15. Operons
1/45
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced |
|---|
No study sessions yet.
46 Terms
trp operon is a classic example of operon
negatively regulated by a repressor
trp is a biosynthetic (anabolic) operon
synthesis of tryptophan when needed
repressor protein is aporepressor - not active in absence of corepressor
five structural genes code for enzymes responsible for synthesizing L-tryptophan transcribed from
a single promoter negatively regulated by the TrpR aporepressor
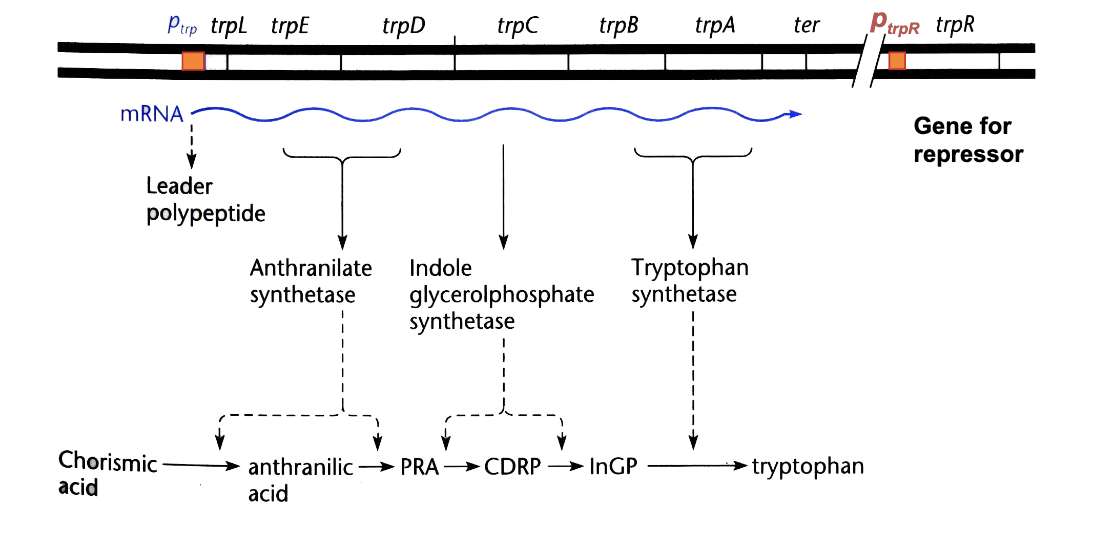
when Trp is needed, has to be synthesized, to do so:
promoter available to RNAP structural genes are transcribed
aporepressor cannot bind to operator site
OUTCOME: structural genes TRANSCRIBED
when trp is not needed anymore because there is enough of it, synthesis has to stop
tryptophan (corepressor) binds to aporepressor dimer to form functional repressor dimer
OUTCOME: autoregulation repressor dimer inhibits transcription of trp operon and transcription of the TrpR gene
active repressor + inducer (lactose)
inactive repressor
OUTCOME: genes transcribed (hence inducing transcription)
aporepressor + corepressor (trp)
= active repressor
OUTCOME: no genes transcribed
L-ara operon was the first example of
positive regulation in bacteria
L-ara operon summary
utilization of 5-carbon sugar L-arabinose - catabolism
three structural genes (araA, araB, araD) transcribed from a single promoter PBAD
two operators araO1 and araO2 - repression
activated through cAMP-CAP as well
AraC regulator - repressor
binds at a one set of cis elements upstream of promoter
Arac regulator - activator upon binding of arabinose
binds at a different set of cis elements upstream from promoter
