3.8.4 Gene technologies allow the study and alteration of gene function allowing a better understanding of organism function and the design of new industrial and medical processes + 3.8.4.2 Differences in DNA between individuals of the same species can be exploited for identification and diagnosis of heritable conditions + 3.8.4.3 Genetic fingerprinting
1/63
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
64 Terms
What is Non-Coding DNA?
Non-coding DNA refers to regions of DNA that do not code for proteins.
What is the 3 functions of Non-Coding DNA?
Regulatory Roles:
Non-coding DNA contains promoters, enhancers, and silencers that control when and how genes are transcribed.
These regions influence which genes are switched on or off in different cell types or under different conditions.
RNA Molecules:
Some non-coding regions are transcribed into functional RNA molecules, such as tRNA, rRNA, and microRNA (miRNA), which regulate gene expression or assist in protein synthesis.
Structural DNA:
Non-coding regions contribute to the structure of chromosomes, such as telomeres (protective caps at the ends of chromosomes) and centromeres (important during cell division).
What does regulatory genes code for? Why are they important?
Regulatory genes code for proteins such as transcription factors, which control the activity of other genes by binding to DNA.
These genes determine whether a specific gene is transcribed into mRNA, affecting protein production.
What is the Challenges in Translating Genomes into Proteomes
Selective Gene Expression:
Not all genes are active in every cell at all times.
Genes are selectively expressed depending on the cell type, developmental stage, or environmental conditions.
Complexity in Eukaryotes:
In higher organisms, a significant proportion of the genome is non-coding DNA, making it difficult to predict the proteome (all proteins a genome can code for).
Alternative Splicing:
A single gene can produce multiple proteins through alternative splicing, where different combinations of exons are joined.
What is the aim of sequencing projects
Sequencing projects aim to determine the complete sequence of nucleotides in an organism’s genome.
Genome sequencing in simpler organisms
In simpler organisms (e.g., bacteria), genome sequencing helps determine the sequences of proteins derived from their genetic code.
