MCB 2210 L12: Protein Targeting - Nucleus Part 1
1/29
Earn XP
Description and Tags
Slides 1-32
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
30 Terms
Protein Targeting
Biological process cells use to delivery newly made proteins to their correct, functional location to carry out specific functions within cell.
How do proteins get localized to different places in the cell?
Components of organelles in eukaryotic cells
Organelle = compartment within eukaryotic cell
Ex. Nucleus, Golgi apparatus, ER, lysosomes, mitochondria, chloroplasts, etc.
Bound by ≥1 membranes
Proteins define function of organelles
Potential destinations for proteins
Cytosol = fluid around organelles (main part of cytoplasm)
Extracellular/secreted = outside cell
Plasma membrane = part of membrane
Membrane of organelles
Lumen of organelles = space within organelle
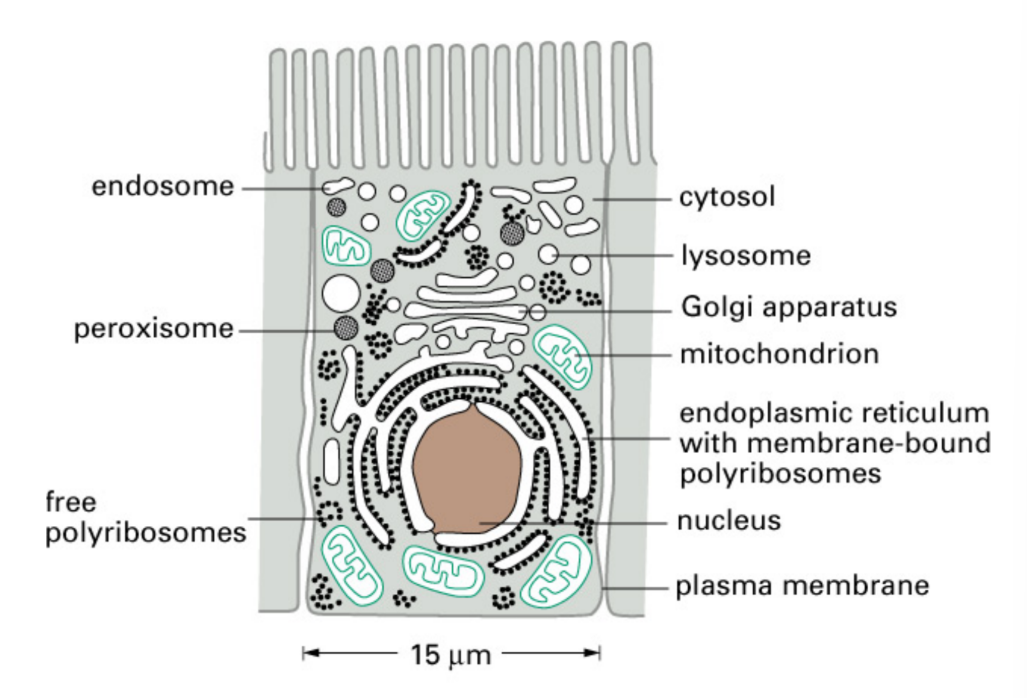
Where do the majority of proteins begin to be synthesized?
Cytosolic ribosomes
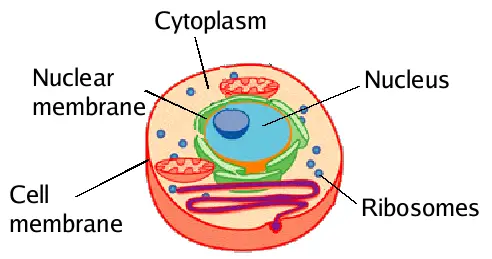
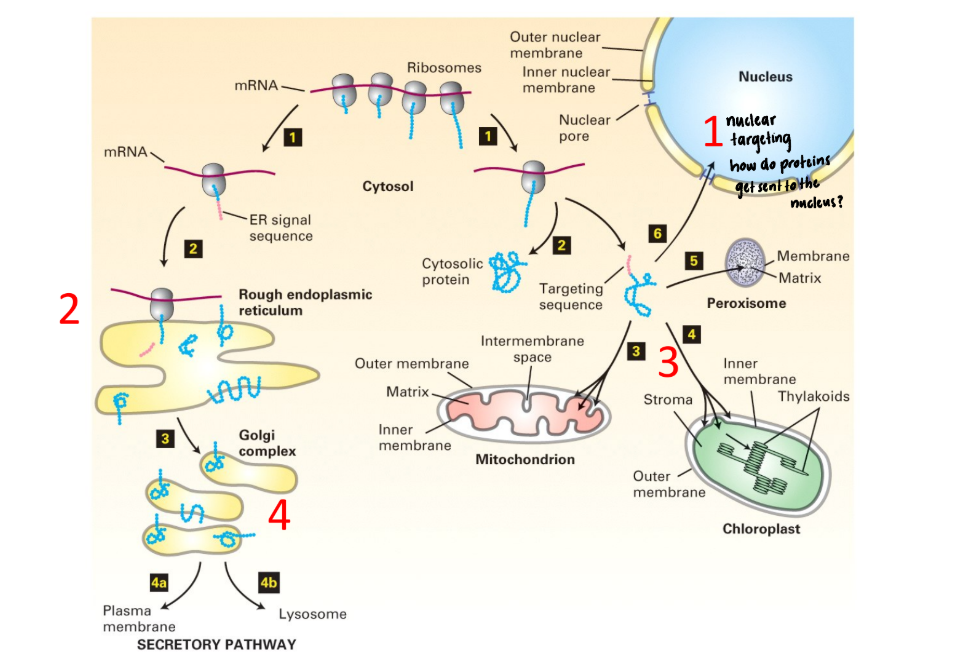
Pathway of protein targeting in eukaryotes
Protein synthesis @ cytosolic/free ribosomes
Has targeting sequence (address)?