when no arabinose present
araC binds to araO2 and araI1, bends DNA and hides araPbad promoter from RNA polymerase
OUTCOME: no synthesis
when arabinose present
arabinose binds to araC, this changes the conformation of the protein, so arabinose/araC now binds to araI1, and araI2 and not to araO2
OUTCOME: araPbad promoter is available to RNAP
ara operon also subject to activation by cAMP - CAP
too much arac protein (autoregulation)
binds to araO1 → prevents furhter transcription - RNAP binds to araPc
allosteric change in regulatory molecules as a result of small molecule binding is not the only way in which regulator proteins are controlled
OxyR (transcriptional activator of genes induced by H2O2) is directly activated by oxidation
phosphorylation of regulatory proteins is a common mechanism
multiple operators - auxiliary operators in prokaryotes
strongest - most commonly shown O1
downstream O2
upstream O3
if all three O1,2,3 are occupied - trancription suppressed 1000-fold
if either O2 or O3 are open - 500 fold
both O2 and O3 open (O1 only suppressed) - 20 fold
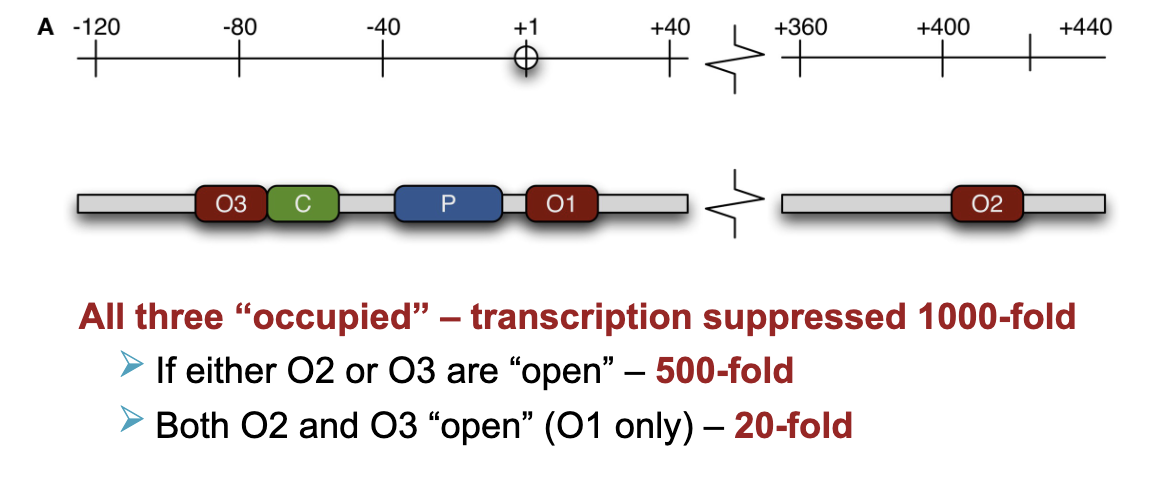
repressor binds as a tetramer - could bind to all four of them (prokaryotes)
DNA looping - RNAP prevented from binding to promoter
presence of auxiliary operators (O2 and O3) near the functional operator O2 increases the local concentration of the repressor, so that it can occupy the functional operator O1
if a molecule cannot be transported across the membrane
two component regulatory system
one protein is the SENSOR-TRANSMITTER protein
monitors specific changes int he environment (level of nutrients. pH, solvent concentration-osmolarity etc) kinase