This is useful for identifying potential antigens for use in vaccine production.
Genome sequencing in complex organisms
In more complex organisms, the presence of non-coding DNA and regulatory genes means that understanding the genome cannot directly predict the proteome.
The proteome refers to all the proteins a genome can code for. However, not all proteins are produced in every cell due to selective gene expression.
Benefits of genome sequencing
Genome-Wide Comparisons:
Between Species:
Identifies evolutionary relationships and how species have diverged.
Between Individuals:
Detects genetic differences that can guide the development of personalised medicine and improve the understanding of human diseases.
Predicting Protein Structure:
Enables scientists to predict the amino acid sequences of polypeptides, aiding in the study of protein function.
Synthetic Biology:
Facilitates the design and synthesis of novel biological systems and molecules using gene sequences.
Recombinant DNA technology
Recombinant DNA technology involves the transfer of fragments of DNA from one organism, or species, to another
Transgenic organisms can express the transferred gene to produce what? Explain why this occurs
Transgenic organisms can express the transferred gene to produce the encoded protein and express new genetic traits. This process is possible due to the universal nature of the genetic code and the similarity of transcription and translation across different organisms.
Steps to Producing fragments of DNA in Recombinant DNA technology
1. Using Reverse Transcriptase to Make DNA
2. Using Restriction Endonucleases to Cut DNA Fragments
3. Gene Machine
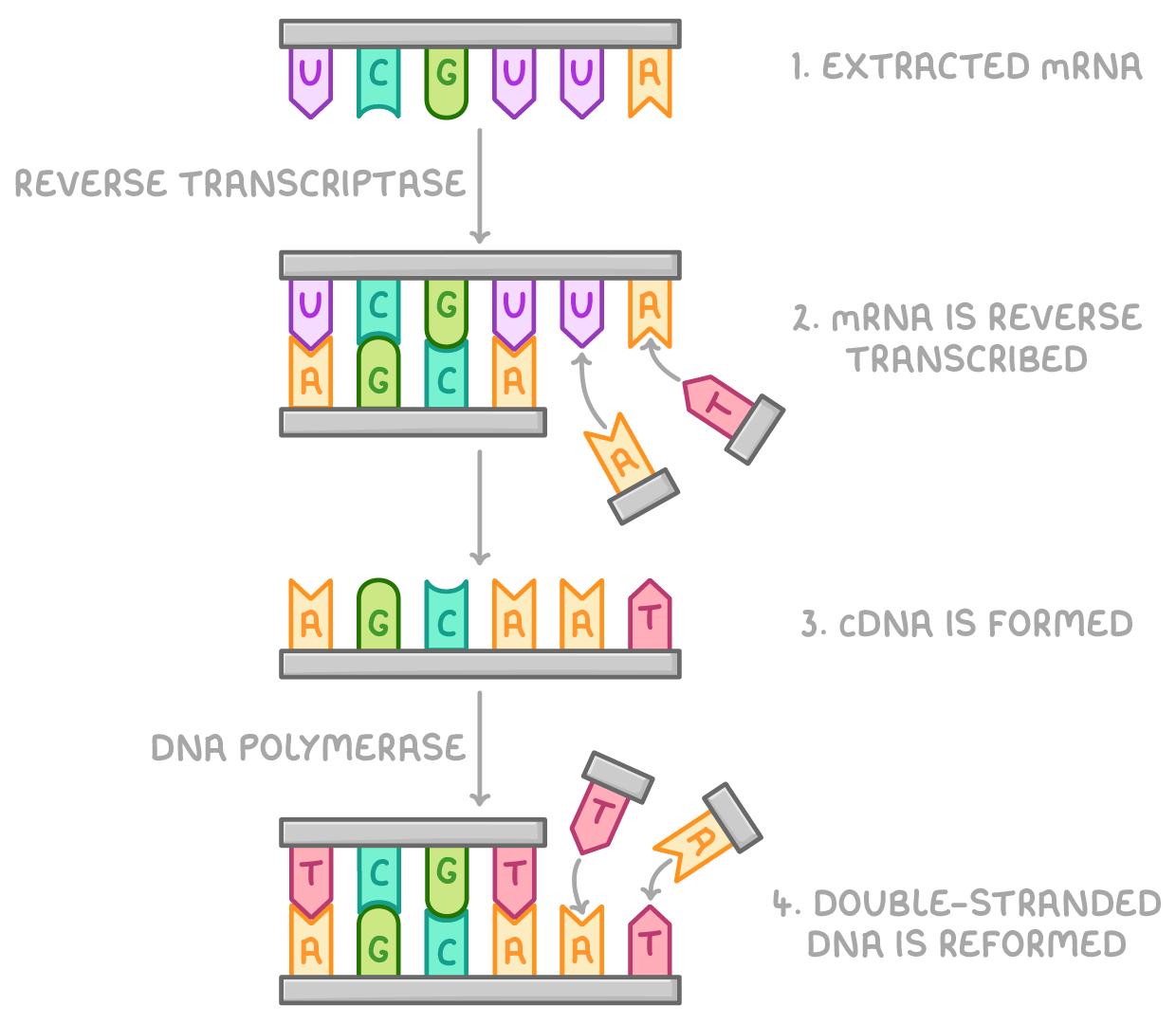
1. Using Reverse Transcriptase to Make DNA
mRNA is extracted from cells.
mRNA is reverse transcribed using the reverse transcriptase enzyme and DNA nucleotides.
This makes a cDNA strand identical to the original DNA strand, and cDNA is isolated from the mRNA strand.
cDNA, free nucleotides, and DNA polymerase can then be used to form the other strand of DNA, reforming the desired gene.
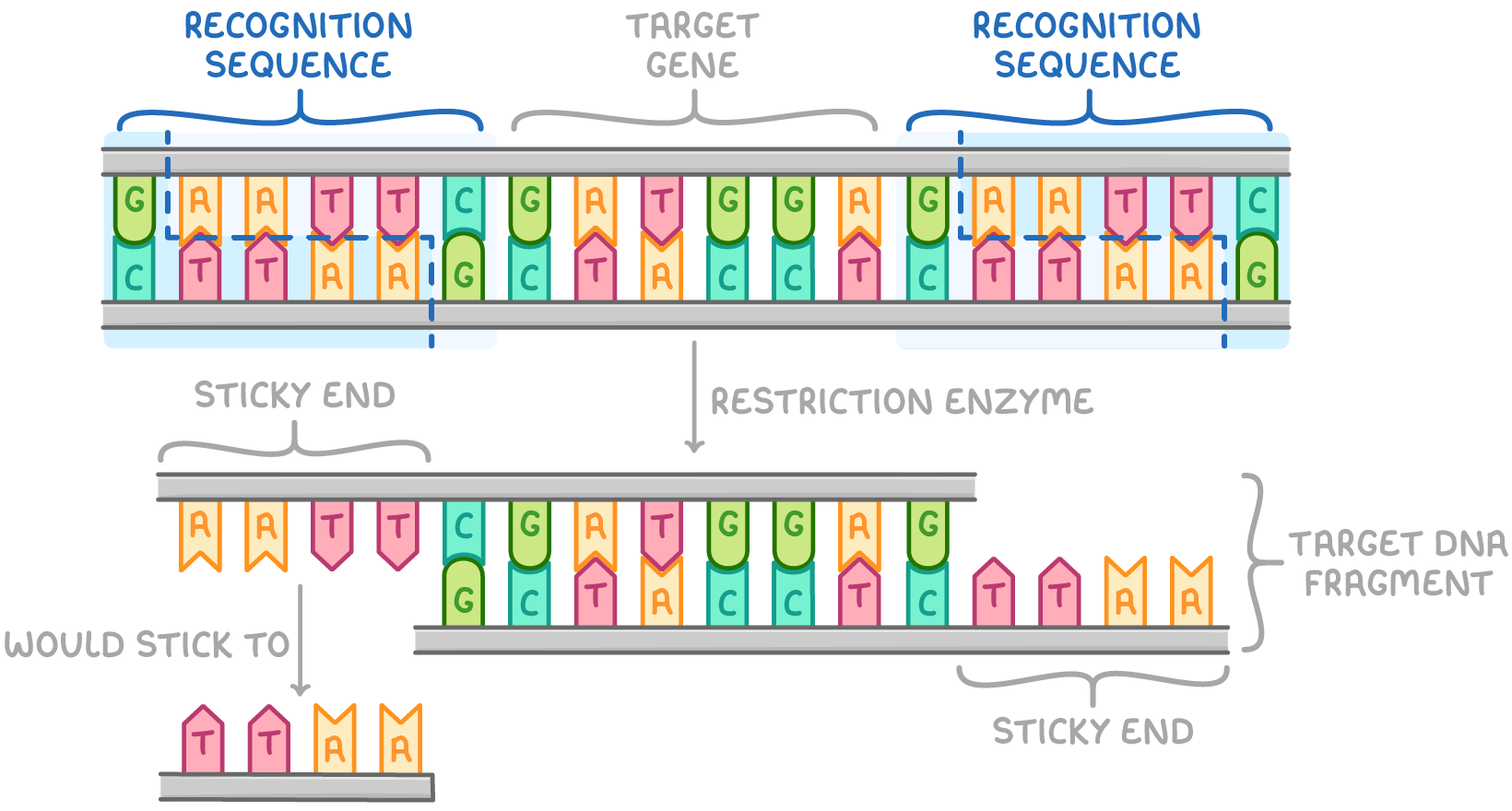
2. Using Restriction Endonucleases to Cut DNA Fragments
DNA is incubated with chosen restriction enzyme(s).