NO → stays in cytosol
YES → guided to specific destination (Mitochondrion, nucleus, chloroplast, endoplasmic reticulum)
Main pathway = ER/Secretion
HAS signal peptide (address)
Co-translational process
Ribosome → ER WHILE protein synthesis occurs
Protein pushed through pore (translocator) into ER lumen
Protein fold inside ER → vesicles → Golgi apparatus
Golgi sorts protein to final destination (Plasma membrane/lysosome)
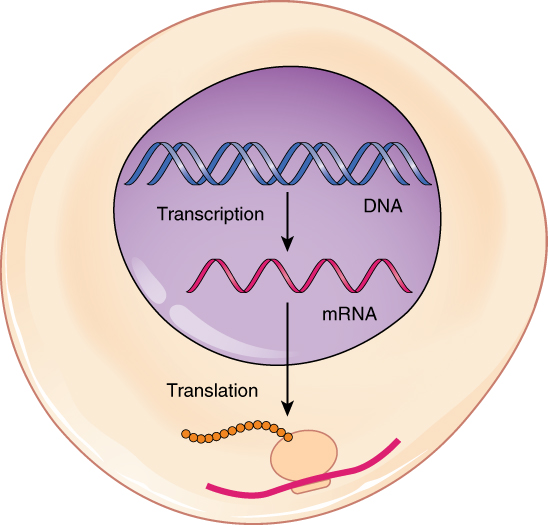
Production of cytosolic proteins
mRNA leaves nucleus
mRNA binds to cytosolic/free ribosome
Protein = translated & folds (potentially w/ chaperones/chaperonins)
Which is more complex?
Production of cytosolic proteins
Production & targeting organellar proteins
Production and targeting organellar proteins = MORE COMPLEX
Cellular topology
Structure connectivity, neighborhood relationships, & arrangement of components within cell or b/w cells in a tissue.
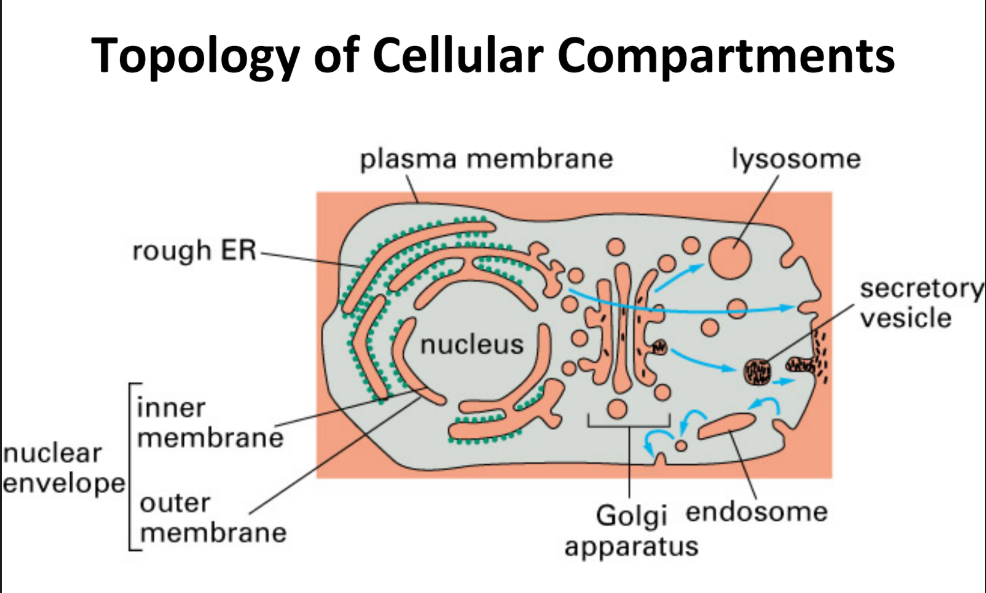
What are the 3 types of transport of proteins across organelle membranes?
