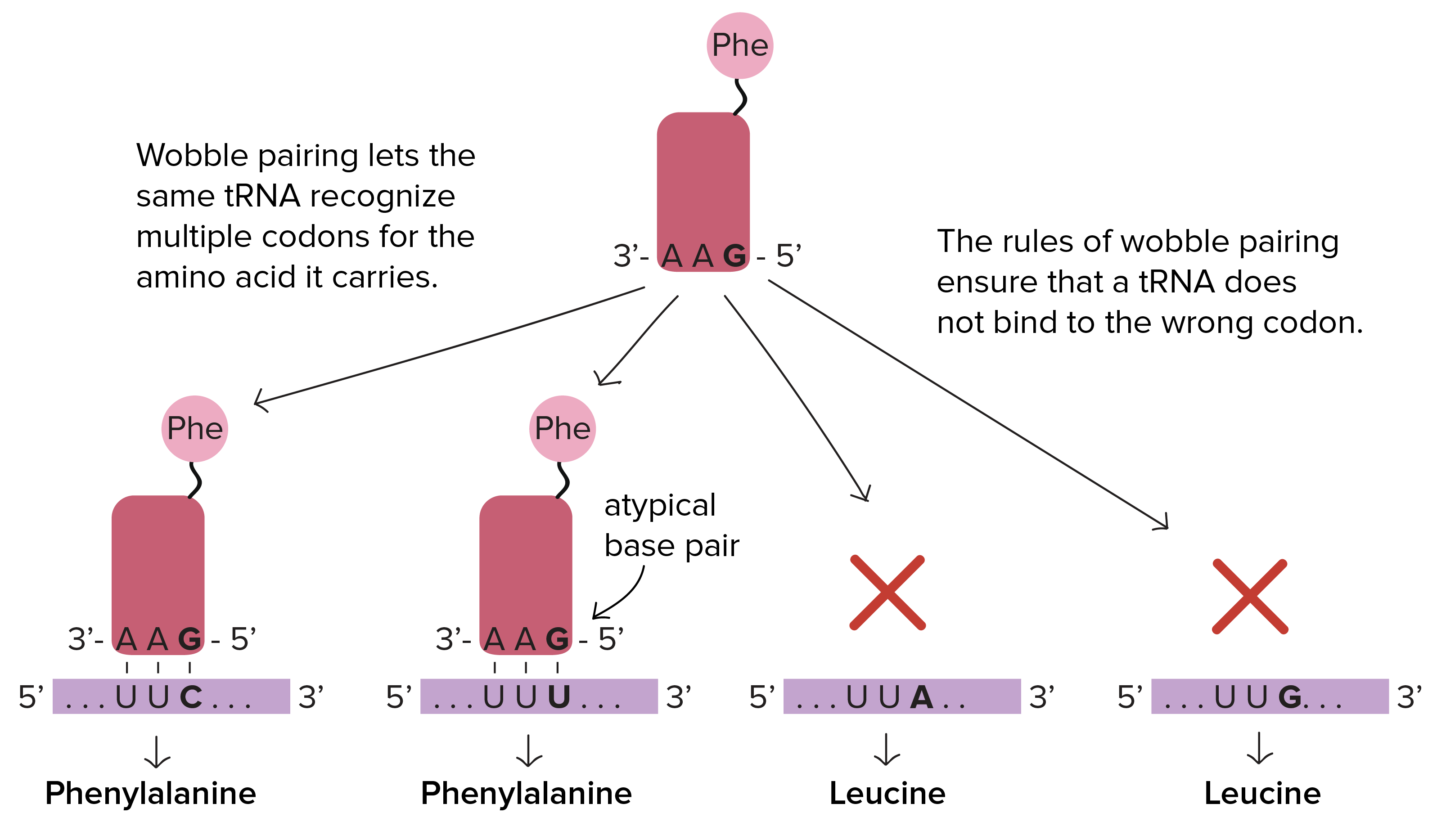
Naming convention for anticodon
3’-5’
mRNA 5’-GCC’3
tRNA written 3’-CGG5’ → antiparallel bp
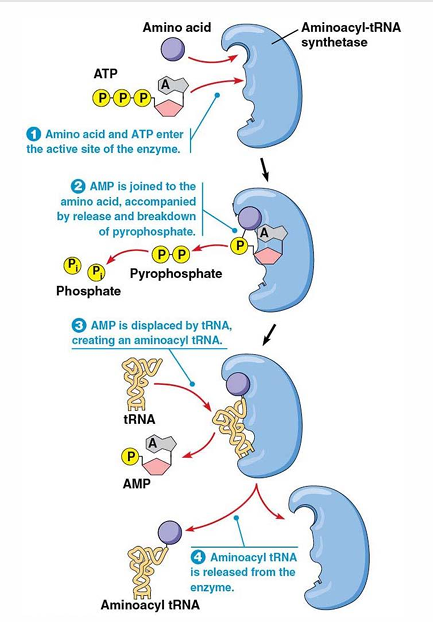
Aminoacyl-tRNA synthetases
Enzymes that attach appropriate aa to correct tRNA
amino acid activation
Cells have 20 diff tRNA synthetases
“super specificity”= synthetases recognizing tRNAs involves
Anticodon loop
3’ end
acceptor stem
What energy is required to add amino acid to tRNA?