Restriction enzymes identify palindromic sequences in the DNA double helix and cut double-stranded DNA if their recognition sequence is present.
Recognition sequences at either end of a desired DNA fragment allow restriction enzymes to separate the fragment from the rest of the DNA to obtain the desired gene.
Enzymes cut target gene fragment out via hydrolysis reaction.
Different restriction enzymes cut at different sequences based on their active site shape.
Difference between sticky ends and blunt ends
Sticky Ends:
Some restriction endonucleases make staggered cuts, leaving single-stranded overhangs.
These overhangs are called sticky ends because they can easily bond with complementary sequences on other DNA fragments.
Sticky ends enable easier recombination.
Blunt Ends:
Other restriction enzymes cut straight across the DNA, leaving blunt ends with no overhangs.
Blunt ends are less efficient for recombination as they lack complementary overhangs.
What is the importance of Sticky Ends
When DNA fragments are cut with the same restriction endonuclease, the sticky ends of both fragments will be complementary.
This means the fragments can base-pair with each other.
DNA ligase is then used to form strong covalent phosphodiester bonds between the fragments, creating a stable recombinant DNA molecule.
3. Gene machines (DNA synthesisers) can now synthesise completely new DNA sequences.
Choose codons for the desired amino acid sequence from a known protein structure.
Use a computer to direct the synthesis of short fragments of DNA (oligonucleotides) in gene machines.
Join the fragments together to make a longer sequence of nucleotides, forming the desired gene.
Polymerase chain reaction (PCR) constructs a complementary DNA strand and amplifies the gene to produce multiple copies.
This method produces custom DNA without needing a natural template.
Define Reverse transcriptase
Makes DNA from RNA.
Define Restriction enzymes
Cut DNA at specific sites, producing sticky ends for recombination
Define Vectors
Carry DNA into host cells; plasmids are common vectors
Define Gene markers
Detect successful plasmid uptake
Define Electroporation
Enhances plasmid uptake by bacterial cells
What is in vivo cloning
A process that involves replicating cells or organisms within a living organism
Steps for in vivo cloning
Inserting a DNA fragment into a vector
Using vectors to transfer DNA into host cells
Identifying transformed host cells
The need for promoter and terminator regions
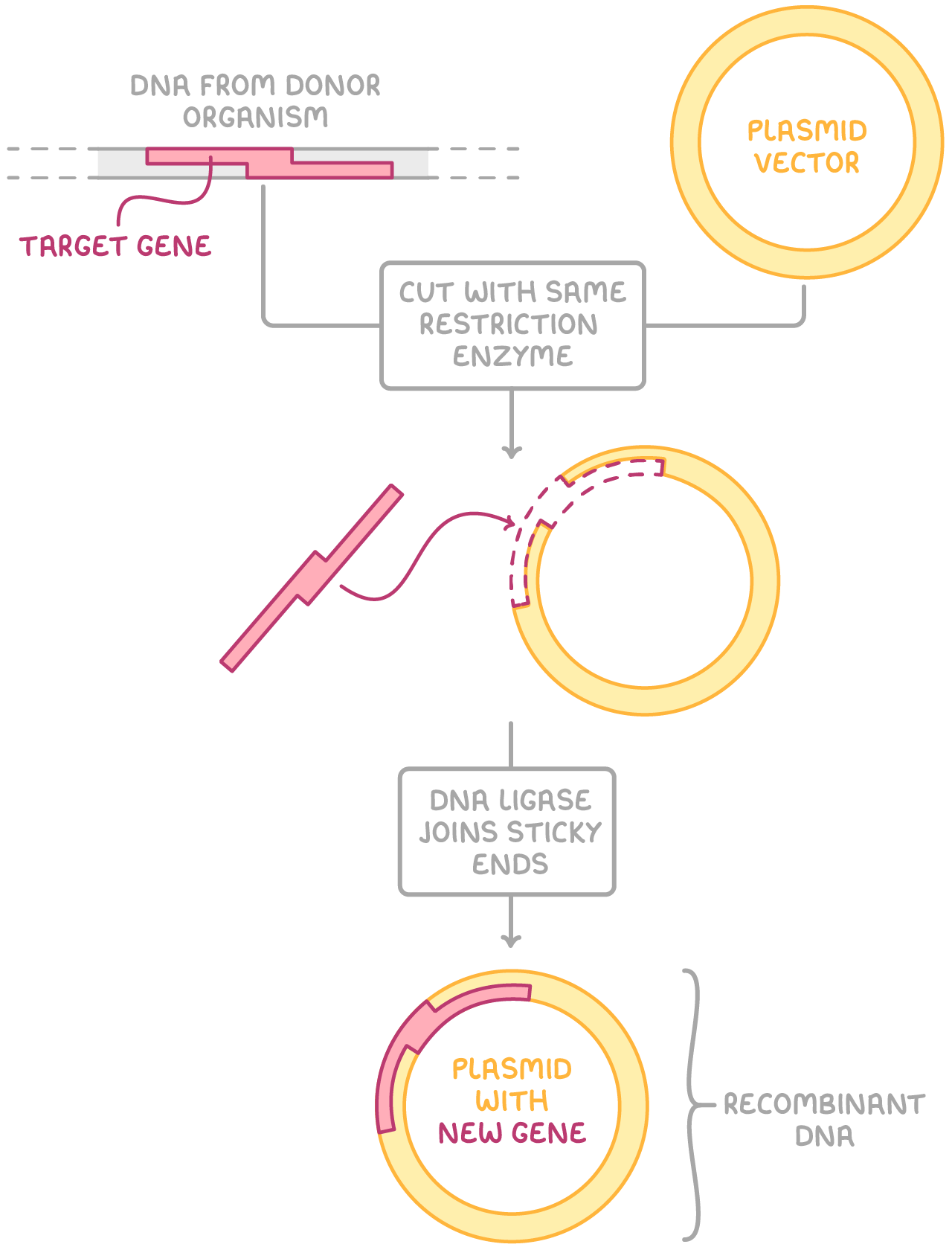
Step 1 - Inserting DNA fragments into vectors
(The process of producing large quantities of a target DNA fragment in living cells is sometimes called 'in vivo cloning' or 'in vivo DNA amplification'. It begins with inserting a target DNA fragment into a vector. This vector is then capable of transferring the DNA into a host cell.)
A vector is cut open at a specific site using a restriction enzyme, creating sticky ends.
The same restriction enzyme is used to cut the target DNA fragment, creating complementary sticky ends.
DNA ligase forms phosphodiester bonds between the sugar and phosphate groups on the two strands of DNA, joining the sticky ends of the vector and DNA fragment together.
The newly formed combined DNA molecule is known as recombinant DNA.
Step 2 - Transferring recombinant DNA into host cells
