HLSC322 exam 3 woohoo yippee
1/24
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
25 Terms
Restriction enzyme
One of bacteria’s antiviral defense mechanisms
dsbreaks DNA at specific ‘palindromic’ sequences
eg. EcoR1’s recognition sequence:
5’ – GAATTC – 3’
3’ – CTTAAG – 5’
^ reading 5’ to 3’ on both strands gets u the same sequence
Sticky vs. Blunt cuts
Sticky = staggered = cohesive cuts will leave overhangs, while blunt cuts do not
3 features of a cloning vector
Origin of replication recognized by bacteria
so recombinant plasmid can get replicated
Selectable marker (often antibiotic resistance)
allows isolation of transgenic bacteria from nontransgenic bacteria
Polylinker aka Multiple Cloning Site (MCS)
contains many restriction sites — allows piece of DNA to be inserted into that region

Steps of DNA insertion into vector
Digest plasmid
Phosphatase treatment
Digest DNA of interest
Purify both fragments
Allow fragments to anneal
Seal nicks in bonds
Transform bacteria
Spread colonies on plate with ampicillin
Isolate colonies
Determine if correct vector was inserted into correct vector
Screening for success of DNA insertion - Disruption of LacZ
So u got ur Plasmid with LacZ and restriction site within the LacZ
If u place untreated bacteria in media with ampicillin and X-gal, they appear blue — this is cuz LacZ gene codes for beta-galactosidase, which cleaves x gal to produce blue pigment.
When ur DNA of interest gets inserted in middle of LacZ, it effectively kills the gene: so DNA successfully inserted → colony remains white.
Polymerase Chain Reaction
what are the ingredients?
what is the procedure?
Technique to amplify — produce many many copies of — DNA
Ingredients: heat-resistant Taq polymerase, dNTPs, primers, thermocycler machine
since primers are used, u must have some knowledge about sequence u are PCRing
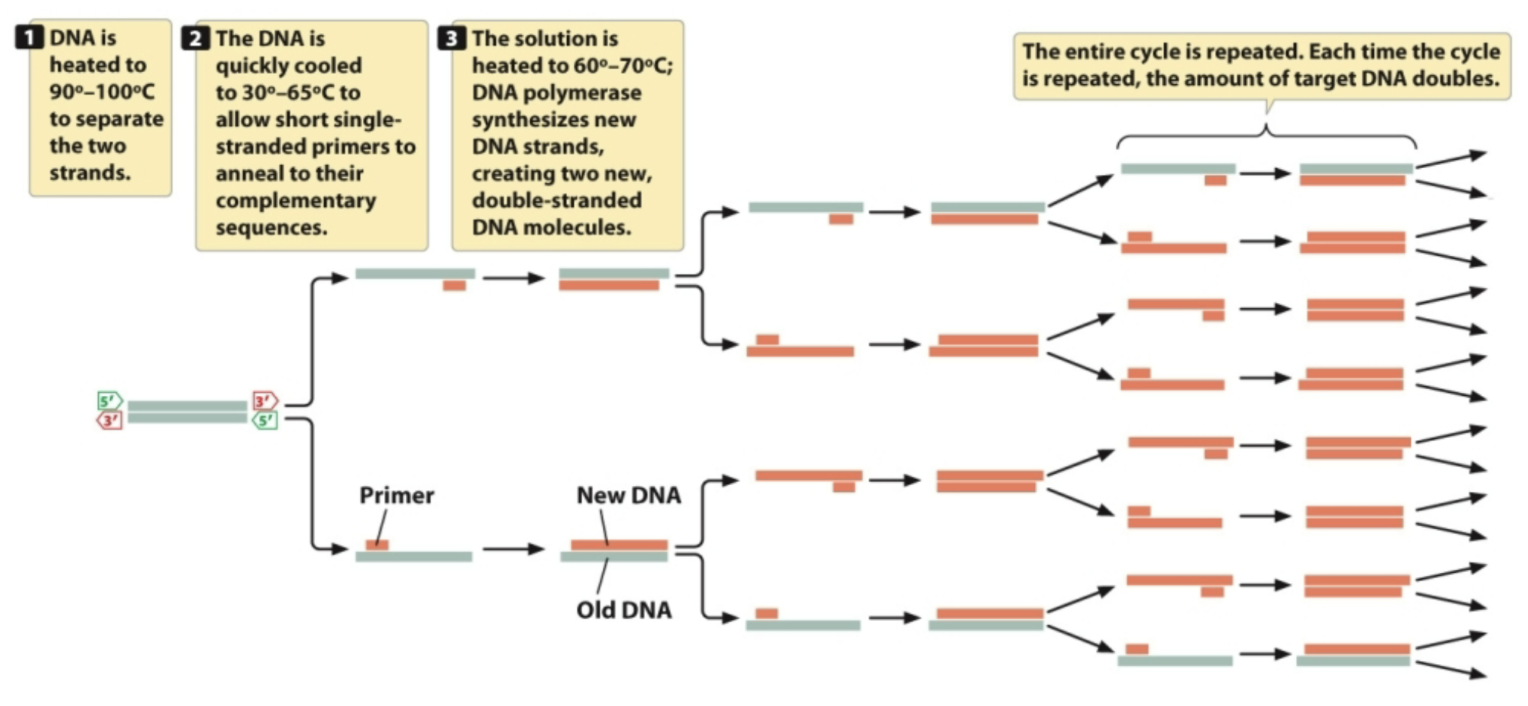
Procedure
Heat to 90 - 100oC to separate 2 strands
Cool to 30 - 65oC so primers can anneal
Heat to 60 - 70oC so Taq polymerase can do its thing
(exact temperatures depend on specific sequence, prolly more GC% content means higher melting point)
Types of genomic differences between hoomans
Single Nucleotide Polymorphisms (SNPs)
Indels
Short Tandem Repeats (STRs)
Copy Number Variants (CNVs)
Combined DNA Index System (CODIS)
uses standard set of 13 primers for 13 unlinked loci as a genetic profile
used for forensics, paternity tests and the like
Haplotype