Gated = nuclear ↔ cytosol
Folded protein moved through aqueous pores → compartment (topologically similar)
Transmembrane = ER transport
Folded/unfolded protein through non-aqueous transport complexes → compartment (topologically similar)
Vesicular
Protein → lumen or membrane of small vesicles (bubbles) fuse w/ compartment (topology conserved)
Topologically similar compartment = spaces w/ same connectivity & structure (connected by vesicles)
Topology conserved = stay within membranes, never cross membrane
move through budding/fusion
Preserve orientation
What are two ways that transport of proteins can occur? + Understandings
Post-translationally = after protein synthesis = complete
Co-translationally = @ same time as protein synthesis
Understandings:
Gated transport = post-translational
Transmembrane transport = post-/co-translational
Vesicular transport ONLY OCCURS AFTER transmembrane transport
Post-translational transmembrane transport = Cytosolic chaperones
Keep protein unfolded to ensure protein can pass through
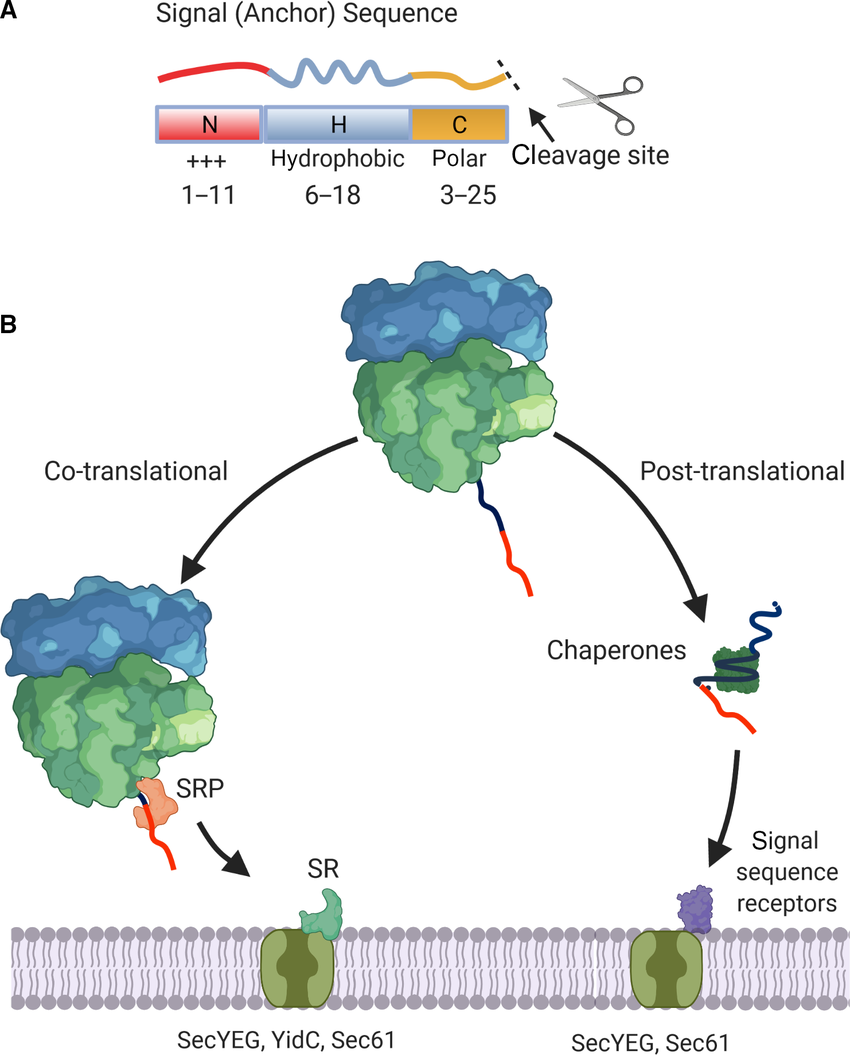
Explain how amino acid sequences on proteins work?
No label = Cytosolic protein → stays in cytoplasm
Targeting signal = short sequences of amino acids
Specific labels for specific organelle (Nucleus, mitochondria, ER)
Contiguous amino acid sequences = combinations of labels/multiple sequences
Location of label of protein depends on function of organelle protein
Recognized by other proteins (receptors) → initiate sequence → protein directed to correct place
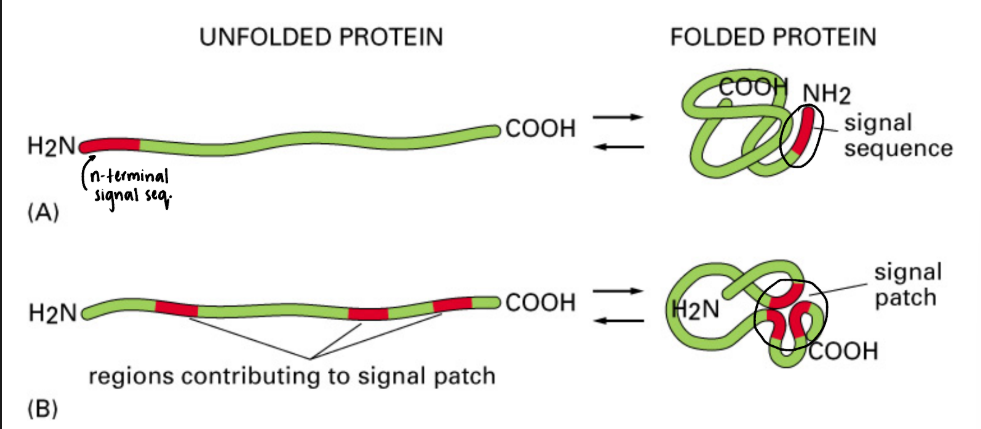
What are two forms that targeting signals can take after the protein folds?
