microbio exam 3 (start from lec. 16)
1/246
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
247 Terms
chromosome segragation in bacteria
prokaryotes segregate new replicated DNA during the replication process
par proteins
in Bacillus sp, Vibrio sp, and Caulibacter sp
dedicated partitioning systems that segregate sister chromosomes
quinolones and fluoroquinolones
bind topo IV and DNA gyrase, inhibiting their activity (bactericidal)
w/o activity of these protenis, cell is unable to unwind DNA ahead of the replication fork
existing replication forks stall and collapse
e.g ciprofloxacin and levofloxacin
eukaryotic chromosomes
contain multiple origins of replication
replication origins are activated in what?
clusters
new units are activated at diff. times during S-phase of cell cycle
gene
segment of DNA (at a specific site in the genome) that specifies the structure of an RNA molecule
codes for a mRNA which is then translated into a protein
parts of a gene
promoter, coding sequence, termination sequence
genes are organized into what?
operons
operon
unit of genetic material that functions in a coordinated manner by means of an operator, promoter, and one or more structural genes
what does a single promoter control?
expression of multiple genes
polycistronic RNA
multiple messages
bacterial RNA polymerase
5 proteins in core complex (holoenzymes:
2 (alpha)
beta
beta’
w
doesn’t stay bound to transcriptional machinery all the time: sigma factor
2 alpha subunits
interacts w/ regulatory proteins
beta subunit
catalytic activity
add nucleotides in 5’ to 3’ direction, always needs 3’ hydroxyl group to function
beta’ subunit
involved binding and staying bound to DNA
w subunit
stabilizes the complex
sigma factor
recognizes promoter region of gene
released soon after start of transcription
can have multiple sigma factors
each sigma factor recognizes what?
a specific consensus sequence
sequence in the promoter
sequences are generally at the -35bp and -10bp upstream of the transcription start site
mutation in consensus sequences downstream can cause what? ***double check answer
decrease in transcription
mutation in consensus sequences upstream can cause what? ***double check answer
increase in transcription
steps of transcription initiation
sigma factor binds core RNA pol., forming RNA pol. holoenzyme
RNA pol. binds promoter
pol. unwinds DNA at promoter → open complex
transcription begins and sigma factor is released at 10 bp
rifampin/rifampicin
bind beta subunit of RNA pol., inhibiting pol. activity
bacteriostatic, not often used on its own
termination
pol. slows at pause site, GC-rich sequence that forms stem loop
rho factor binds to mRNA
GC rich region followed by series of U residues
GC residues form stem loop structure + RNA pol. binds stem loop structure, causing a pause in transcription
rho factor
slides along mRNA to pol.
dissociates pol., mRNA off of RNA
differences btwn rho-dependent + rho-independent termination?
***** ask
which type of RNA is the least stable? (shortest half-life?)
mRNA
mRNA
encodes protein (transcription of DNA code)
rRNA
synthesizes protein
tRNA
shuttles amino acids
RNA
transcription of the DNA code
things common to both types of mRNA
5’ untranslated section, 3’ untranslated section
ribosome binding site **** (ask: does eukaryotic mRNA have a ribosome binding site?)
translation start site (start codon)
translation stop site (stop codon)
what do eukaryotes have that bacterial mRNA doesn’t?
eukaryotes cap their mRNA at 5’ (methyl guanosine cap) and 3’ (poly adenosine tail)
what is the function of a methyl guanosine cap and poly A tail?
allows a eukaryotic to assess whether an mRNA is intact before export from the nucleus and translation by the ribosome
stop codons
UUAA, UAG, UGA
wobble position
degenerate nature of the codons due to a different third base position and the same first two bases
how many codons code for 20 amino acids?
61 but 64 possible codons (3 of them are for stop codons)
start codon
AUG
how is the sequence of nucleotides in an mRNA molecule read?
5’ to 3’
a signal at the beginning of each mRNA sets the correct reading frame
do mammalian or bacterial cells have more types of different tRNA?
mammalian
anticodon loop will base pair (interact) w/ what?
complimentary mRNA codon
each tRNA, except those complimentary to the stop codon, is what?