group of closely linked genes / genetic markers on a single chromosome that tend to be inherited together
used to trace ancestry
Transposable elements
aka jumping genes / transposons / mobile DNA
44.4% of human genome
code for transposase enzyme. Transposase does staggered cut → transposable element inserts → DNA polymerase fills in gaps, forming flanking direct repeats
4 mechanisms of transposition
DNA transposition
Replicative transposition = new copy inserts itself, original copy stays behind
Nonreplicative transposition = original copy moves to new site
RNA intermediate transposition
How do genomes retain stability with so many transposable elements potentially messing things up?
Methylation
piRNAs
Inhibition of transposase expression
Polymorphism
natural occurrence of multiple alleles of gene or DNA sequence within population
Allo vs. Auto polyploidy
having more than 2 complete sets of chromosomes from 2 or more different species (i.e. vs. the same species
more common in plants than animals
Aneuploidy & its 4 types (in humans)
Having different than 2 complete sets of chromosomes
Nullisomy = 0 copies (2 deleted)
Monosomy = 1 copy (1 deleted)
Trisomy = 3 copies (1 duplicated)
Tetrasomy = 4 copes (2 duplicated)
Primary vs. Familial Down Syndrome
Primary DS (95% of cases) = Trisomy 21 ft. meiotic nondisjunction
Familial DS = Parent carries a balanced rearrangement of chromosome 21 and another chromosome (e.g., 14), which does not affect them but leads to unbalanced, inheriting three 21st chromosome copies in their child.
4 types of major chromosomal rearrangements
Duplication
Deletion
Inversion
Translocation
Operon
Parts of the Lac operon
Promoter = RNA Polymerase binding site
Operator = Repressor binding site
Constitutive expression
Gene is constantly expressed under normal cellular conditions
Positive vs Negative control
increase vs decrease gene expression
Upregulation vs downregulation
Cis vs trans-acting regulatory elements
Cis-acting = regulates adjacent gene on same chromosome
Trans-acting = regulates genes anywhere in genome via production of diffusible molecules
DNA-binding proteins aka Transcription factors
Inducible vs Repressible
Inducible = gene expression usually off
Repressible = gene expression usually on
How does the lac operon allow for environment-dependent digestion of lactose in bacteria?
LacI is constitutively expressed = repressor is constitutively produced
When no lactose present, inhibitor