Signal sequence: formed from n-terminal signal sequence
Signal patch: formed from regions contributing to signal patch
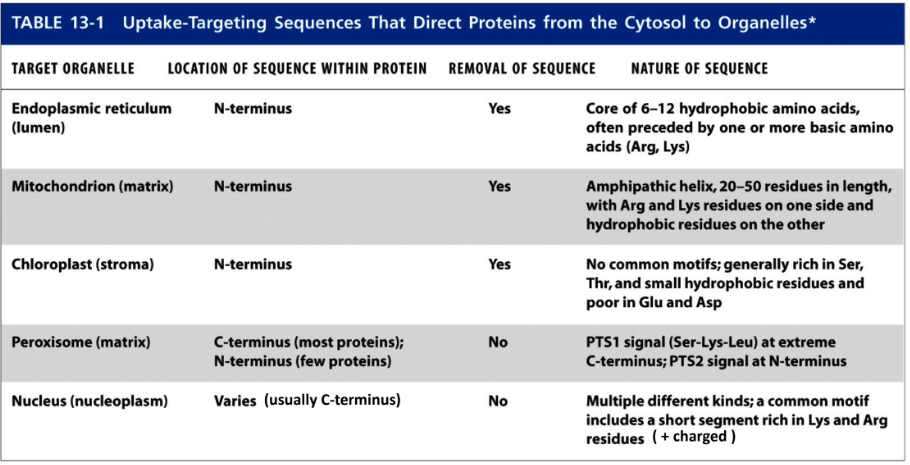
What are the 5 types of targeting signals?
Endoplasmic reticulum @ N-terminus, REMOVED
6-12 hydrophobic amino acids
Preceded by 1+ basic amino acid (Arg, Lys)
Mitochondrion (matrix) @ N-terminus, REMOVED
20-50 residues form amphipathic helix
Basic amino acid (Arg, Lys) and hydrophobic residues on opposite sides
Chloroplast (stroma) @ N-terminus, REMOVED
No common motifs
Rich in Ser, Thr, & small hydrophobic residues
Poor in Glu & Asp
Peroxisome (matrix) most @ C-terminus, few @ N-terminus NOT REMOVED
PTS1 signal @ extreme C-terminus
PTS2 signal @ N-terminus
Nucleus (nucleoplasm) usually @ C-terminus, NOT REMOVED
Multiple kinds
Short segment of Lys & Arg residues (+ charged)
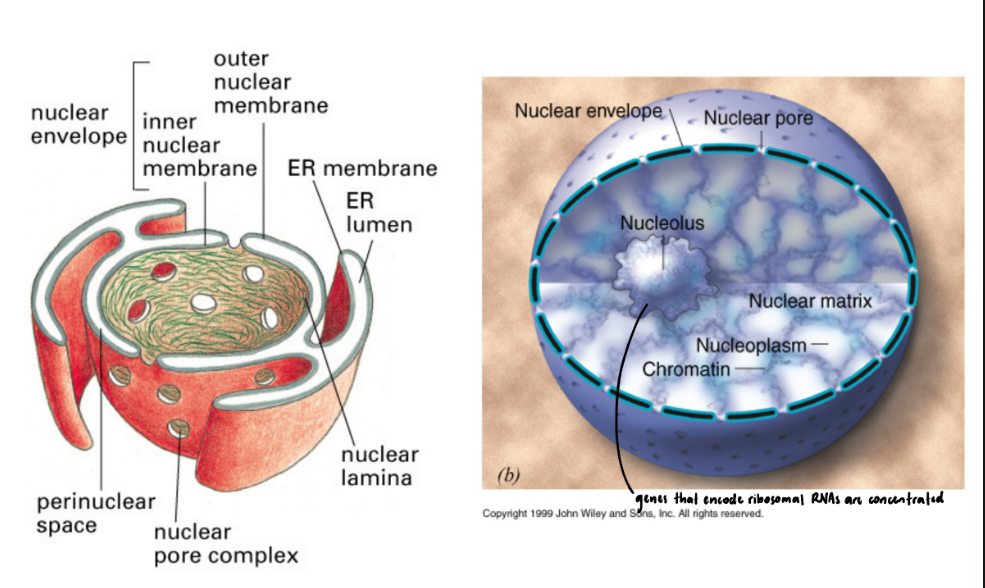
5 components of Nuclear Structure
Nuclear envelope = double membrane around nucleus
Outer membrane ↔ Rough Endoplasmic Reticulum
Space b/w membranes topologically = outside/interior of ER
Nuclear pore complex controls entry & exit
Nuclear lamina = network of intermediate filaments (Lamins) under nuclear envelope → support
Nucleoplasm = fluid in nucleus
Chromatin = DNA/protein complex
Nucleolus = DNA/protein complex hold rRNA genes for ribosomal RNA production
Where ribosomal subunits assembly
Where are nuclear proteins from?