“charged” w/ an amino acid at acceptor end (phenylalanine)
are there tRNAs for stop codons?
no
tRNA charging
carried out by a specific aminoacyl-tRNA synthetase
the enzyme recognizes tRNA structure in the D-Loop and Anti-codon stem of tRNA and links acceptor end of tRNA to the appropriate aa
charged tRNA
has amino acid
uncharged tRNA
doesn’t have amino acid
multiple aminoacyl tRNA synthetases are required to what?
charge the 20 diff. aa’s to the appropriate tRNAs
ribosomes
protein machine responsible for protein synthesis (known as protein polymerase)
2 subunits, 30S and 50S
what does the S in 30S and 50S stand for?
Svedberg units: measure of sedimentation rate
30S (small) and 50S (large) ribosome subunits
52 proteins, 3 rRNAs (mostly rRNA)
can bind 1 mRNA + 3 tRNAs
what do ribosomes facilitate?
proper reading of code
creation of a peptide bond btwn adjacent aa’s
dehydration synthesis
chemical reaction facilitated by a ribosome
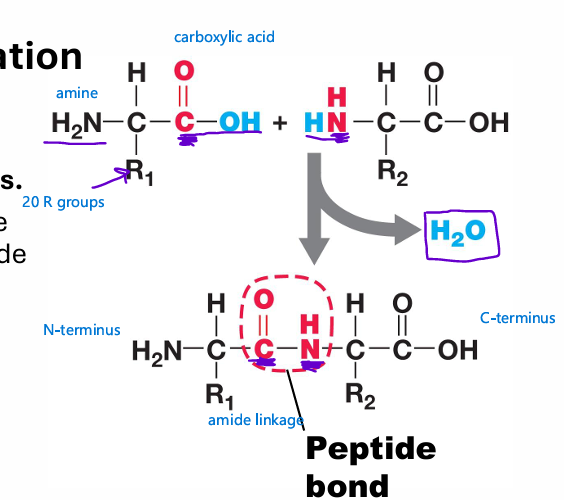
peptide bond
bond linking the aa’s
nascent proteins
peptides or polypeptides
transcription creates what?
mRNA → multiple mRNAs made from a single gene
bacterial ribosome can readily assemble on start codons in interior of mRNA
bacterial mRNAs are often polycistronic. what does this mean?
encode several diff. proteins translated from same mRNA molecule
ribosomes bind mRNA while what? (in prokayotes)
while mRNA is still being created
multiple proteins made rapidly from each mRNA
advantage of not having a nucleus
peptide bond formation
translation of RNA to protein
initiation
elongation
termination
initiation
initiation factors (IF’s) bind ribosome to ribosome binding site (upstream of start site), known as Shine Delgarno Sequence
translation begins at AUG
elongation
polymerization, movement of ribosome along mRNA
EF-Ts, EF-Tu, EF-G bring GTP energy
termination - stop codons
releasing factors undock ribosome from mRNA
in-depth translation initiation
30s subunit binds mRNA (spectinomycin reversibly interferes w/ mRNA interaction w/ 30S ribosome)
IF2 interacts w/ initiator tRNA
fMet-tRNA binds to start codon
association of initiator tRNA w/ 30S mRNA releases IF3 and allows IF1 to bind
50S subunit enters complex, 50S binding to 30S complex and GTP hydrolysis releases IF1 and IF2
in-depth translation elongation
EF-Tu-GTP binds to tRNA and guides it to A site
once A site is filled, peptidyltransferase activity makes peptide bond btwn amino acid or peptide in P site and aa in A site
formation of peptide bond results in transfer of aa or peptide from tRNA in P site to tRNA in A site
EF-G-GTP complex binds to ribosome, causing 50S subunit to advance one codon. tRNA in A site moves into P site (translocation)
aminoglycoside antibiotics like kanamycin, gentamycin, streptomycin
thought to bind the initiation complex (30s-mRNA-tRNA) and block construction of complete ribosome
spectinomycin
reversibly interferes w/ mRNA interaction w/ 30S ribosome
tetracycline
binds the 30s subunit and inhibits binding of amino-acyl tRNA to the A-site of the ribosome
chloramphenicol
binds the 50s subunit and inhibits the peptidyltransferase activity
in-depth translation termination
uncharged tRNA leaves ribosome
stop codon on mRNA enters A site. since there is no corresponding tRNA, protein release factor (RF1 or 2) enters the site
peptidyltransferase is activated and releases completed protein from tRNA in P site
RF3 enters and ejects RF1 or RF2
ribosome recycling factor (RRF) and EF-G enter A site. GTP hydrolysis undocks 50S from 30S
IF3 enters 30S subunit to remove uncharged tRNA and mRNA
primary structure of a protein
unique sequence of amino acids
ribosome generates primary structure
secondary structure of a protein