ATP→ AMP (hydrolyzed) this is catalyzed by aminoacyl tRNA synthetases
“high energy bond” formed bw tRNA and aa
aminoacyl tRNA aka charged tRNA
Be able to draw mature mRNA for proks and euks
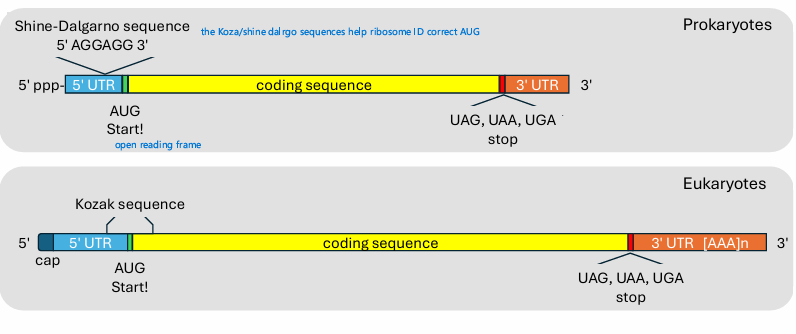
Differences bw proks and euks
Prokaryotes | Eukaryotes |
|---|---|
Direct recruitment of ribosome to AUG start codon | Cap-dependent recruitment followed by scanning for AUG start codon |
|
|
Special formylated tRNA (fMet) initiator | Methionine (Met) binds to AUG start codon |
Much of what we know about termination is based on prokaryotes | Much of what we know in eukaryotes is based on yeast |
— | Nucleus — transcription and translation are spatially and temporally separated |
— | WAY more protein factors (trans) and mRNA structural features (cis) regulate the process |
translation is primarily regulated at the initiation step. Why do you think that is?
Bc its extreme energy expensive so cells avoid wasting resources by controlling intitiation
What helps ribosome find start codon in proks vs euks
Proks
Shine dalgarno seq
Base pairs w/ anti-SD in 16S
Euks
Kozak seq
Cap-dependent scanning
Bacterial translation initiation regulated through
coordinated action of initiation factors
IFs
Step 1 of bacterial initiation
Initiation factors (trans) IF1, IF2, IF3 bind to SSU
GTP binds IF2
IF3 blocks 50S joining (LSU)
roadblock
Step 2 bac intiation
tRNA and mRNA recruitment
fmet tRNA binds P site
IF2 assists specificity
Shine-Dalgarno on mRNA
binds anti-SD in 16S rRNA (which is part of 30S subunit)
Start codon is now positioned correctly
IF3 dissociates= roadblock lifted
Step 3 bac initiation
LSU recruitment
After IF3 released
50S binds→ 70S initiation complex formed
GTP hydrolysis releases IF1 and IF2 during complex formation
Initiation in euks regultaed through
coordinated action of euk initiation factors
eIFs
5 main events that initiate translation in euks
Initiatior met-tRNAi met recruitment= ternary complex formation
mRNA activation= cap binding eIF4F complex + PABP
formation of preinitiation complex (PIC)= tRNA + 40S and bunch of eIFs
recruitment of mRNA to PIC + scanning for AUG start codon
recruitment of 60S= licensed to translate
Ternary complex formation
eIF2-GTP bind to Met-tRNAi Met
ternary bc there’s 3 items : EIF2, GTP, Met-tRNAiMet
mRNA activation
eIF4F complex binds 5’m7G cap
PolyA binding protein (PABP) binds 3’polyA tail
eIF4F complex binds PABP
circularized mRNA bc now a loop
Formation of 43S PIC
eIF3, eIF1, eIF1A binds to 40S
ternary complex joins party
eIF5 jumps on too
roadblocks
eIF1 sits in P site
eIF2-GTP blocks 60S binding (prevents it coming in too early)
Recruitment of mRNA to PIC= formation of 48S
interactions bw PIC + eIF4F brings mRNA → 40S
ATP dependent scanning for AUG start codon
correct AUG det by Kozak Seq
triggers removal of eIF1→ roadblock removed that was blocking P site
Met-tRNAiMet moves into P site
Recruitment of 60S to complete initiation
60S and eIF5B recruited
eIF5B GTPase activity promotes eIF2-GTP→ eIF2-GDP + Pi
Releases eIF2→ roadblock removed
60S can now bind
80S formed and licensed to translate
Non canonical translation- IRESs (cap independent)
IRESs= internal ribsome entry sites
highly struc RNA seq that can recruit ribosome directly in cap-independent manner (ribs can bind mRNA without cap)
might still need some initiation factors
Found in some viral RNA, and the struc of RNA can mimic tRNA
this tRNA like structure helps recruit the ribosome to the mRNA.