Imported bc protein synthesis = cytoplasm
Nuclear proteins = DNA polymerase, histones, lamins, etc.
All nuclear proteins MUST be reacquired → targeting signals
Nucleus breaks down & reforms after each cell division
What happens to many RNAs produced in nucleus?
Exported
Some reimported w/ associated proteins
mRNA → nucleus → cytoplasm
Ribosomal subunits assembled in nucleolus = in nucleus → out to cytoplasm
Small nuclear RNAs (snRNAs) → cytoplasm → assembled w/ proteins → nucleus
SnRNPs = regulate mRNA splicing
Nuclear Pore Complex = controls entry & exit
Nuclear Pore Complex
Large aqueous channel/pore though double nuclear membrane
Movement = selective barrier in both directions
Structure = ring, basket, plug
50 proteins
3000-4000 per nucleus in typical cell
30x mass of ribosome
Can help cell transport ribosome through it
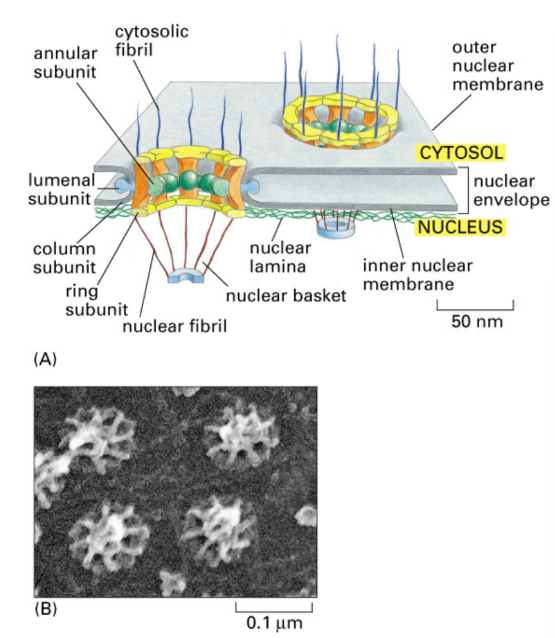