found in most proteins, consists of coils and folds in the polypeptide chain
hydrogen bonding of polypeptide backbone
composed of beta pleated sheet and alpha helix
tertiary structure of a protein (3)
determined by interactions among various side chains (R groups → 3D)
covalent disulfide bond
quaternary structure of a protein
a protein consists of multiple polypeptide chains
alt. proteins
have 1,2, and 3 prime
only multisubunit have 4 prime
operons are organized into what?
regulons
regulon
system of genes, formed by one or more operons, that have a common regulatory element
common transcription factor protein to promote expression
how are genes in prokaryotes designated?
in italics w/ the first letter lowercase and NOT capitalized
e.g lacZ - refers to the gene or DNA sequence
how are proteins in prokaryotes designated?
capitalizing first letter of name and NOT using italics
e.g LacZ - refers to the protein encoded by the lacZ gene
why do cells not express every gene at maximal level under all conditions?
physical space limitations
energy and resources conservation
contradictory functions
most proteins are enzymes that carry out what?
biochemical reactions
constitutive proteins
needed at the same level all the time
microbial genomes encode many proteins that are what?
that are NOT needed all of the time
how does regulation of protein production and function help?
conserves cell’s energy and resources
post-translational regulation
regulation of gene expression
post-translation regulation
controls the activity of preexisting enzymes
a protein is modified to change its activity
very rapid process (seconds)
gene expression
transcription of gene into mRNA followed by translation of mRNA into protein
regulation of gene expression
controls amount of an enzyme
generally occurs at level of transcription by controlling the amnt of mRNA
sometimes can occur at translation level by controlling whether an mRNA is translated
slower process (minutes)
where does a DNA-binding domain fit in?
major grooves and along sugar-phosphate backbone
how do most DNA-binding proteins interact w/ DNA?
in a sequence-specific manner
interactions are btwn aa side chains and chemical groups on bases and sugar-phosphate backbone of DNA
what is the main site of protein binding?
major groove of DNA
what is frequently the binding site for regulatory proteins?
inverted repeats
what influences the binding of regulatory proteins to DNA?
small molecules
homodimeric proteins
proteins composed of two identical polypeptides
regulation of transcription typically requires what?
proteins that can bind to DNA
proteins that bind DNA can alter what?
transcription rates
binding event can block transcription (neg. regulation)
binding event can activate transcription (pos. regulation)
negative transcriptional regulation
native state of RNA allows RNA pol complex to be recruited, and transcription takes place
if repressor protein (downstream of promoter) binds to DNA, it inhibits recruitment of RNA pol, and transcription does not occur
positive transcriptional regulation
RNA pol can bind to promoter only if an activator protein binds to a site near the promoter
activator binding site is upstream of promoter
if activator does not bind to DNA, RNA pol cannot bind and transcription does not occur
negative regulator/repressor
bind to regulatory sequences in DNA and prevent transcription of target genes
most often repressors block what from binding promoter?
sigma factor of RNA polymerase
bind DNA at sequence called operator sequence
repressors can require what to repress or be released from binding site?
co-factors (ligand)
co-repressors
cofactors required for repressor binding
inducers
cofactors that relieve repression