bio unit 7
1/92
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced |
|---|
No study sessions yet.
93 Terms
nucleic acid elements (5)
C, H, O, N, P
nucleic acid monomer
nucleotide
nucleic acid polymer
polynucleotide
what bond connects nucleic acid monomers
phosphodiester bonds (covalent)
nucleic acid examples (3)
DNA, RNA, ATP (modified)
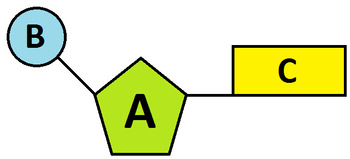
label the parts
A - ribose sugar
B - phosphate group
C - nitrogenous base (A, T, U, G, C)
what differentiates RNA from DNA structurally
RNA: OH at 2nd carbon
DNA: H at 2nd carbon
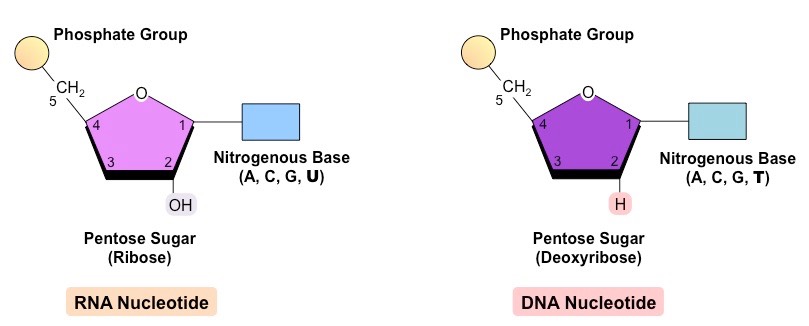
where are phosphodiester bonds created (2)
between the 3’ carbon on the sugar of one nucleotide and the phosphate group of the next nucleotide
created by condensation reactions
nitrogenous bases (5)
adenine, cytosine, guanine, thymine, uracil
purine bases (2)
adenine and guanine
pyrimidines bases (3)
uracil, cytosine, thymine
what keeps DNA double helix the same width
purine-pyrimidine pairs
A-T base pairs have ___ hydrogen bonds
two
G-C base pairs have ___ hydrogen bonds
three
DNA replication purpose (3)
process of copying the genome within a cell
creates two identical DNA molecules each with 2 complementary strands
needed for cell division (each cell needs it’s own copy)
meselson and stahl experiment significance (2)
determined how DNA is replicated
replication is semi conservative
meselson and stahl experiment process (4)
1) grew ecoli in N15 (heavy)
2) switched the ecoli to N14 (light)
3) round 1 replication: half heavy half light → rules out conservative
4) round 2 replication: half heavy half light, then half light → rules out dispersive
initiation of DNA replication (4)
helicase binds to the origin of replication and unzips the double helix by breaking the hydrogen bonds
creates replication fork (region where original DNA splits into two)
single strand binding proteins bind to the single stranded DNA to keep the strands separate by preventing the bonds from reforming
gyrase/topoisomerase moves in front of the helicase to relieve tension that could damage the DNA
DNA replication synthesizing complementary strand (4)
DNA polymerase III reads the template to build the complementary strand in 5’ → 3’ direction
primase creates an existing strand for DNA polymerase III to add to the 3’end by adding a primer made of a short sequence of RNA nucleotides
after the primer, DNA polymerase III can attach to the next strand
DNA proofreading (2)
can proofread as the new strand is being built (can remove incorrect ones)
not 100% accurate → mistakes are mutations for natural selection
how are primers removed
DNA polymerase I removes the RNA nucleotides and replaces them with the correct DNA nucleotides
leading vs lagging strands (3)
leading needs 1 primer, each Okazaki fragment needs a primer for lagging
leading is synthesized continuously, lagging is synthesized discontinously
leading goes in the same direction as helicase, lagging goes in the opposite direction
which strand has more primase and DNA polymerase I activity
lagging
ligase
catalyzes formation of phosphodiester bonds between the Okazaki fragments forming a continuous bond
polymerase chain reaction (PCR) (2)
used to amplify (create millions/billions of copies) small fragments of DNA
basically DNA replication in a test tube
taq polymerase (2)
DNA polymerase enzyme that’s heat stable
originally found in a prokaryote that lives in hot springs
PCR steps (5)
setup: many free DNA nucleotides, primers, and taq polymerase in test tube
1) denaturation: DNA is heated to break the hydrogen bonds between the strands
2) annealing: sample is cooled to allow primers to bind to the complementary DNA strand
3) extension: at room temperature, taq polymerase replicates the DNA
process is repeated 100 times or so
gel electrophoresis (3)
often done after PCR
uses an electrical current to move DNA fragments through a gel
fragments are separated based on size
restriction enzymes (2)
typically DNA molecules are too long to travel through a gel so they need to be cut into pieces first
restriction enzymes (endonuclease) will cut DNA molecules at very specific sequences
DNA fingerprints
restriction enzymes cut-sites create a unique pattern of bands when a sample is run through a gel “DNA Fingerprint”
SNPS can change cut sites which causes no enzyme activity at the location → changes the bonding pattern
PCR and gel electrophoresis applications
PCR COVID-19 testing, paternity testing, forensic investigations
the central dogma (3)
describes the flow of genetic information from DNA - RNA - protein
DNA is transcribed into mRNA
mRNA is transcribed into a polypeptide chain
transcription (2)
producing mRNA using DNA as a template
allows for only a portion of the genome to be copied (resource efficiency) and DNA to remain protected in the nucleus
where does transcription occur
eukaryotes: nucleus, prokaryotes: cytoplasm
RNA polymerase (2)