(Transformation involves introducing vectors with recombinant DNA into host cells, transforming these cells. The vectors are usually either plasmids or bacteriophages.)
1.) Plasmid vectors:
These are small, circular DNA molecules, typically found in bacteria.
Host cells are treated to enhance the uptake of plasmids that have recombinant DNA.
For instance, applying calcium ions and temperature shifts can make bacterial membranes more permeable to plasmids.
2.) Bacteriophage vectors:
These are viruses that infect bacteria.
Bacteriophages inject their DNA into host bacterial cells during infection.
The phage DNA, now carrying the recombinant DNA, inserts into the host's DNA.
Step 3 - Identifying transformed host cells
(As only around 5% of host cells may uptake DNA, it's important to identify which cells have been successfully transformed.)
Marker genes indicate which host cells took up recombinant DNA:
They are inserted into vectors alongside target genes.
Transformed cells are cultivated on selective agar plates.
Only transformed cells display the characteristics encoded by marker genes.
These transformed cells can then be cultured to mass-produce the target DNA fragment through cellular replication.
Examples of marker genes include
A marker gene for a specific trait, like antibiotic resistance, ensures that only transformed cells form colonies.
A marker gene that is visible under UV light like green fluorescent protein (GFP).
Inserting a marker gene within the GFP gene inhibits fluorescence if it is successfully incorporated.
A marker gene coding for an enzyme that alters the colour of a specific substrate.
Step 4: Adding promoter and terminator regions
(For the protein coded by an inserted DNA fragment to be produced, vectors require certain control elements.)
Control elements that a vector needs:
Promoter regions - Provide a binding area for RNA polymerase, signalling where the transcription of the gene should start.
Terminator regions - Indicate where transcription should end, releasing RNA polymerase and terminating transcription.
Without these essential regions, the production of proteins from recombinant DNA would not occur. These promoter and terminator regions might already be present in the vector or may need to be added separately.
What is the polymerase chain reaction?
The polymerase chain reaction (PCR) is a method for amplifying DNA fragments rapidly and efficiently. 'Amplifying' means that a large number of DNA fragments are being produced.
PCR is an automated technique that does not require living cells to rapidly replicate specific DNA fragments. It is sometimes called 'in vitro cloning'.
Components required for PCR
DNA fragment - This is the specific target DNA template sequence that needs to be replicated.
Primers - These are short sequences of nucleotides that attach to the start and end of the DNA fragment to be copied.
DNA polymerase - This is an enzyme that creates new DNA strands by adding DNA nucleotides.
Free nucleotides - The building blocks (A, T, C, and G) that are used to construct the new DNA strands.
Thermocycler - This is a device that precisely heats and cools the PCR mixture to facilitate the reaction.
(A type of DNA polymerase that can withstand high temperatures without denaturing is used to ensure it remains active throughout the process. An example is Taq polymerase from thermophilic bacteria.)
