Genetic Mechanisms: Conjugation, Transformation, and CRISPR
1/82
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced |
|---|
No study sessions yet.
83 Terms
CRISPR
Cluster of Regularly Interspaced Short Palindromic Repeats.
CRISPR immunity
An adaptive immune system that can establish immunity to a phage or transmissible plasmid after being exposed to it.
Adaptive immune system for bacteria
Cells can remember the phage that was infected before.
CRISPR response to phage
If the same phage comes back in the cell, the CRISPR will get rid of it.
F plasmid fertility inhibition
Involves the traJ gene, a transcriptional activator needed for traY, traX, and finO expression from ptraY.
Antisense RNA FinP
Blocks TraJ translation and is stabilized by FinO.
oriT site identification
Identified by cloning random plasmid fragments into a nonmobilizable vector, transforming cells with a self-transmissible plasmid, and testing for mobilization.
Griffith experiment
Showed bacterial transformation where nonencapsulated Type R bacteria convert to pathogenic Type S when mixed with heat-killed Type S.
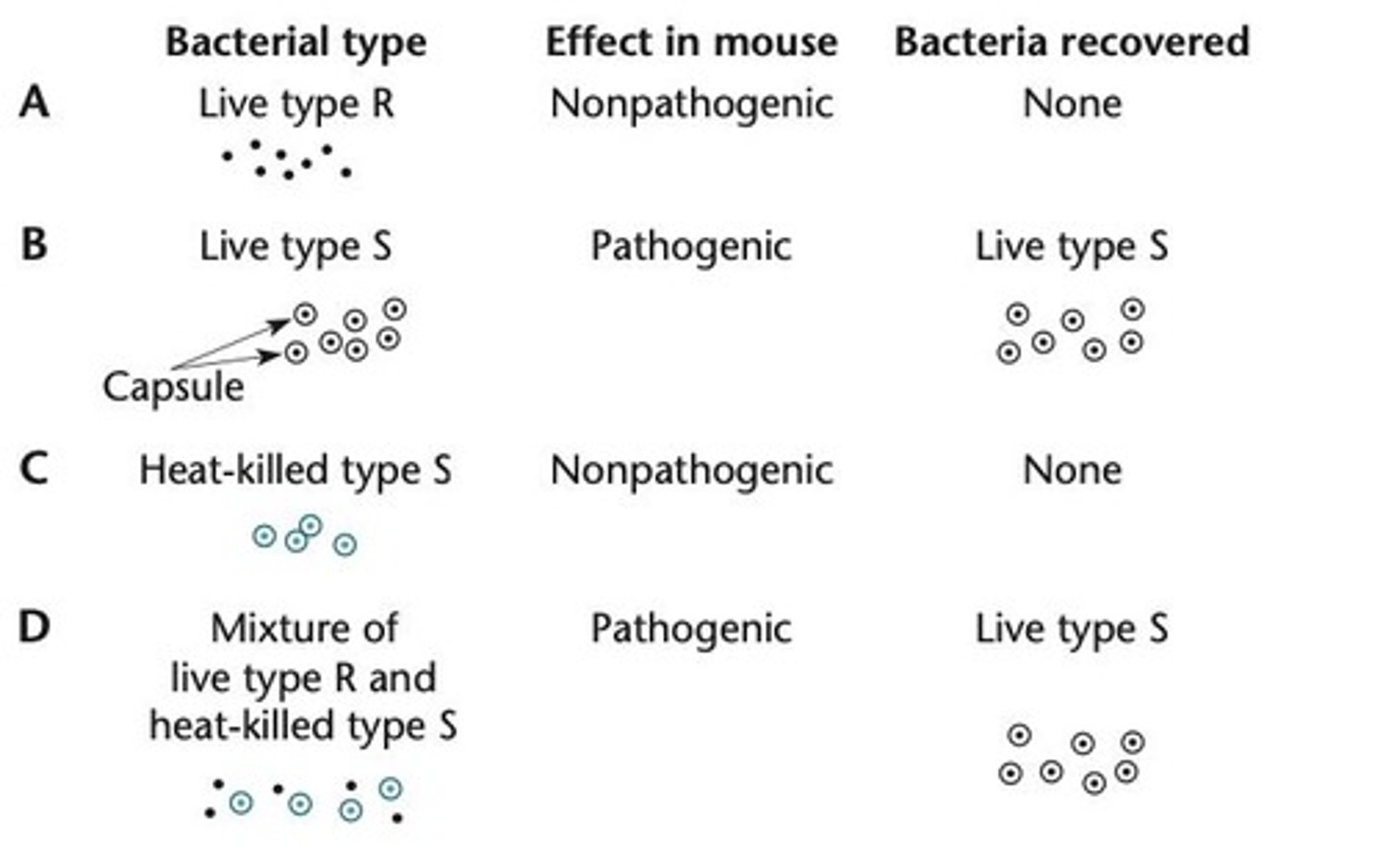
Efficiency of DNA uptake during transformation
DNA in the cell is insensitive to DNase; DNA that did not enter the cell is degraded and passes through a filter.