performs transcription (elongating mRNA strand) using DNA template as a guide
must synthesize mRNA in 5’ → 3’ direction
phases of transcription (3)
initation, elongation, termination
transcription initation (2)
promoter: non-coding region of DNA in front of the gene of interest that begins with the TATA box
transcription factors (proteins that bind to the promoter) recruit RNA polymerase — begins to temporarily unzip a small section of the double helix to expose the bases
transcription elongation (3)
RNA polymerase “reads” the template strand (antisense strand) to synthesize the mRNA in 5’ → 3’ direction
as its synthesized, the RNA nucleotides will temporarily hydrogen bond with the template strand
the growing mRNA strand exits RNA polymerase and the DNA rezips
template strand
antisense stand, RNA polymerase reads it
coding strand
sense strand, complementary DNA strand
transcription termination
signals for RNA polymerase to release the mRNA and detaches from the DNA → transcription now finished
regulation of transcription (4)
noncoding regions of DNA that don’t code for a protein
some parts help with regulation
enhancers: increases rate of transcription
silencers: decrease rate of transcription
non-coding DNA regions (3)
telomers: repetitive sequences at the end of linear (eukaryotic) chromosomes → protects the ends of the chromosomes
genes for rRNA and tRNA: RNA is synthesized from these genes, but they don’t code for proteins
introns: base sequences that get removed from the mRNA after transcription (only in eukaryotes)
mRNA processing (3)
done in eukaryotic cells after transcription and before it leaves the nucleus
converts pre-mRNA into mature mRNA
includes mRNA splicing and addition for the 5’ cap and poly-A tail
5’ cap (3)
modified nucleotide that is added to the 5’ end of mRNA
helps with ribosome binding during translation
aids in the export of the mature RNA from the nucleus and protects the mRNA from degradation in the cytoplasm
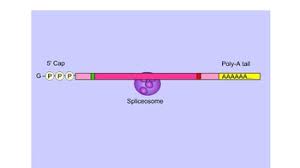
poly-A tail (2)
string of adenines attached to the 3’ end of mRNA
aids in the export of the mature of mRNA from the nucleus and protects the mRNA from degradation in the cytoplasm
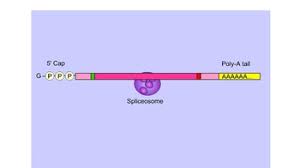
mRNA splicing (4)
pre-mRNA has exons (expressed base sequences) and introns
introns are removed from the mRNA during splicing (stays in the nucleus)
snRNPs bind to either side of the introns and then assemble into spliceosomes
spliceosomes remove the introns and ligate the exons together
snRNPs
small nuclear ribonucleoproteins
alternative splicing (2)
different introns are removed creating unique mature mRNAs — creating unique polypeptides
one gene (and so one pre-mRNA) can provide instructions for several different polypeptides due to alternative splicing
elements in proteins (4)
C, H, O, N
proteins monomer
amino acid
how many amino acids to proteins have
20 bc 20 different side chains
proteins polymer
polypeptide
protein bond
peptide bond (covalent)
protein structure levels (4)
1o: polypeptide chain
2o: local folding (alpha helics and beta pleated sheets)
3o: 3D structure determined by side chains
4o: 2+ polypeptide chains interacting
protein synthesis in prokaryotes (2)
translation can occur immediately after transcription
causes protein synthesis to be faster than in eukaryotes
protein synthesis in eukaryotes (3)
transcription occurs in the nucleus
mRNA processing occurs before the mature mRNA can leave the nucleus
transcription occurs in the cytoplasm either by free or attached (on the rough ER) ribosomes
codon (2)
mRNA is ready in triplets of bases
each codon codes for a specific amino acid
start codon
AUG
stop codons (3)
UGA, UAA, UAG
characteristics of genetic code (3)
universal - basis of several biotechnology techniques (ex GMOs)
redundant/degenerate - some condones code for the same AA
unambiguous - no codon specifies more than one AA
implications of universal genetic code (2)
allows for genetically modified organisms
splice the gene(s) of interest into another organism - allows for protein synthesis of the gene(s) of interest by the new organism
uses of genetically modified organisms (2)
insulin gene in bacteria → allows for mass production of the insulin protein
pesticide gene in crops → allows for the crops to produce a protein that acts as a pesticide so bugs don’t eat them
genetic modification process (5)
A - isolate gene of interest and amplify it using PCR
B- isolate bacterial plasmid and amplify it using PCR
C - use a restriction enzymes to cut the gene of interest and then insert it into plasmid (creating a recombination plasmid)
D - insert recombinant plasmid into host organism: creating a transgenic organism
E - (optional) if the goal was to create/harvest a protein, allow the transgenic organism to grow in culture, then extract the protein of interest
types of RNA (3)
mRNA: messenger - takes the message from DNA and brings it to the ribosomes