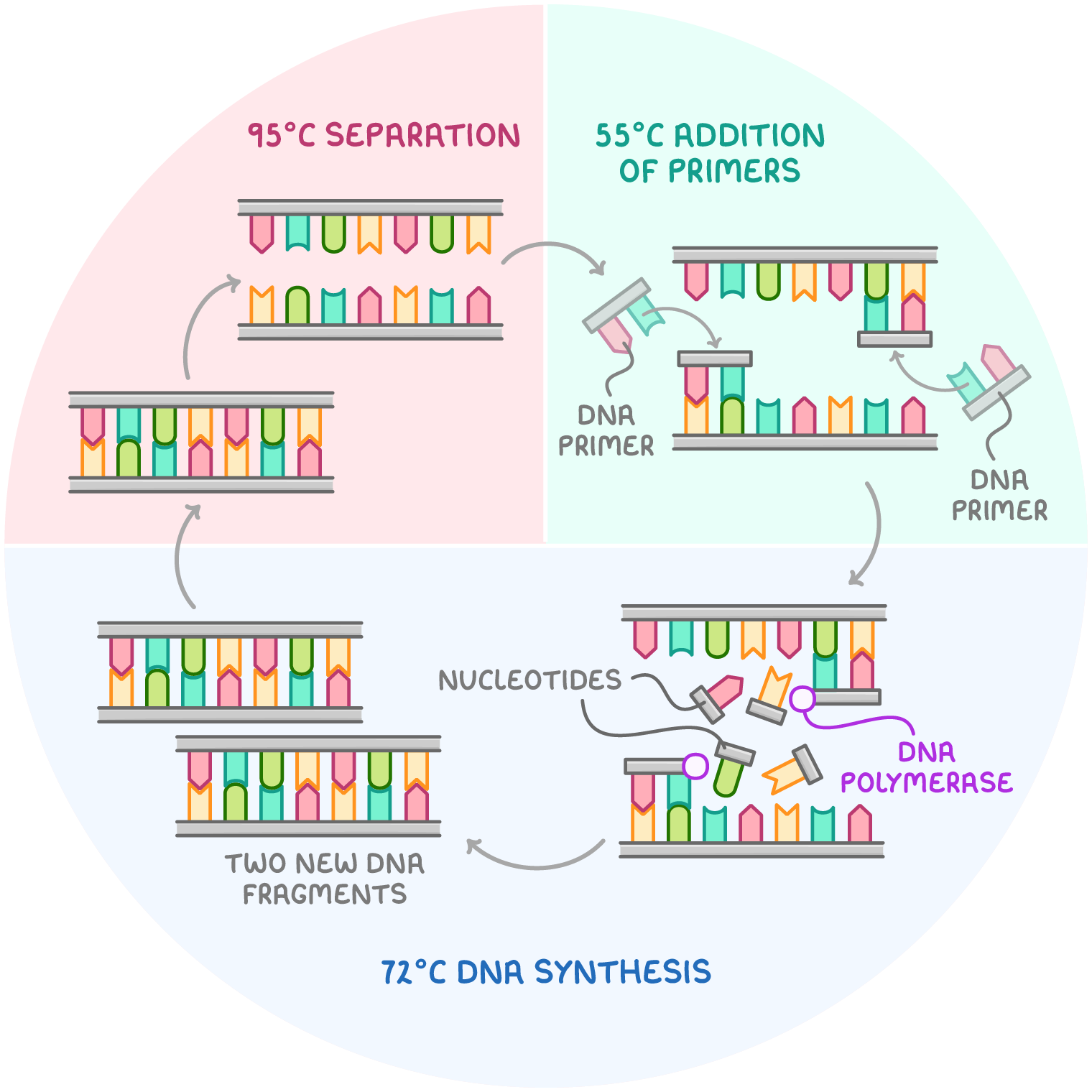
The three main stages of the polymerase chain reaction
Stage | Temperature | Process |
|---|---|---|
1. Separation (denaturation) | 95°C | Heating the DNA separates the hydrogen bonds between its two complementary strands. |
2. Addition of primers (annealing) | 55°C | The primers attach to the specific starting points on each of the separated DNA strands by forming hydrogen bonds. |
3. DNA synthesis (extension) | 72°C | DNA polymerase adds free nucleotides to the ends of the primers, extending the DNA strand to form a complete copy. |
The advantages of the polymerase chain reaction
Rapid speed - It can produce approximately 100 billion copies in just a few hours, which is not possible with in vivo cloning.
Precision - The automated nature of thermocycling ensures accurate amplification of the DNA fragment.
Low DNA needs - Even tiny samples can be amplified to produce a large quantity for analysis.
No cells needed - This method is simpler and more straightforward than traditional in vivo techniques, which are typically slower and more complex.
What is Genetic engineering (Crops)
Genetic engineering is a tool that can enhance crop plants by making them resistant to herbicides or insects. Competition with weeds and insect pests can significantly reduce yields, impacting global food security so genetically modified (GM) crops can be very useful.
E.g.
(Soybeans are a vital food source worldwide, and insect pests feeding on these plants can drastically lower yields.)
(Through genetic engineering, soybeans have been modified to include a gene from the bacterium Bacillus thuringiensis (Bt).)
(The Bt gene produces a protein that is toxic to many insect pests but is harmless to humans and other organisms.)