Uptake sequences on DNA
Only DNA with specific sequences is taken up by certain bacteria.
Genetic assay for DNA state during transformation
Only double-stranded DNA binds to initiate transformation; it becomes single-stranded after entering the cell.
Recombination in transformation
Donor DNA must be incorporated into the chromosome by recombination to transform new recipient cells.
Artificially induced competence
Treatment with calcium (CaCl2) and rubidium (RbCl2) ions can make some bacteria competent.
Competent cells
Can take up both double-stranded (Ds) and single-stranded (Ss) DNA.
Transformants
The presence of transformants in step 4 confirms that the DNA was double-stranded during DNase treatment.
Artificially induced competence
Treatment with calcium (CaCl2) and rubidium (RbCl2) ions can make some bacteria competent, not efficient so there is a need for a selectable marker.
Competent cells
Ds and Ss DNA can be taken up by the competent cells. Circular DNA is more efficient, though. Linear DNA is more susceptible to DNA degradation by exonucleases.
Transfection
The introduction of coral DNA or RNA.
Electroporation
Exposure to a strong electric field.
Washing cells
Important to wash cells to remove salts. Wash with 10% glycerol.
Universal transformation method
Most universal transformation method.
Protoplast
Cells without a cell wall.
EG exposure
Exposure of DNA in the presence of EG results in the trapping of DNA in the cytoplasm.
Regeneration of cell wall
The cell wall must be regenerated for the transformed cells to become viable.
Circular DNA
Circular DNA is a better substrate as it is more stable during the process.
Transformation efficiency
Transformation efficiency is expressed as the number of CFU per ug of DNA.
CFU calculation
You got 100 cfu from the transformation done with 1 ng of plasmid DNA, so 100/0.001 ug = 100,000 cfu/ug.
pET vector
Strategy for regulating the expression of genes cloned into a pET vector.
T7 RNA polymerase
The gene for T7 RNA polymerase (gene 1) is inserted into the chromosome of E. coli and transcribed from the lac promoter.
IPTG
The inducer IPTG is added to express the T7 RNA polymerase.
T7 late promoter
The T7 RNA polymerase transcribes the gene cloned into the pET vector downstream of the T7 late promoter.
T7 lysozyme
The T7 lysozyme encoded by a compatible plasmid, pLysS, binds to any residual T7 RNA polymerase made in the absence of induction and inactivates it.
lac operators
The presence of lac operators between the T7 promoter and the cloned gene further reduces transcription of the cloned gene in the absence of IPTG.
Polyacrylamide gel electrophoresis
Polyacrylamide gel electrophoresis of proteins synthesized during the development of phage T4.
Radioactive amino acids
The proteins were labeled by adding radioactive amino acids at various times after the phage was added to the bacteria.