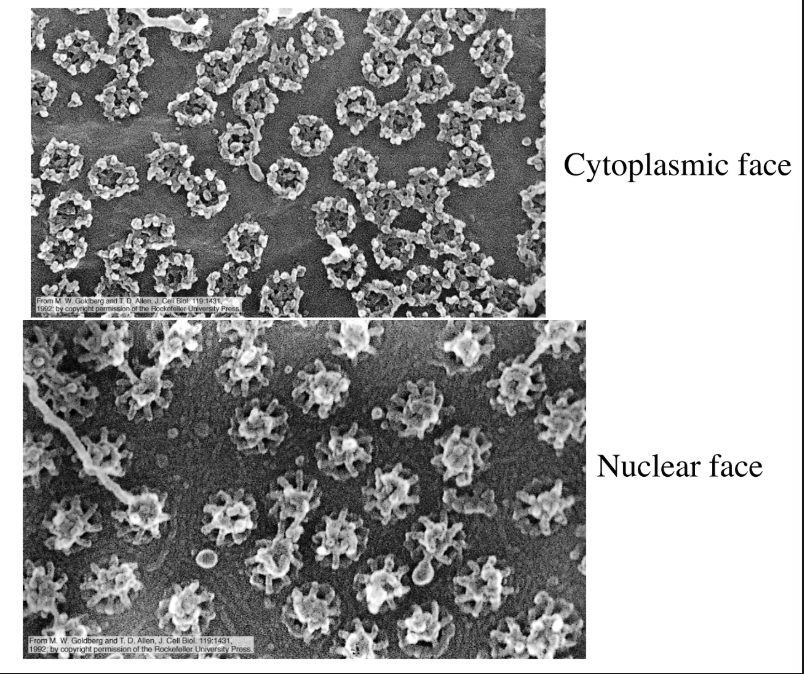
SEM of nuclear membranes
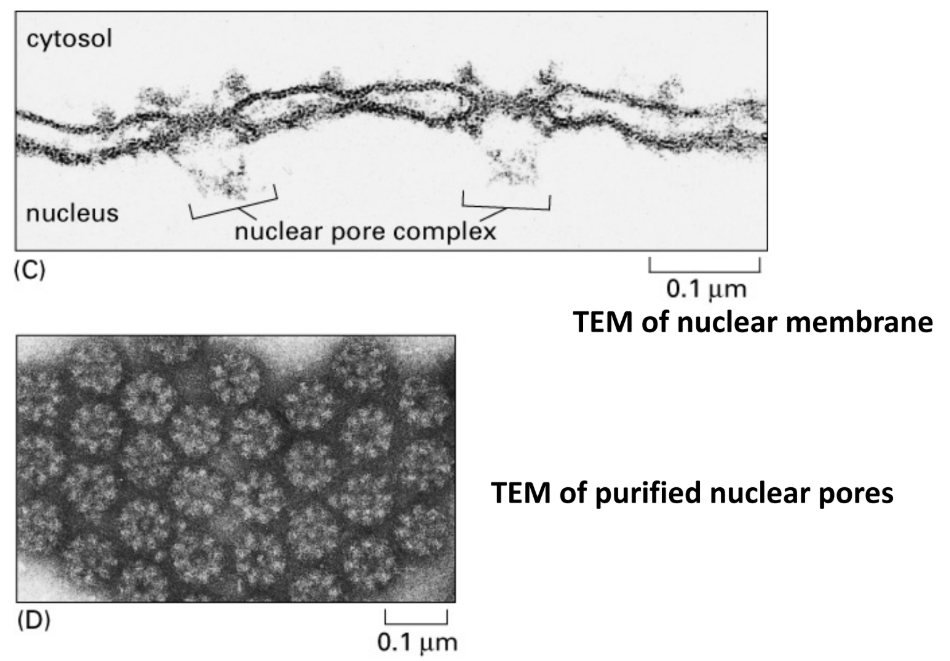
TEM of nuclear membrane
Nuclear Pore Diffusion & size of molecule
Molecule < 5000 MW (small) → passive diffusion
Molecule = large → slower passive diffusion
Molecule > 40-60,000 MW (600 amino acids) cannot freely cross
Size of average protein = 40,000 MW → active transportation
Import/export = gated transport → proteins transported(post-translation)
Nuclear transport = bidirectional
NPC selectivity + nuclear targeting signals = different proteins in cytoplasm & nucleus
What diameter of particles can diffuse passively through nuclear pores?
Particles < 9 nm = 9 × 10-6 mm
Particles > 9 nm → active transport
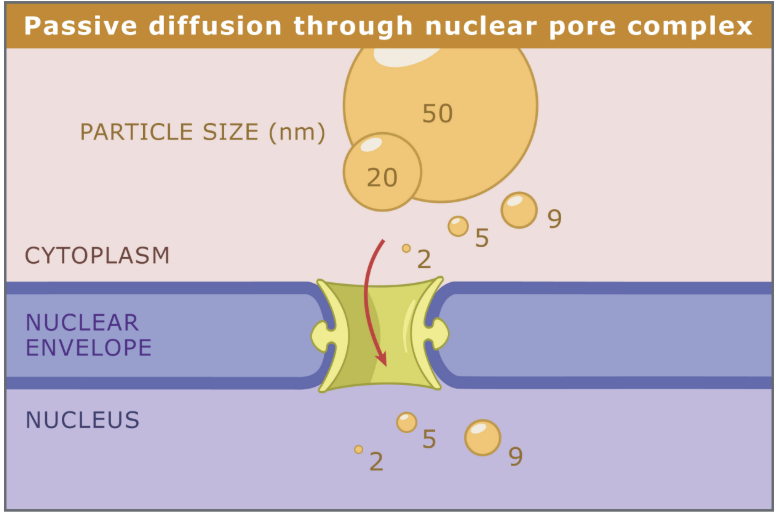
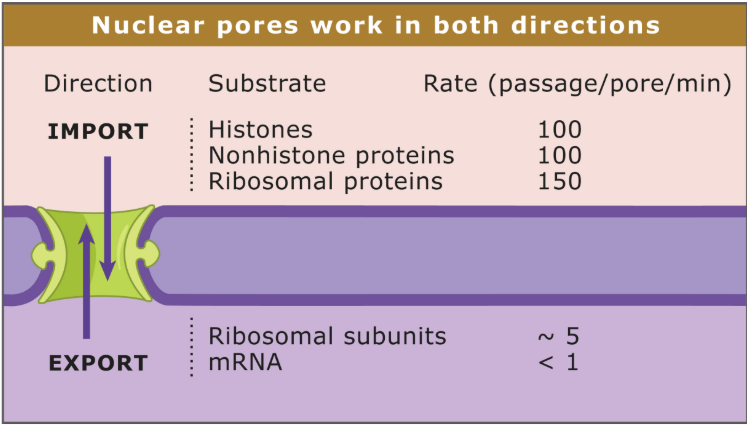
What is the direction of different substrates through the nuclear membrane?