Advantages/Disadvantages of GM crops
Advantages of GM crops | Disadvantages of GM crops |
|---|---|
Involves less labour than traditional herbicides and pesticides. | Encourage monocultures, reducing biodiversity. |
Cheaper than traditional herbicides and pesticides. | Herbicide-resistant GM plants may crossbreed with wild plants, creating herbicide-resistant weeds. |
Herbicide-resistant GM crops allow the use of herbicides to kill weeds without harming the crop, increasing crop yield. | Loss of traditional crop varieties reduces genetic diversity, which makes crops vulnerable to disease and climate changes. |
Insect-resistant GM crops are less likely to be killed by pests, increasing crop yield. | Non-target insects could be harmed by the toxin coded for by insect-resistant GM crops. |
Insect-resistant GM crops reduce the need for pesticides, benefiting the environment. | Insect pests may evolve resistance to the toxin coded for by insect-resistant GM crops. |
Advantages of GM farmed animals
Improves quality e.g. by enhancing disease resistance.
Improves quantity e.g. by enabling year-round reproduction.
Improves productivity e.g. through faster growth.
What is Pharming
Pharming involves the production of pharmaceuticals and human medicines by inserting human genes into other animals. This process allows for the extraction of pharmaceutical proteins from GM animals' milk or blood at high yields.
For example, goats can be genetically engineered to produce antithrombin:
Antithrombin deficiency puts individuals at risk of excessive blood clotting, requiring regular antithrombin treatment.
GM goats can produce the antithrombin protein in their mammary glands, offering a new source of this crucial treatment.
Evaluate Pharming
Advantages of pharming | Disadvantages of pharming |
|---|---|
Enables mass production of rare treatments. | Raises animal welfare concerns. |
Makes drugs more accessible. | Can lead to animals being viewed solely as commodities. |
Creating genetically engineered microorganisms
GM microorganisms have many applications, including the development of new treatments for diseases. By genetically altering pathogens, researchers can turn traditional disease-causing organisms into tools for medicine.
For example, poliovirus can be engineered to target cancer:
Cancer cells often possess receptors for the poliovirus.
Disabling specific genes within the poliovirus can make a version that targets and kills cancer cells without causing polio.
This innovative approach holds promise for developing new cancer treatments.
Evaluate GM pathogens
Advantages of GM pathogens | Disadvantages of GM pathogens |
|---|---|
Offers potential treatments for diseases previously deemed incurable. | Carries a risk of accidental infections and disease outbreaks. |
Can produce essential medicines, antibiotics, and enzymes, such as insulin. | There's a danger that engineered pathogens could revert to their original, harmful form. |
Useful in creating vaccines. | Could be misused in biological warfare. |
Facilitates the creation of DNA libraries for research. |
To minimise these risks, researchers adhere to strict safety protocols.
Ethical issues surrounding patents in genetic engineering
Patenting GMOs introduces ethical concerns:
The legal patenting of engineered organisms raises questions about global access.
This especially affects smallholder farmers in developing countries who may be unable to afford patented seeds due to high costs and legal barriers.
There is a need for a balance between rewarding innovation and ensuring ethical access for those in need, highlighting the importance of equitable patent policies.
What is a DNA probe?
A DNA probe is a short, labelled, single-stranded DNA sequence that can identify a specific allele. It does this by binding to a complementary DNA sequence.
(For example, a DNA probe can determine whether an individual carries a mutated allele responsible for a genetic disorder.)
What needs to be done to create DNA probe
To create a DNA probe, the target allele's base sequence is first identified using DNA sequencing so that a complementary probe can be made. Then, the sequence is replicated using PCR.
The process of screening DNA for a specific allele using a DNA probe:
Denaturation - An individual's DNA strands are heated causing them to separate into two single strands.
Fragmentation - Digest the DNA sample into fragments using restriction enzymes.
Mixing - Add fluorescently or radioactively labelled DNA probes.
Hybridisation - DNA probes bind with the target sequence.
Identification - Visualise any DNA probe that has hybridised to DNA fragments under UV light (for fluorescently labelled DNA probes) or X-ray (for radioactively labelled probes) to signal whether the allele is present.
What is DNA microarrays
DNA microarrays allow for the simultaneous screening of many genes using multiple DNA probes through a process called high-throughput screening. Glass microarray slides have many different immobilised DNA probes arranged in specific places on a grid.
