BIOL300 Exam 2
1/34
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
35 Terms
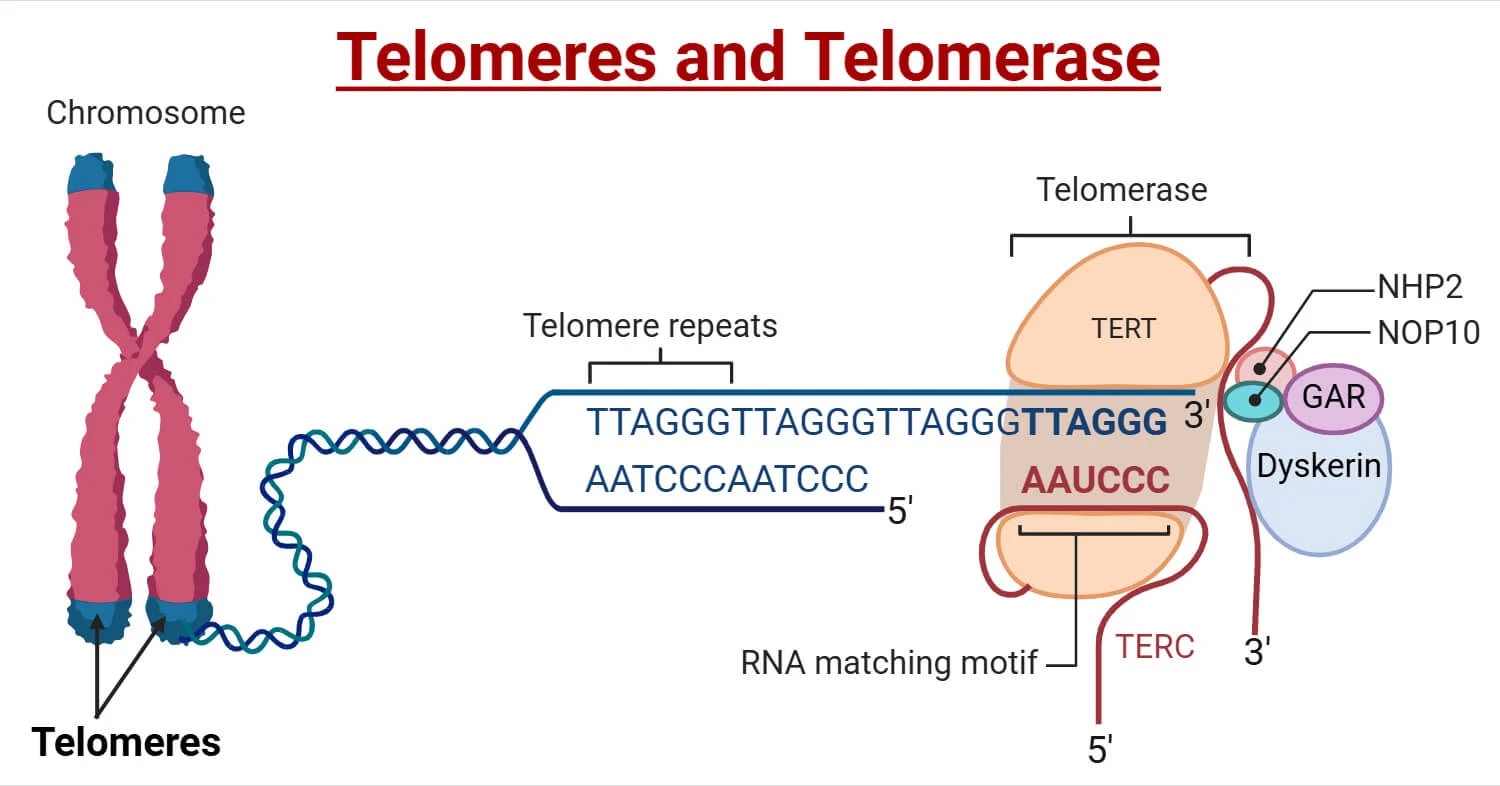
SA - Explain in detail the structural features of telomerase and how it works to prevent chromosome shortening; include a sketch/diagram
Telomerase is an enzyme that functions to lengthen telomeres and prevent shortening
It is composed of a catalytic protein subunit (TERT) and a functional RNA template (TER) arranged in a ring-like structure. The telomerase would bind to the sequence at the end of a telomere and synthesize DNA using the enzyme telomerase reverse transcriptase.