SDS-PAGE
The proteins were denatured with sodium dodecyl sulfate (SDS) and run on an SDS-polyacrylamide gel electrophoresis (PAGE) gel.
Autoradiography
The gel was then exposed to autoradiography.
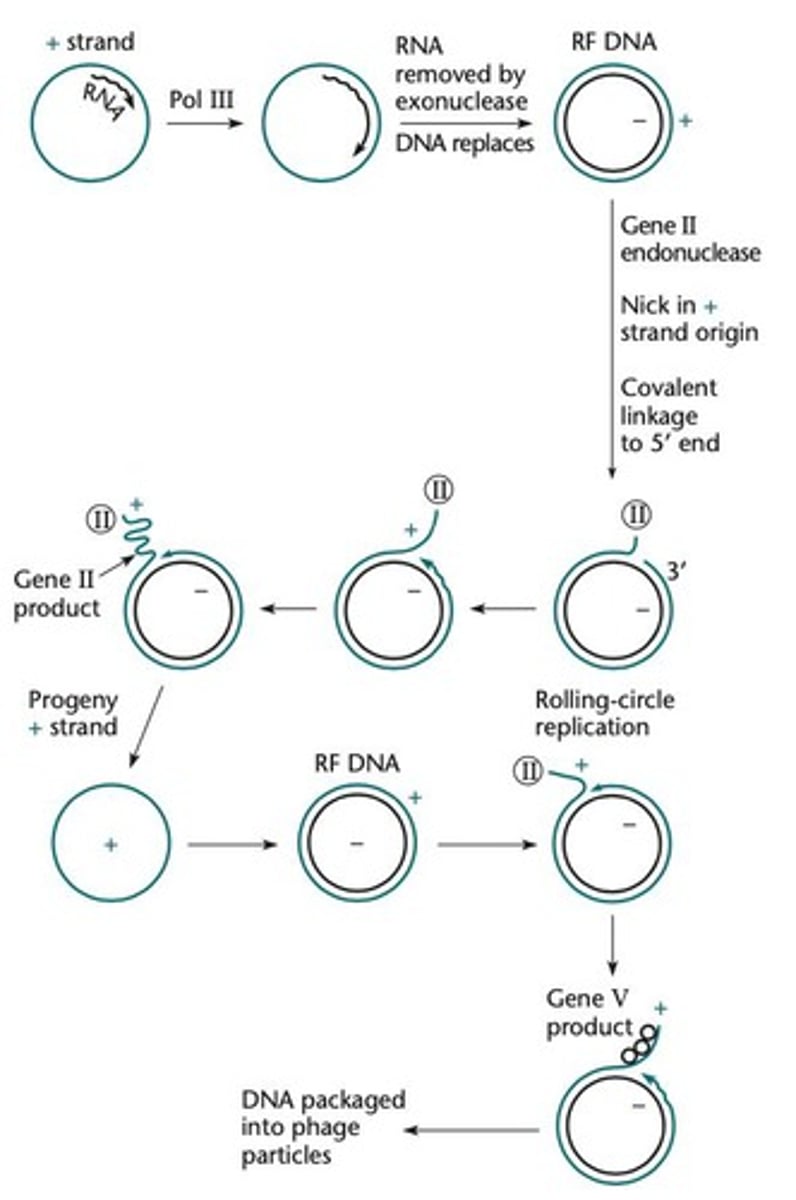
M13 phage replication
Replication of the circular single-stranded DNA phage M13.
Rolling-circle replication
More plus strands are synthesized via rolling-circle replication.
Gene II endonuclease
Nicks the plus strand, allowing rolling-circle replication to produce more plus strands.
Gene V
Binds the plus strands to prevent further RF synthesis and aids in packaging them into phage heads.
Holin
Becomes active, forming a pore that allows lysozyme to traverse the membrane and then degrade the cell wall and lyse the cell.