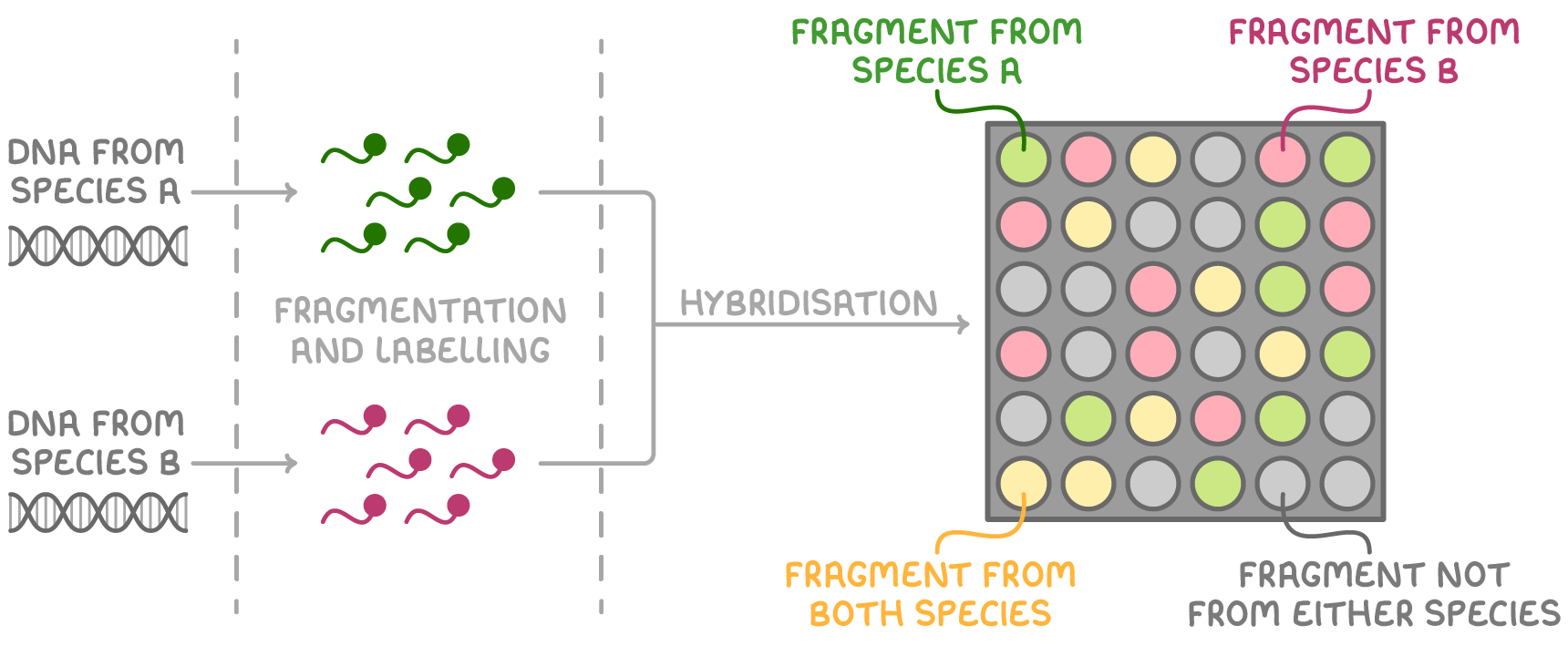
How does DNA microarrays work:
Extract DNA from the species or cells of interest.
Fragment and fluorescently label these DNA samples.
Mix the labelled DNA and wash it over the microarray probes.
DNA fragments that match any of the probes will hybridise.
The array is then viewed under UV light.
Any fluorescent spots show that the sample contains those fragments.
Applications of DNA probes in genetic screening
Identifying individuals carrying specific alleles that may lead to genetic disorders like Huntington's disease and cystic fibrosis.
Determining how patients will respond to certain drugs, which is crucial for targeted cancer therapies.
Evaluating susceptibility to diseases due to inherited or mutated alleles, aiding in preventive healthcare decisions.
There are concerns that probes might lead to discrimination in insurance or employment by revealing high-risk alleles.
Genetic counselling
Genetic counselling uses the outcomes of genetic screening tests and research into family history to provide advice on the implications of genetic disorders in individuals and the risks of them being inherited by offspring.
Process of genetic counselling
Discussing the significance and risks of screening tests.
Clarifying the results and the likelihood of associated diseases developing in individuals and their offspring.
Offering information on preventive measures or treatments if mutations are detected.
Importance of genetic councelling
Genetic counselling helps individuals make informed decisions about family planning, risk-reducing surgeries, and other health-related choices.
Examples:
Counselling an individual with a BRCA1 mutation about their elevated risk of breast cancer and preventive options such as mastectomy.
Advising a couple, both carriers of the sickle cell anaemia allele, on the probability of having an affected child and possible interventions like embryo screening.
What does genetic makeup influence
Genetic makeup influences how a person reacts to medications.
Personalised medicine
Personalised medicine tailors healthcare for optimal treatment based on an individual's genetic makeup, ensuring the most effective drugs are chosen.
How does gene therapy seeks to correct genetic disorders
Silencing defective dominant alleles that could lead to dominant genetic disorders.
Adding a working dominant allele to silence defective recessive alleles that could cause recessive genetic disorders.
Using vectors such as viruses or liposomes to transport healthy alleles into cells.
Focusing on somatic cells for treating individuals or germline cells to prevent passing the disease to offspring.
What is gel electrophoresis?
Gel electrophoresis is a technique used to separate molecules such as DNA, RNA, or proteins based on size by using an electric current applied to an agarose gel matrix.
Describe how to set up gel electrophoresis:
Insert a gel tray with solidified agarose gel into a gel tank.
Ensure the wells are close to the negative electrode to position the gel correctly.
Pour a buffer solution over the gel until it is submerged to maintain a constant, suitable pH throughout the experiment.
How to run electrophoresis on DNA samples
Mix the DNA samples with loading dye to make them visible.
Carefully deposit equal volumes of each sample into the wells using a micropipette.
Touch the micropipette tip to the buffer, not the bottom of the gel, to prevent damaging the gel.
Keep a record of which sample is in each well for later analysis.
DNA molecules carry a negative charge due to their phosphate groups.