Bidirectional
Import: Histones, nonhistone proteins, ribosomal proteins
Export: Ribosomal subunits, mRNA
Nuclear Localization Signal (NLS) + Experiment used to determine
Targeting sequences allowing for nuclear transport through the nuclear pore
Xenopus oocyte (frog egg) = LARGE CELLS (1 mm)
Inject substances into nucleus/cytoplasm → see Δ compartments
Protein = nucleoplasmin = pentameric nuclear protein
Describe steps of preparing Nucleoplasmin targeting signal + 4 types of experiments
Purify nucleoplasmin protein
(Fluorescently/radioactively) Tag protein to determine localization
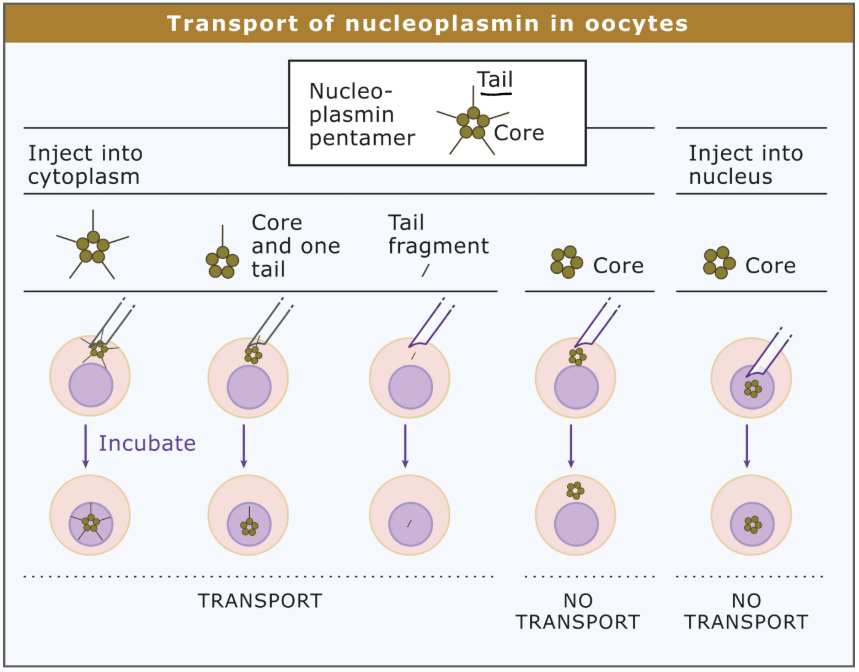
Experiment 1
Inject nucleoplasmin pentamer → oocyte (frog egg) cytoplasm
Nucleoplasmin taken up into nuclei
Experiment 2
Inject pentameric core + 1 tail
Nucleoplasmin taken up into nuclei
Experiment 3
Inject tail
Tail taken up into nucleus
Experiment 4
Inject pentameric heads into cytoplasm/nucleus
No movement
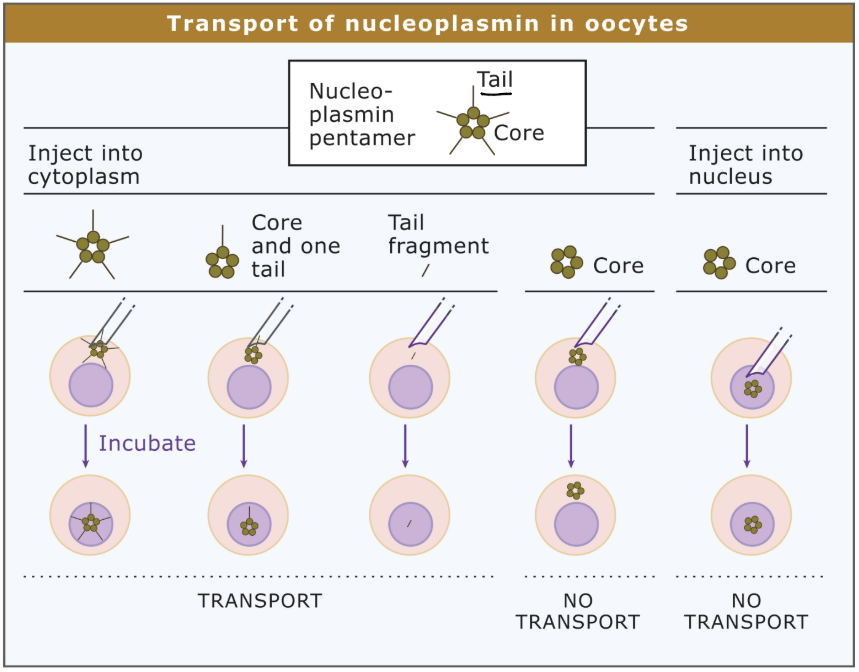
What are 3 conclusions that can be made from the nucleoplasmin injection experiments
Entire protein can be taken up into nuclei
Tail is required AND sufficient for uptake
Head is NEITHER required nor sufficient
Start cutting off amino acids @ end of tail (C-terminal) until the shortened protein no longer enters the nucleus (= NLS = REMOVED → protein no longer enters nucleus)
Defines nuclear localization signal (NLS)
What is the role of the Nuclear Localization Signal in nuclear import? + Where is it attached on the cytoplasmic protein?