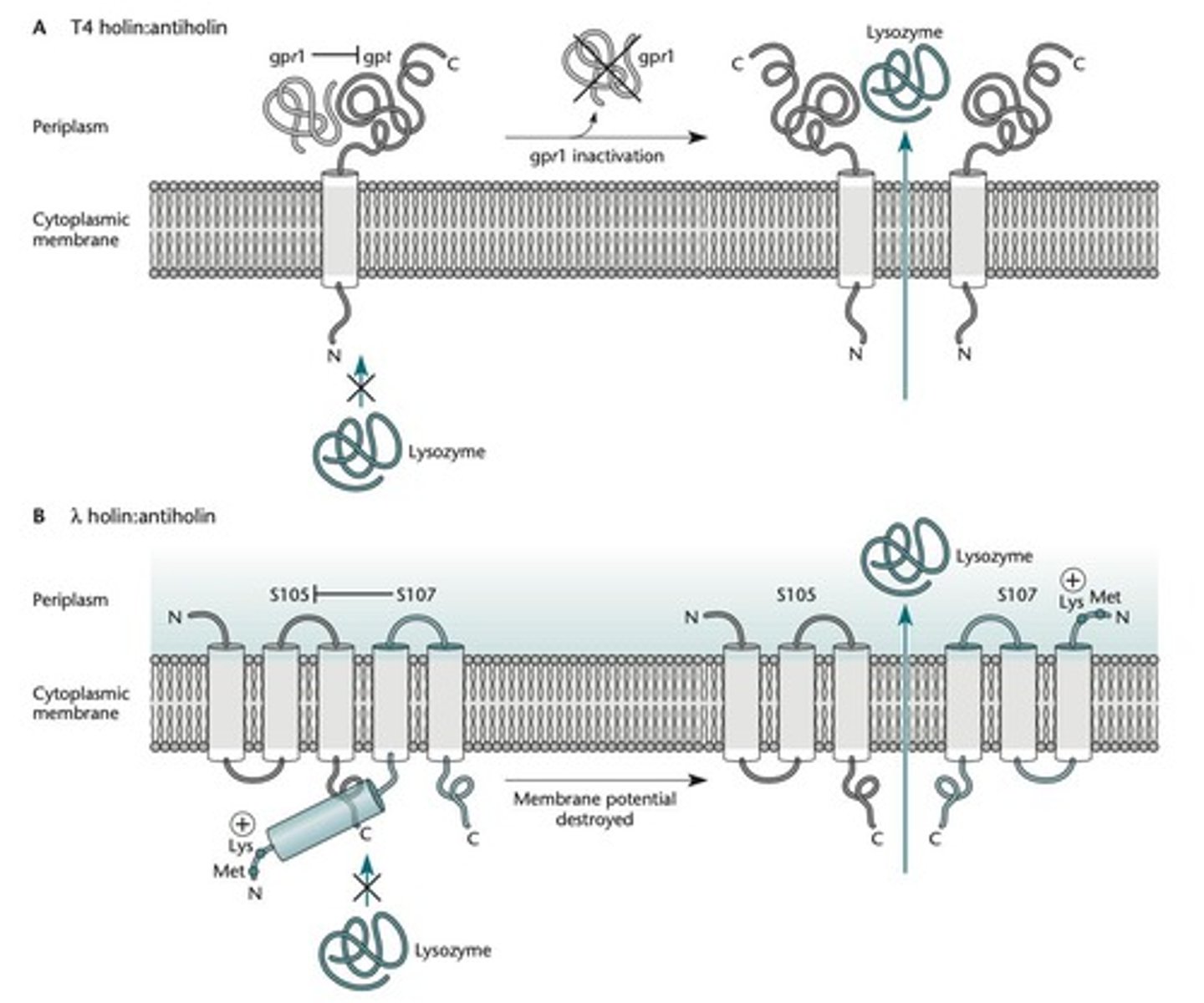
Anti-holin
Keeps the holin inactive until the time of lysis.
T4 phage model
If a phage is bound to the outside of the cell, the antiholin (gpr1) binds to the periplasmic domain of the holin (gpt), preventing it from forming pores.