SA - Explain the steps of DSBR by homologous recombination
Spo11 endonuclease actively produces DSB
Endonuclease and exonuclease digest 5’ end of break and create 3’ overhang
Strand invades and creates a heteroduplex and D-loop
Invading strand is extended. Movement of recombination joint along the duplex is called branch migration
D-loop converts into two Holliday junctions
Holliday junctions are resolved with two cuts. Result is noncrossover DNA if cut in same direction, and is recombinant DNA if in opposite directions
SA - Explain two features of DNA polymerase that prevent it from replicating one of the DNA strands at the 3’ end of linear chromosomes.
DNA polymerase requires a pre-existing DNA primer to initiate synthesis and it can only extend strands from the 5’ to 3’ direction
SA - Explain why translesion-replicating polymerases are more error-prone than replicative polymerases.
Translesion DNA polymerases add any base along the DNA sequence to get past distortive lesions, making translesion repair error-prone
SA - What are Okazaki fragments, and why is this strategy necessary for replicating the lagging strand?
Okazaki fragments are sequences synthesized by DNA polymerase and covalently linked together by DNA ligase. They form on the lagging strand and are discontinuous segments
It’s necessary because DNA polymerase only synthesizes DNA from the 5’ to 3’ direction. Polymerase must wait for DNA helicase to expose new segments of DNA, then synthesize small fragments moving away from the fork with several primers placed
SA - In one sentence each, describe RecA, RecB, RecC, and RecD.
RecA enables homologous strands to find one another and promotes D-loop formation
RecB degrades DNA and loads RecA onto ssDNA
RecC recognizes Chi sites and generates ss 3’ end
RecD functions as a 5’ to 3’ helicase that unwinds dsDNA
SA - Why is recombination important for both DNA repair and meiosis?
Recombination serves to repair DSBs and stalled replication forks and create chiasmata, which ensure proper chromosome segregation during meiosis
SA - What is an OriC versus ORC? How many do eukaryotes have, and how many do prokaryotes have? What is their function?
OriC is a prokaryotic DNA sequence where replication begins, using DnaA to recognize the site and start splitting. ORC is a eukaryotic complex that binds to dispersed origins to start replication and manage cell division
Prokaryotes have one OriC per chromosome, and eukaryotes have thousands
SA - What are Okazaki fragments? Why are they necessary for DNA replication?
Okazaki fragments are short DNA sequences that are synthesized discontinuously on the lagging strand during DNA replication
They are necessary as DNA polymerase only travels in the 5’ to 3’ direction and, with several primers laid out, is capable of synthesizing on the lagging strand. Without forming these fragments, the cell wouldn’t be able to replicate the lagging template strand and genetic duplication would be incomplete
SA - Explain what DnaA is.
DnaA is a protein that binds to DnaA boxes in the OriC region, causing the AT-rich DNA to unwind. It also acts as a transcriptional regulator for genes involved in cell cycle control
Parts of the “human hand” model of DNA polymerases and their functions?
Palm - catalytic active site
Fingers - position the template into the active site
Thumb - binds DNA as it exits, processivity
Explain the differences between DNA synthesis in prokaryotes and eukaryotes.
Prokaryotic DNA synthesis is rapid, occurs at one OriC, and uses DNA polymerase III
In eukaryotes, it occurs in the nucleus, has multiple origins or replicons, and uses several specialized polymerases (α, ε, δ)
What is needed to prime DNA synthesis; and explain why.
A primer. It provides a free 3’-OH group, which DNA polymerase needs to add dNTPs
What does it mean that DNA polymerase III is a highly processive enzyme?
DNA polymerase III can add thousands of nucleotides to a growing DNA strand without it falling off. This is driven by a sliding β-clamp, which keeps it securely attached to the template
What are licensing factors and when are they expressed?
These are proteins, ORC, Cdc6, Cdt1, and MCM helicase, which are loaded onto DNA during the late mitosis/early G1 phase
What are the sequence elements at OriC, and what purpose does each serve?