NLS = necessary and sufficient for nuclear targeting
Attaches to C-TERMINUS → transported to nucleus
Necessary = NEEDED for protein → nucleus
Sufficient = ONLY SIGNAL NEEDED for protein → nucleus
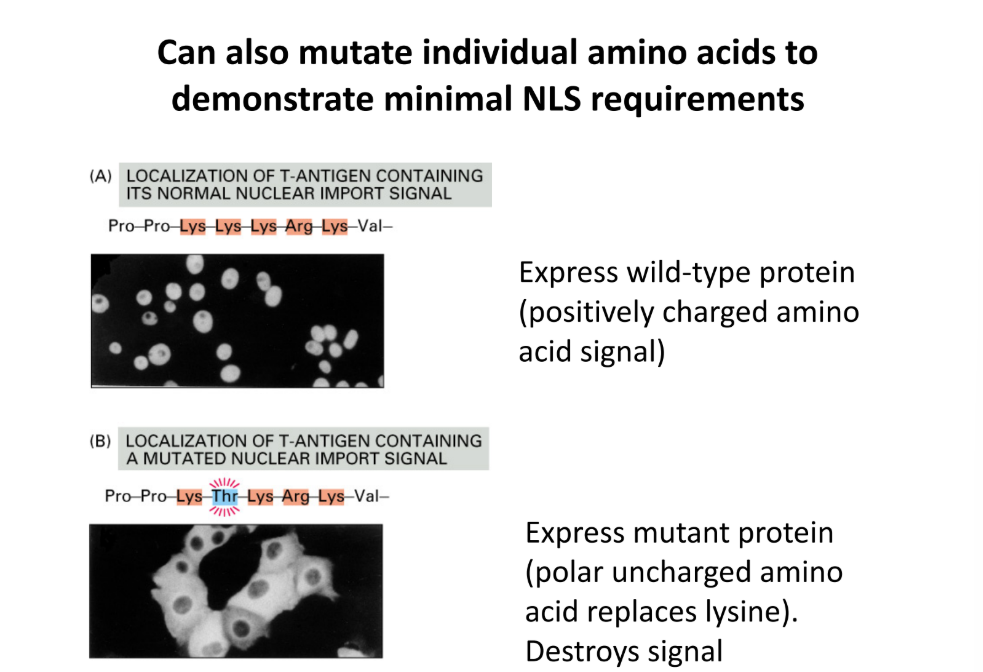
What are type of mutation in a protein can demonstrate that NLS is necessary?
Compare wild-type protein (positively charged amino acid - Lys, Arg)
Replace Lys for polar uncharged amino acid (Thr) → DESTROYS signal
Proves that NLS is necessary
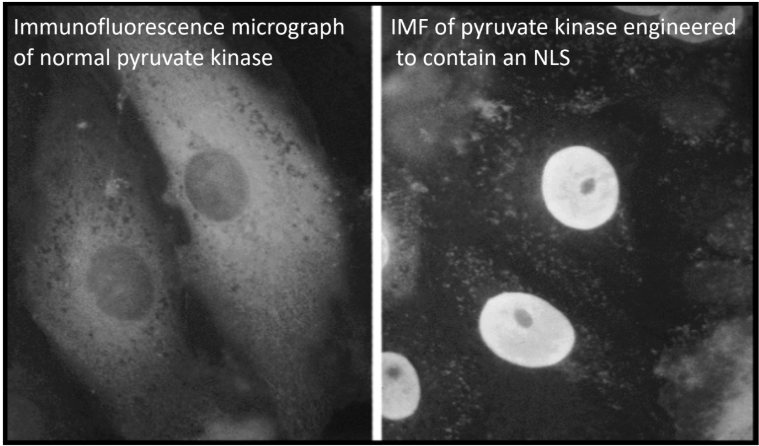
What protein can be targeted to nucleus to prove that NLS is sufficient?
Pyruvate Kinase = enzyme involved in last step of glycolysis
Summary of Nuclear Localization Signals (NLS)
Proteins imported to nucleus ALL HAVE NLS = (+) charged group of 5-6 amino acids near C-terminal end of protein
NLS sequence ≠ same for all nuclear proteins (similar, + charged)
Transport after translation (post-translational) & proteins = folded
(+) charged region of protein MUST be on surface of folded protein → interact with transport proteins/pore
Signal = necessary + sufficient → protein → nucleus
Signal recognized by transport-regulating proteins
Signal ≠ removed upon entering nucleus, stay attached, REUSED
Reused after each cell division (localization lost during cell division)
Nuclear envelope breaks down completely → proteins → cytoplasm
NLS remaining attached allows nuclear proteins to return to nucleus
Transported through aqueous pore (passive or active depending on size)
Nuclear Export Signal (NES)
Leucine-rich short amino-acid sequence that acts as a signaling directs protein to be transported to cytoplasm