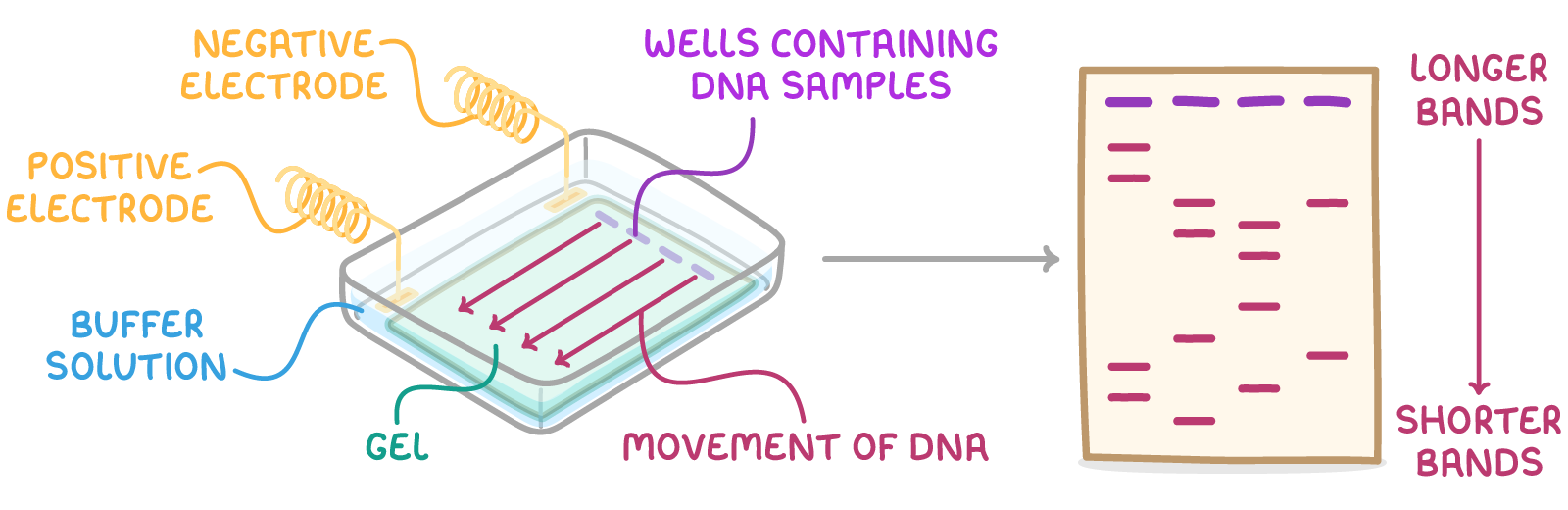
Explain what happens during electrophoresis
A voltage (around 100 V) is applied across the gel.
Fragments of DNA move towards the positive electrode (anode).
The smaller fragments travel faster and thus separate by size.
Continue the process for about 30 minutes or until the dye approaches the end of the gel.
(The size-based separation occurs because the gel's mesh matrix slows down larger fragments more than the smaller ones. The pore size of the gel affects the rate at which fragments move.)
How to visualise the result
Switch off the voltage and remove the gel from the tank.
Apply a stain to the DNA to reveal the bands of fragments.
Assess the migration distances of the bands to approximate the sizes of the fragments.
What is genetic fingerprinting?
Genetic fingerprinting, sometimes called DNA profiling, is a technique used to identify unique DNA patterns in individuals, to help identify individuals in forensics or family relationships.
Why does genetic fingerprinting work?
Genetic fingerprinting relies on the fact that, with the exception of identical twins, every person's DNA sequence is distinct. This is due to variation in the sequence and length of unique non-coding, repetitive DNA segments called variable number tandem repeats (VNTRs, sometimes called minisatellites).
Key features of VNTRs:
They are present across the genomes of most eukaryotes.
They are not involved in protein coding.
They have extensive variability in sequence and length among individuals.
Their length and location are heritable.
The chance of two individuals who are not identical twins sharing identical VNTR patterns is very low.
A high similarity in VNTR patterns indicates two individuals may be closely related.
(Short tandem repeats (STRs, sometimes called microsatellites) are repeated sequences of nucleotides that are smaller than VNTRs and can also be used in genetic fingerprinting.)
The process of creating a genetic fingerprint
DNA extraction - DNA is extracted from a tissue sample and amplified using PCR.
DNA digestion - Restriction enzymes are used to cut the DNA into fragments at points near the VNTR sequences.
Fragment separation - Gel electrophoresis separates the fragments by size, and they are denatured to produce single strands.
Hybridisation - Specific radioactive or fluorescent probes bind to complementary VNTR sequences.
Development - The positions of the probes are revealed, creating a barcode-like pattern of DNA bands unique to each individual.
What if radioactive probes are used
X-ray images are taken of the paper or membrane during the development stage. If fluorescent probes are used, the paper or membrane is placed under UV light causing the fluorescent probes to glow.
(The probability of a match can then be determined by computer analysis of the band patterns. The higher the number of matching bands, the greater the likelihood that the fingerprints are from the same individual.)
Key uses of genetic fingerprinting technology
Establishing paternity by matching bands on a child's genetic fingerprint to those of potential parents and assessing the similarities between VNTR patterns.
Identifying suspects from crime scene DNA (e.g. in blood, semen, saliva, skin cells, or hair roots).
Supporting criminal convictions with match probability calculations.
Identifying the risk of genetic disorders and predicting their onset and severity.
Selecting desirable traits in plants and animals for selective breeding while preventing severe inbreeding.
Evaluating genetic diversity by comparing the variety of genetic fingerprints within a population.
Limitations of genetic fingerprinting
There are a few limitations of genetic fingerprinting.
These limitations include:
Environmental contamination may compromise results.
Close genetic relatives could have similar fingerprints.
Assumptions about genetic variation in populations underpin probability calculations, meaning genetic fingerprinting does not prove guilt or causation and other evidence must also be taken into account.
Despite these limitations, genetic fingerprinting remains an invaluable tool, with its accuracy continually enhanced as DNA sequencing technologies advance.