second protein is RESPONSE regulator protein
either stimulates or represses regulation of specific genes
changes in gene expression necessary for bacterium to adapt to environmental change
sensor-transmitter protein spans across the cell membrane
snesor is an outer part which detects specific changes in environment
transmitter is an inner part which usually acts as a kinase
changes in the environment - detected by sensor causing change of conformation of sensor domain causing
activates transmitter’s kinase activity in the cell
activated kinase - autophosphorylation of the transmitter domain (transfers usually gamma phosphate from atp to itself)
change of conformation of transmitter domain
phosphate is transferred to receiver domain of response regulator → change of conformation - activation of the effector domain
response regulator binds to DNA regulator sequences
transcription factor responsive to environmetnal change
direct contact between RNAP and regulator protein could
happen even when binding sites are far apart
RNAP - NtrB/NtrC nitrogen regulation system
RNAP binds promoter - closed complex + omega 54 factor
signal for low nitrogen/glutamine
triggers two-component regulator system
NtrB = sensor kinase → phosphorylates NtrC (the activator protein)
phophorylation changes NtrC to its active form
NtrC binds to enhancers far upstream (-140 and -108) from promoter
DNA bends so NtrC can physically contact RNAP at the promoter
The ATPase activity of NtrC stimulates the polymerase to unwind the template and form open complex
RNA polymerase initiates transcription at a unique site promotoer - asymmetrical
RNAP is positioned so it can transcribe only one strand from one promoter