tRNA: transfer - carries amino acids to the ribosomes
rRNA: ribosomal - structural component of ribosomes
tRNA structure (5)
single strand folded into 3D structure
2D representation resembles clover
held together by hydrogen bonds
anticodon: group of 3 bases in the tRNA that will bind to the mRNA codon
amino acid binding site: where the amino acid gets attached
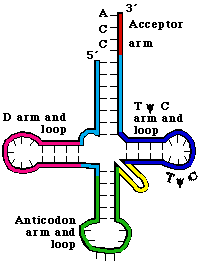
tRNA enzyme and function (3)
aminoacyl-tRNA synthetase
attaches the correct amino acid to the tRNA
the tRNA brings the amino acid to the ribosome and binds to the mRNA codon
ribosome structure (4)
made of rRNA and proteins
2 subunits: small and large
mRNA attaches to the small subunit
large subunit has 3 binding sites for tRNA (A, P, and E sites)
ribosome APE binding sites (3)
A site: aminoacyl-tRNA binding site (for incoming tRNA)
P site: peptidyl-tRNA binding site (for tRNA “holding” the growing polypeptide chain)
E site: exit site (for discharging/”empty” tRNA to leave)
ribosomes facilitate binding of ______________ with _______ AND the formation of the __________________ between amino acids and the peptide chain
mRNA codon, tRNA, peptide bond
RNA translation initiation (4)
5’ end of the mRNA binds to the small ribosomal subunit
subunit moves from 5’ to 3’ and scans for the start codon
at the start codon, the initiator tRNA binds to the start codon
large ribosomal subunit assembles, placing the initator tRNA into the P site
RNA translation elongation (2)
the ribosomes begins to read the mRNA in codons
cycle repeats until the stop codon is reached
RNA translation elongation codon recognition (2)
incoming tRNA’s anticodon binds to the mRNA’s codon in the A site
ensures that the correct amino acid is added
RNA translation elongation peptide bond formation (2)
peptide bond is formed between the polypeptide chain and the new amino acid
transfers the chain from the P site to the A site
RNA translation elongation translocation (3)
ribosome shifts over one codon
discharged tRNA moves to the E site and exits, tRNA in the A site moves into the P site
the A site is now empty for the next tRNA
translation termination (4)
when a stop codon is in the A site, no new tRNA will enter
release factor will enter and perform a hydrolysis reaction to break the bond between polypeptide chain and tRNA in the P site
after the polypeptide chain is released, the translation complex dissolves
the polypeptide chain will then be modified and folded into its final structure/functional form
RNA post-translation modifications (2)
often occurs in the golgi
can involve adding chemical groups (phosphates or sugars) or cleaning specific peptide bonds
pre-proinsulin to prosulin
in the RER, the signal peptide is removed → proinsulin
pre-proinsulin (2)
110 amino acid length
4 sections (signal peptide, A chain, B chain, C-peptide)
proinsulin to insulin (3)
in the RER: form disulfide bridges between the A chain and B chain
the bridges are packaged into vesicles to move to the golgi
in the golgi: removed C-peptide → insulin
preteomes (3)
the total of all proteins made and used by the body
dynamic: constantly synthesizing and hydrolysing proteins
done by proteasomes
proteasomes
protein complex that hydrolyzes damaged or unused proteins
cell differentiation/specialization (3)
all cells in the body have the same DNA
cells have differentiated into different types (ex muscle cells, neurons, heart cells, skin cells, etc.)
different genes are expressed (proteins produced) in those different cell types → one major thing making them different
mutations (2)
structural changes or alterations in the DNA sequence of an organism
have zero impact, decrease, or increase fitness (adaptation)
types of mutations (4)
substitutions (point mutations): single nucleotide is changed (SNPs)
synonymous mutation (silent) amino acid stays the same
non-synonymous mutation: amino acid is changed
insertions or deletions: one or more nucleotides are added/removed → results in a frameshift mutation where the reading frame is altered, drastic impact bc all codons are changed
synonymous mutations _______ impact the polypeptide while nonsynonymous mutations _______ impact the polypeptide
do not, can
causes of mutations (2)
errors in DNA replication/repair (DNA polymerase III or breakdown in repair process)
exposure to mutagens (chemical mutagens and radiation)
which base is more likely to be mutated
cytosine (into uracil)
consequences of mutations in somatic cells
causes issues to the individual but is not passed on to offspring
somatic cells
all the cells in the body excluding germ cells
germ cells
cells that produce sperm and eggs
consequences of mutations in germ cells
can be passed on and inherited
impact of mutations on fitness (3)
neutral/silent mutation: doesn’t change fitness (ex synonymous or non-coding regions)
harmful mutation: reduces fitness and selected against (ex causes disease)
beneficial mutation: increases fitness and selected for (adaptation)