initiates translation at non-AUG start codon
During viral infection, host cells often shut down normal cap-dependent translation.
Viral RNAs with IRES elements can still recruit ribosomes, allowing viral proteins to be produced even when host translation is inhibited.
Elongation proks
Repetitive 3 steps coordinated by 3 elongation factors (EF-Tu, EF-Ts, EF-G)
Binding of aminoacyl tRNA
Specified tRNA enters A site
Peptide bond formation
AA on tRNA in P site hydrolyzed + peptide bonded to aa on A site tRNA
this catalyzed by ribozyme site in 23/28S rRNA
Translocation
Motion of ribosomal subunits moves the complex 3 nucleotides on mRNA, so tRNA in P site to enter E site, A site tRNA→ P site
Proks elongation Step 1-Aminoacyl tRNA binding
Begins with AUG start codon in P site
binding of tRNA to A site requires EFs
EF-Tu (GTPase turns GTP into GDP) binds GTP + then aa-tRNA^aa
EF-Tu-GTP-aa-tRNAaa binds to mRNA
GTP hydrolysis releases EF-Tu-GDP
EF-Ts (GEF) recycles EF-Tu
GDP→ GTP
Prok Elongation Step 2-Peptide bond formation
AA in P site cleaved from connection at 3’ end of tRNA
AAs in P and A site located in peptidyl transferase center (PTC=catalytic core of the ribosome and acts as a ribozyme)
covalent peptide bond formed bw aa on a tRNA in A site + aa on a tRNA in P site
Elongation step 3-translocation
EF-G-GTP binds to 50S (LSU)
Ef-G= G protein
GTP hydrolysis causes conformation change in ribosome, moving it 3 nucleotides along mRNA
• EF-G-GTP →EF-G-GDP + Pi
everything shifts over 1 site
“A” site now free to receive next aa-tRNA
REPEAT till stop codon reached
Elongation is the same for proks and euks but switch
Prokaryotes Eukaryotes
EF-Tu eEF1A
EF-Ts eEF1B
EF-G eEF2
Termination proks
Mediates through release factor (RF) which mimics a tRNA
RF-GTP binds stop codon in A site
Hydrolysis of RF-GTP→ RF-GDP + Pi
releases polypeptide
Ribosome/mRNA complex dissociates
Termination and recyling in euks
Similar to proksL euk release factors (eRFs) recognize stop codon + release polypeptide and tRNA
eRF1 mimics a tRNA and binds the stop codon
eRF3 through GTP hydrolysis helps release the polypeptide.
ABCE1 uses ATP splits the subunits and recycling factors remove mRNA/tRNA so the 40S can be reused.
eRF1 mimics a tRNA and binds to any of the 3 stop codons (UAA, UAG, UGA) via its N-terminal domain
eRF3 is a GTPase that binds eRF1, GTP hydrolysis moves part of eRF1 into the PTC where it can promote polypeptide release
eRF1 recruits ABCE1 which catalyzes subunit splitting via ATP hydrolysis = “power stroke” -
Additional recycling factors remove the mRNA and tRNA from the 40S
40S is now able to be recycled for a new round of translation (recall circularization of mRNA thanks)
How is mRNA translation energetically expensive
use as much as 80% of cells energy
10 aa> 300 kcal/mol
40 aa/sec by polyribosome ( multiplt ribosomes simultanosuely translating mRNA)
Polypeptide folding via molecular chaperones
protein need to fold into correct 3D shapes before they can function
Spon fold based on primary aa seq but cytoplasm BUSY place
Hsp70→ new proteins
Hsp60→ aka chaperonin→ unfolded proteins (heat shock)