regulatory proteins and RNA polymerase work together to regulate transcription initiation
distribution and actual nucleotide sequence of binding sites for regulatory proteins
Binding sites
usually not perfect inverted repeats - proteins bind as dimers
two basic methods that have led to understanding of transcriptional initiation and control in bacteria
gel shift or electrophoretic mobility shift assay (EMSA)
DNase I/DMS footprinting
principle of gel shift assays
electrophoretic mobility of naked DNA fragment and DNA fragment carrying bound proteins will differ, need proteins to retain 3D structure to bind to DNA
DNase I footprinting
DNase I cleaves DNA randomly, but not where a protein is bound - that bound region is “protected” and leaves a blank space on a gel
the missing bands in the protein lane indicate the exact nucleotides protected by the protein
NOTE: DNA is run on Polyacrylamide gel finer resolution = sequencing gels
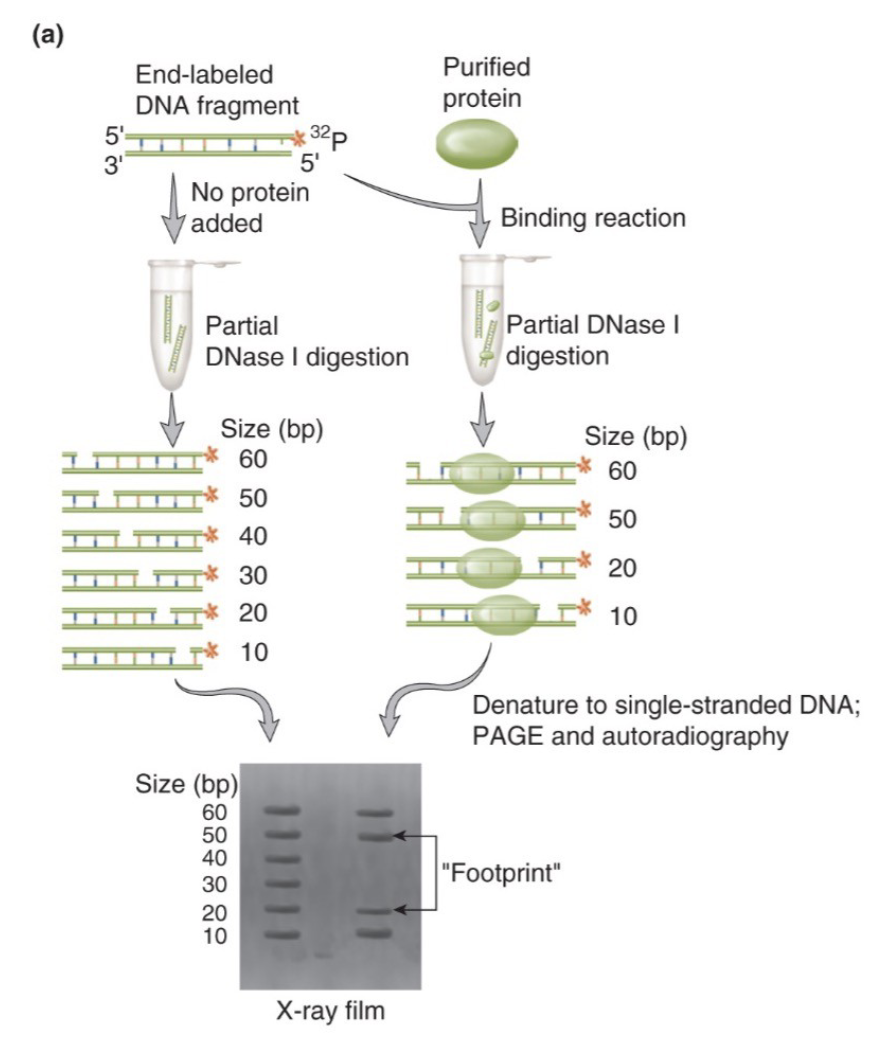
DNA foot printing
part of DNA carrying protein will be protected from modification
→ carry out modification (limited DNAse digestion)
→ DNA within the protein’s footprint will be unaffected (protected)
Domains
protein primary, secondary and tertiary structures allow amino acid side chains to form regions that “read” DNA sequence
Domains recognize hydrogen bond
acceptors (oxygen and nitrogen ions) and donors (hydrogen bound to acceptor)
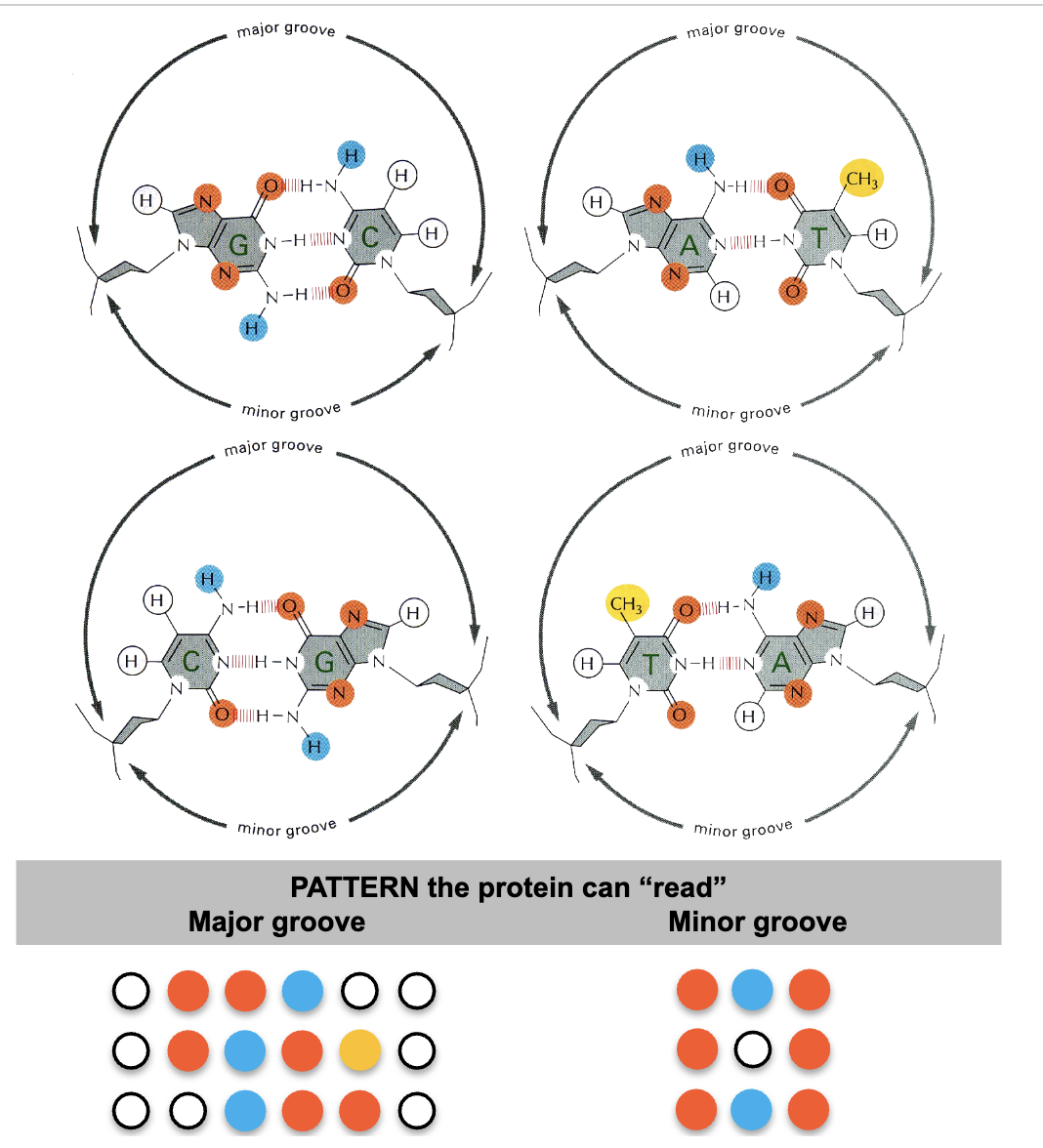
proteins have to recognize base pairs while
DNA is double strandedn
usually 10-20 contacts
specific H-bond donors and acceptors on protein and DNA complement each other
DNA and protein interxn not optimum simply due to sequence recognition by regulatory proteins