DnaA boxes are found throughout OriC and serve as binding sites for the DnaA initiator protein
GATC methylation sites are recognized by Dam. Newly synthesized DNA is hemimethylated
Know what the proteins are that are involved in the formation of the replication fork at an Ori.
DnaA binds to OriC, and DnaC (helicase holder) loads DnaB (helicase) to melt the DNA. Single-stranded binding proteins (SSB), DNA primase (DnaG), and DNA polymerase III then assemble to begin replication
What are the ways that replication origins in eukaryotic cells are different from in prokaryotic cells
Eukaryotic origins have multiple, less-defined sites per chromosome that activate during S-phase
Prokaryotes use a single, specific origin called an OriC
How are RNA primers removed in prokaryotic cells and in eukaryotic cells?
DNA polymerase I in prokaryotic cells
Flap endonuclease I and DNA polymerase δ in eukaryotic cells
What are the various functions of homologous recombination?
It is engaged in critical DNA repair and genetic exchange mechanisms. It repairs DSBs and facilitates genetic diversity via crossover of chromosomes
What proteins are be involved in the resolution of Holliday junctions?
Endonucleases such as resolvases cleave DNA intermediates to finalize recombination. RuvAB and helicases are involved
What are CHI sites and what is their role?
Chi sites stimulate recombination and create hotspots. The function to signal RecBCD to stop degrading DNA and instead imitate repair of broken replication forks, facilitating repair
What does the RecBCD complex do?
It initiates repair of DNA DSBs and homologous recombination. It loads RecA onto ssDNA
What are the proteins involved in Base excision repair, and explain their functions.
Glycosylase (cleaves hydrogen bonds between base and DNA), lyase (endonuclease opens sugar-ring), DNA polymerase, and ligase
Explain the basis of Xeroderma Pigmentosum.
Xeroderma Pigmentosum is an autosomal disorder caused by mutations in genes that disrupt the nucleotide excision repair pathway. The result is an inability to repair DNA damage caused by UV radiation
Which type(s) of recombination systems result in loss (deletion) of genetic material, and which do not?
The loss of genetic material is the result of non-homologous end joining (NHEJ), site-specific recombination between direct repeats, single-strand annealing, etc
Which DNA repair systems are most error-prone?
Non-homologous end joining (NHEJ) and translesion synthesis (TLS)
What processes may lead to gene conversion?
Double-strand break repair (DSBR) during meiosis, synthesis-dependent strand annealing (SDSA) during mitosis, and mismatch repair
Understand the NER system in E. coli and describe the functions of the Uvr proteins.
Nucleotide excision repair (NER) repairs bulky DNA damages, such as UV-induced thymine dimers
UvrABC cuts out the damaged segment, and UvrD helicase removes it to be replaced via DNA polymerase I and ligase
Explain the role of the ruv genes.
They play a role in DNA repair and homologous recombination by processing Holliday junctions
RuvA binds to the Holliday junction
RuvAB promotes branch migration
RuvC endonuclease resolves Holliday junctions via cuts
What is the structure of DNA transposons? What do they all contain?
Structure consists of a central coding region (ORF), terminal inverted repeats, and flanking direct repeats
What are the structures of class I transposons? What property do they all share?
Long terminal repeat (LTR) retrotransposons have long LTRs at both ends in the same orientation. Between both LTRs are genes that engage in transposition
Non-LTR retrotransposons lack LTRs and are divided into two types
LINEs are autonomous
SINEs are nonautonomous
They all share an RNA intermediate (copy-and-paste)
Distinguish between autonomous and non-autonomous transposable elements.
Autonomous transposable elements encode their own genomes and jump on their own (containing transposase and reverse transcriptase)
Non-autonomous transposable elements lack these genes and rely on machinery produced by autonomous elements to move
Know the percentages of the human genome represented by each category of transposable elements.
Transposons make up 45% of the human genome
DNA transposons (3%)
LTR retrotransposons (8%)
SINEs (15%)
LINEs (20%)
What percentage of transposons are active (and which ones)?
Less than 0.05%, including LINE-1 and Alu