λ phage model
The anti-holin (S107) has an extra 2 amino acids, Lys-Met, at its N terminus due to a second upstream translation initiation region (TIR).
S107 anti-holin
Inactive as a holin but still binds to the S105 holin, interfering with its ability to form pores.
Lysis timing
At the time of lysis, the membrane loses its potential, allowing TMD1 of S107 to enter the membrane and participate with S105 in the formation of pores.
CRISPR array
Transcribed into a single long RNA, which is cut in the repeat sequences to make guide CRISPR RNAs (crRNAs).
Protospacer sequence
The sequence on the phage DNA targeted by the crRNA during the immunity phase.
Conjugation
Involves the transfer of genetic material between bacterial cells through direct contact.
F plasmid
Carries the tra region, which is essential for conjugation.
TraJ
A transcriptional activator necessary for tra gene expression, regulated by antisense RNA (FinP) and FinO.
Plasmid mobilization
Some plasmids are not self-transmissible and rely on co-resident self-transmissible plasmids for mobilization.
oriT site
Crucial for plasmid mobilization.
Transformation
Only double-stranded DNA (dsDNA) binds to recipient cells to initiate transformation.
Eclipse phase
Single-stranded DNA (ssDNA) isolated during transformation cannot restart the process.
DNA uptake efficiency
Determined by DNA resistance to DNAse.
Chemical Induction
Treatment with CaCl2 or RbCl2 can make bacteria competent, though inefficient; a selectable marker is required.
Electroporation
Exposes cells to a strong electric field, requiring salt removal (washed with 10% glycerol) to prevent damage.
Protoplast Transformation
Cells without a wall uptake DNA in the presence of PEG; cell walls must regenerate for viability.
Transformation efficiency
Measured as CFU/µg of DNA.
M13 phage
A circular ssDNA phage.
RNA primer
Initiates minus-strand synthesis, forming a double-stranded replicative form (RF).
Gene II product
Nicks the plus-strand, facilitating rolling-circle replication.
Gene V
Binds plus-strand DNA, inhibiting further RF synthesis and aiding phage packaging.
DNA polymerase I (Pol I)
Removes RNA primers, fills gaps, and DNA ligase seals them.
T4 Phage Holins
Create pores in the membrane to allow lysozyme to degrade the cell wall.
gpr1 antiholin
Binds gpt holin, preventing premature lysis.
S107 antiholin
Prevents membrane insertion due to an extra Met-Lys charge.
Loss of membrane potential
Triggers lysis by allowing S107 insertion and pore formation.
F Plasmid
Governed by the tra region, which involves the regulatory action of the TraJ protein.
TraJ protein
Activated by other plasmid elements and regulates plasmid transfer.
RNA interference via antisense RNA (FinP)
Inhibits TraJ translation, regulating plasmid transfer.
oriT site
Critical for mobilization of non-self-transmissible plasmids.
Double-stranded DNA (dsDNA)
Competent for transformation, while single-stranded DNA (ssDNA) cannot be used after the 'eclipse phase.'
DNA uptake efficiency
Measured using radioactive-labeled DNA to track the process.
Chemical induction methods
Calcium chloride (CaCl₂) and rubidium chloride (RbCl₂) can induce competence in bacteria.
Electroporation
A method where a strong electric field makes bacterial membranes permeable to DNA.
Protoplasts
Cells without cell walls that can be transformed by DNA exposure in the presence of polyethylene glycol (PEG).
Transformation efficiency
Calculated by determining the number of colony-forming units (CFU) per microgram of DNA.
CFU calculation
Efficiency is calculated by dividing the number of CFUs by the amount of DNA used in the experiment.
Replication of Circular ssDNA Phage (M13)
Explains the rolling-circle replication mechanism using RNA primers and gene II endonuclease.
Holin activation
Describes the role of holins in the timing of phage lysis, with antiholins preventing pore formation.