maximum protein to DNA contact when DNA is distorted
these proteins can bind and ‘bend” DNA: toward the bound dimer or away from the bound dimer
e.coli domains
the tertiary structure of large proteins is organized in distinct regions of the protein; each domain is responsible for different function (s)
motifs
specific combinations of secondary structures, which are organized into specific 3D structure inside the domains that account for the function
Transcription factors
dna binding domain
effector binding domain
oligomerization domain
DNA binding domain
part of the protein responsible for binding DNA, with structural motifs that read DNA sequence (the part of DNA binding domain which actually comes in contact with DNA)
effector binding domain
region of protein can be altered by binding a small molecule or covalent modification at effector site (star) causes conformational change in TF
oligomerization domain
structure to allow specific dimerization with similar TFs
DNA binding domains of regulatory proteins share
similar structural motifs
most frequent motif in DNA binding domain of bacterial repressors is
helix-turn-helix motif, about 20 amino acids long
2 short alpha helices (7-9 aa long) connected with a short turn
DNA recognition helix (binds specific DNA sequence) makes most of the contact with the DNA and the other stabilizes the interaction
recognition heli and stablizing helix (clier to N terminus) forms ~ 90 degree angle
first recognized in prokaryotes
in helix -turn-helix motfi which helix is stabilizing and recognixing
recognizing helix is #3 and stabliizing is #2,
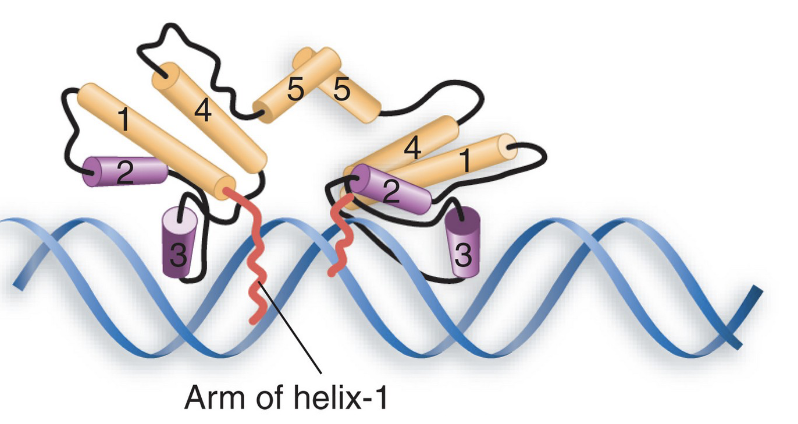
the N-terminal arms of the dimer’s monomers wrap around to the other face